ParB spreading on DNA requires cytidine triphosphate in vitro
Figures
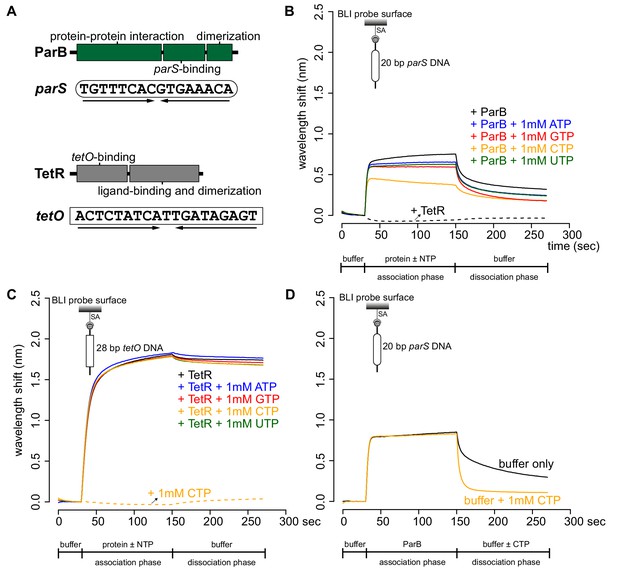
CTP reduces the nucleation of Caulobacter ParB at parS.
(A) The domain architecture of ParB (dark green) and TetR (grey), and their respective DNA-binding sites parS and tetO. Convergent arrows below DNA-binding sites indicate that parS and tetO are palindromic. (B) Bio-layer interferometric (BLI) analysis of the interaction between a premix of 1 μM ParB-His6 dimer ± 1 mM NTP and a 20 bp DNA duplex containing parS. Biotinylated DNA fragments were immobilized onto the surface of a Streptavidin (SA)-coated probe (See Materials and methods). The BLI probe was dipped into a buffer only solution (0–30 s), then to a premix of protein ± NTP (30–150 s: association phase), and finally returned to a buffer only solution (150–270 s: dissociation phase). Sensorgrams were recorded over time. BLI analysis of the interaction between 1 μM TetR-His6 and a 20 bp parS probe was also recorded (a negative control). (C) BLI analysis of the interaction between a premix of 1 μM TetR-His6 ± 1 mM NTP and a 28 bp DNA duplex containing tetO. BLI analysis of the interaction between 1 mM CTP and a 28 bp tetO probe was also recorded (a negative control). (D) BLI analysis of the interaction between 1 μM Caulobacter ParB-His6 (without CTP) and a 20 bp parS DNA. For the dissociation phase, the probe was returned to a buffer only or buffer supplemented with 1 mM CTP. All buffers used for experiments in this figure contained Mg2+. Each BLI experiment was triplicated and a representative sensorgram was presented.
-
Figure 1—source data 1
Data used to generate Figure 1.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig1-data1-v2.zip
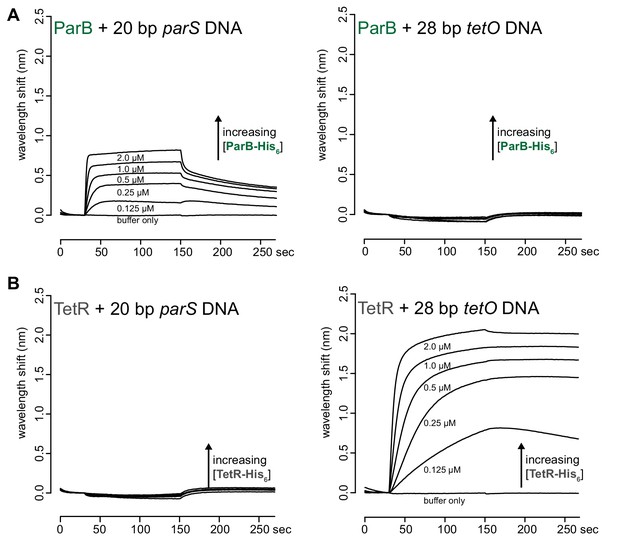
ParB and TetR bind specifically to their cognate-binding sites parS and tetO, respectively.
(A) BLI analysis of the interaction between purified Caulobacter ParB-His6 (0.125 to 2 µM) and a 20 bp parS DNA probe or a 28 bp tetO DNA probe. (B) BLI analysis of the interaction between purified TetR-His6 (0.125 to 2 µM) and a 20 bp parS DNA probe or a 28 bp tetO DNA probe.
-
Figure 1—figure supplement 1—source data 1
Data used to generate Figure 1—figure supplement 1.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig1-figsupp1-data1-v2.zip
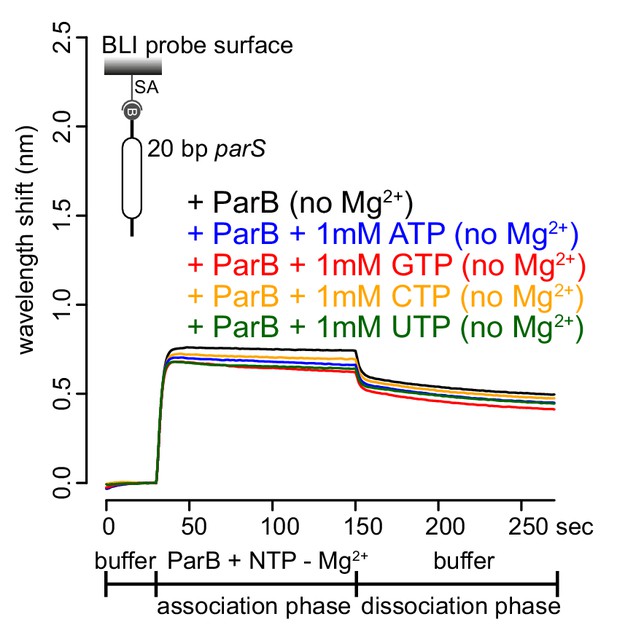
BLI analysis of the interaction between purified Caulobacter ParB and NTP in buffers lacking Mg2+.
Caulobacter ParB-His6 (1 µM dimer)±1 mM NTP, and a 20 bp DNA duplex containing parS were used. Mg2+ was omitted from all buffers in this experiment. A schematic of the DNA substrate is shown above the sensorgram.
-
Figure 1—figure supplement 2—source data 1
Data used to generate Figure 1—figure supplement 2.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig1-figsupp2-data1-v2.zip

BLI analysis of the interaction between purified Caulobacter ParB and cytidine mono-, di-, or triphosphate.
(A) BLI analysis of the interaction between purified Caulobacter ParB and an increasing concentration of CTP-Mg2+. ParB-His6 (1 µM dimer) + 0–10 mM CTP, and a 20 bp DNA duplex containing parS were used for this experiment. (B) BLI analysis of the interaction between a premix of 1 μM Caulobacter ParB-His6 ± 1 mM cytidine mono-, di-, or triphosphate, and a 20 bp parS DNA. A schematic of the DNA substrate is shown above the sensorgram. Mg2+ was included in all buffers for experiments in this figure.
-
Figure 1—figure supplement 3—source data 1
Data used to generate Figure 1—figure supplement 3.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig1-figsupp3-data1-v2.zip
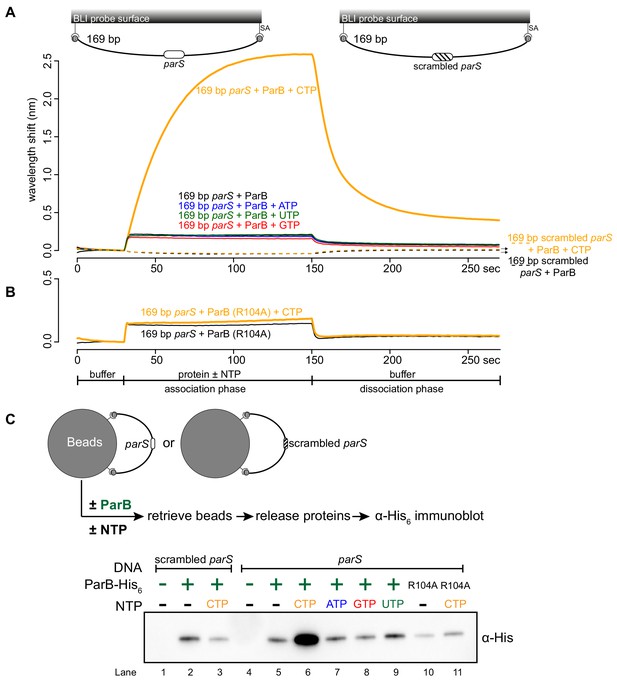
CTP facilitates ParB association with a closed DNA substrate beyond nucleation.
(A) BLI analysis of the interaction between a premix of 1 μM Caulobacter ParB-His6 ± 1 mM NTP and a 169 bp dual biotin-labeled DNA containing a parS or a scrambled parS site. Interactions between a dual biotinylated DNA and streptavidin (SA)-coated probe created a DNA substrate where both ends were blocked (a closed DNA substrate) (see the schematic of the BLI probes above the sensorgram). (B) Interactions between a nucleation-competent but spreading-defective ParB (R104) variant with a 169 bp parS DNA fragment in the presence or absence of CTP were also recorded. Each BLI experiment was triplicated and a representative sensorgram was presented. (C) A schematic of the pull-down assay and immunoblot analysis of pulled-down Caulobacter ParB-His6. The length of bound DNA is ~2.8 kb. Beads were incubated with ParB protein for five minutes before being pulled down magnetically. All buffers used for experiments in this figure contained Mg2+.
-
Figure 2—source data 1
Data used to generate Figure 2.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig2-data1-v2.zip
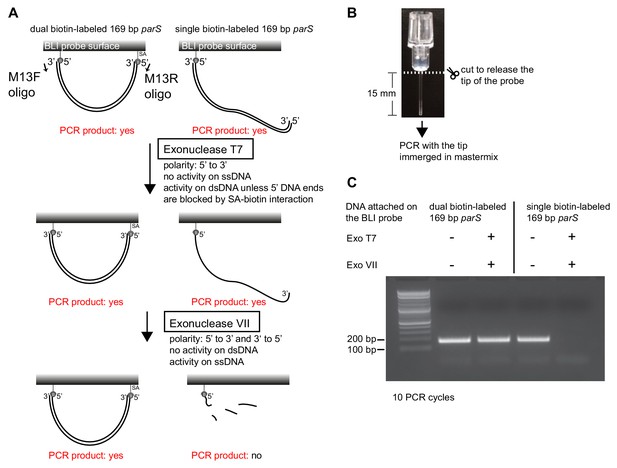
Dual biotin-labeled DNA fragments form a closed substrate on the surface of the BLI probe.
(A) A schematic of a double digestion assay using Exonuclease T7 + Exonuclease VII and PCR. PCR was performed using M13F, M13R oligos, and DNA attached to the BLI surface as a template. (B) The BLI probe was severed from the plastic adaptor and immerged into a PCR master mix. (C) Dual biotin-labeled DNA fragments on the BLI surface were resistant to Exo T7 + Exo VII digestion while single biotin-labeled DNA fragments on the BLI surface were not.
-
Figure 2—figure supplement 1—source data 1
Data used to generate Figure 2—figure supplement 1.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig2-figsupp1-data1-v2.zip
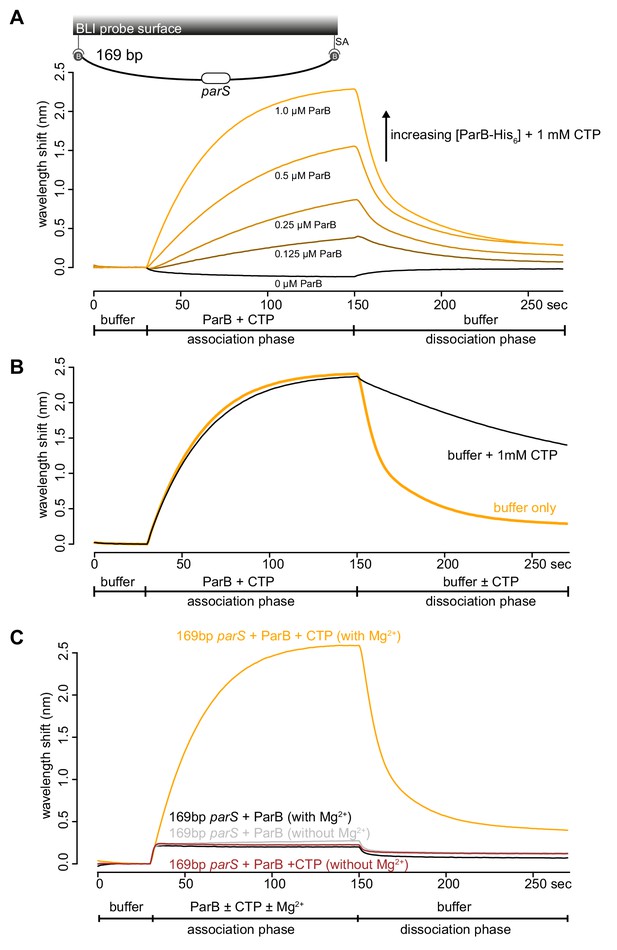
CTP-Mg2+ enhances ParB accumulation on a closed DNA substrate.
(A) BLI analysis of the interaction between an increasing concentration of Caulobacter ParB-His6 (0.125 to 1.0 µM) + 1 mM CTP and a 169 bp dual biotin-labeled parS DNA. A schematic of the DNA substrate is shown above the sensorgram. (B) BLI analysis of the interaction between 1 μM Caulobacter ParB-His6 (with 1 mM CTP) and a 169 bp dual biotin-labeled parS DNA. For the dissociation phase, the probe was returned to a buffer only or buffer supplemented with 1 mM CTP solution. All buffers used for experiments in panel A-B contained Mg2+. (C) BLI analysis of the interaction between a premix of 1 μM Caulobacter ParB-His6 ±1 mM CTP and a 169 bp dual biotin-labeled DNA containing a parS site in buffer with or without MgCl2.
-
Figure 2—figure supplement 2—source data 1
Data used to generate Figure 2—figure supplement 2.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig2-figsupp2-data1-v2.zip

CTP facilitates the association of eight ParB orthologs with DNA.
BLI analysis of the interaction between a premix of 1 μM His6-MBP-tagged ParB proteins from a set of diverse bacterial species ± NTPs and a 169 bp dual biotin-labeled parS DNA. NTPs at 1 mM (final concentration) was used, except for BLI experiments with Zymomonas mobilis and Xanthomonas campestris ParBs where 0.01 mM CTP was used instead. Mg2+ was included in all buffers for this experiment. A schematic of the DNA substrate is shown above the sensorgrams.
-
Figure 2—figure supplement 3—source data 1
Data used to generate Figure 2—figure supplement 3.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig2-figsupp3-data1-v2.zip
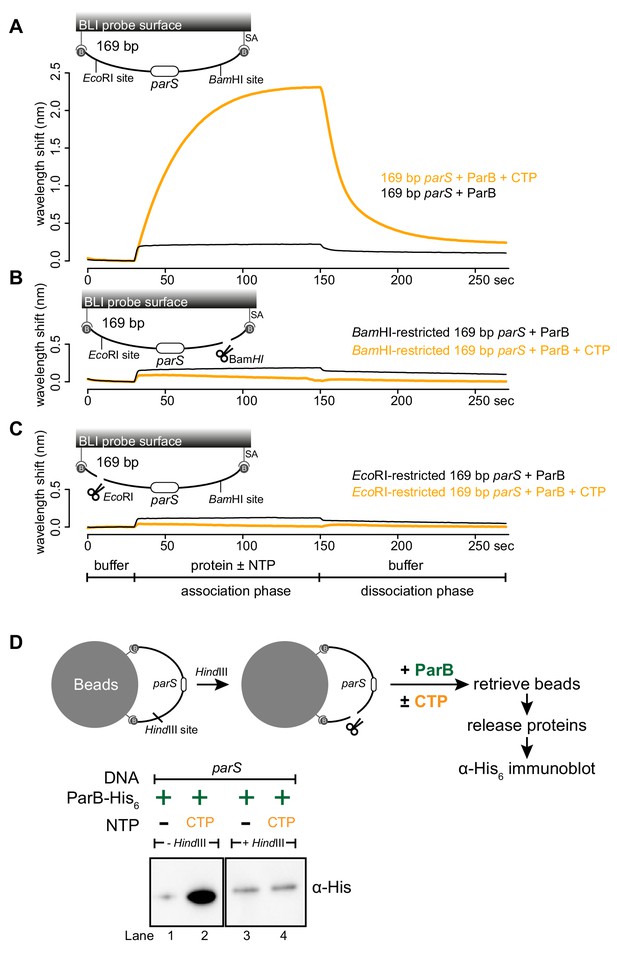
A closed DNA substrate is required for an increased association of ParB with DNA.
(A) BLI analysis of the interaction between a premix of 1 μM Caulobacter ParB-His6 ± 1 mM CTP and a 169 bp dual biotin-labeled parS DNA. (B) Same as panel A but immobilized DNA fragments have been restricted with BamHI before BLI analysis. (C) Same as panel A but immobilized DNA fragments have been restricted with EcoRI before BLI analysis. Schematic of DNA fragments with the relative positions of parS and restriction enzyme recognition sites are shown above the sensorgram. Each BLI experiment was triplicated and a representative sensorgram was presented. (D) A schematic of the pull-down assay and immunoblot analysis of pulled-down Caulobacter ParB-His6. For lanes 3 and 4, DNA-bound beads were incubated with HindIII to linearize bound DNA. Samples in lanes 1–4 were loaded on the same gel, the immunoblot was spliced together for presentation purposes. All buffers used for experiments in this figure contained Mg2+.
-
Figure 3—source data 1
Data used to generate Figure 3.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig3-data1-v2.zip
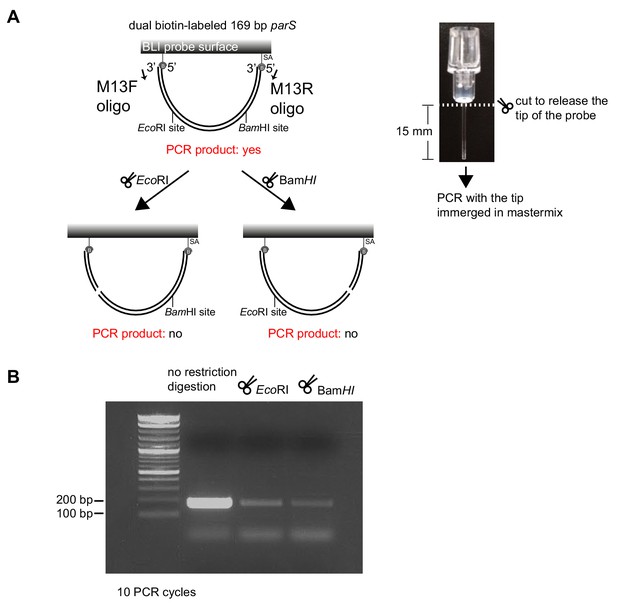
Restriction enzymes linearize dual biotin-labeled DNA fragments on the surface of the BLI probe.
(A) (Left panel) A schematic of a digestion assay using BamHI/EcoRI and PCR. PCR was performed using M13F, M13R oligos, and DNA attached to the BLI surface as a template. (Right panel) The BLI probe was severed from the plastic adaptor and immerged into a PCR master mix. (B) Detection of intact 169 bp DNA fragments that were bound on the surface of the BLI probe before and after treatment with restriction enzymes.
-
Figure 3—figure supplement 1—source data 1
Data used to generate Figure 3—figure supplement 1.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig3-figsupp1-data1-v2.zip
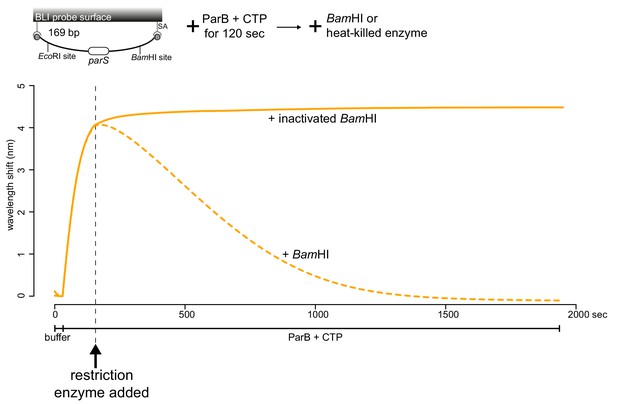
Linearization of a closed DNA substrate by BamHI liberates pre-bound ParB from DNA.
BamHI was added after 1 μM Caulobacter ParB-His6 + 1 mM CTP and a 169 bp dual biotin-labeled parS DNA were preincubated together for 120 s. As a negative control, heat-inactivated BamHI was added instead. Mg2+ was included in all buffers for this experiment. A schematic of the DNA substrate is shown above the sensorgram.
-
Figure 3—figure supplement 2—source data 1
Data used to generate Figure 3—figure supplement 2.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig3-figsupp2-data1-v2.zip
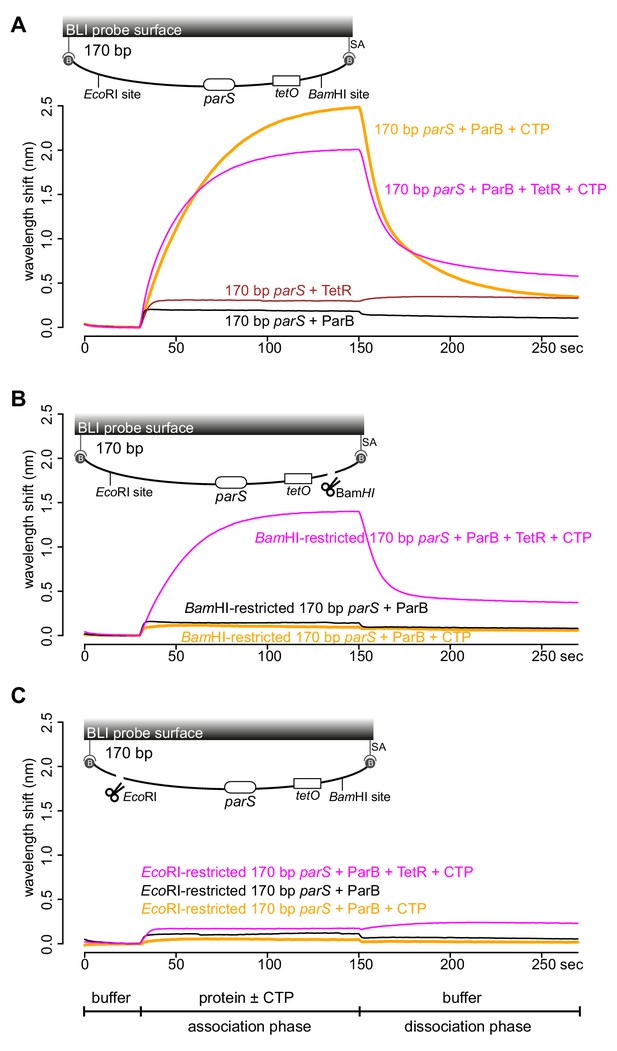
TetR-tetO binding restores ParB association with an open DNA substrate.
(A) BLI analysis of the interaction between a premix of 1 μM Caulobacter ParB-His6 ± 1 mM CTP ± 1 μM TetR-His6 and a 170 bp dual biotin-labeled DNA containing a parS site. (B) Same as panel A but immobilized DNA fragments have been restricted with BamHI before BLI analysis. (C) Same as panel A but immobilized DNA fragments have been restricted with EcoRI before BLI analysis. Schematic of DNA fragments together with the relative positions of parS, tetO, and restriction enzyme recognition sites are shown above the sensorgram. Each BLI experiment was triplicated and a representative sensorgram was presented. All buffers used for experiments in this figure contained Mg2+.
-
Figure 4—source data 1
Data used to generate Figure 4.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig4-data1-v2.zip
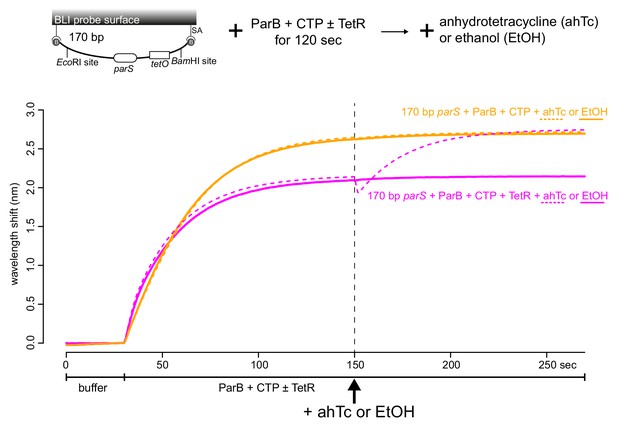
Anhydrotetracycline removes DNA-bound TetR roadblock and allows ParB to accumulate further on DNA.
Anhydrotetracycline (3 μM) or ethanol (0.01%) was added after 1 μM Caulobacter ParB-His6 + 1 mM CTP ± 1 μM TetR and a 170 bp dual biotin-labeled parS DNA were preincubated together for 120 s. Mg2+ was included in all buffers for this experiment. A schematic of the DNA substrate is shown above the sensorgram.
-
Figure 4—figure supplement 1—source data 1
Data used to generate Figure 4—figure supplement 1.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig4-figsupp1-data1-v2.zip
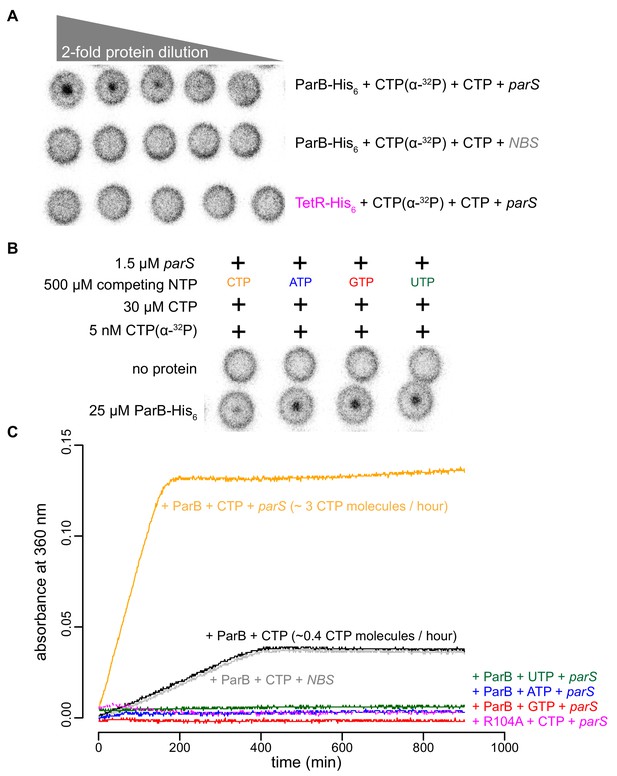
parS DNA increases the CTP binding and hydrolysis rate by Caulobacter ParB.
(A–B) CTP binding as monitored by DRaCALA assay using radiolabeled CTP α-P32. The bulls-eye staining indicates CTP binding due to a more rapid immobilization of protein-ligand complexes compared to free ligands alone. The starting concentration of proteins used in panel A was 25 μM. The same concentration of radioactive CTP, unlabeled CTP, and DNA was used in experiments shown in panels A and B. (C) A continuous monitoring of inorganic phosphate (Pi) released by recording absorbance at 360 nm overtime at 25°C. The rates of CTP hydrolysis were inferred from a Pi standard. The NTP hydrolysis of Caulobacter ParB was also monitored in the presence of ATP, GTP, or UTP, with a 22 bp parS DNA duplex or a non-cognate 22 bp NBS DNA duplex (a DNA-binding site of Noc protein Wu and Errington, 2004). The nucleation-competent but spreading-defective ParB (R104A) mutant did not hydrolyze CTP in the presence of parS DNA. All buffers used for experiments in this figure contained Mg2+.
-
Figure 5—source data 1
Data used to generate Figure 5.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig5-data1-v2.zip
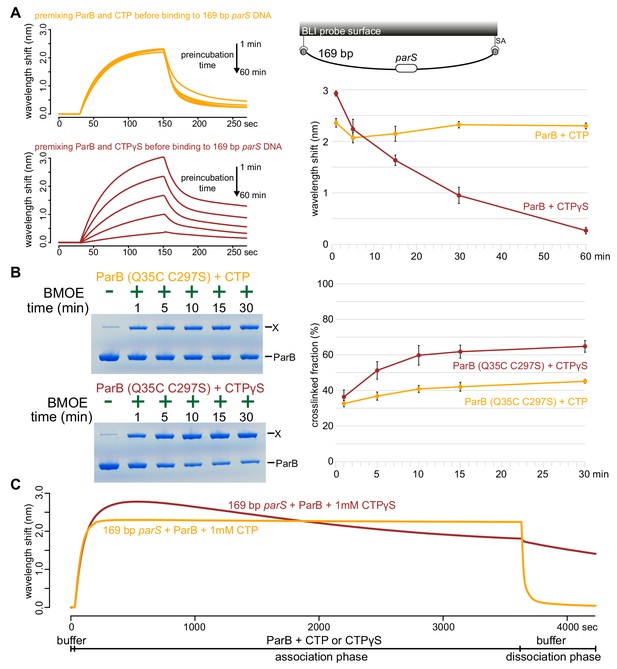
ParB accumulation on DNA is unstable in the presence of a non-hydrolyzable analog CTPγS.
(A) (Left panel) BLI analysis of the interaction between a premix of 1 μM Caulobacter ParB-His6 ± 1 mM CTP or CTPγS and a 169 bp dual biotin-labeled parS DNA. Purified ParB was preincubated with CTP or CTPγS for 1, 5, 15, 30, or 60 min before BLI analysis. (Right panel) Quantification of ParB-DNA interaction at the end of each association phase (150th sec). Error bars represent SD from three replicates. (B) (Left panel) Time course of Caulobacter ParB (Q35C C297S) cross-linking with CTP or CTPγS in the absence of parS DNA. Purified ParB-His6 (Q35C C297S) (10 µM) were preincubated with 1 mM CTP or CTPγS for 1, 5, 10, 15, or 30 min before BMOE was added. X indicates a cross-linked form of ParB. (Right panel) Quantification of the cross-linked (X) fractions. Error bars represent SD from three replicates. (C) BLI analysis of the interaction between a premix of 1 μM Caulobacter ParB-His6 ± 1 mM CTP or CTPγS and a 169 bp dual biotin-labeled parS DNA. CTP or CTPγS was added to purified ParB, and the mixture was immediately used for BLI experiments. All buffers used for experiments in this figure contained Mg2+.
-
Figure 6—source data 1
Data used to generate Figure 6.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig6-data1-v2.zip
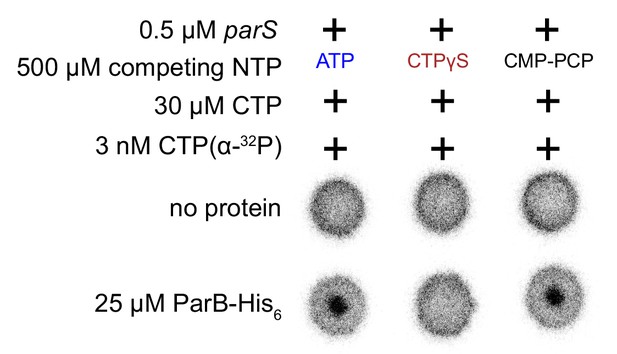
Caulobacter ParB binds CTP and a non-hydrolyzable analog CTPγS but not CMP-PCP.
CTPγS (but not CMP-PCP) outcompeted radioactive CTP α-P32 for binding to ParB-parS complex, indicating that Caulobacter ParB can bind to CTPγS. Binding to radioactive CTP α-P32 was monitored by DRaCALA assay. The bulls-eye staining indicates CTP binding due to a more rapid immobilization of protein-ligand complexes compared to free ligands alone.
-
Figure 6—figure supplement 1—source data 1
Data used to generate Figure 6—figure supplement 1.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig6-figsupp1-data1-v2.zip
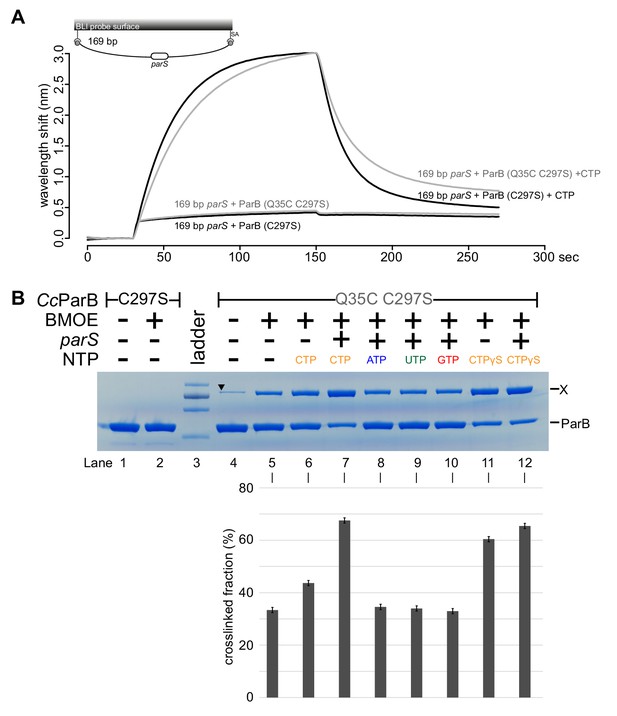
BLI analysis and gel analysis of cross-linking products of purified Caulobacter ParB (Q35C C297S).
(A) BLI analysis of the interaction between a premix of 1 μM His-tagged ParB (C97S) or (Q35C C297S) ± 1 mM CTP and a 169 bp dual biotin-labeled parS DNA. In the presence of CTP, both variants of ParB were able to spread, indicating that the engineered mutations Q35C and C297S did not impair the function of ParB in vitro. (B) Purified Caulobacter ParB-His6 (Q35C C297S) ± NTPs ± parS DNA were incubated for 15 min at room temperature before cross-linking with BMOE. A cysteine-less ParB (C297S) was used as a negative control. X indicates a cross-linked form of ParB. A solid black triangle on lane four indicates a contaminated protein from the purification of ParB-His6 (Q35C C297S). A lower panel shows the quantification of the cross-linked (X) fractions. Error bars represent SD from three replicates.
-
Figure 6—figure supplement 2—source data 1
Data used to generate Figure 1—figure supplement 2.
- https://cdn.elifesciences.org/articles/53515/elife-53515-fig6-figsupp2-data1-v2.zip
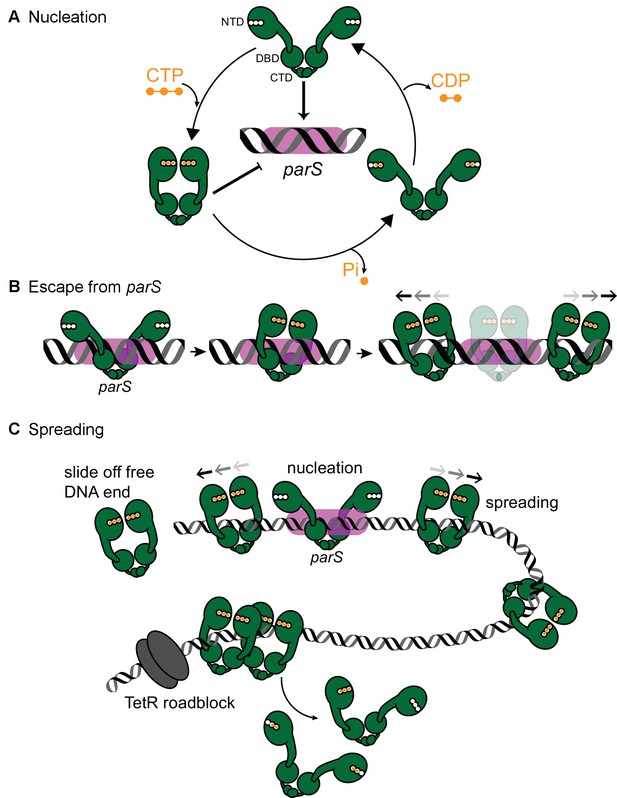
A model for Caulobacter ParB nucleation and spreading.
(A) Caulobacter ParB nucleation at parS. CTP (orange) reduces Caulobacter ParB (dark green) nucleation at parS (magenta box), presumably by inducing conformational changes that are incompatible with a site-specific parS binding (Soh et al., 2019). Only apo- or CDP-bound ParB can nucleate on parS. ParB hydrolyzes CTP at a faster rate in the presence of parS. The domain architecture of ParB is also shown: NTD: N-terminal domain, DBD: DNA-binding domain, and CTD: C-terminal domain. (B) Caulobacter ParB escapes from the nucleation site parS. Apo-ParB at parS binds CTP and slides laterally away from the nucleation site parS while still associating with DNA. (C) Caulobacter ParB sliding and spreading on DNA. CTP-bound ParBs diffuse from the nucleation site parS and can run off the free DNA end unless they are blocked by DNA-bound roadblocks such as transcriptional regulators for example TetR. CTP hydrolysis is not required for ParB to escape from the nucleation parS site but might contribute to ParB recycling. It is not yet known whether both CTP molecules on a ParB dimer are concertedly hydrolyzed/dissociated for ParB to escape from the chromosome or a heterodimer state of ParB with a single CTP bound also exists in vivo.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Escherichia coli) | Rosetta (DE3) | Merck | Cat# 70954 | Electro-competent cells |
| Recombinant DNA reagent | See Supplementary file 1 | This paper | See Supplementary file 1 | |
| Sequence-based reagent | See Supplementary file 1 | This paper | See Supplementary file 1 | |
| Antibody | Anti-6xHis tag antibody (HRP) (Rabbit polyclonal) | Abcam | Cat# ab1187 | Western blot (1:5,000) |
| Commercial assay or kit | EnzChek Phosphate Assay Kit | ThermoFisher | Cat# E6646 | |
| Commercial assay or kit | Gibson Assembly Master Mix | NEB | Cat# E2611S | |
| Commercial assay or kit | Gateway BP Clonase II enzyme mix | ThermoFisher | Cat# 11789020 | |
| Commercial assay or kit | Dip-and-Read Streptavidin (SA) biosensors | Molecular Devices | Cat# 18–5019 | |
| Commercial assay or kit | HisTrap High Performance column | GE Healthcare | Cat# GE17524801 | |
| Commercial assay or kit | HiTrap Heparin High Performance column | GE Healthcare | Cat# GE17040601 | |
| Commercial assay or kit | HiLoad 16/600 Superdex 75 pg column | GE Healthcare | Cat# GE28989333 | |
| Commercial assay or kit | Dynabeads MyOne Streptavidin C1 | ThermoFisher | Cat# 65001 | |
| Commercial assay or kit | Amersham Protran supported Western blotting membranes, nitrocellulose | GE Healthcare | Cat# GE10600016 | pore size 0.45 μm, for DRaCALA assay |
| Peptide, recombinant protein | BamHI-HF | NEB | Cat# R3136S | 20,000 units/mL |
| Peptide, recombinant protein | BamHI | NEB | Cat# R0136S | 20,000 units/mL |
| Peptide, recombinant protein | EcoRI-HF | NEB | Cat# R3101S | 20,000 units/mL |
| peptide, recombinant protein | HindIII-HF | NEB | Cat# R3104S | 20,000 units/mL |
| Peptide, recombinant protein | Exonuclease VII | NEB | Cat# M0379S | 10,000 units/mL |
| Peptide, recombinant protein | Exonuclease T7 | NEB | Cat# M0263S | 10,000 units/mL |
| Chemical compound, drug | Benzonase nuclease | Sigma-Aldrich | Cat# E1014 | |
| Chemical compound, drug | NTP | ThermoFisher | Cat# R0481 | 100 mM solution |
| Chemical compound, drug | CTPγS | Jena Bioscience | A gift from S. Gruber and custom synthesis (purity ≥ 96%) | |
| Chemical compound, drug | CMP-PCP | Jena Bioscience | Cat# NU-254 | |
| Chemical compound, drug | P32-α-CTP | Perkin Elmer | Cat# BLU008H250UC | 3000 Ci/mmol, 10 mCi/ml, 250 µCi |
| Chemical compound, drug | Anhydrotetracycline hydrochloride (ahTc) | Abcam | Cat# ab145350 | Dissolved in ethanol |
| Chemical compound, drug | Bismaleimidoethane (BMOE) | ThermoFisher | Cat# 22323 | Dissolved in DMSO |
| Software, algorithm | BLItz Pro | Molecular Devices | Cat# 50–0156 | Version 1.2 https://www.moleculardevices.com/ |
| Software, algorithm | R | R Foundation for Statistical Computing | RRID: SCR_001905 | Version 3.2.4 |
| Software, algorithm | Image Studio Lite | LI-COR Biosciences | RRID: SCR_013715 | Version 5.2 |
| Software, algorithm | Excel 2016 | Microsoft | RRID: SCR_016137 |
Additional files
-
Supplementary file 1
Plasmids, DNA, and protein sequences used in this study.
- https://cdn.elifesciences.org/articles/53515/elife-53515-supp1-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/53515/elife-53515-transrepform-v2.docx





