Epigenetics: A memory of longevity
It is commonly accepted that genetic sequences coded within DNA are passed down through generations and can influence characteristics such as appearance, behavior and health. However, emerging evidence suggests that some traits can also be inherited ‘epigenetically’ from information that is independent of the DNA sequence.
One of the ways characteristics may be epigenetically passed down is through the temporary modification of histone proteins which help to package DNA into the cell. Histones are adorned with chemical marks that can regulate how and when a gene is expressed by changing how tightly the DNA is wrapped. These marks are typically removed before genetic information is passed on to the next generation, but some sites escape erasure (Heard and Martienssen, 2014; Kelly, 2014; Miska and Ferguson-Smith, 2016).
It has previously been reported that genetic mutations in an enzyme complex called COMPASS increase the lifespan of tiny worms called Caenorhabditis elegans (Greer et al., 2010). This complex acts on histones and creates a chemical mark called H3K4me, which is typically associated with less compact DNA and higher gene expression. When these mutants mate with wild-type worms they generate descendants that no longer have COMPASS mutations. Although these wild-type offspring recover normal levels of H3K4me, they still inherit the long-lived phenotype which they sustain for several generations (Greer et al., 2011). This observation suggests that an epigenetic mechanism that is independent from the gene mutation causes this inherited longevity. Now, in eLife, David Katz and co-workers at the Emory University School of Medicine in Atlanta – including Teresa Lee as first author – report a possible mechanism to explain how this longer lifespan is epigenetically inherited across multiple generations (Lee et al., 2019).
Previous work showed that one of the COMPASS complex mutants, known as wdr-5, has increased levels of another histone mark called H3K9me2 (Kerr et al., 2014). This epigenetic mark generally promotes DNA compaction and appears to antagonize the action of H3K4me. This led Lee et al. to question whether the elevated levels of H3K9me2 may be important for the inheritance of this extended lifespan in wdr-5 worms.
To test their hypothesis, the team carefully monitored the levels and patterns of H3K9me2 in the mutants. Surprisingly, they found that homozygous wdr-5 mutants, which had descended from ancestors carrying one copy of the mutated wdr-5 gene and one wild-type copy for multiple generations, did not live for longer than their non-mutant counterparts. This indicates that the mutation carried by wdr-5 worms did not immediately cause a lifespan change. However, future generations of worms that maintained the homozygous wdr-5 mutation had an increasingly longer lifespan, suggesting that the accumulation of an epigenetic signal across generations promotes longer living. These late generation wdr-5 mutants had higher levels of H3K9me2, and they were able to pass on this extended longevity to their progeny following mating with wild-type worms as previously reported (Greer et al., 2011).
Next, Lee et al. manipulated the levels of H3K9me2 to see how this affected the phenotype of the late generation, long-lived wdr-5 worms. First, they blocked the gain in H3K9me2 levels in the mutants by introducing a defective version of an enzyme called MET-2 which normally promotes the addition of H3K9me2 (Figure 1). As a result, neither the wdr-5 mutants nor their descendants experienced a longer lifespan. Lee et al. reasoned that if higher H3K9me2 levels are responsible for longevity, then increasing the amount of H3K9me2 by a different mutation should result in the same phenotype as the wdr-5 worms. They found that worms with defects in the enzyme JHDM-1, which is predicted to remove H3K9me2, not only lived longer but also passed on this trait to their wild-type progeny for several generations. Together, these data strongly suggest that increased H3K9me2 levels contribute to extended longevity and its inheritance.
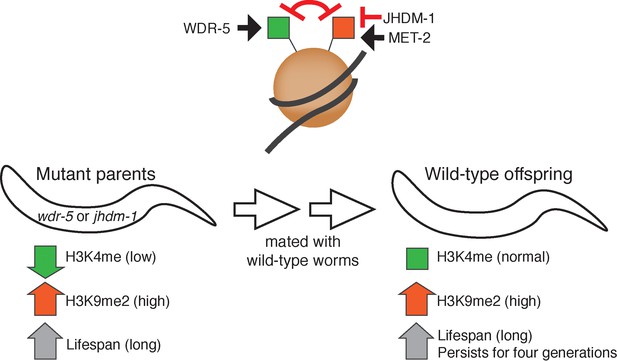
Certain epigenetic changes are linked to the inheritance of extended lifespans in worms.
Top: The WDR-5 enzyme helps to place the H3K4me mark (green), which promotes gene expression, on proteins called histones (brown circle) that package DNA (grey ribbon). In parallel, the MET-2 enzyme places the H3K9me2 mark (red), which represses gene expression. The two marks functionally antagonize each other. An enzyme called JHDM-1 is predicted to remove H3K9me2. Bottom: Worms with mutations in wdr-5 or jhdm-1 (left) that have low levels of H3K4me (green arrow), also show higher levels of H3K9me2 (red arrow) and an increased lifespan (grey arrow). When these long-lived mutants are mated to wild-type worms with a normal lifespan, their genetically wild-type offspring (right) are still long-lived for several generations (grey arrow). These worms show normal levels of H3K4me mark (green square) and regions of sustained increase in H3K9me2 (red arrow) inherited from their mutant ancestors.
Image credit: Cheng-Lin Li.
To build on these findings, Lee et al. explored where the H3K9me2 marks were deposited in the genomes of the worms. As expected, long-lived wdr-5 and jhdm-1 mutants have more H3K9me2 marks spread across their genomes. Critically, Lee et al. found that specific genes in the wild-type offspring of jhdm-1 mutants had higher levels of H3K9me2. These results are intriguing and suggest that increasing H3K9me2 levels in certain genes may be the key to passing on this long living phenotype to future generations. Exciting future investigations will be to identify all the gene regions associated with the inherited increase in H3K9me2, and to understand how changes to DNA packaging and gene expression in those regions influence longevity.
A handful of previous studies in C. elegans have demonstrated that specific histone modifications can be inherited across generations (Katz et al., 2009; Kerr et al., 2014; Kaneshiro et al., 2019). However, the paper by Lee et al. is the first to tie together the inheritance of a histone mark to longer lifespan. H3K9me2 is an evolutionarily conserved histone mark which is known to preserve spatial organization during cell division in organisms ranging from humans to worms (Poleshko et al., 2019). Going forward, it will be interesting to study whether H3K9me2 also participates in how traits are inherited across multiple generations in mammals.
References
Article and author information
Author details
Publication history
- Version of Record published: January 24, 2020 (version 1)
Copyright
© 2020, Emerson et al.
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 3,192
- views
-
- 272
- downloads
-
- 1
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Download links
Downloads (link to download the article as PDF)
Open citations (links to open the citations from this article in various online reference manager services)
Cite this article (links to download the citations from this article in formats compatible with various reference manager tools)
Further reading
-
- Cancer Biology
- Chromosomes and Gene Expression
The chromatin-associated protein WD Repeat Domain 5 (WDR5) is a promising target for cancer drug discovery, with most efforts blocking an arginine-binding cavity on the protein called the ‘WIN’ site that tethers WDR5 to chromatin. WIN site inhibitors (WINi) are active against multiple cancer cell types in vitro, the most notable of which are those derived from MLL-rearranged (MLLr) leukemias. Peptidomimetic WINi were originally proposed to inhibit MLLr cells via dysregulation of genes connected to hematopoietic stem cell expansion. Our discovery and interrogation of small-molecule WINi, however, revealed that they act in MLLr cell lines to suppress ribosome protein gene (RPG) transcription, induce nucleolar stress, and activate p53. Because there is no precedent for an anticancer strategy that specifically targets RPG expression, we took an integrated multi-omics approach to further interrogate the mechanism of action of WINi in human MLLr cancer cells. We show that WINi induce depletion of the stock of ribosomes, accompanied by a broad yet modest translational choke and changes in alternative mRNA splicing that inactivate the p53 antagonist MDM4. We also show that WINi are synergistic with agents including venetoclax and BET-bromodomain inhibitors. Together, these studies reinforce the concept that WINi are a novel type of ribosome-directed anticancer therapy and provide a resource to support their clinical implementation in MLLr leukemias and other malignancies.
-
- Chromosomes and Gene Expression
- Immunology and Inflammation
Ikaros is a transcriptional factor required for conventional T cell development, differentiation, and anergy. While the related factors Helios and Eos have defined roles in regulatory T cells (Treg), a role for Ikaros has not been established. To determine the function of Ikaros in the Treg lineage, we generated mice with Treg-specific deletion of the Ikaros gene (Ikzf1). We find that Ikaros cooperates with Foxp3 to establish a major portion of the Treg epigenome and transcriptome. Ikaros-deficient Treg exhibit Th1-like gene expression with abnormal production of IL-2, IFNg, TNFa, and factors involved in Wnt and Notch signaling. While Ikzf1-Treg-cko mice do not develop spontaneous autoimmunity, Ikaros-deficient Treg are unable to control conventional T cell-mediated immune pathology in response to TCR and inflammatory stimuli in models of IBD and organ transplantation. These studies establish Ikaros as a core factor required in Treg for tolerance and the control of inflammatory immune responses.






