Regulation of BMP4/Dpp retrotranslocation and signaling by deglycosylation
Figures
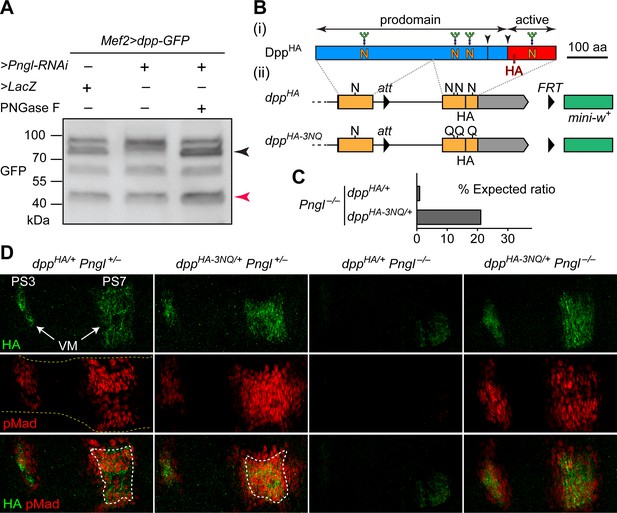
Deglycosylation of Dpp by Pngl is essential for BMP signaling during Drosophila midgut development.
(A) Western blot with α-GFP on protein lysates from embryos of indicated genotypes. The shift in band size upon treatment with PNGase F (black arrowhead) shows that full-length Dpp-GFP retains N-glycans upon RNAi-mediated Pngl knock-down in the embryonic mesoderm. The mature Dpp-GFP is indicated by the red arrowhead. (B) (i) Schematic representation of DppHA, which contains an HA tag in the active domain (red box). (ii) Schematic representation of dppHA knock-in allele and its mutant version dppHA-3NQ, in which three of Dpp’s four N-glycosylation sites are ablated by N-to-Q mutations. (C) Eclosion tests of Pngl–/– flies harboring a copy of dppHA (1.3% of the expected Mendelian ratio, n = 153 total progeny scored) or dppHA-3NQ (20.8% of the expected Mendelian ratio, n = 158 total progeny scored). One copy of dppHA-3NQ partially rescues the lethality of Pngl mutant flies. (D) Immunofluorescence staining of parasegment 3 (PS3) to PS7 region of embryonic midguts (marked by the dashed yellow lines) of animals with the indicated genotypes. Dpp is expressed in PS3 and PS7 of the embryonic visceral mesoderm (VM) and induces BMP signaling (evidenced by pMad expression) in PS3 and PS7, both in the VM (paracrine/autocrine) and in the endoderm. pMad staining (red) indicates that one copy of dppHA-3NQ, but not dppHA, restores the Dpp signaling during Pngl–/– midgut development. n = 5 for each genotype. Scale bar, 50 μm.
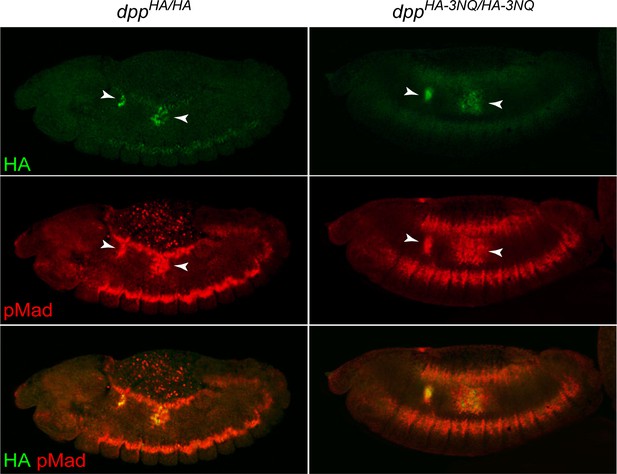
The dppHA-3NQ knock-in allele does not impair Dpp signaling in the midgut.
Confocal microscopy images of embryos homozygous for dppHA and dppHA-3NQ alleles at stage 14 stained for HA and pMad. Anterior is to the left, dorsal is up. Projection views are shown. The HA and especially pMad expression are broadened upon 3NQ mutation. Arrowheads indicate embryonic parasegment (PS) three and PS7. The red puncta in the dorsal (amnioserosa) region of the dppHA/HA embryo are artifacts due to nonspecific background staining.
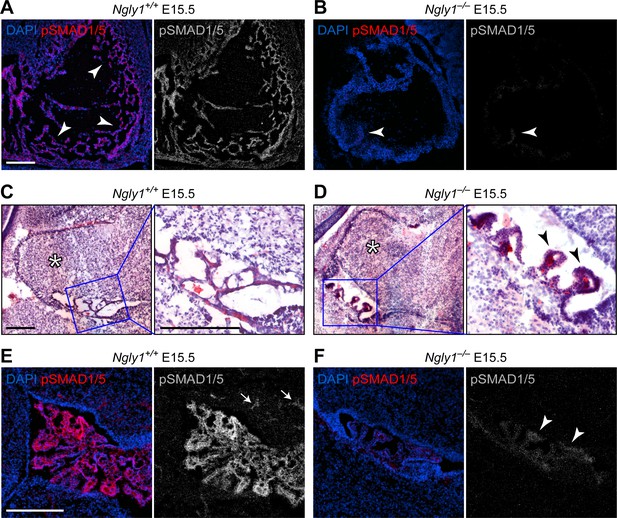
Ngly1 is required for BMP signaling in embryonic heart and cerebellum.
(A,B) Immunofluorescence staining in sagittal sections of the embryonic heart of control (Ngly1+/+) and Ngly1–/– animals at E15.5. Nuclei (DAPI) are in blue and pSMAD1/5 in red. n = 5 for each genotype. pSMAD1/5 staining is severely reduced in mutant embryos (compare grey scale images), suggesting that loss of Ngly1 leads to reduced BMP signaling in the heart. Arrowheads mark the myocardial trabeculae. (C–F) H&E (C,D) and immunofluorescence (E,F) staining in sagittal sections of embryonic cerebellum of control (Ngly1+/+) and Ngly1–/– animals at E15.5. n = 5 for each genotype. Loss of Ngly1 leads to significant morphological abnormalities in the 4th ventricle choroid plexus (close-up images from blue boxes) and severe reduction in pSMAD1/5 staining (in grey scale). Scale bars are 100 µm in A,B,E,F and 200 µm in c,d (both low magnification and close-up images). These data suggest that Ngly1 has a role in BMP signaling in developing choroid plexus.
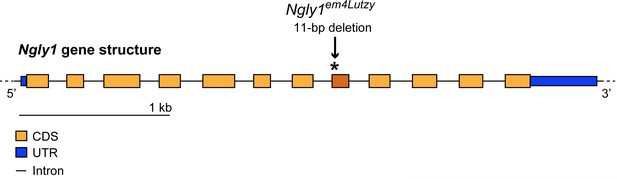
Schematic of the mouse Ngly1 locus and the location of the 11 bp deletion in the Ngly1em4Lutzy allele.
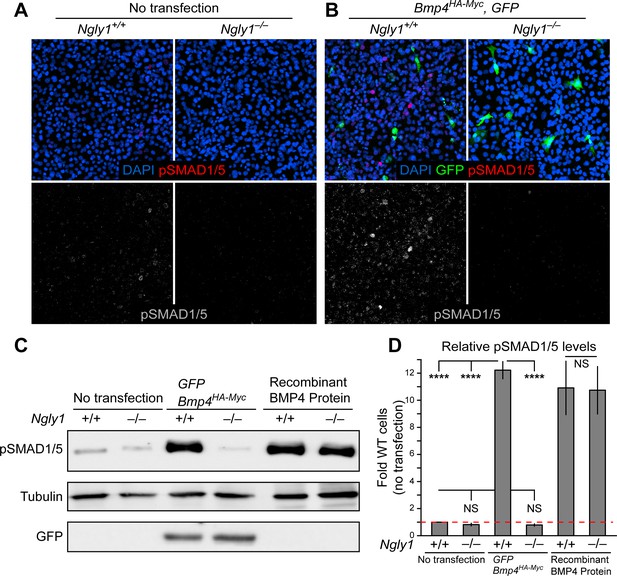
Loss of Ngly1 leads to impaired BMP4 signal-sending in mouse embryonic fibroblasts.
(A) Representative immunofluorescence staining of DAPI (blue) and pSMAD1/5 (red) in untransfected WT and Ngly1 mutant (–/–) MEFs used as a baseline for pSMAD1/5 expression (grey scale). (B) WT and Ngly1 mutant MEFs co-transfected with expression vectors for BMP4HA-Myc and GFP. GFP expression is used as control for transfection efficiency. pSMAD1/5 expression (grey scale) is used as readout of BMP signaling. Scale bars, 50 μm. n = 6 biologically independent samples. Loss of Ngly1 severely reduces pSMAD1/5 levels. (C) Representative western blot of pSMAD1/5 from WT and Ngly1 mutant MEF cell lysates. The first two lanes are without transfection (control); the middle two lanes are co-transfected with expression vectors for BMP4HA-Myc and GFP (for transfection efficiency); the last two lanes are treated with recombinant BMP4 in the culture media. (D) Quantification of the data shown in C. For each blot, the pSMAD1/5 expression was normalized to tubulin and the value of Ngly1+/+ cells without transfection was set as 1. n = 3 biologically independent samples. Two-way ANOVA with Tukey's multiple comparisons test was used for statistical analysis. Mean ± s.d. are shown. ****p<0.0001, NS, not significant. These data exhibit that BMP4 signal-sending, but not signal-receiving, is impaired in MEFs upon loss of Ngly1, similarly to the impaired Dpp signal-sending in Drosophila embryonic midguts (Galeone et al., 2017).
-
Figure 3—source data 1
Raw data and statistical analysis for panel D.
- https://cdn.elifesciences.org/articles/55596/elife-55596-fig3-data1-v2.xlsx
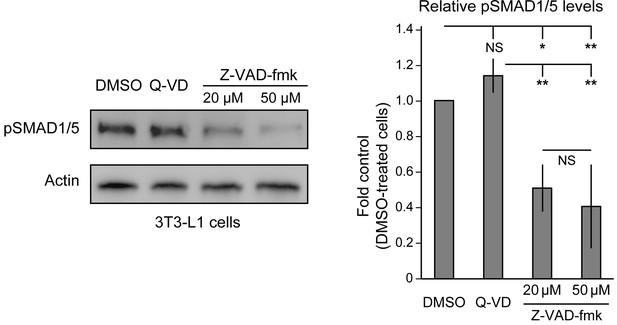
Endogenous BMP signaling is impaired upon NGLY1 inhibition in 3T3-L1 cells.
Representative pSMAD1/5 western blot on cell lysates of 3T3-L1 cells treated with DMSO, the broad caspase inhibitor Q-VD-OPh at 20 μM, and Z-VAD-fmk at 20 μM and 50 μM. The first two lanes are used as negative controls. For each blot, the pSMAD1/5 expression was normalized to tubulin. n = 3 biologically independent samples. Two-way ANOVA with Tukey's multiple comparisons test was used for statistical analysis. Mean ± s.d. are shown. *p<0.05; **p<0.01; NS, not significant.
-
Figure 3—figure supplement 1—source data 1
Raw data and statistical analysis for the graph.
- https://cdn.elifesciences.org/articles/55596/elife-55596-fig3-figsupp1-data1-v2.xlsx
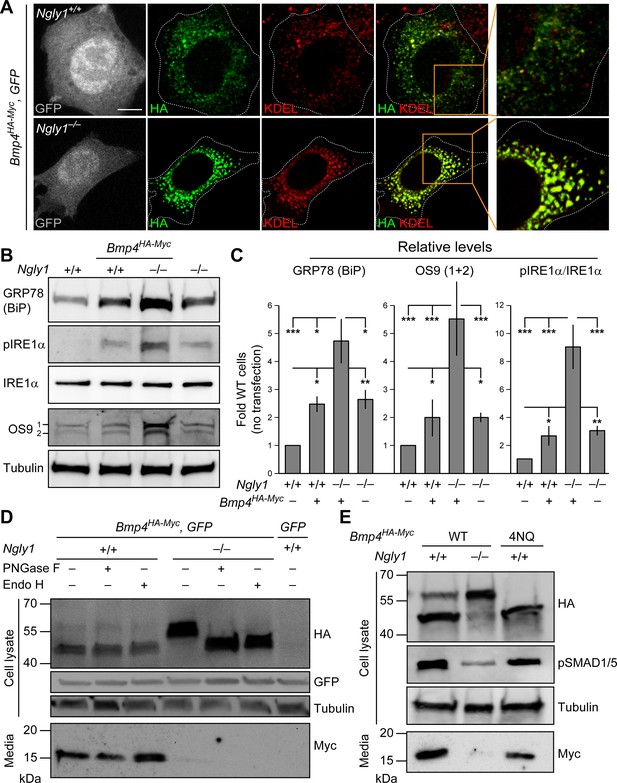
In Ngly1 mutant MEFs, BMP4 retains N-glycans and is trapped in the ER.
(A) Confocal sections of WT and Ngly1 mutant MEFs co-transfected with Bmp4HA-Myc and GFP (for transfection efficiency) and stained for HA (to mark BMP4HA-Myc, green) and the ER marker KDEL (red). HA-tagged BMP4 almost fully colocalizes with KDEL/GRP78 in Ngly1 mutant cells, suggesting impaired exocytic trafficking. Scale bars, 10 μm. n = 8 biologically independent samples. (B,C) Western blot with α-KDEL antibody, which marks GRP78 (BiP), and α-pIRE1α, α-IRE1α, and α-OS9 antibodies on protein lysates from WT and Ngly1 mutant MEFs with or without Bmp4HA-Myc transfection. ER stress is induced in Ngly1–/– MEFs and is significantly enhanced upon Bmp4 transfection, likely due to the accumulation of misfolded BMP4. A representative image of three independent blots is shown. For each blot, the protein expression was normalized to tubulin and the value of Ngly1+/+ cells without transfection was set as 1. Two-way ANOVA with Tukey's multiple comparisons test was used for statistical analysis. Mean ± s.d. is shown. *p<0.05; **p<0.01; ***p<0.001; NS, not significant. (D) Western blot for HA and Myc on cell lysates and media from WT and Ngly1 mutant MEFs transfected with expression vectors for BMP4HA-Myc and GFP or GFP only (last lane). Protein extracts were treated with PNGase and Endo H for glycosylation profiling. Note the accumulation of a slow-migrating HA+ band in Ngly1–/– MEFs which returns to normal size upon treatment with both enzymes, indicating the retention of N-glycans on BMP4 upon loss of Ngly1. Note also the severe decrease in the level of active BMP4 (Myc blot) secreted by Ngly1–/– MEFs into the culture media. (E) Western blot for HA, pSMAD1/5 and Myc on cell lysates and media from WT MEFs transfected with an expression vector for BMP4HA-Myc-4NQ compared with WT and Ngly1–/– MEFs expressing wild-type BMP4HA-Myc. n = 3 independent biological samples. These data indicate that N-glycans on BMP4 are not necessary for its secretion (Myc) and signaling (pSMAD1/5) and that NGLY1 promotes BMP4 signaling by removing one or more N-glycans from BMP4, similar to their Drosophila counterparts.
-
Figure 4—source data 1
Raw data and statistical analysis for panel C.
- https://cdn.elifesciences.org/articles/55596/elife-55596-fig4-data1-v2.xlsx
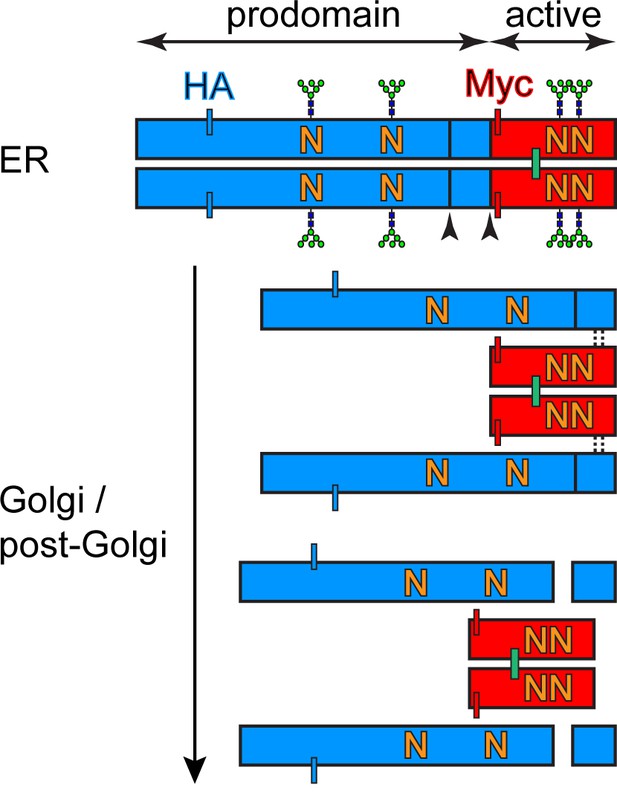
BMP4 processing in sending cells.
Schematic representation of BMP4HA-Myc glycosylation sites, cleavages sites, location of the tags and maturation from ER to Golgi. The prodomain is marked in blue, the active domain is marked in red.
Mutations in VCP-binding sites abolish the recruitment of NGLY1 to ER.
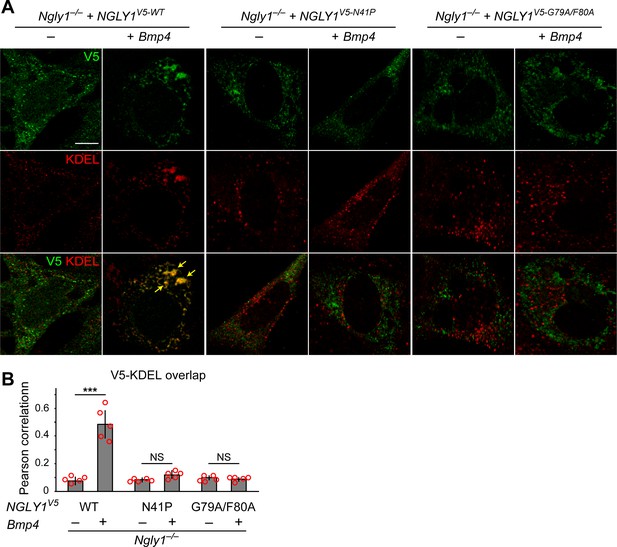
(A) Confocal sections of Ngly1–/– MEFs transfected with expression vectors for wild-type or VCP-binding mutant versions of human NGLY1 tagged with V5 and stained with V5 (green) and KDEL (ER marker, red). Note the ER recruitment of wild-type NGLY1 upon Bmp4 transfection, which is likely due to the accumulation of misfolded BMP4 in the ER. Mutant NGLY1 proteins do not show ER recruitment. (B) Analysis of relative fluorescence overlap of V5-tagged NGLY1 (green) and KDEL signals (red) in A. The signal overlap is quantified by Pearson correlation analysis of five images from three independent experiments. Scale bars, 10 μm. Mean ± s.d. is shown. ***p=0.00011; NS, not significant. These results show that mutations in VCP-binding sites lead to a failure of the NGLY1 recruitment to ER.
-
Figure 5—source data 1
Raw data and statistical analysis for panel B.
- https://cdn.elifesciences.org/articles/55596/elife-55596-fig5-data1-v2.xlsx
ER recruitment of NGLY1 is required for BMP4 deglycosylation and signaling in MEFs and Dpp signaling in Drosophila embryonic midgut.
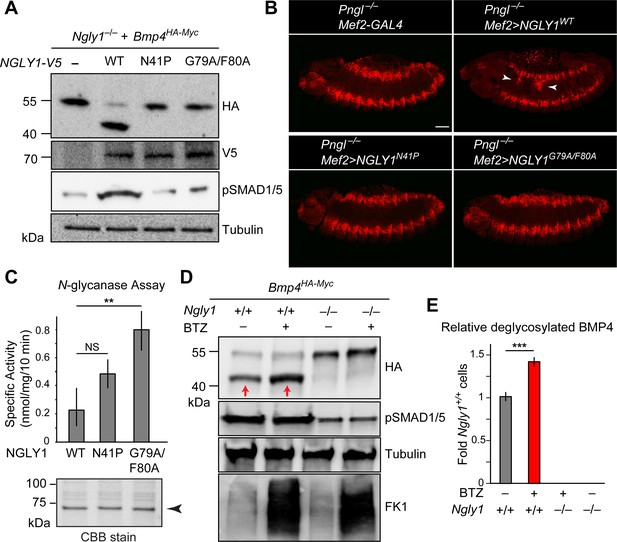
(A) Western blots with V5, HA and pSMAD1/5 antibodies on cell lysates from Ngly1–/– cells expressing BMP4HA-Myc alone or with human NGLY1V5-His6 (wild-type or VCP-binding mutants). Wild-type NGLY1 efficiently deglycosylates BMP4 (downward shift of the HA+ band) and induces robust BMP signaling (pSMAD1/5), but NGLY1 with mutations in its VCP-binding sites fail to deglycosylate BMP4 and induce BMP signaling. (B) Projection views of confocal image stacks of stage 14 embryos with indicated genotypes stained for pMad. Expression of wild-type human NGLY1 in mesoderm rescues BMP signaling in the midgut region (arrowheads), but the VCP-binding mutant versions of human NGLY1 do not rescue this phenotype. (C) Coomassie Brilliant Blue (CBB) stain of recombinant wild-type and VCP binding mutant (N41P and G79A/F80A) NGLY1V5-His6 proteins generated in wheat germ cell-free system and deglycosylation activity assays on these proteins. Note that the VCP-binding mutants are stably expressed (arrowhead) and do not decrease NGLY1’s enzymatic activity. One-way ANOVA with Tukey's multiple comparisons test was used. Mean ± s.d. **p<0.01; NS, not significant. (D) Western blot of HA-tagged BMP4 from cell lysates of WT and Ngly1–/– MEFs treated with bortezomib (BTZ, 10 nM for 6 hr). (E) Quantification of deglycosylated HA-tagged BMP4 expression. n = 3 biologically independent samples. Two-way ANOVA test with Tukey's multiple comparisons test was used for statistical analysis. Mean ± s.d. ***p<0.001. These results indicate that deglycosylated BMP4 is degraded by proteasome as result of ER-to-cytosol retrotranslocation. The proteasome impairment of BMP4 degradation does not block BMP signaling.
-
Figure 6—source data 1
Raw data and statistical analysis for panels C and E.
- https://cdn.elifesciences.org/articles/55596/elife-55596-fig6-data1-v2.xlsx
Transgenic expression of human NGLY1 in Drosophila wing discs.
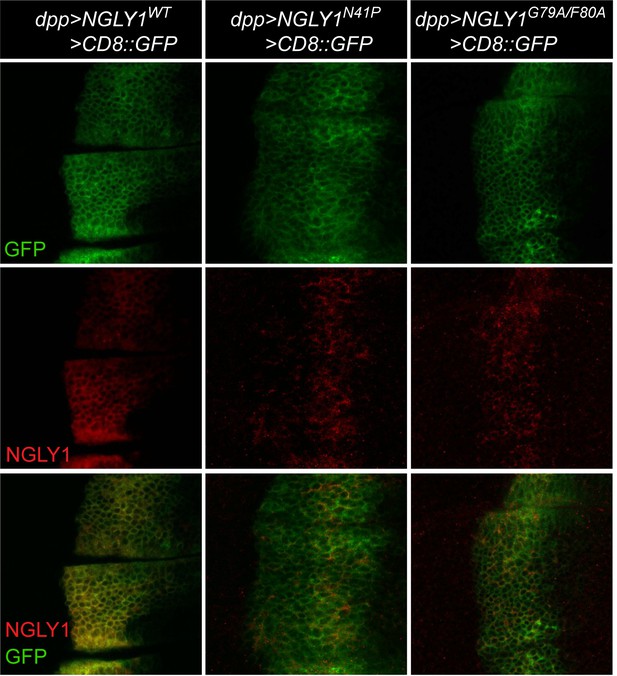
Shown are third instar wing imaginal discs that express wild-type human NGLY1 and NGLY1 with mutations in VCP-binding sites (N41P and G79A/F80A) using the dpp-GAL4 driver. NGLY1 is marked in red; the dpp expression domain is marked in green by using UAS-CD8::GFP in the background. n = 5 for each genotype. Scale bar, 25 μm. Note that VCP-binding mutant versions of NGLY1 are expressed in vivo but show a pattern different from that of wild-type NGLY1.
Misfolded BMP4 binds the ER lectin OS9 and is a substrate of ERAD.
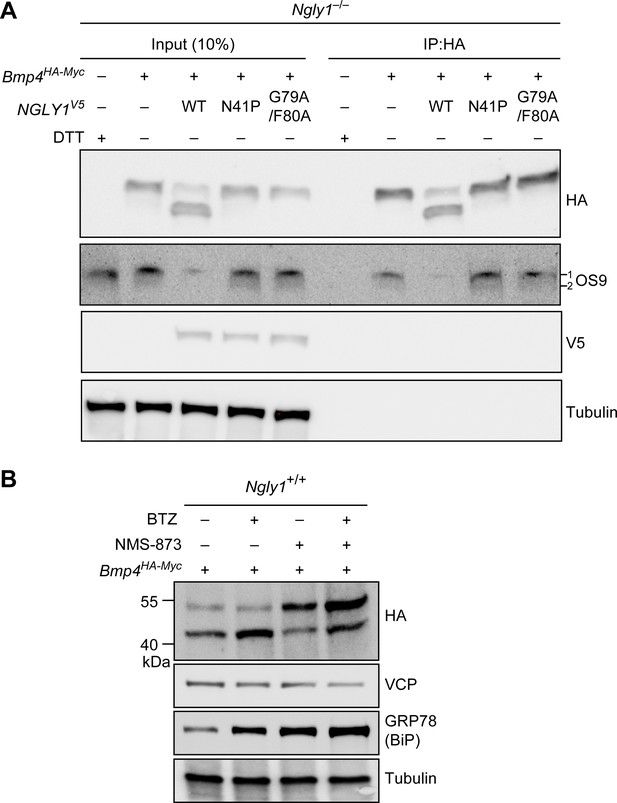
(A) Co-IP analysis of HA-tagged BMP4 and OS9 from Ngly1–/– MEFs. The lysates were immunoprecipitated with HA and analyzed by immunoblotting with α-HA and α-OS9 antibodies. Untransfected cells treated with 2 mM DTT were used as control. Input represents 10% of the total cell extract used for IP. Note that BMP4 binds OS9 when NGLY1 is missing or fails to be recruited to the ER. (B) Western blot of HA-tagged BMP4 from Ngly1+/+ MEF cell lysates treated with NMS-873 (5 μM for 6 hr) and/or BTZ (10 nM for 6 hr). Note that upon NMS-873 treatment, glycosylated BMP4 accumulates in the cell and the level of ER chaperone GRP78 (BiP) is increased, suggesting that misfolded BMP4 accumulates in the ER due to impaired ERAD.
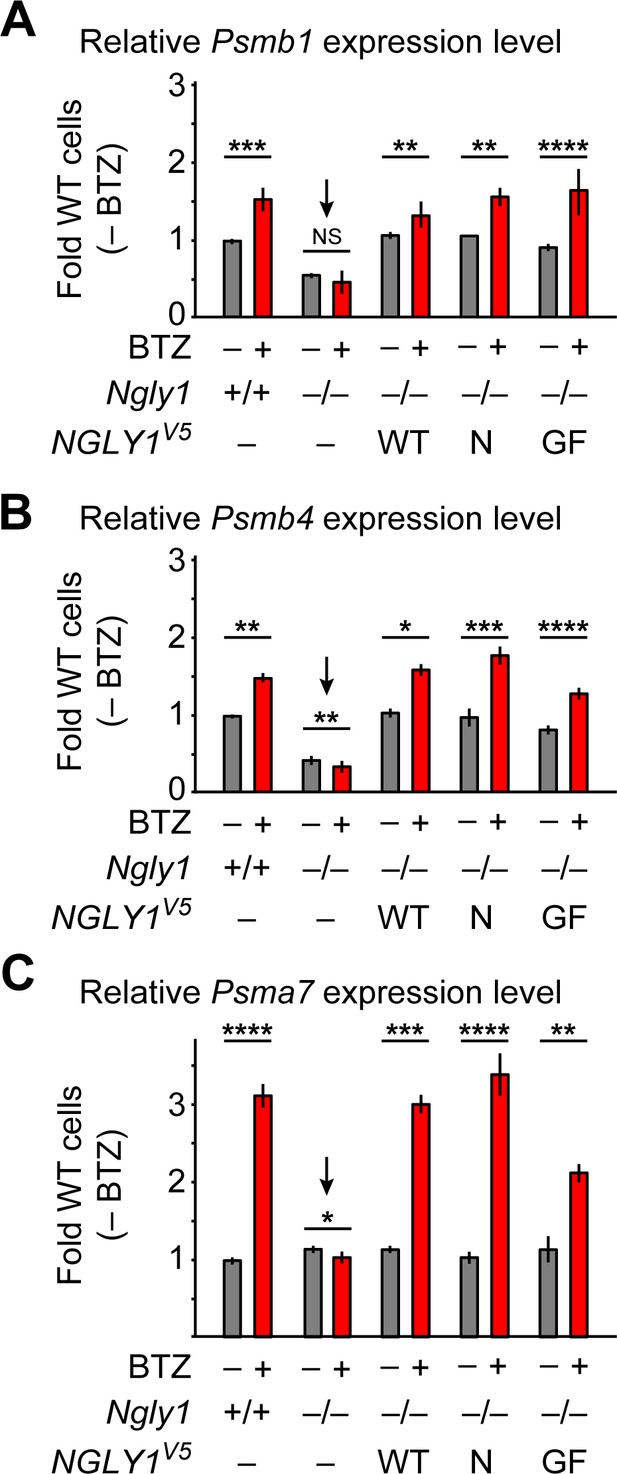
ER-associated NGLY1 is not required for NFE2L1-mediated proteasome bounce-back.
(A–C) qRT-PCR assays for proteasome subunit genes in WT and Ngly1 mutant MEFs with or without bortezomib (BTZ) treatment. Ngly1 mutant MEFs were transfected with NGLY1V5-WT and NGLY1V5 with mutations in VCP-binding sites (N41P and G79A/F80A). n = 3 biologically independent samples. Two-way ANOVA test with Tukey's multiple comparisons tests was used for statistical analysis. Mean ± s.e.m. is shown. *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001; NS, not significant. Despite impaired ER recruitment, the mutant NGLY1 proteins retain the ability to promote the proteasome bounce-back response, which depends on NFE2L1 deglycosylation by NGLY1 (Tomlin et al., 2017).
-
Figure 8—source data 1
Raw data and statistical analysis for panels A-C.
- https://cdn.elifesciences.org/articles/55596/elife-55596-fig8-data1-v2.xlsx
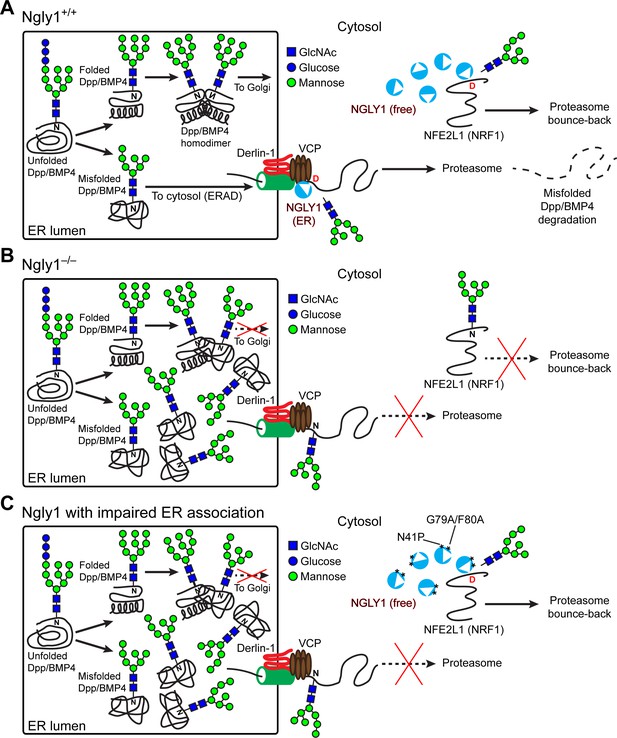
Cytosolic NGLY1 is dynamically recruited to retrotranslocon machinery for misfolded BMP4 retrotranslocation.
(A) Proposed model: Upon ER stress due to the generation of misfolded BMP4/Dpp in the ER, cytosolic NGLY1 is recruited to the ER in a VCP-dependent manner and deglycosylates BMP4 molecules to promote their retrotranslocation and proteasomal degradation. Deglycosylation and N-to-D sequence editing of NFE2L1 by NGLY1 allows the proteasome bounce-back response. (B) Loss of Ngly1 leads to impaired BMP4 deglycosylation and retrotranslocation and thereby results in the accumulation of misfolded BMP4 molecules in the ER and a severe decrease in BMP4 signaling. It also impairs NFE2L1 deglycosylation/sequence editing, which leads to the loss of the proteasome bounce-back response. (C) The cytosolic NGLY1 pool is sufficient for NFE2L1 deglycosylation/sequence editing and proteasome bounce-back but not for BMP4 deglycosylation and retrotranslocation. Asterisks mark NGLY1 mutations which abolish VCP binding and ER recruitment.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (D. melanogaster) | Pngl | GenBank | FLYB:FBgn0033050 | |
| Gene (Mus musculus) | Ngly1 | GenBank | Gene ID: 59007 | |
| Gene (Homo sapiens) | NGLY1 | GenBank | Gene ID: 55768 | |
| Strain, strain background (Escherichia coli) | TOP10. Genotype: F- mcrA Δ(mrr-hsdRMS-mcrBC) φ80lacZΔM15 ΔlacX74 recA1 araD139 Δ(araleu)7697 galU galK rpsL (StrR) endA1 nupG | Life Technologies | C404010 | Chemically Competent Cells |
| Genetic reagent (D. melanogaster) | Mef2-GAL4 | Bloomington Drosophila Stock Center | BDSC: 27390; FLYB: FBti0115746 | FlyBase symbol: P{GAL4-Mef2.R}3 |
| Genetic reagent (D. melanogaster) | UAS-dpp-GFP | Bloomington Drosophila Stock Center | BDSC: 53716; FLYB: FBti0157003 | FlyBase symbol: P{UAS-dpp.GFP.T}3 |
| Genetic reagent (D. melanogaster) | dpp-GAL4 | Bloomington Drosophila Stock Center | BDSC: 1553; FLYB: FBti0002123 | FlyBase symbol: P{GAL4-dpp.blk1}40C.6 |
| Genetic reagent (D. melanogaster) | Mi{[MIC]}dppMI03752 | Bloomington Drosophila Stock Center | BDSC: 36399; FLYB: FBti0145223 | FlyBase symbol: Mi{MIC}dppMI03752 |
| Genetic reagent (D. melanogaster) | PBac{[RB]} | Exelixis at Harvard Medical School | FLYB:e00178 FBti0046265 | FlyBase symbol: PBac{RB}e00178 |
| Genetic reagent (D. melanogaster) | Pnglex14 | Funakoshi et al., 2010 | FLYB: FBal0244826 | FlyBase symbol: Pnglex14 |
| Genetic reagent (D. melanogaster) | UAS-PnglRNAiKK101641 | Vienna Drosophila Resource Center | VDRC: v103607 FLYB: FBst0475465 | FlyBase symbol: P{KK101641}VIE-260B |
| Genetic reagent (D. melanogaster) | UAS-attB-NGLY1WT-VK31 | Galeone et al., 2017 | Fly strain carrying human UAS-cDNA of wild-type NGLY1 | |
| Genetic reagent (D. melanogaster) | UAS-attB-NGLY1N41P-VK31 | This paper | Fly strain carrying human UAS-cDNA of NGLY1 with mutation in VCP binding site | |
| Genetic reagent (D. melanogaster) | UAS-attB-NGLY1F70A/G80A-VK31 | This paper | Fly strain carrying human UAS-cDNA of NGLY1 with mutation in VCP binding site | |
| Genetic reagent (D. melanogaster) | dppHA | This paper | Fly strain carrying endogenous HA-tag of dpp active domain | |
| Genetic reagent (D. melanogaster) | dppHA-3NQ | This paper | Fly strain carrying endogenous HA-tag of dpp active domain and N to Q mutations in N-glycosylation sites | |
| Strain, strain background Mus musculus | Ngly1em4Lutzy kept on a C57BL/6J background | The Jackson Laboratory | Stock #027060 | Mouse strain carrying an 11 bp deletion in exon 8 of the mouse Ngly1 |
| Cell line (Mus musculus) | Mouse Embryonic Fibroblast (MEF) cells | Huang et al., 2015 | Established from C57BL/6J background | |
| Cell line (Mus musculus) | 3T3-L1 cells | American Type Culture Collection | ATCC, CL-173 | From Dr. Sandhya Thomas |
| Recombinant DNA reagent | Human NGLY1 cDNA in pCMV6-AC (plasmid) | OriGene | clone SC320763 | Human untagged cloning vector |
| Recombinant DNA reagent | pIRES-DsRed2-Express2 | Clontech | Cat# 632540 | Expressing vector backbone construct |
| Transfected construct (Homo sapiens) | pIRES-NGLY1WT-V5-His6-DsRED | This paper | transfected NGLY1 construct (human) | |
| Transfected construct (Homo sapiens) | pIRES-NGLY1N41P-V5-His6-DsRED | This paper | transfected NGLY1 construct (human) | |
| Transfected construct (Homo sapiens) | pIRES-NGLY1 G79A/F80A-V5-His6-DsRED | This paper | transfected NGLY1 construct (human) | |
| Transfected construct (Homo sapiens) | pCS2+Bmp4HA-Myc | Gift from Dr. Jan Christian | mouse BMP4 protein (NP_031580) | Double-tagged mouse BMP4 construct |
| Transfected construct (Homo sapiens) | pCS2+Bmp4HA-Myc-4NQ | This paper | mouse BMP4 protein (NP_031580) | Double-tagged mouse BMP4 construct with N to Q mutations in N-glycosylation sites |
| Transfected construct (Homo sapiens) | pUAST-attB-NGLY1WT | This paper | Fly expressing vector carrying human UAS-cDNA of wild-type NGLY1 | |
| Transfected construct (Homo sapiens) | pUAST-attB-NGLY N41P | This paper | Fly expressing vector carrying human UAS-cDNA of wild-type NGLY1 with mutation in VCP binding site | |
| Transfected construct (Homo sapiens) | pUAST-attB-NGLY F70A/G80A | This paper | Fly expressing vector carrying human UAS-cDNA of wild-type NGLY1 with mutation in VCP binding site | |
| Antibody | anti-HA (Rat monoclonal) | Roche | Cat# 11867423001 RRID:AB_390918 | IF (1:250) |
| Antibody | anti- pSMAD3 (Rabbit monoclonal) | Abcam | Cat# ab52903 RRID:AB_882596 | IF (1:250) |
| Antibody | anti-NGLY1 (Rabbit polyclonal) | Sigma-Aldrich | Cat# HPA036825 RRID:AB_10672231 | IF (1:200) |
| Antibody | anti-GFP (Mouse monoclonal) | Sigma-Aldrich | Cat# G1546 RRID:AB_1079024 | WB (1:1000) |
| Antibody | anti-pSMAD1/5 (Rabbit monoclonal) | Cell Signaling | Cat# 9516 RRID:AB_491015 | IF (1:500) WB (1:500) |
| Antibody | anti-HA (Mouse monoclonal) | Sigma-Aldrich | Cat# SAB2702217 RRID:AB_2750919 | IF(1:500) WB (1:1000) |
| Antibody | anti-myc (Mouse monoclonal) | Cell Signaling | Cat# 2276 RRID:AB_331783 | WB (1:500) |
| Antibody | Anti-FK1(Mouse monoclonal) | Sigma-Aldrich | Cat# 04–262 RRID:AB_612094 | WB (1:500) |
| Antibody | anti-KDEL (Mouse monoclonal) | Santa Cruz Biotechnology | Cat# sc-58774 RRID:AB_784161 | WB (1:500) |
| Antibody | anti-VCP (Mouse Polyclonal) | Santa Cruz Biotechnology | Cat# sc-20799 RRID:AB_793930 | WB (1:500) |
| Antibody | anti-IRE1α (Rabbit monoclonal) | Cell Signaling | Cat# 3294 RRID:AB_823545 | WB (1:1000) |
| Antibody | anti-pIRE1α (Rabbit polyclonal) | Novus Biologicals | Cat# 100–2323 RRID:AB_10145203 | WB (1:1000) |
| Antibody | anti-OS-9 (Rabbit monoclonal) | Abcam | Cat# ab109510 RRID:AB_2848681 | WB (1:1000) |
| Antibody | anti-V5 (Mouse monoclonal) | FUJIFILM Wako Pure Chemical Corporation | Cat# 4548995010711 | WB (1:1000) |
| Sequence-based reagent | hNG1-N41P-for | This paper | PCR primers | CTCACCTATGCTGACCCCATCCTCAGAAACCC |
| Sequence-based reagent | hNG1-N41P-rev | This paper | PCR primers | GGGTTTCTGAGGATGGGGTCAGCATAGGTGA |
| Sequence-based reagent | hNG1-G79F80-for | This paper | PCR primers | GTTTATTTGAAATGGCCGCTGAAGAGGGAGAAAC |
| Sequence-based reagent | hNG1-G79F80-rev | This paper | PCR primers | GTTTCTCCCTCTTCAGCGGCCATTTCAAATAAAC |
| Sequence-based reagent | pIRES-EcoRI-huNGLY1-F | This paper | PCR primers | CTCAAGCTTCGAATTCTCAAGCATGGCGGCGGCG |
| Sequence-based reagent | v5His6-tga-SalI-pIRES-R | This paper | PCR primers | CCGCGGTACCGTCGACTCAATGGTGATGGTGGTGATGCGTAGAATCGAGACCGAG |
| Sequence-based reagent | NGLY1-N41P-F | This paper | PCR primers | ATTAGGGTTTCTGAGGATGGGGTCAGCATAGGTGAGCAGC |
| Sequence-based reagent | NGLY1-N41P-R | This paper | PCR primers | GCTGCTCACCTATGCTGACCCCATCCTCAGAAACCCTAAT |
| Sequence-based reagent | NGLY1-G79A_F80A-F | This paper | PCR primers | GATGAGATGTGTTTCTCCCTCTTCAGCGGCCATTTCAAATAAACATTCAACAGC |
| Sequence-based reagent | NGLY1-G79A_F80A-R | This paper | PCR primers | GCTGTTGAATGTTTATTTGAAATGGCCGCTGAAGAGGGAGAAACACATCTCATC |
| Sequence-based reagent | m-Psmb1-F | Yang et al., 2018 | PCR primers | CCTTCAACGGAGGTACTGTATTG |
| Sequence-based reagent | m-Psmb1-R | Yang et al., 2018 | PCR primers | GGGCTATCTCGGGTATGAATTG |
| Sequence-based reagent | m-Psmb4-F | Yang et al., 2018 | PCR primers | CGAGTCAACGACAGCACTAT |
| Sequence-based reagent | m-Psmb4-R | Yang et al., 2018 | PCR primers | ATCTCCCAACAGCTCTTCATC |
| Sequence-based reagent | m-Psma7-F | Yang et al., 2018 | PCR primers | CGAGTCTGAAGCAGCGTTAT |
| Sequence-based reagent | m-Psma7-R | Yang et al., 2018 | PCR primers | AGTCTGATAGAGTCTGGGAGTG |
| Peptide, recombinant protein | Recombinant Human BMP-4 Protein | R and D Systems | Cat# 314 BP | |
| Commercial assay or kit | In-Fusion HD Cloning | TaKaRa | Cat# 638920 | |
| Commercial assay or kit | Site-Directed Mutagenesis Kit | Agilent Technologies | Cat# 200523 | |
| Commercial assay or kit | ENDEXTTechnology, Protein Research Kit (H) | Cell Free Science | CFS-PRK-H16 | |
| Commercial assay or kit | HA-tag IP/Co-IP kit | PIERCE | Cat# 26180 | |
| Chemical compound, drug | NMS-873 | Sigma-Aldrich | Cat# 5310880001 | |
| Chemical compound, drug | Bortezomib | Cayman Chemical | Cat# 10008822 | |
| Software, algorithm | GraphPad Prism | Prism | RRID:SCR_002798 |





