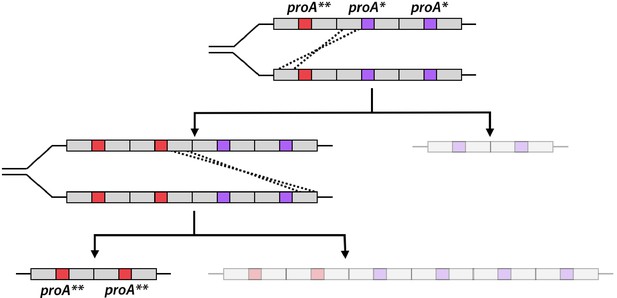
Mutations that improve efficiency of a weak-link enzyme are rare compared to adaptive mutations elsewhere in the genome
Figures
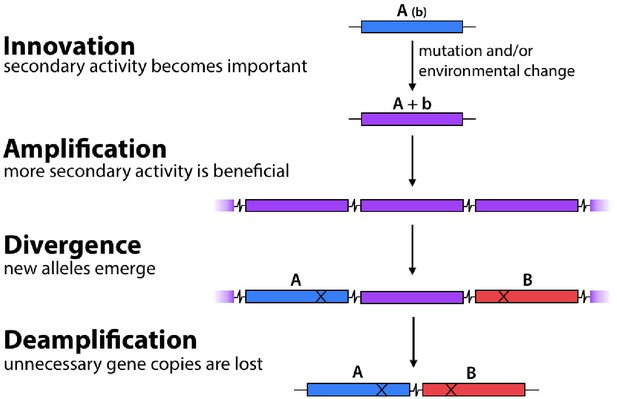
The Innovation-Amplification-Divergence (IAD) model of gene evolution.
A promiscuous activity B of an enzyme may become physiologically relevant due to a mutation or environmental change. Gene amplification increases the abundance of the weak-link enzyme. Mutations can improve the efficiency of the newly important activity B. Once sufficient B activity is achieved, selection is relaxed and extra gene copies are lost, leaving behind two paralogs.
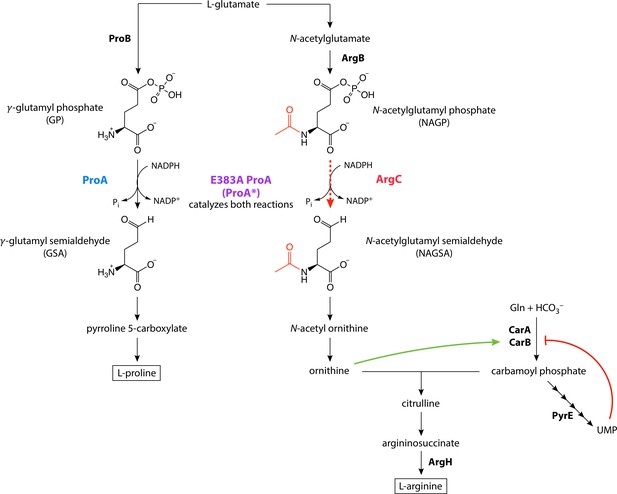
E383A ProA (ProA*) replaces ArgC in the arginine synthesis pathway in ∆argC proA* E. coli, but is the bottleneck in the pathway due to its poor catalytic activity.
The reaction normally catalyzed by ArgC and replaced by ProA* in the parental strain is indicated by the red dotted line. The green and red lines indicate allosteric activation and inhibition, respectively.
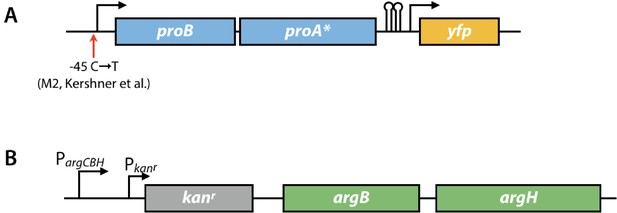
Genome modifications in the parental AM187 strain.
(A) Modifications to the proBA operon. proA*; A1148→C proA (changes Glu383 to Ala); M2, a −45 C→T promoter mutation that increases expression of the operon more than 4-fold (Kershner et al., 2016). (B) Modifications to the argCBH operon. argC was replaced with a gene encoding kanamycin resistance in the Keio strain created by Baba et al. (2006).
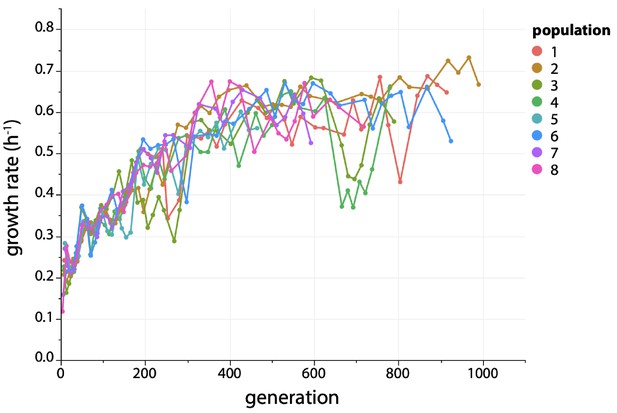
Growth rate increases ~ 3 fold during evolution of ∆argC M2-proA* E. coli in M9 minimal medium containing 0.2% glucose (w/v), 0.4 mM proline and 20 µg/mL kanamycin.
M2 is the C to T mutation at −45 in the promoter for the proBA operon (Kershner et al., 2016).
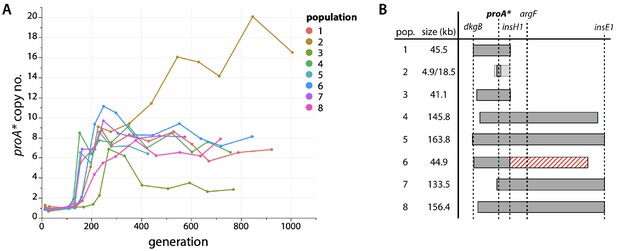
proA* is amplified during evolution.
(A) proA* copy number in each evolved population as measured by qPCR. (B) Regions of amplification in each evolved population based on population genome sequencing. Population 2 had two overlapping regions of amplification, both of which included proA* (shown as differently shaded bars). Population 6 had a 95.1 kb deletion (shown as a red striped bar) immediately downstream of the amplified region.
-
Figure 4—source data 1
Mutations found during the evolution experiment.
Sheet 1: mutations that were present at frequencies > 30% or that appeared in different populations. Sheet 2: genomic positions of the amplified regions surrounding proA*. Sheet 3: other amplified or deleted genomic regions in the evolved populations. Sheet 4: times during the adaptation when the turbidostat failed and was restarted either from an aliquot of a previously stored culture or from the full population preserved at the point of the failure (Materials and methods).
- https://cdn.elifesciences.org/articles/53535/elife-53535-fig4-data1-v2.xlsx
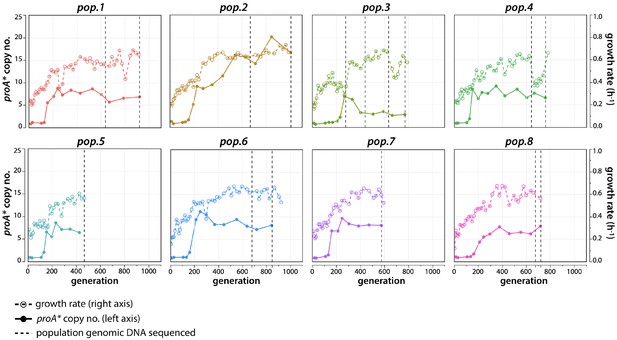
Growth rate (right axis, dotted lines) and proA* copy number (left axis, solid lines) for each evolved population.
Vertical dotted lines indicate when population genomic DNA was sequenced.
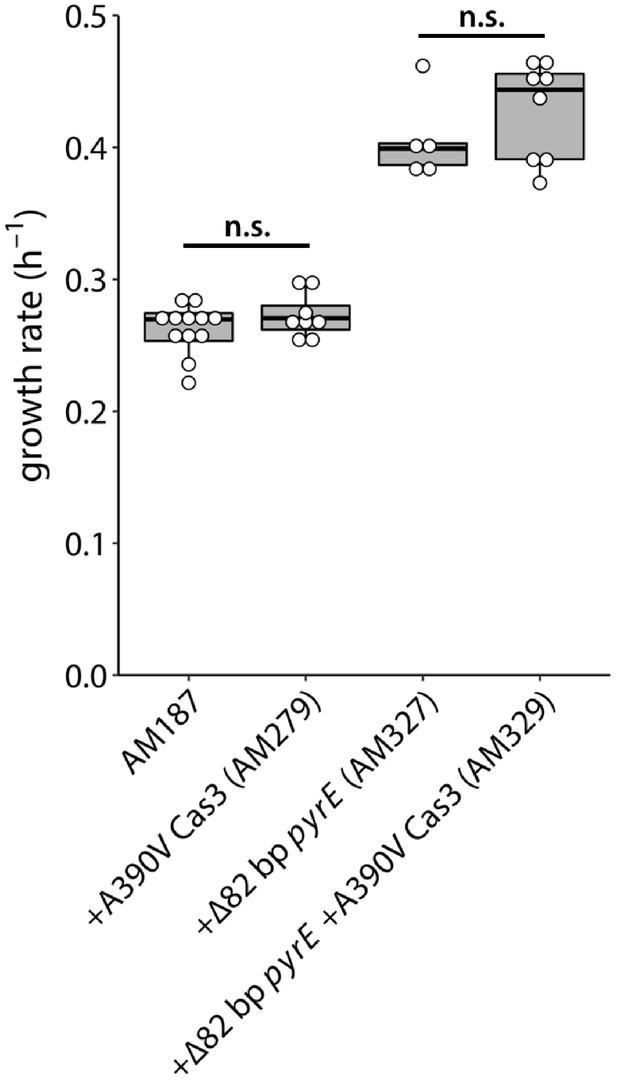
Effects of the A390V Cas3 and rph-pyrE mutations on growth rate of the parental strain AM187.
AM187, N = 12; AM279, N = 8; AM327, N = 5; AM329, N = 8. (All p values are > 0.2 based on a two-tailed, unequal variance Student’s t-test. n.s., not significant).
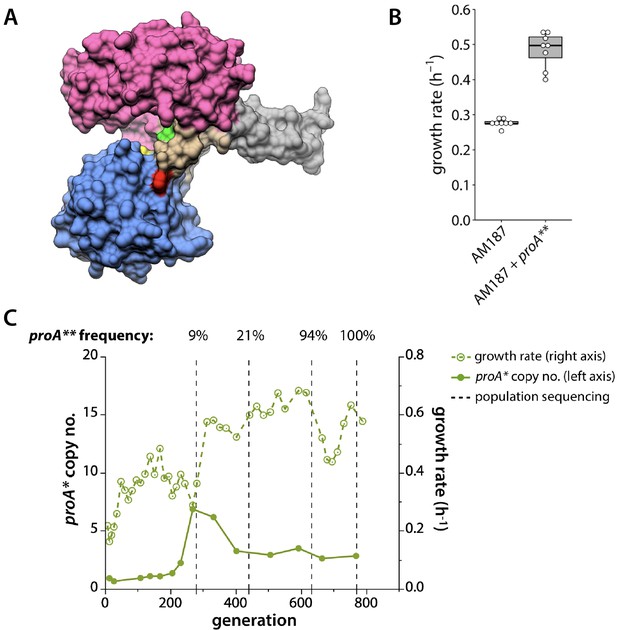
proA* acquired a beneficial mutation in population 3.
(A) Crystal structure of Thermotoga maritima ProA (PDB 1O20) (Page et al., 2003). Yellow, catalytic cysteine; green, equivalent of E. coli ProA Glu383; red, equivalent of E. coli ProA Phe372; magenta, NADPH-binding domain; blue, catalytic domain; beige, hinge region; gray, oligomerization domain. (B) Change in growth rate when the mutation changing Phe372 to Leu (proA**) is introduced into the genome of AM187. P value = 4.5 × 10−6 by a two-tailed, unequal variance Student’s t-test, N = 8. (C) proA* copy number (left axis, solid lines) and growth rate (right axis, dotted lines) for population 3. Vertical dotted lines indicate when population genomic DNA was sequenced. Sequencing depth was 130x, 122x, 70x and 81x at the four points, respectively. The frequency of the proA** allele at each time point is noted above the plot.
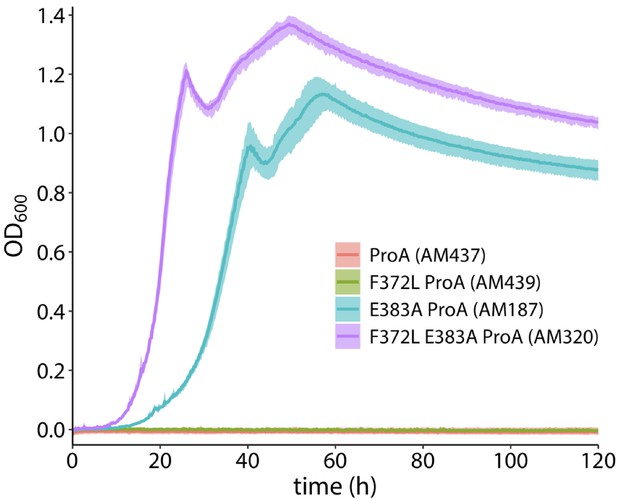
Growth of the parental strain (AM187) and comparable strains with various proA alleles in the genome.
Each strain has the M2 proBA promoter mutation that was in AM187 at the start of the evolution. The solid line shows the average values of four replicate populations. The shaded area shows the minimum and maximum values of the replicates. The strains containing wild-type ProA and F372L ProA showed no detectable growth by eight days.
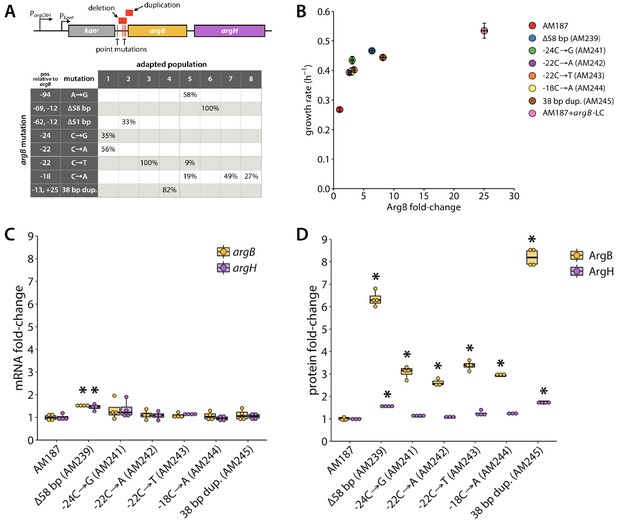
Several adaptive mutations occurred upstream of argB.
(A) Locations of adaptive mutations (red) upstream of argB and argH. argC was replaced with kanr in the parental strain AM187, giving this operon two promoters, one native to the operon (PargCBH), and the other introduced with the kanr gene (Pkanr). The table shows the percentages of each evolved population that contained a given argB mutation at the final time point. Six of the argB mutations were introduced into the genome of the parental AM187 strain and changes in growth rate (B), gene expression (C), and protein abundance (D) were determined (N = 4). Asterisks indicate values that were statistically different from those of the parental strain with p values ≤ 0.005 by a two-tailed, unequal variance Student’s t-test. In (B), error bars represent ± SE. argB-LC denotes argB expression on a low-copy plasmid under control of argB’s native promoter.
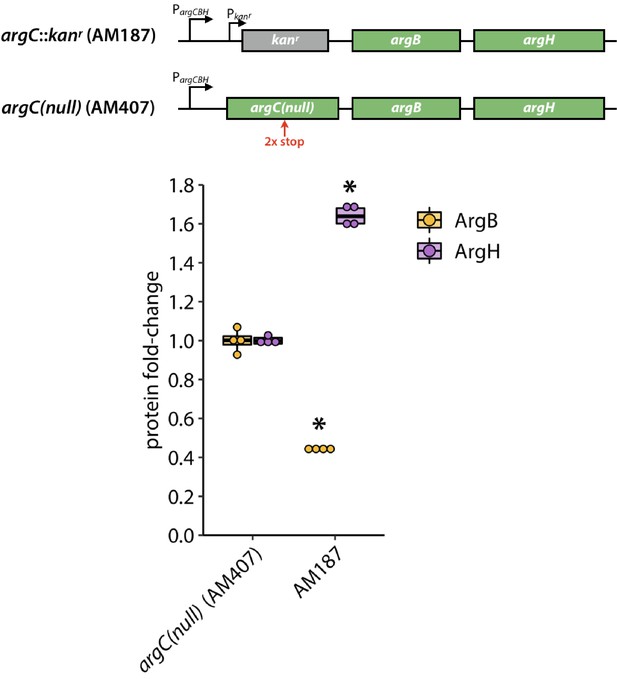
The argC::kanr construct in AM187 was replaced with a loss-of-function argC allele (top).
Levels of ArgB and ArgH levels in AM187 relative to those in AM407 determined by label-free mass spectrometry, N = 4 (bottom). Asterisks indicate values that were statistically different from those from AM407 with p values ≤ 0.0005 by a two-tailed, unequal variance Student’s t-test.
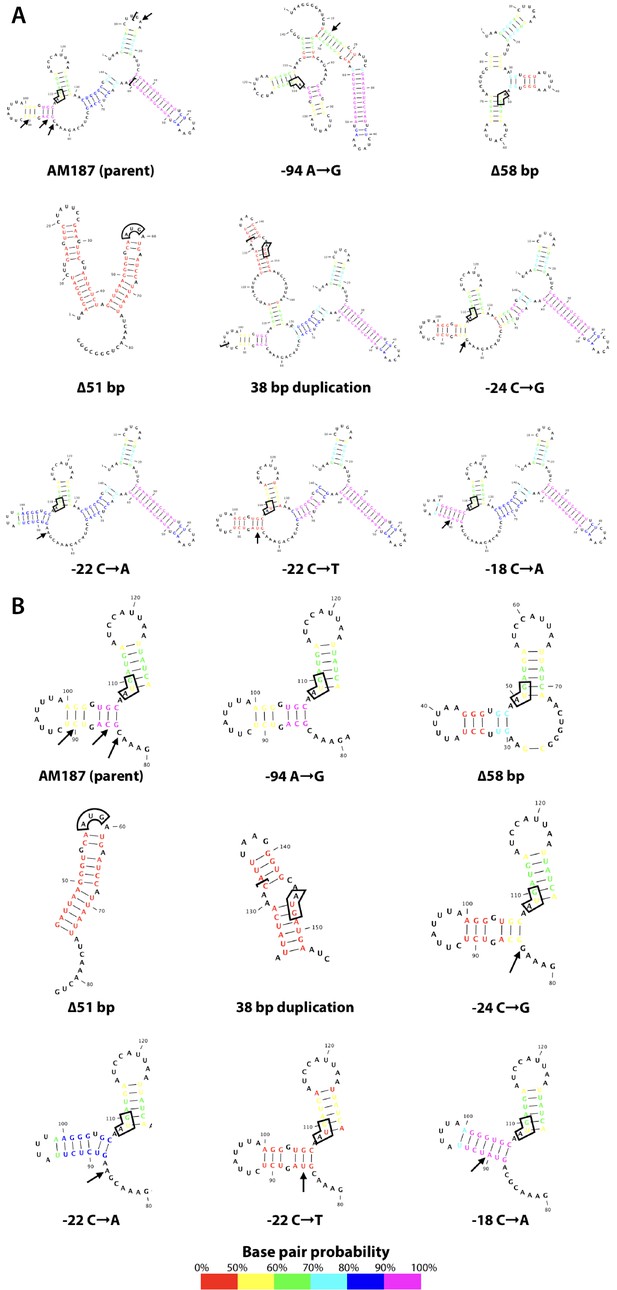
argB mRNA secondary structure comparison.
(A) The entire intergenic region between kanr and argB plus 33 nucleotides of the argB coding region was used for RNA structure prediction. The start codon for argB is boxed in each structure. Arrows in the AM187 and wild-type structure indicate locations of adaptive point mutations. Brackets in the AM187 structure contain the FLP recognition target (FRT) site. Brackets in the 38 bp duplication mutant structure contain the 38 inserted nucleotides. (B) Same as in (A), but showing only the region surrounding the argB start codon.
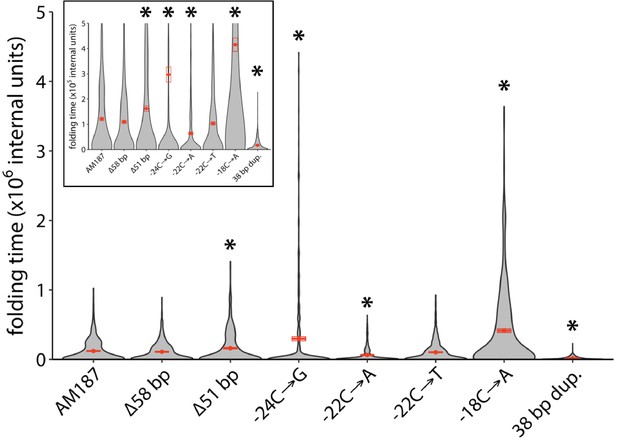
Calculated folding times of regions encompassing 30 nucleotides upstream of the argB start codon and 33 nucleotides of the argB coding region based on Kinfold simulations (Wolfinger et al., 2004).
Red points indicate the mean folding speed, and crossbars indicate one standard error from the mean, N = 500. Asterisks indicate calculated mean folding times that were statistically different from that of the AM187 sequence with p values ≤ 0.001 by a two-tailed, unequal variance Student’s t-test. Inset plot shows the same plot zoomed in 10-fold.
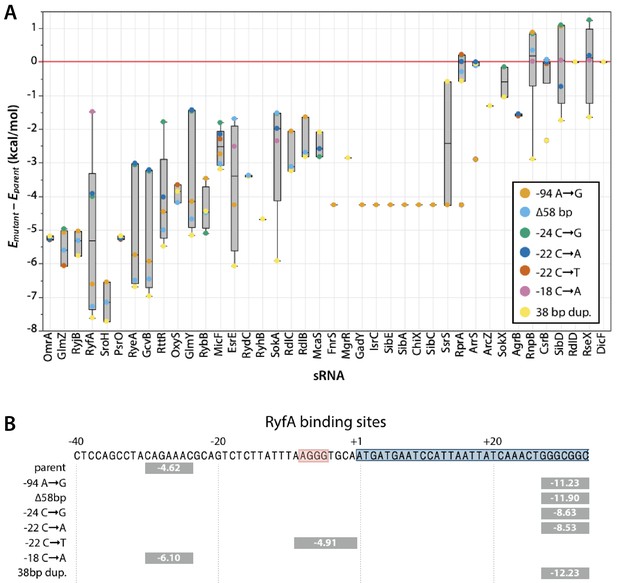
Difference in calculated energies for hybridization of sRNAs to mutant and parental argB sequences.
(A) Each box plot represents the calculated ∆E for sequences with mutations upstream of argB (the mutants shown in Figure 6A except for the ∆51 bp mutant) for each annotated sRNA in the E. coli genome. Points were excluded if Emutant > −5 kcal/mol, as those interactions are likely too weak to be physiologically relevant. ∆E < 0 indicates a stronger sRNA-mRNA interaction for the mutant sequence. ∆E > 0 indicates a weaker sRNA-mRNA interaction for the mutant sequence. (B) Predicted locations of binding of RyfA to the region upstream of argB. Predicted E (kcal/mol) is displayed for each binding site. Blue, 5’ region of argB; red, expected ribosome binding site.
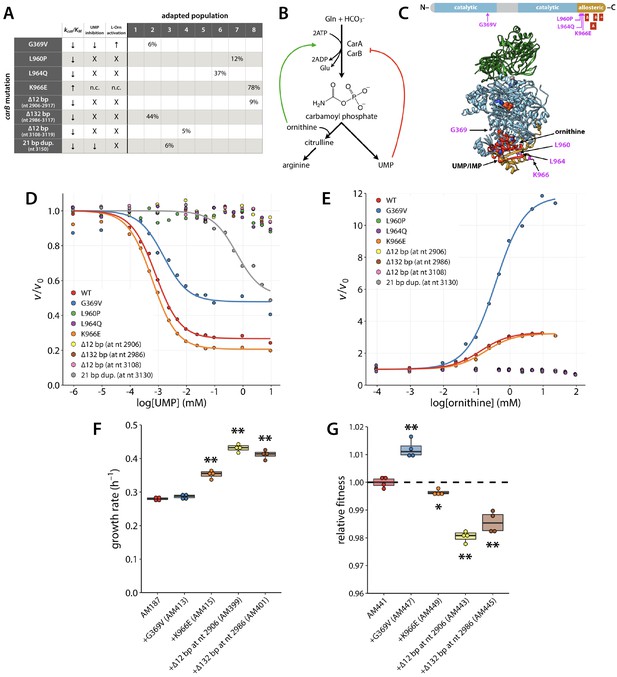
Several adaptive mutations occurred in carB.
(A) Maximum percentage of each carB mutation found in the population at any time during the evolution. Nucleotide (nt) numbers below mutant descriptions indicate where deletions or duplications occurred in the 3222 nt carB. Arrows indicate that a kinetic parameter was either increased or decreased in the variant enzyme relative to the wild-type enzyme. X, loss of activity; n.c., no change. (B) Allosteric regulation of carbamoyl phosphate synthetase. CarA and CarB are the small and large subunits of carbamoyl phosphate synthetase, respectively. (C) CarB functional domains (top) and crystal structure of E. coli CarAB (PDB 1CE8, bottom) (Thoden et al., 1999). Green, CarA; blue, CarB; gold, allosteric domain of CarB; red, residues that are deleted or duplicated in the adapted strains; magenta, point mutations that occur in the adapted strains. IMP and ornithine bound to the allosteric domain are shown as spheres. One of the two bound ATP molecules can be seen as spheres in the center of CarB. (D–E) Influence of UMP and L-ornithine on the ATPase activity of CarAB. v0; reaction rate in the absence of ligand. Each point represents the average of three technical replicates. (F) Growth rates of the parental AM187 strain and strains in which carB mutations had been introduced into the genome of AM187. (G) Relative fitness of AM441 (E. coli BW25113 containing the ∆82 bp mutation upstream of pyrE) and strains in which the carB mutations had been introduced into the genome of AM441. Asterisks in (F) and (G) indicate differences with p values < 0.03 (single asterisk) or ≤ 0.001 (double asterisk) by a two-tailed, unequal variance Student’s t-test, N = 4.
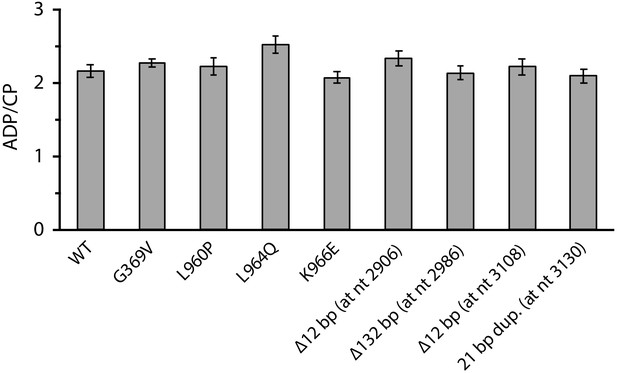
Ratio of ADP and carbamoyl phosphate (CP) produced by wild-type and mutant carbamoyl phosphate synthetases.
Values reported were averaged from three replicates. Error bars represent one standard error from the mean.
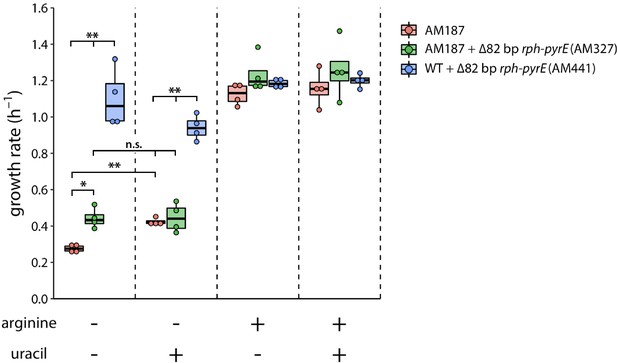
Effects of arginine (5.2 mM) and uracil (0.2 mM) on growth rates of AM187, AM327 and AM441.
Concentrations of arginine and uracil were chosen based on those in EZ rich defined medium (Neidhardt et al., 1974). *, p value < 0.01. **, p value < 0.005; n.s. indicates not significant based on a two-tailed, unequal variance Student’s t-test, N = 4.
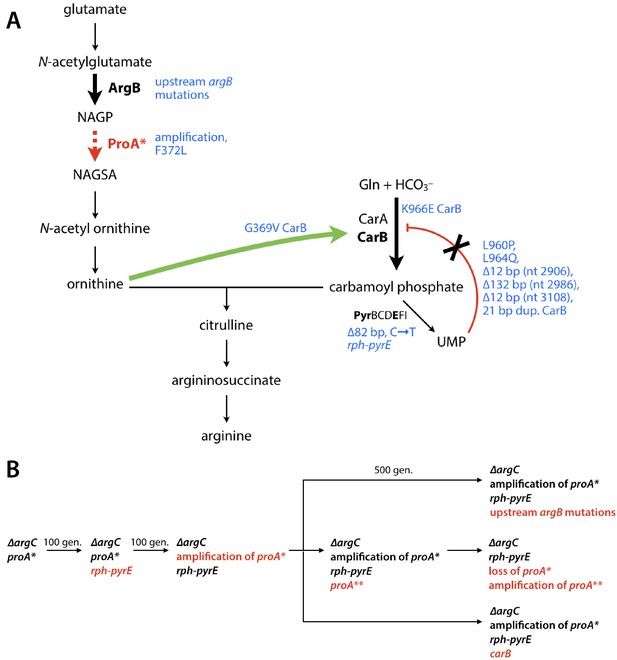
Adaptive mutations are predicted to increase flux through the arginine synthesis pathway.
(A) The pathway bottleneck enzyme ProA* is shown in red. Steps in the arginine synthesis pathway that are affected by adaptive mutations (blue) are highlighted by bold arrows. (B) Trajectories for adaptive mutations observed during 470–1000 generations of evolution. Red text denotes a new mutation.
Tables
Strains used in this work.
| strain | genotype | notes |
|---|---|---|
| E. coli BW25113 | GenBank accession number CP009273 (Grenier et al., 2014) | |
| AM008 | E. coli BW25113; argC::kanr | Keio strain (Baba et al., 2006) |
| AM187 | AM008 + −45 C→T upstream of proB (M2 promoter mutation from Kershner et al., 2016) + A1148→C in proA (changes Glu383 to Ala) + yfp construct downstream of proBA consisting of (from 5’to 3’) BBa_B0015 terminator, P3 promoter, synthetic RBS, yfp (see Materials and methods); ∆fimAICDFGH; ∆csgBAC | parental strain for adaptation, GenBank accession number CP037857.1 |
| AM209 | E. coli BL21(DE3); argC::kanr; proA::cat | used for expression and purification of wild-type and mutant ProAs |
| AM239 | AM187 + 58 bp deletion upstream of argB (pos. 4145856–4145913)a | |
| AM241 | AM187 + C4145901→G (24 bp upstream of argB start codon) | |
| AM242 | AM187 + C4145903→A (22 bp upstream of argB start codon) | |
| AM243 | AM187 + C4145903→T (22 bp upstream of argB start codon) | |
| AM244 | AM187 + C4145907→A (18 bp upstream of argB start codon) | |
| AM245 | AM187 + 38 bp duplication upstream of argB (pos. 4145912–4145949) | |
| AM267 | E. coli BL21; carAB::kanr | used for expression and purification of wild-type and mutant carbamoyl phosphate synthetases |
| AM279 | AM187 + C1169→T in ygcB (changes Ala390 to Val in Cas3) | |
| AM320 | AM187 + T1116→G in proA* (changes Phe372 to Leu) | |
| AM327 | AM187 + 82 bp deletion upstream of pyrE (pos. 3808881–3808962) | |
| AM329 | AM187 + 82 bp deletion upstream of pyrE (pos. 3808881–3808962); C1169→T ygcB (changes Ala390 to Val in Cas3) | |
| AM399 | AM187 + ∆2906–2917 carB | |
| AM401 | AM187 + ∆2986–3117 carB | |
| AM407 | AM187 + kanr::argC(null) (Gly153 and Cys154 changed to TAA stop codons) | |
| AM413 | AM187 + G1106→T carB (changes Gly369 to Val) | |
| AM415 | AM187 + A2896→G carB (changes Lys966 to Glu) | |
| AM437 | AM187 + C1148→A in proA* (reverts E383A mutation) | |
| AM439 | AM320 + C1148→A in proA** (reverts E383A mutation) | |
| AM441 | E. coli BW25113 + 82 bp deletion upstream of pyrE (pos. 3808881–3808962) | |
| AM443 | AM441 + ∆2906–2917 carB | |
| AM445 | AM441 + ∆2986–3117 carB | |
| AM447 | AM441 + G1106→T carB (changes Gly369 to Val) | |
| AM449 | AM441 + A2896→G carB (changes Lys966 to Glu) |
-
a Genome positions refer to the sequence of strain AM187 (GenBank accession number CP037857), which was modified from the E. coli BW25113 sequence (GenBank accession number CP009273; Grenier et al., 2014) based on the mutations that had been introduced.
Kinetic parameters for GSA and NAGSA dehydrogenase activities of ProA, ProA*, and ProA**.
| GSA activity (ProA) | NAGSA activity (neo-ArgC) | |||||
|---|---|---|---|---|---|---|
| kcat (s−1) | KM (mM) | kcat/KM,GSA (M−1 s−1) | kcat (s−1) | KM (mM) | kcat/KM, NAGSA (M−1 s−1) | |
| WT | 16 ± 0.3 | 0.22 ± 0.01 | 72000 ± 2000 | 0.0083 ± 0.0009 | 0.30 ± 0.09 | 28 ± 9 |
| ProA* (E383A) | 0.0076 ± 0.0008 | 0.20 ± 0.04 | 37 ± 8 | 0.046 ± 0.002 | 0.076 ± 0.009 | 610 ± 74 |
| ProA** (E383A F372L) | 0.023 ± 0.005 | 0.42 ± 0.14 | 55 ± 22 | 0.21 ± 0.01 | 0.095 ± 0.011 | 2200 ± 260 |
-
a Values reported were calculated from a nonlinear least squares regression of three replicates at each substrate concentration ± standard error.
Kinetic parametersa for the glutamine- and bicarbonate-dependent ATPase reaction of wild-type and variant carbamoyl phosphate synthetases.
| Enzyme | KM, ATP (mM) | kcat (s−1) | kcat/KM,ATP (M−1 s−1) | UMP Kd (µM) | UMP a | ornithine Kd (µM) | ornithine a |
|---|---|---|---|---|---|---|---|
| WT | 1.05 ± 0.08 | 13.5 ± 0.3 | 12.9 (±1.0)×103 | 0.81 ± 0.04 | 0.27 ± 0.01 | 130 ± 7 | 3.28 ± 0.03 |
| G369V | 3.31 ± 0.25 | 21.5 ± 0.7 | 6.51 (±0.54)×103 | 1.53 ± 0.29 | 0.48 ± 0.02 | 372 ± 20 | 11.8 ± 0.13 |
| L960P | 1.12 ± 0.05 | 9.10 ± 0.15 | 8.12 (±0.41)×103 | na | na | na | na |
| L964Q | 1.25 ± 0.08 | 8.04 ± 0.17 | 6.41 (±0.42)×103 | na | na | na | na |
| K966E | 0.97 ± 0.06 | 20.6 ± 0.4 | 21.2 (±1.4)×103 | 0.61 ± 0.04 | 0.21 ± 0.01 | 181 ± 34 | 3.23 ± 0.08 |
| ∆12 bp (at nt 2906)b | 1.09 ± 0.06 | 4.80 ± 0.09 | 4.39 (±0.25)×103 | na | na | na | na |
| ∆132 bp (at nt 2986) | 1.16 ± 0.06 | 6.47 ± 0.11 | 5.57 (±0.31)×103 | na | na | na | na |
| ∆12 bp (at nt 3108) | 1.30 ± 0.10 | 5.86 ± 0.16 | 4.51 (±0.37)×103 | na | na | na | na |
| 21 bp dup. (at nt 3130) | 1.40 ± 0.12 | 9.70 ± 0.30 | 6.94 (±0.64)×103 | 597 ± 133 | 0.51 ± 0.03 | na | na |
-
a Values reported ± standard error. Values for Kd and a for UMP and ornithine were determined by fitting the data to the following equation: v/v0 = (aL + Kd)/(L + Kd), where L is the ligand concentration, v is the initial reaction rate, v0 is the initial reaction rate in the absence of ligand, a is v/v0 at infinite L, and Kd is the apparent dissociation constant. No Kd or a values are given (indicated by na) when inhibition by the allosteric ligand was too weak to measure.
b Nucleotide (nt) numbers refer to the position of deletions or duplications in carB.
Additional files
-
Source code 1
R script used to calculate growth rate for each population from turbidostat OD650 readings.
- https://cdn.elifesciences.org/articles/53535/elife-53535-code1-v2.r
-
Source code 2
Example R script used to calculate Michaelis-Menten parameters.
- https://cdn.elifesciences.org/articles/53535/elife-53535-code2-v2.r
-
Source code 3
Example R script used to calculate Kd and a for CPS.
- https://cdn.elifesciences.org/articles/53535/elife-53535-code3-v2.r
-
Supplementary file 1
Plasmids used in this study.
- https://cdn.elifesciences.org/articles/53535/elife-53535-supp1-v2.docx
-
Supplementary file 2
Primers used in this study.
- https://cdn.elifesciences.org/articles/53535/elife-53535-supp2-v2.docx
-
Supplementary file 3
Protospacers used for Cas9-mediated scarless genome editing.
- https://cdn.elifesciences.org/articles/53535/elife-53535-supp3-v2.docx
-
Supplementary file 4
Mutation cassettes used for Cas9-mediated genome editing.
- https://cdn.elifesciences.org/articles/53535/elife-53535-supp4-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/53535/elife-53535-transrepform-v2.docx