Herpesviral lytic gene functions render the viral genome susceptible to novel editing by CRISPR/Cas9
Figures
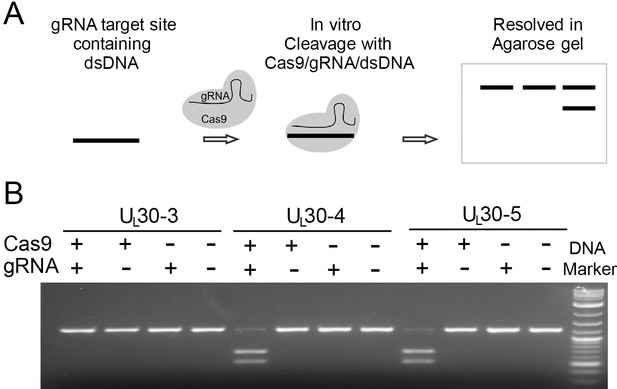
In vitro cleavage assay.
(A) Schematic diagram of in vitro cleavage assay (B) Results are shown for three sgRNAs targeting UL30 (UL30-3, -4, and -5). T7 in vitro transcribed sgRNA was combined with SpCas9 protein and a PCR template containing the CRISPR sequence, incubated 1 hr at 37°C and run on an agarose gel. Lane (1) SpCas9+sgRNA, lane (2) Cas9 only, lane (3) sgRNA only, lane (4) no Cas9/sgRNA. Efficient cutting is seen for UL30-4 and -5 but not UL30-3.
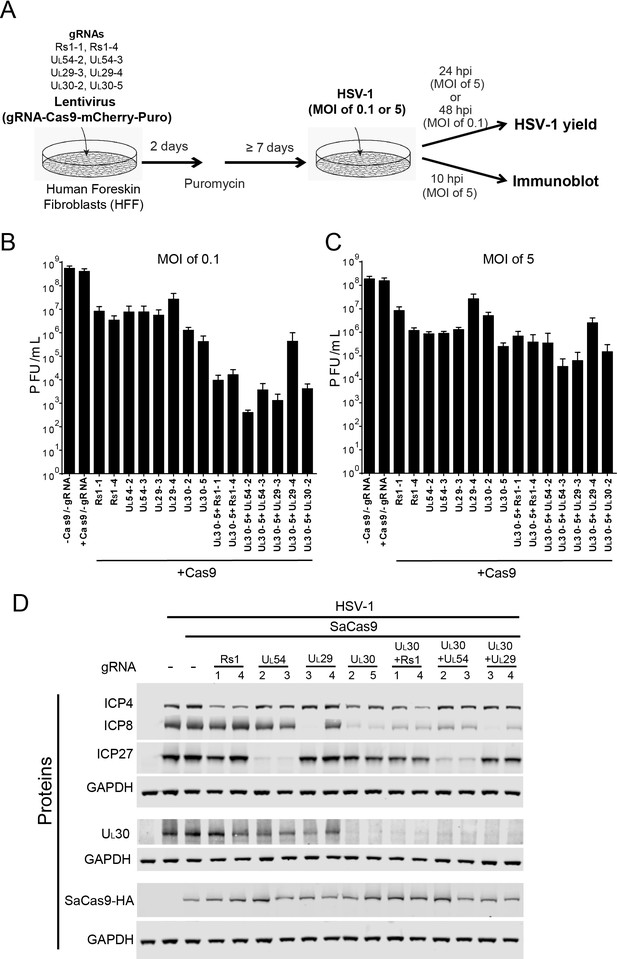
Effect of CRISPR-Cas9 on HSV-1 lytic infection.
(A) Experimental scheme of SaCas9/sgRNA-mediated inhibition of HSV lytic infection. (B and C) HFFs transduced with lentivirus expressing SaCas9 and sgRNAs were infected with HSV-1 at an MOI of 0.1 (C) or 5 (D) and harvested at 48 hpi or 24 hpi, respectively. Viral yields were determined by plaque assays. The histogram shows the mean values and standard deviations of biological replicates at an MOI of 0.1 (N = 3) or at an MOI of 5 (N = 4). All the sgRNA added conditions showed statistical significance compared to +Cas9 /-gRNA (one-way ANOVA with Dunnett’s multiple comparisons test, p<0.0001 (MOI of 0.1) and p<0.01 (MOI of 5, except for UL29-4 (p<0.05))). (D) HFFs transduced with lentivirus expressing SaCas9 and sgRNA were infected with HSV-1 at an MOI of 5 and harvested at 10 hpi. Proteins were detected using immunoblotting with antibodies specific for the indicated proteins. Three SDS-PAGE gels loaded with the same amount of proteins were used to detect multiple proteins. Immunoblots of GAPDH are shown as a control under the individual immunoblots.
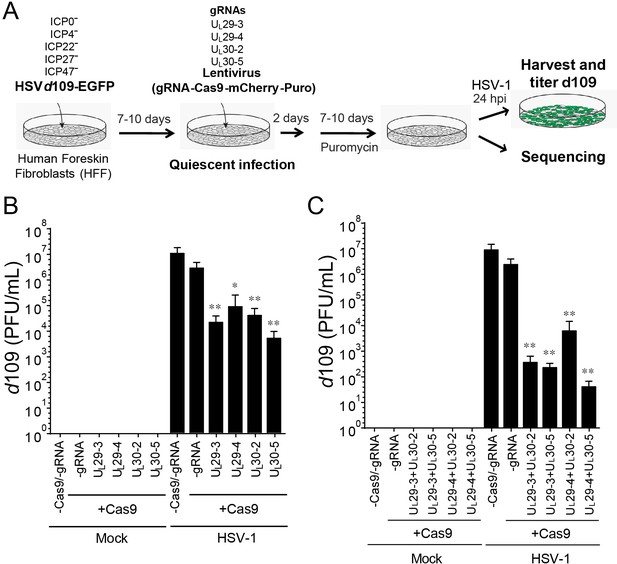
CRISPR/Cas9-induced mutagenesis of quiescent d109 genomes and effect on reactivation.
(A) Experimental scheme of SaCas9/sgRNA-mediated inhibition of reactivation of quiescent d109 genomes in HFFs. HFFs were infected with HSV-1 d109 virus to establish quiescent infection for 7–10 d and transduced with lentivirus expressing SaCas9 and sgRNAs for 7–10 d. (B and C) HFF were infected with d109 to establish quiescent infection for 7–10 d and transduced with lentivirus expressing SaCas9 and sgRNAs for 7–10 d. To reactivate quiescent d109 genomes, HFFs were superinfected with WT HSV-1 at an MOI of 5 and harvested at 24 hpi. GFP-positive viral yields were determined by plaque assays on FO6 and V27 cells. The histogram shows the mean values and standard deviations of biological replicates (B and C: N = 5 and N = 7 respectively). All the sgRNA added conditions showed statistical significance compared to +Cas9 /-gRNA (Ratio paired t test, *p<0.05 and **p<0.01).
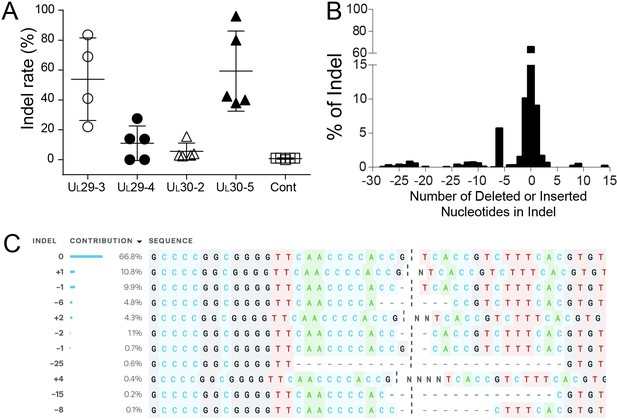
Indel mutations in the HSV-1 genome during the quiescent infection.
(A) Indel mutation frequencies of quiescent d109 genomes are shown at the indicated sgRNA target sites. (B) Histogram representing the frequency (count) of indel lengths induced by SaCas9/UL30-5 in quiescent d109 genomes. (<0) deletions (>0) insertions. (C) Examples of sequences that show mutations induced by SaCas9/UL30-5 in quiescent d109 genomes.
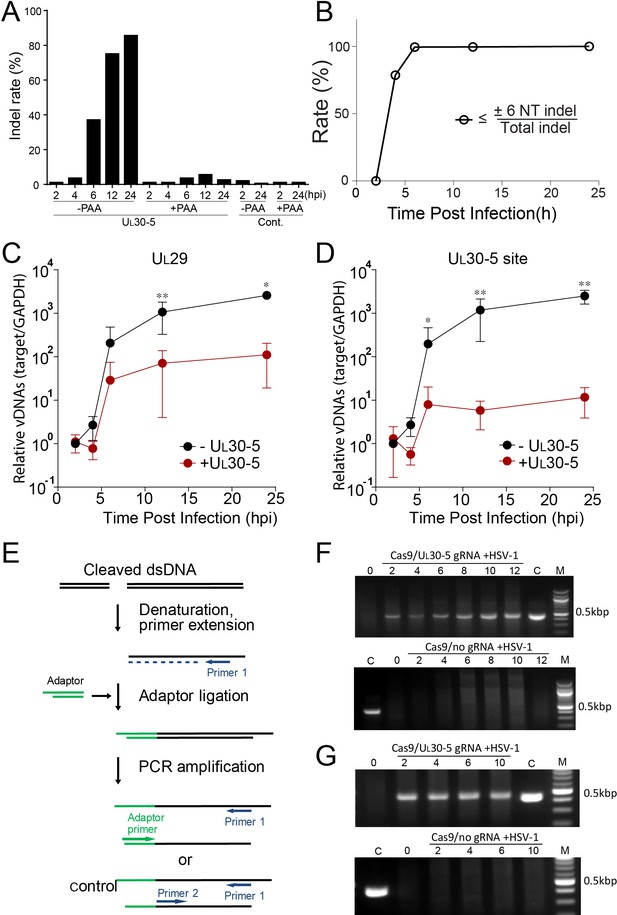
Effect of CRISPR-Cas9 on input and replicating HSV genomes.
(A) Kinetics of indel mutations in the HSV-1 genome during lytic infection. HFFs transduced with lentivirus expressing SaCas9 and sgRNA were infected with HSV-1 at an MOI of 3 in the presence or absence of phosphonoacetic acid (PAA) and harvested at the indicated times post infection. Indel mutation frequencies are shown at the indicated times post infection at the sgRNA target sites by MiSeq. (B) Calculated portion of short indels (≤±6 nucleotide (NT) indel) out of total indel during the lytic replication. (C and D) HFFs transduced with lentivirus expressing SaCas9 and sgRNA were infected with HSV-1 at an MOI of 3 and harvested at the indicated times post infection. The accumulated DNAs were detected by real time qPCR amplification within the UL29 gene (C) or over the UL30-5 sgRNA (D) target site. The histogram shows the mean values and standard deviations of biological replicates (N ≥ 3, Ratio paired t test, *p<0.05 and **p<0.01). Ligation-Mediated -PCR (LM-PCR) of HSV-1 viral DNA during lytic replication. (E) Schematic diagram of LM-PCR (adapted and modified from Brinkman et al., 2018). To make blunt end dsDNA, primer extension was performed using cleaved dsDNA and a primer (primer 1) about 500 bp away from the sgRNA target site. An annealed adaptor was ligated to the blunt end dsDNA and PCR amplification was performed using the pair of either the primer 1/adaptor primer or the primer 1/primer 2 for control. (F and G) HFFs transduced with lentivirus expressing SaCas9 and UL30-5 sgRNA were infected with HSV-1 (MOI of 3) in the absence (F) or presence (G) of PAA and harvested at the indicated times post infection. Total DNA was purified, and a primer extension reaction was performed using a complementing primer downstream of the UL30-5 site to convert all the cleaved DNA into blunt end dsDNA. An adaptor was ligated to the blunt end of dsDNA, and PCR was performed using a primer complementing adaptor and the primer used for the extension reaction. Top panel: +UL30-5 sgRNA, bottom: control. C: control PCR product generated using the primer for the extension reaction and a primer complementing near the UL30-5 target site. The right lane (M): DNA ladder.
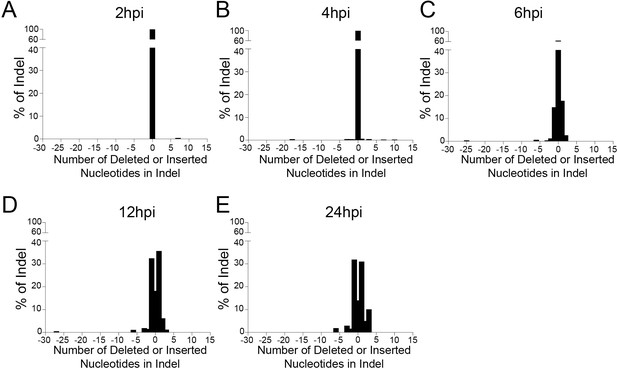
Histogram of indel length of HSV-1 and short indel accumulation during lytic replication.
The lengths of SaCas9/UL30-5 sgRNA induced indel mutations in HSV-1 genome during lytic replication are analyzed using the Inference of CRISPR Editing (ICE) tool (https://ice.synthego.com) at indicated times post infection. (A–E) HFF cells were infected with HSV-1 and harvested at the indicated time post infection.
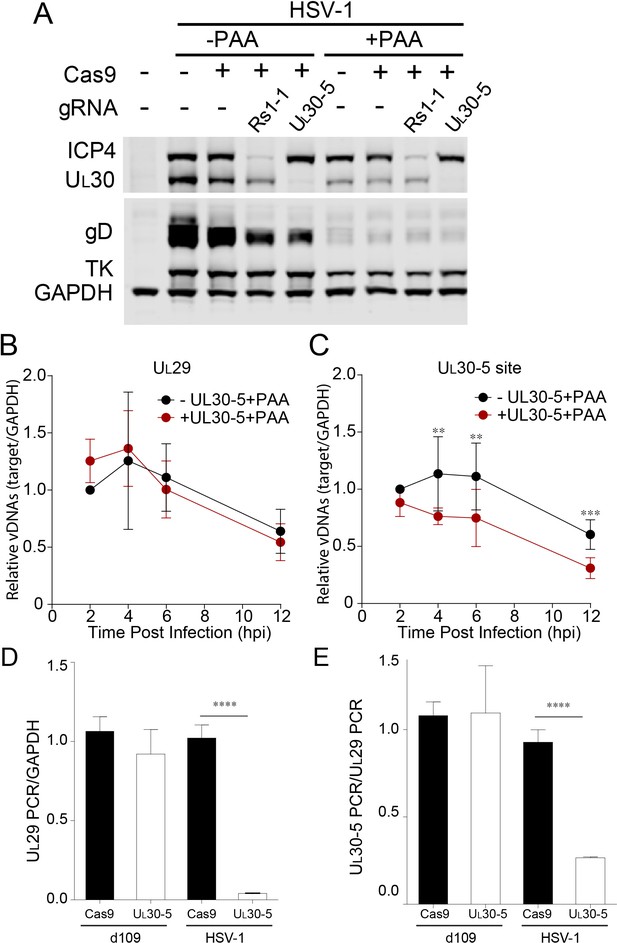
Effect of CRISPR-Cas9 on input HSV genomes.
(A) HFFs transduced with lentivirus expressing Cas9 and sgRNA were infected with HSV-1 at an MOI of 1 in the presence or absence of PAA and harvested at 10 hpi. Proteins were detected by immunoblotting with antibodies specific for the indicated proteins. Immunoblots of GAPDH are shown as a control. (B and C) HFFs transduced with lentivirus expressing SaCas9 and sgRNA were infected with HSV-1 at an MOI of 3 in the presence of PAA and harvested at the indicated times post infection. The accumulated DNAs were detected by real time qPCR amplifying within the UL29 gene (B) or over the UL30-5 sgRNA (C) target site. The histogram shows the mean values and standard deviations from biological replicates (N ≥ 3, Ratio paired t test, **p<0.01 and ***p<0.001). (D and E) PCR amplification across the UL30-5 target site in quiescent d109 genomes and replicating HSV-1 genomes. Quiescently infected HFF cells were transduced with lentivirus expressing Cas9/UL30-5 sgRNA as described in Figure 2A and analyzed at the UL30-5 sgRNA target site by qPCR. The qPCR across the Cas9/UL30-5 sgRNA target site (UL30-5 PCR) was normalized to GAPDH (D) or a remote site of qPCR (E, UL29 PCR) with (UL30-5) or without (Cas9) UL30-5 sgRNA expression. As a control, Cas9 ± UL30-5 sgRNA transduced HFF cells were infected with HSV-1, harvested at 12 hpi and analyzed as described above. Cas9: no sgRNA, UL30-5: Cas9 with UL30-5 sgRNA. The histogram shows the mean values and standard deviations from biological replicates of d109 (N = 6) or HSV-1 (N = 3) infected cells (t-test: ****p<0.0001).
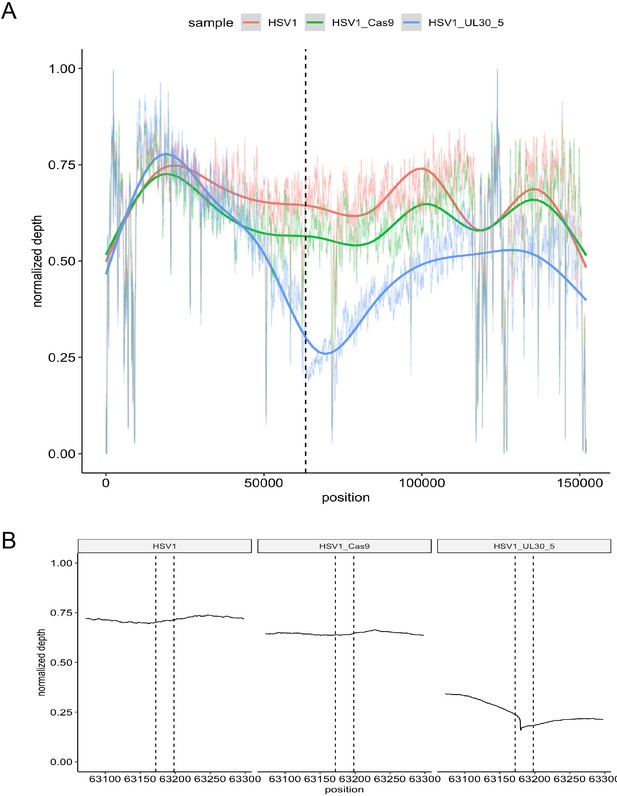
On-target activity of UL30-5 sgRNA within the HSV genome sequence during lytic replication.
Per-base plot of WGS coverage over each specific base in HSV during lytic replication in HFFs for UL30-5 against untreated controls (no sgRNA, no SaCas9/sgRNA). (A) Per-base WGS coverage across the entire HSV genome with fitted LOESS curves obtained by local regression of per-base data (B) Zoomed in view of the WGS coverage in the genomic area around the UL30-5 cleavage site within UL30 for UL30-5 treated against untreated controls. The vertical lines demarcate the UL30-5 target sequence in each sample.
-
Figure 7—source data 1
List of 437 possible UL30-5 off-target sites within the human genome (GRCh38/hg38).
The 437 possible UL30-5 off-target sites are shown together with the off-target sequence and the number of mismatches at each position. Off-target sites were identified through sequence analysis using Cas-OFFinder to identify all predicted off-target sites for SaCas9/UL30-5 with ≤6 mismatches within GRCh38/hg38.
- https://cdn.elifesciences.org/articles/51662/elife-51662-fig7-data1-v2.xlsx
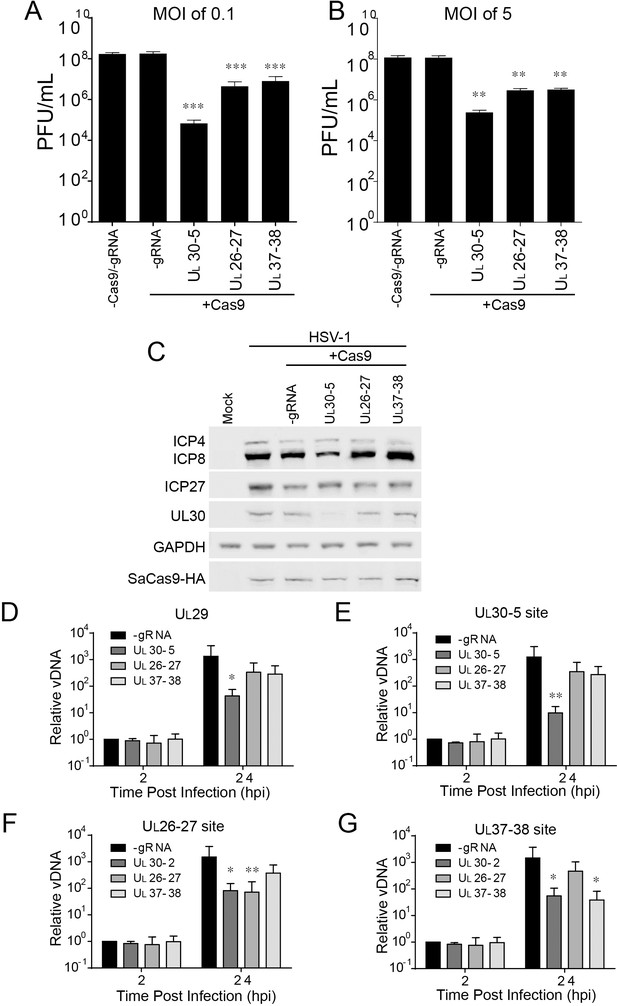
Effect of CRISPR-Cas9 on non-coding and non-essential regions of the HSV-1 genome.
(A and B) HFFs transduced with lentivirus expressing SaCas9 and sgRNAs were infected with HSV-1 at an MOI of 0.1 (A) or 5 (B), harvested at 48 hpi or 24 hpi respectively. Viral yields were determined by plaque assays. The histogram shows the mean values and standard deviations from biological replicates (N = 4). (C) HFFs transduced with lentivirus expressing SaCas9 and sgRNA were infected with HSV-1 at an MOI of 5 and harvested at 10 hpi. Proteins were detected using immunoblotting with antibodies specific for the indicated proteins. GAPDH was shown as a control. (D–G) HFFs transduced with lentivirus expressing SaCas9 and sgRNA were infected with HSV-1 at an MOI of 3 and harvested at the indicated time post infection. The accumulated DNAs were detected by real time PCR amplifying in the UL29 gene (ICP8, (D) or over the UL30-5 (E), UL26-27 (F), and UL37-38 (G) sgRNA target sites. The histogram shows the mean values and standard deviations from biological replicates (N = 4, (A and B) one-way ANOVA with Dunnett’s multiple comparisons test, (D–G) Ratio paired t test, *p<0.05, **p<0.01, and ***p<0.001).
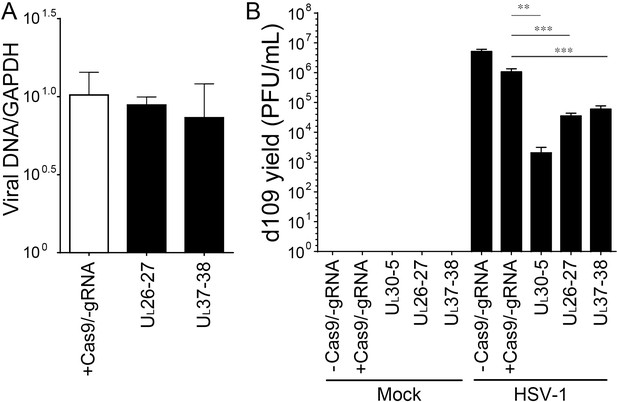
PCR amplification across the UL26-27 or UL37-38 target site in quiescent d109 genomes and reactivation.
(A) Quiescently infected HFF cells were transduced with lentivirus expressing SaCas9/UL26-27 or SaCas9/UL37-38 sgRNA as described in Figure 2A and analyzed at the indicated sgRNA target sites by qPCR. qPCRs across the indicated sgRNA target sites were normalized to GAPDH. The histogram shows the mean values and standard deviations of biological replicates (N = 3). (B) Reactivation assay as described in Figure 3 using indicated sgRNAs (N = 6, Ratio paired t test, **p<0.01 and ***p<0.001).
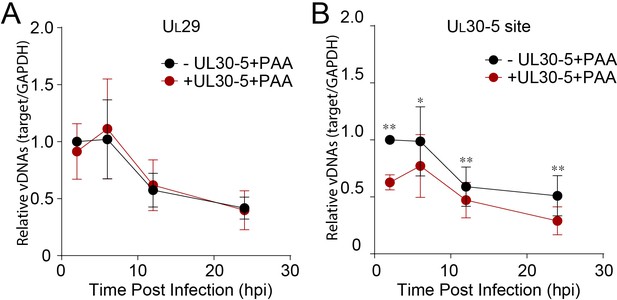
Effect of ICP0 on CRISPR-Cas9-induced DNA repair of input HSV genome.
HFFs transduced with lentivirus expressing SaCas9 and sgRNA were infected with ICP0-null mutant HSV-1 at an MOI of 3 in the presence of PAA and harvested at the indicated time post infection. The accumulated DNAs were detected by real-time qPCR amplifying within the UL29 gene (A) or over the UL30-5 sgRNA (B) target site. The histogram shows the mean values and standard deviations biological replicates (N ≥ 3, Ratio paired t test, *p<0.05 and **p<0.01).
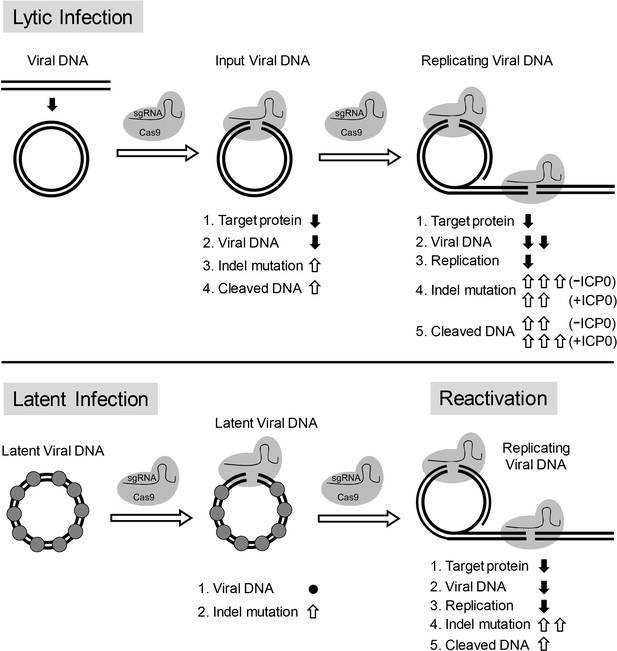
Model for CRISPR/Cas9 mediated inhibition of HSV lytic replication, editing of latent HSV genomes, and inhibition of reactivation of latent HSV.
Lytic infection: Cas9/sgRNA cleaves input viral DNA. In the absence of viral DNA replication, either prior to the onset of viral replication or in the presence of PAA, the expression of Cas9/sgRNA targeting viral gene encoded protein is reduced and the input viral DNAs decrease. Cas9/sgRNA induces low levels of indel mutations at the sgRNA target site of the input viral DNA, and cleaved input viral DNA is accumulated. During viral DNA replication, expression of Cas9/sgRNA targeting the viral gene encoded protein is reduced, non-mutated template and its replicated viral DNAs are targeted by SaCas9/sgRNA, which results in a decrease of viral DNA, an increase in indel mutations and accumulation of cleaved viral DNA. Viral protein ICP0 contributes to Cas9/sgRNA-mediated editing/cleavage by removing histones and preventing DNA repair. Quiescent infection: Cas9/sgRNA induces indel mutations to viral DNA without significant change in latent viral DNA levels. (⬤: nucleosome) Reactivation: Cas9/sgRNA induces more indel mutations in non-mutated viral DNA and accumulation of cleaved viral DNA, which results in decrease of the expression of SaCas9/sgRNA targeted gene encoded protein, viral DNA, and viral replication. (⇧: increase, ⬇: decrease, ●: no change).
Tables
CRISPR/Cas9 target sequences.
| Name | Efficiency of cleavage | SaCas9 sgRNA + PAM (g was added as needed) | Target sequences (GenBank: KT899744) |
|---|---|---|---|
| UL30 | |||
| UL30-1 | + | GCGTCCCGACTGGGGCGAGGT AGGGGT | 62811–62831 |
| UL30-2 | ++ | gAAGTTTTGCCTCAAACAAGGC GGGGGT | 62779–62799 |
| UL30-3 | - | GCGGCGTGGACCACGCCCCGG CGGGGT | 63060–63080 |
| UL30-4 | ++ | gTGCCCCCCCGGAGAAGCGCG CCGGGGT | 62923–62942 |
| UL30-5 | ++ | gACACGTGAAAGACGGTGACG GTGGGGT | 63097–63116 |
| UL30-6 | + | gACCAGCCGAAGGTGACGAAC CCGGGGT | 63595–63614 |
| UL30-7 | ++ | GGCCATCAAGAAGTACGAGGG TGGGGT | 63532–63552 |
| UL30-24* | ++ | gAAACCCCAAAAGCCGCTTGGG TGGGAT | 62589–62609 |
| UL30-25* | + | gCCACCCGAACCCCTAAAGAGG GGGGAT | 62637–62657 |
| UL30-26* | - | gCATGCCGGCCCGGGCGAGCCT GGGGGT | 62542–62562 |
| UL30-27* | ++ | gCCATCCCACCCAAGCGGCTTT TGGGGT | 62581–62601 |
| UL29 | |||
| UL29-1 | ++ | gTCAAGGTCCCCCCCGGGCCCC TGGGAT | 61861–61881 |
| UL29-2 | + | GTGTTTGAGGTCGCCGGGCCG GGGGGT | 61502–61522 |
| UL29-3 | ++ | GCCAGCCAGGGTAAGACCCCG CGGGGT | 61028–61048 |
| UL29-4 | ++ | GCCGCCGTCGCGCCCACCCCG CGGGGT | 61007–61027 |
| UL29-14* | ++ | GAGGGTGGGAGACCGGGGTTG GGGAAT | 62029–62049 |
| UL29-15* | ++ | GTCGGGCGTCCGTCGTCGTGC CGGGAT | 61952–61972 |
| UL29-16* | ++ | gCGGGGGTTGTCTGTGAAGGGT AGGGAT | 62064–62084 |
| UL29-17* | ++ | gATCGGCACCCCGTGGTTACCC GGGGGT | 62084–62104 |
| UL29-18* | ++ | gCAGACAACCCCCGGGTAACCA CGGGGT | 62072–62092 |
| UL29-19* | ++ | GGACCCCGCGTTGCCAGCCGC CGGGGT | 62113–62133 |
| UL29-20* | ++ | GAACCCCGGCGGCTGGCAACG CGGGGT | 62105–62125 |
| UL29- 21 | ++ | GGTTCTCGCACGACGGGGCTC GGGGGT | 61685–61705 |
| UL54 | |||
| UL54-1* | ++ | GCTGTCGGCTGCCGTCGGGGC TGGGGT | 113541–113561 |
| UL54-2 | ++ | gACCTGGAATCGGACAGCAAC GGGGAGT | 113667–113687 |
| UL54-3 | ++ | GCTCCGGTCCGTCCTCTCCGT GGGGGT | 113728–113748 |
| UL54-4 | ++ | GCGTCTGGGTGCTGGGTACGC CGGGGT | 113803–113823 |
| UL54-5 | ++ | GGCGGACGCCGTGGGCGTCGC AGGGGT | 113982–114002 |
| UL54-6 | ++ | gTGGTTCTGGGGGCACGCCGGC GGGGGT | 114055–114075 |
| UL54-7 | ++ | GCAGGCTGGGCTTTGGTCGGT GGGGGT | 113957–113977 |
| UL54-8 | ++ | gCGCCGTGGGCGTCGCAGGGGT CGGGGT | 113988–114008 |
| UL54-9 | + | GTCCGTCCACCCCGCCCCGGGG CGGGGT | 114098–114119 |
| UL54-14* | ++ | gCGCTTCCGCGGGGACCCGGGC GGGGGT | 113234–113254 |
| UL54-15* | ++ | gCGCCCGGGGGGCGGAACTAGG AGGGGT | 113347–113367 |
| UL54-16 | ++ | GGCGGCTCTCCGCCGGCTCGG GGGGGT | 113641–113661 |
| Rs1 | |||
| Rs1-1 | ++ | GCCGGGCGTCGTCGAGGTCGT GGGGGT | 130775–130795, 146992–147012 |
| Rs1-2 | - | gCCGCTCGTCGCGGTCTGGGCT CGGGGT | 130866–130886, 146901–146921 |
| Rs1-3 | - | GGGGGTGGTCGGGGTCGTGGT CGGGGT | 130796–130816, 146971–146991 |
| Rs1-4 | ++ | gATCGTCGTCGGCTAGAAAGGC GGGGGT | 130599–130619, 147168–147188 |
| Rs1-5 | ++ | GGCGCGGCGACAGGCGGTCCG TGGGGT | 130475–130495, 147292–147312 |
| Rs1-6 | ++ | GCGAGGCCGCGGGGTCGGGCGT CGGGAT | 130634–130655, 147132–147153 |
| Rs1-7 | + | GGGTCCGGGGCGGCGAGGCCG CGGGGT | 130622–130642, 147145–147165 |
| Rs1-8 | ++ | gCGCGAGGCGCGGGCCGTCGGG CGGGGT | 130290–130310, 147477–147497 |
| Rs1-9 | ++ | GCGGACGACGAGGACGAGGACC CGGAGT | 130378–130399, 147388–147409 |
| Rs1-15* | ++ | GCCGATGCGGGGCGATCCTCC GGGGAT | 130954–130974, 146813–146833 |
| Rs1-16* | - | gTACGCGGACGAAGCGCGGGAG GGGGAT | 131142–131162, 146625–146645 |
| Rs1-17* | + | gCGCGTCGACGGCGGGGGTCGT CGGGGT | 131061–131081, 146706–146726 |
| Rs1-18* | ++ | GCGCTAGTTCCGCGTCGACGGC GGGGGT | 131070–131091, 146696–146717 |
| UL26-27 | ++ | GAGGAAATCGGCACTGACCAA GGGGGT | 52742–52762 |
| UL37-38 | ++ | GTATAACACCCCGCGAAGACG CGGGGT | 84066–84086 |
-
++:full cleavage (no residual substrate DNA on agarose gel), +: partial cleavage (some residual substrate DNA on agarose gel), -: no cleavage, *: non-coding region
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Staphylococcus aureus) | SaCas9 | Addgene | pX601, Cat. #: #61591 | |
| Peptide, recombinant protein | SpCas9 | NEB | Cat. #: M0386 | |
| Cell line (Homo-sapiens) | HFF (Hs27) | ATCC | Cat# CRL-1634, RRID:CVCL_0335) | |
| Cell line (Homo-sapiens) | U2OS | ATCC | Cat# HTB-96, RRID:CVCL_0042 | |
| Cell line (Chlorocebus sabaeus) | Vero | ATCC | Cat# CCL-81, RRID:CVCL_0059 | |
| Antibody | Anti-ICP8 (Rabbit serum) | Knipe et al., 1987 | 1:5000 | |
| Antibody | Anti-ICP4 (Mouse monoclonal, purified from hybridoma cell line 58S (ATCC HB8183)) | Showalter et al., 1981 | 1:2000 | |
| Antibody | Anti-ICP27 (Mouse monoclonal) | Eastcoast Bio | Cat. #: P1119 | 1:5000 |
| Antibody | Anti-GAPDH ([6C5], Mouse monoclonal) | Abcam | Cat. #: ab8245 | 1:10000 |
| Recombinant DNA reagent | Addgene | lentiCRISPRv2, Cat #: 52961 | Cloning vector | |
| Recombinant DNA reagent | pX601-AAV-CMV-SaCas9-T2A-mCherry | This paper | Template of SaCas9-T2A-mCherry for lentiSaCas9-mCherry-Puro | |
| Recombinant DNA reagent | lentiSaCas9-mCherry-Puro | This paper | Cloned SaCas9 gene into lentiCRISPRv2 | |
| Software, algorithm | ICE analysis toolbox | https://ice.synthego.com | ||
| Software, algorithm | bcbio-nextgen | https://github.com/bcbio/bcbio-nextgen | v1.15 | |
| Software, algorithm | MuTect2 | https://www.ncbi.nlm.nih.gov/pubmed?term=20644199 | v2 | |
| Software, algorithm | bwa-mem | https://arxiv.org/abs/1303.3997 | v0.7.17 | |
| Software, algorithm | Cas-OFFinder | https://www.ncbi.nlm.nih.gov/pubmed/24463181 | v2.4 |





