Quantitative analysis of 1300-nm three-photon calcium imaging in the mouse brain
Figures
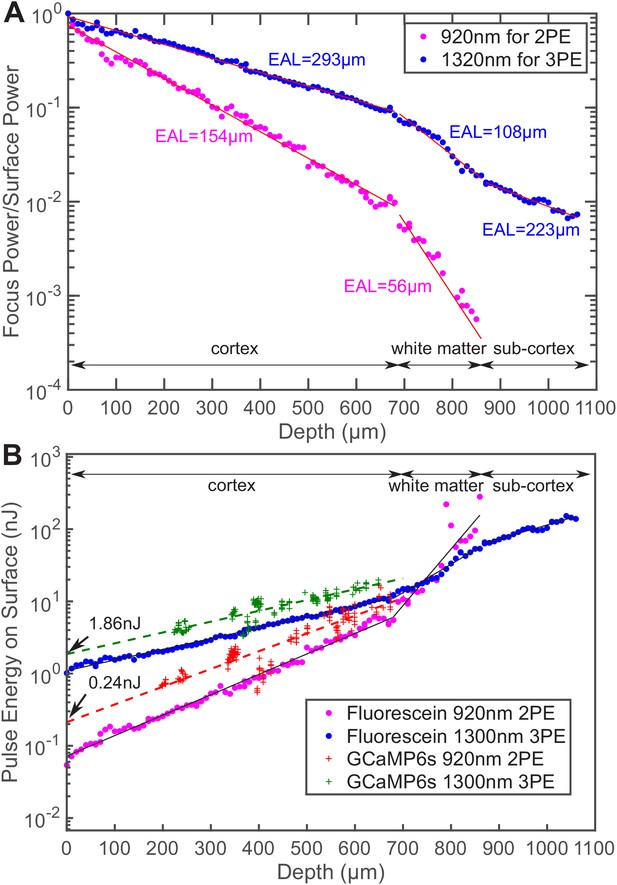
Comparison of the power attenuation of 1320 nm and 920 nm excitation light and their respective 3-photon and 2-photon excitation efficiency in the mouse brain.
(A) Power attenuation of 920 nm and 1320 nm excitation light in the mouse brain. The mouse brain vasculature was uniformly labeled with fluorescein dextran and imaged simultaneously by 920 nm 2PM and 1320 nm 3PM at precisely the same location. The fraction of excitation power reaching the focus from the brain surface (Focus Power/Surface Power) was calculated based on the attenuation of 2PE and 3PE signal with the imaging depth (Materials and methods). (B) The pulse energy required at the brain surface to generate the same 2PE and 3PE signal strength (0.1 signal photon detected per laser pulse) at different imaging depths, measured in fluorescein-labeled blood vessels (n = 1 shown) and GCaMP6s-labeled neurons (CamKII-tTA/tetO-GCaMP6s; n = 5 mice). The signal strength is scaled to 60 fs pulse duration for both 2PE and 3PE.
-
Figure 1—source data 1
A typical power attenuation with depth curve of 1320 nm and 920 nm excitation light in the mouse brain, plotted in Figure 1A.
- https://cdn.elifesciences.org/articles/53205/elife-53205-fig1-data1-v2.csv
-
Figure 1—source data 2
The excitation pulse energy at the brain surface required to generate 0.1 signal photon per pulse at different depths in the mouse brain, measured in fluorescein-labeled vasculature and plotted in Figure 1B.
- https://cdn.elifesciences.org/articles/53205/elife-53205-fig1-data2-v2.csv
-
Figure 1—source data 3
The excitation pulse energy at the brain surface required to generate 0.1 signal photon per pulse at different depths in the mouse brain, measured in the neurons of transgenic animals (CamKII-tTA/tetO-GCaMP6s) and plotted in Figure 1B.
- https://cdn.elifesciences.org/articles/53205/elife-53205-fig1-data3-v2.csv
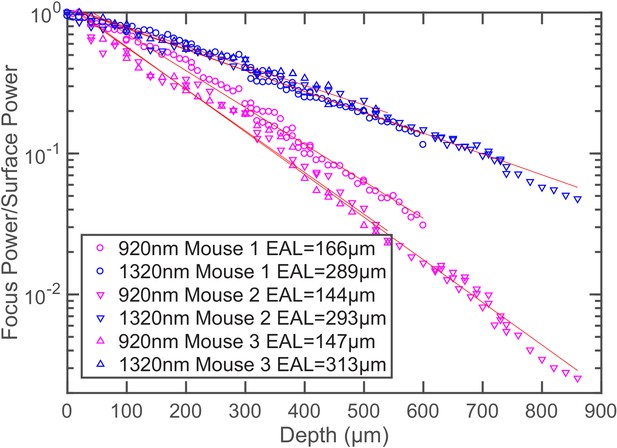
Additional power attenuation curves of 920 nm and 1320 nm excitation light in the mouse neocortex (n = 3, >12 weeks old mice, all males).
-
Figure 1—figure supplement 1—source data 1
Additional power attenuation with depth curves of 1320 nm and 920 nm excitation light in the mouse neocortex, plotted in Figure 1—figure supplement 1.
- https://cdn.elifesciences.org/articles/53205/elife-53205-fig1-figsupp1-data1-v2.xlsx
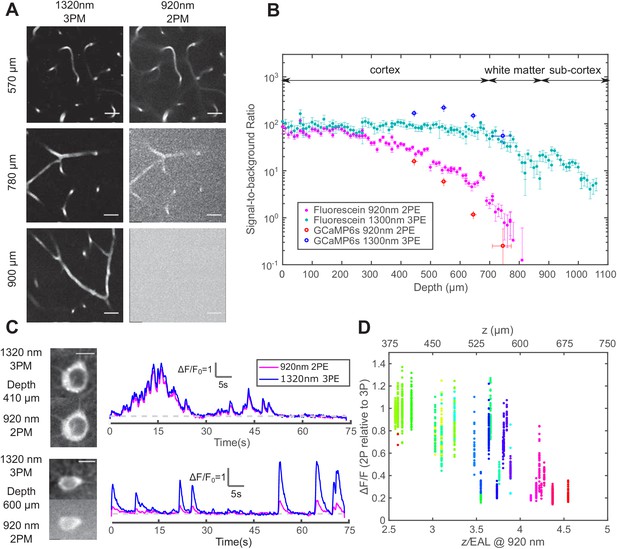
Comparison of signal-to-background ratio for 1320 nm 3PM and 920 nm 2PM in the non-sparsely labeled mouse brain and its effect on calcium imaging sensitivity.
(A) Comparison of 3PM and 2PM images of fluorescein-labeled blood vessels at different depths. Scale bar 30 μm. (B) SBR measured simultaneously by 1320 nm 3PE and 920 nm 2PE on fluorescein-labeled blood vessels and GCaMP6s-labeled neurons. Each set of 3PE and 2PE comparison was performed in the same mouse. All vertical error bars denote the standard deviation of SBR caused by feature brightness variation, and horizontal error bars represent the depth range of the measurement. (C) Images and corresponding calcium traces of a cortical L4 neuron (410 μm) and a L5 neuron (600 μm), simultaneously recorded by 920 nm 2PM and 1320 nm 3PM. Both traces were low-pass filtered with a hamming window of a time constant 0.37 s. Scale bar, 10 μm. (D) The ratio of ΔF/F calculated on simultaneously recorded 3PM and 2PM calcium traces (CamKII-tTA/tetO-GCaMP6s; n = 3 mice). Each point was measured on a single calcium transient. The depth was normalized by the 2PM attenuation length at 920 nm of each animal to collapse the data better onto a single trendline. Data points from different neurons at the same depth are colored differently.
-
Figure 2—source data 1
The change of signal-to-background ratio with depth of 1320 nm 3PM and 920 nm 2PM in the mouse brain, measured in fluorescein-labeled vasculature and plotted in Figure 2B.
- https://cdn.elifesciences.org/articles/53205/elife-53205-fig2-data1-v2.csv
-
Figure 2—source data 2
The change of signal-to-background ratio with depth of 1320 nm 3PM and 920 nm 2PM in the mouse brain, measured in the neurons of transgenic animals (CamKII-tTA/tetO-GCaMP6s) and plotted in Figure 2B.
- https://cdn.elifesciences.org/articles/53205/elife-53205-fig2-data2-v2.csv
-
Figure 2—source data 3
Calcium traces recorded by 920nm 2PM on GCaMP6s-labeled neurons at different depths in transgenic animals (CamKII-tTA/tetO-GCaMP6s), based on which Figure 2—source data 5 is derived.
- https://cdn.elifesciences.org/articles/53205/elife-53205-fig2-data3-v2.csv
-
Figure 2—source data 4
Calcium traces recorded by 1320nm 3PM simultaneously on the same GCaMP6s-labeled neurons as in Figure 2—source data 3 in transgenic animals (CamKII-tTA/tetO-GCaMP6s), based on which Figure 2—source data 5 is derived.
- https://cdn.elifesciences.org/articles/53205/elife-53205-fig2-data4-v2.csv
-
Figure 2—source data 5
The ratio of calcium transient ΔF/F between simultaneously recorded by 1320 nm 3PM and 920 nm 2PM calcium traces, on the same GCaMP6s-labeled neurons as described in Figure 2—source datas 3 and 4.
This data is plotted in Figure 2D.
- https://cdn.elifesciences.org/articles/53205/elife-53205-fig2-data5-v2.csv
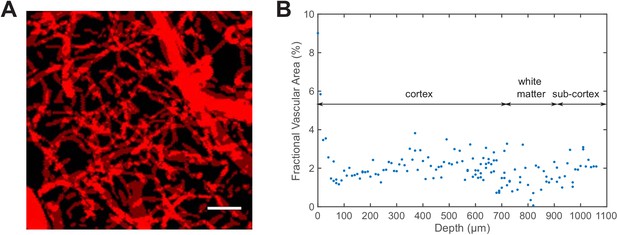
Measurement of the staining density in uniformly labeled vasculature.
(A) Z-projection of the identified blood vessels by summing all the XY-images in a vertical image stack (0–400 μm deep, with 10 μm step size). Scale bar, 30 μm. (B) The fractional vascular area in each XY-image at various depths.
-
Figure 2—figure supplement 1—source data 1
The area fraction of vasculature measured in the mouse brain, plotted in Figure 2—figure supplement 1.
- https://cdn.elifesciences.org/articles/53205/elife-53205-fig2-figsupp1-data1-v2.csv
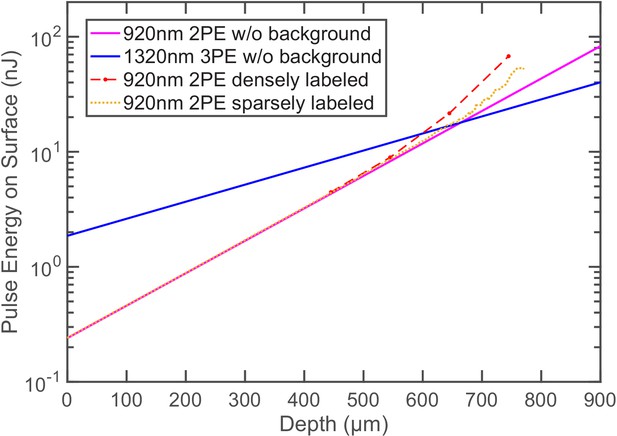
The pulse energy required at the brain surface to generate the same per pulse sampling the neuron for 2PE and 3PE of GCaMP6s at different imaging depths.
In the absence of the background fluorescence, is entirely determined by signal photon counts: when a neuron is sampled by 1000 excitation pulses in 1 s, it generates , which gives . Therefore, Group ‘920nm 2PE w/o background’ and ‘1320nm 3PE w/o background’ exactly reproduce the pulse energy curves for the same signal of 0.1 photon/excitation pulse (Figure 1B). Group ‘920nm 2PE densely labeled’ and ‘920nm 2PE sparsely labeled’ are calculated based on the SBR measured in GCaMP6s-labeled neurons and fluorescein-labeled blood vessels, respectively (Figure 2B). 1320-nm 3PE is assumed to be background-free in the mouse cortex.
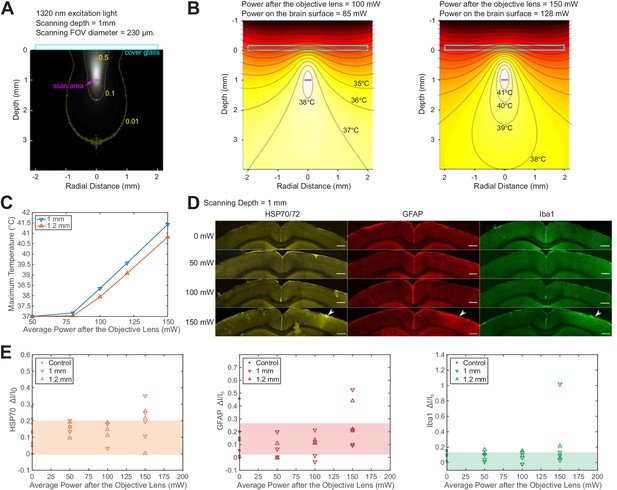
Brain heating and thermal damage induced by continuous scanning by 1320 nm 3PM.
(A) Monte Carlo simulation of light intensity of 1320 nm excitation light. The excitation light is focused at 1 mm below the brain surface in the cortex by an objective of 1.05 NA at ~75% filling of its back aperture and scanned telecentrically in a 230 μm diameter FOV (the horizontal magenta line segment). Three iso-contour lines (from inside to outside) correspond to 0.01, 0.1, and 0.5 of the maximum intensity. (B) Temperature maps under 1320 nm illumination calculated from the power absorption per unit volume in (A) after 60 s of continuous scanning with 100 and 150 mW average power after the objective lens. Temperature is color-coded with isotherms plotted at 1°C increment with the highest four temperature levels labeled. The average imaging power is listed at the top of each plot. (C) The maximum temperature versus imaging power for 1 mm and 1.2 mm focal depth, with other imaging parameters the same as in (A) and (B). The maximum temperature is calculated as the average in the hottest volume around the scanned area (~107 μm3). (D) Immunolabeled brain slices of mouse brains after 1320 nm 3PM scanning with 0, 50 mW, 100 mW, and 150 mW average power (one mouse is shown for each power). The brain was scanned for 20 min continuously at 1 mm below the surface, with 230 μm x 230 μm FOV and 2 Hz frame rate. The location of the damage is indicated by the white arrowheads. Scale bar, 0.5 mm. (E) Quantification of heat-induced damage by the fractional change of immunolabeling intensity relative to the region in the contralateral hemisphere. Shaded areas denote 95% confidence interval of the control group mean (n = 4 mice for control; n = 6 mice for 50 mW; n = 5 mice for 100 mW; n = 7 mice for 150 mW).
-
Figure 3—source data 1
Quantification of the staining intensity of immunolabeld mouse brain slices post mortem after the exporsure to continuous 1320 nm 3PM scanning, plotted in Figure 3E.
- https://cdn.elifesciences.org/articles/53205/elife-53205-fig3-data1-v2.zip
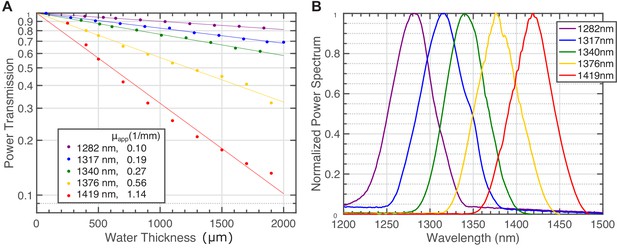
Light attenuation by immersion water for various excitation wavelengths.
(A) Power transmission versus immersion water thickness under the objective (Olympus XLPLN25XWMP2, 25X, NA=1.05, ~ 75% filling of the back aperture) for 5 spectra centered from 1280 nm to 1420 nm using broadband pulses from the NOPA. Each transmission curve is fitted with a single exponential function to calculate the apparent attenuation coefficient (). The inset lists these coefficients. (B) The spectra of the pulses for the transmission measurement in (A).
-
Figure 3—figure supplement 1—source data 1
Power transmission through immersion water of different thicknesses under the objective lens, measured with different excitation spectra and plotted in Figure 3—figure supplement 1.
- https://cdn.elifesciences.org/articles/53205/elife-53205-fig3-figsupp1-data1-v2.xlsx
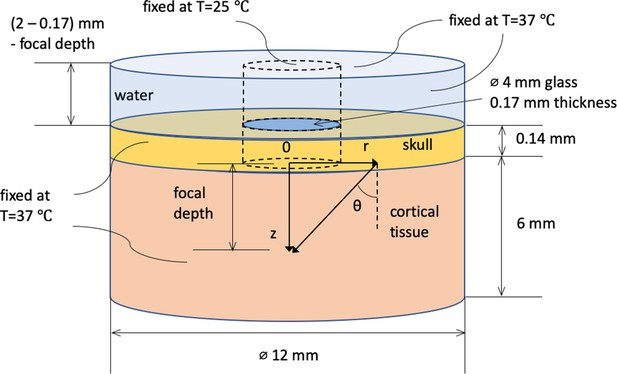
Spatial dimension, coordinate system, and boundary conditions for Monte Carlo and heat conduction simulation.
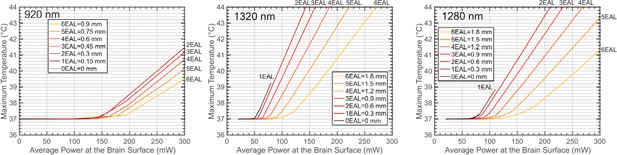
The maximum brain temperature as a function of the average power at the brain surface, shown for different imaging depths at 920 nm, 1320 nm, and 1280 nm.
The imaging depth grows in the increment of the nominal EAL of the cortex, which is 150 μm for 920 nm and 300 μm for both 1320 nm and 1280 nm. The linear scanning FOV diameter is 300 μm, and other parameters used for the simulation are listed in Table 1 and Table 3. For 1320 nm, the power immediately after the objective lens can be calculated by multiplying the power at the brain surface with a factor to correct for immersion water absorption: 1.42, 1.33, 1.27, 1.20, 1.13, 1.07, and 1.01 (from 0 to 6 EALs). For 1280 nm, the correction factors are: 1.20, 1.17, 1.13, 1.10, 1.07, 1.03, and 1.00 (from 0 to 6 EALs).
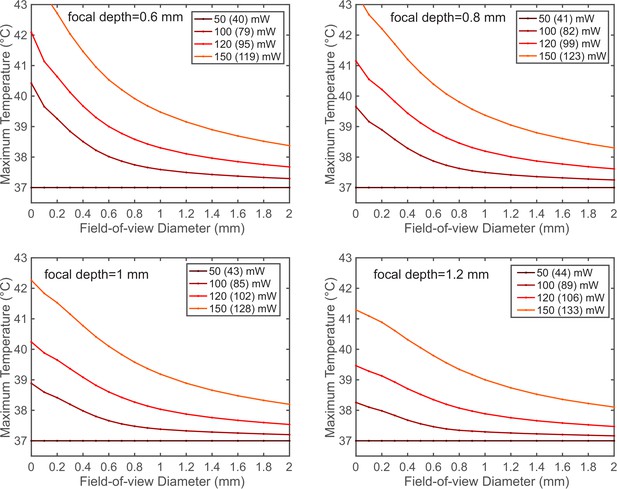
The maximum brain temperature decreases with the scanning field-of-view.
The simulation was performed for 1320 nm excitation wavelength at 1- and 1.2 mm imaging depth with various average powers after the objective lens. The NA and the back-aperture beam size remain the same as in Figure 3A. Each legend key indicates the power immediately after the objective lens, followed by the power at the brain surface after immersion water absorption (in the bracket).
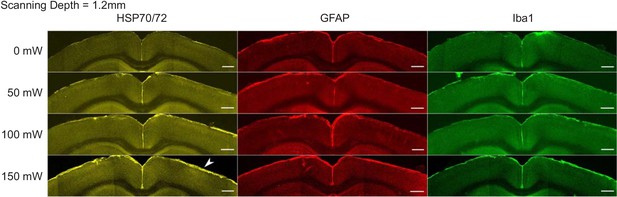
Immunostaining reveals brain tissue damage by 1320 nm 3PM with 150 mW imaging power after the objective lens at 1.2 mm imaging depth.
The scanning was performed with 230 μm FOV for 20 min at various average powers in the mouse brain. The window and scanning area are on the right side of each brain slice, and the location of heating is indicated with white arrowheads. Scale bar, 500 μm.
Tables
Optical parameters of gray matter.
| Variable | Parameter | Wavelength | Value | Units |
|---|---|---|---|---|
| absorption coefficient | 920 | 0.039 | 1/mm | |
| 1280 | 0.078 | 1/mm | ||
| 1320 | 0.12 | 1/mm | ||
| scattering coefficient | 920 | 6.7 | 1/mm | |
| 1280 | 3.2 | 1/mm | ||
| 1320 | 3.2 | 1/mm | ||
| anisotropy coefficient | All | 0.9 | dimensionless | |
| refractive index | All | 1.36 | dimensionless |
Percentage of Excitation Photons for Various Final Destinations.
| 920 nm | 1320 nm | 1280 nm | |
|---|---|---|---|
| Contributing to heating | 20 | 63 | 49 |
| Back scattered to window | 25 | 9 | 10 |
| Back scattered to skull | 20 | 9 | 11 |
| Escaped* | 35 | 19 | 30 |
-
*Photons escape the simulation volume by traveling too far (>6 mm) from the center of the cranial window or too deep (>6 mm) from the tissue surface.
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mus musculus) | B6.Cg-Tg(CamK2a-tTA)1Mmay/J | The Jackson Laborartory Stock: 007004 | MGI:2179066 | |
| Strain, strain background (Mus musculus) | B6;DBA-Tg(tetO-GCaMP6s)2Niell/J | The Jackson Laborartory Stock: 024742 | MGI:5553332 | |
| Antibody | anti-HSP70/72 (mouse monoclonal) | Enzo Life Sciences, Cat# SPA-810PED | RRID:AB-2264369 | IHC(1:400) |
| Antibody | anti-GFAP (mouse monoclonal) | Sigma-Aldrich, Cat# G3893 | RRID:AB_477010 | IHC(1:760) |
| Antibody | anti-Iba1 (mouse monoclonal) | Sigma-Aldrich, Cat# SAB2702364 | RRID:AB_2820253 | IHC(1:1000) |
| Antibody | Goat anti-mouse (polyclonal) | Thermo Fisher Scientific, Cat# A-11003 | RRID:AB_2534071 | IHC(1:500) |
| Software, algorithm | Source code 1: matlab code for simulating the brain temperature distribution under continuous long-wavelength illumination by 3PM using Monte Carlo method and heat equation. | Mathworks, Matlab 2016b | RRID:SCR_001622 |
Thermal and mechanical properties of gray matter (Stujenske et al., 2015).
| Variable | Parameter | Value | Units |
|---|---|---|---|
| Density | 1.04 × 10−3 | g/mm3 | |
| Brain specific heat | 3.65 × 103 | mJ/g°C | |
| Thermal conductivity | 0.527 | mW/mm °C | |
| Blood density | 1.06 × 10−3 | g/mm3 | |
| Blood specific heat | 3.6 × 103 | mJ/g°C | |
| Blood perfusion rate | 8.5 × 10−3 | /s | |
| Metabolic heat | 9.5 × 10−3 | mW/mm3 | |
| Arterial temperature | 36.7 | °C |
Two- and three-photon excitation parameters.
| Objective lens focal length (mm) | 1/e2beam diameter at the back aperture (mm) | Effective NA | Pulse duration (fs) | ||
|---|---|---|---|---|---|
| 920 nm 2PE | 7.2 | 11 | 0.75 | 60 | 0.66 |
| 1320 nm 3PE | 7.2 | 11 | 0.75 | 60 | 0.51 |
Additional files
-
Source code 1
Matlab code for simulating the brain temperature distribution under continuous long-wavelength illumination by 3PM using Monte Carlo method and heat equation, which was used to produce Figure 3B and C, Figure 3—figure supplements 3 and 4.
- https://cdn.elifesciences.org/articles/53205/elife-53205-code1-v2.zip
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/53205/elife-53205-transrepform-v2.docx





