The Wg and Dpp morphogens regulate gene expression by modulating the frequency of transcriptional bursts
Figures
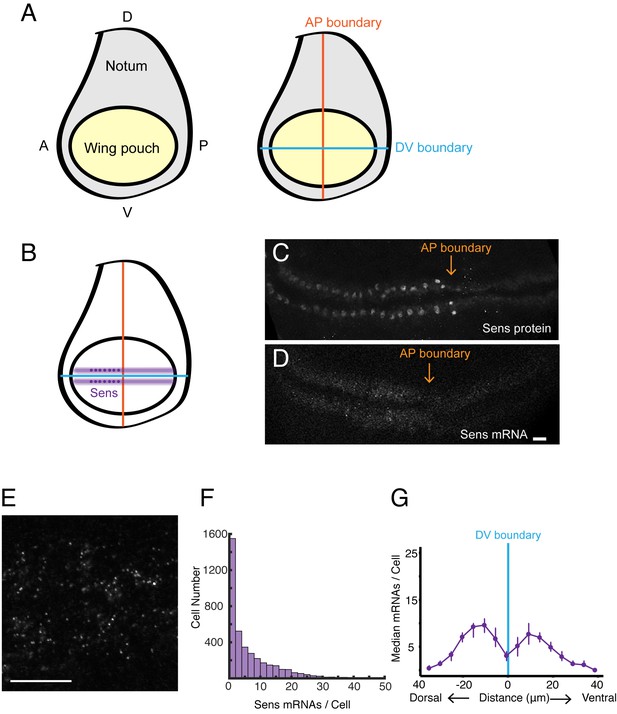
smFISH analysis of sfGFP-sens mRNA levels in wing imaginal discs.
(A) Schematic of a wing disc outlining different regional domains, and the positions of boundaries between Dorsal (D) - Ventral (V) and Anterior (A) - Posterior (P) compartments of the disc. Each wing disc is composed of roughly 50,000 cells organized in a pseudostratified epithelium. (B) Schematized expression pattern for Sens inside the wing pouch centered around the DV boundary. Sens is also expressed in clusters of cells in the notum, which are not shown. (C-E) Confocal sections of wing discs expressing sfGFP-Sens. (C) sfGFP-Sens protein fluorescence. (D) sfGFP-Sens mRNAs as visualized by smFISH using sfGFP probes. Scale bar = 10 μm. (E) Higher magnification of sfGFP-Sens mRNAs as visualized by smFISH using sfGFP probes. Scale bar = 10 μm. (F) Distribution of wing disc cells as a function of the number of Sens mRNA molecules per cell. (G) Sens mRNA number as a function of cell distance from the DV boundary displays a bimodal expression pattern for Sens. Cells were binned according to the shortest path length from its centroid to the DV boundary, and whether they were dorsal or ventral compartment cells. Median mRNA number/cell for each bin is plotted with 95% bootstrapped confidence intervals.
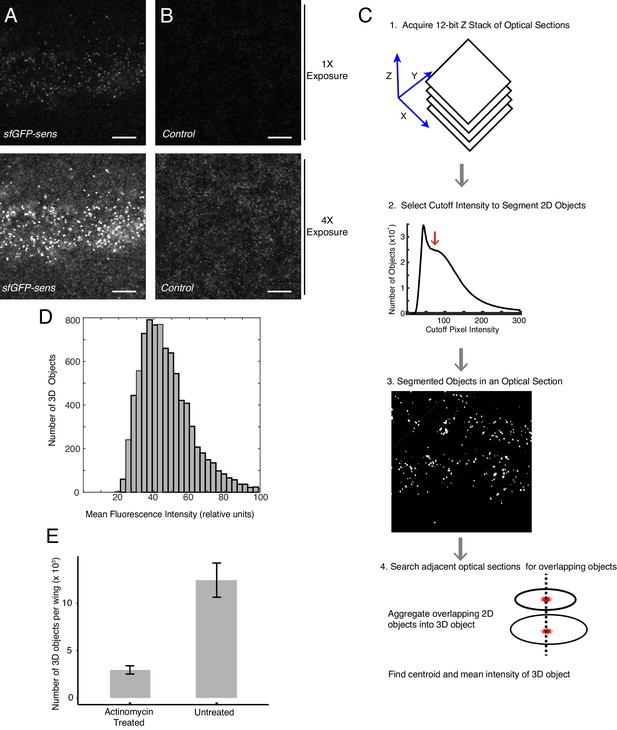
Development of smFISH imaging and analysis.
(A,B) Representative optical sections of wing discs probed for sfGFP mRNAs. Upper panels show 1x exposure of fluorescence from optical sections. Lower panels show same sections with 4x overexposure of fluorescence. Scale bars = 5 μm. (A) A disc from an animal with two copies of the sfGFP-sens transgene. (B) A control disc from an animal without the sfGFP-sens transgene. (C) Imaging and analysis pipeline to quantify mRNAs as 3D fluorescent objects. 1. A stack of 35 optical sections is acquired per sample. 2. RNA spots are segmented by using a pixel intensity value as a cutoff, above which lie true RNA fluorescent spots, and below which lies the background. To select the optimal cutoff for each image stack, a broad range of potential cutoff values are systematically tested, and the number of segmented objects (object >7 contiguous pixels) is counted for each cutoff tested. Object number plateaus over a range of cutoff values (red arrow). This plateau corresponds to the cutoff levels that correctly identify RNA spots (<5% error from ground truth). 3. Such a cutoff value is then applied as a threshold to identify 2D objects in each section. 4. 2D objects must satisfy two criteria in order to be counted as an RNA spot. One, they must have a corresponding object at least one neighboring z-plane within a diffraction limited radius of 4 pixels. Two, they must be larger (contain more pixels) than corresponding objects in neighboring z-planes. This criterion prevents RNA spots from being counted in multiple z-planes. Centroid position and 3D object pixel intensity are then recorded. (D) Distribution of mean fluorescence intensity for all identified 3D fluorescent objects from one wing disc expressing sfGFP-Sens mRNAs. (E) Average number of 3D fluorescent objects per imaged wing disc after a 30 min treatment of the discs in actinomycin-D. Untreated discs were incubated in medium for an identical period of time, and all discs were fixed and imaged for sfGFP-Sens mRNAs. Error bars are SEM.
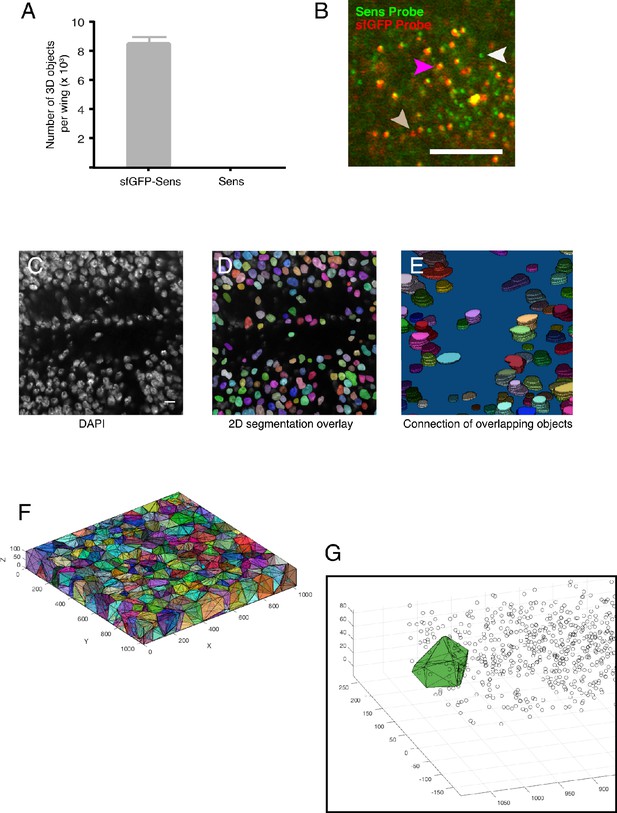
Determination of false-positive and false-negative rates for smFISH.
(A) Wing discs were imaged and scored for 3D fluorescent objects using the sfGFP probe set. Discs were either from animals with two copies of the sfGFP-sens transgene and two copies of the endogenous sensE1 gene, or from animals with just two copies of the endogenous sensE1 gene. Error bars are SEM. (B) A representative optical section taken from a wing disc expressing the sfGFP-sens transgene and endogenous sensE1 gene. The disc was probed for sfGFP (red) and Sens (green) RNA using independent probe sets. Spots that fluoresce both green and red are presumptive sfGFP-Sens mRNAs that have annealed to both probe sets (purple arrow). Spots that only fluoresce with the Sens probe set (white arrow) are presumptive Sens mRNAs that are generated from the endogenous sens gene. Although these sens alleles are mutant for protein output, they still produce mRNA. The occasional spot (beige arrow) that only fluoresces with the sfGFP probe set are presumptive sfGFP-Sens mRNAs that failed to hybridize with the Sens probe set. These are false-negatives. Scale bar = 5 μm. (C-E) Pipeline for 3D segmentation of cell nuclei. (C) An optical section showing DAPI fluorescence. (D) 2D segmentation of this image. (E) Five contiguous z-sections of segmented nuclei are colored and viewed laterally. Note the three-dimensional ‘stack of pancakes’ nature of the nuclear objects in the wing disc 3D rendering. (F) 3D Voronoi tessellation of an image stack of wing disc cells. The centroids of the 3D nuclei (shown as circles) were used to tessellate the image stack, creating virtual cells. Cells are represented with different colors. Numbers in the x-y plane refer to pixel positions in the 1024 × 1024 sections. Please see the Materials and methods for a detailed description of tessellation and its meaning. (G) An image stack showing the centroid positions of 3D mRNA objects as circles. One tessellated cell (green) is superimposed to show the mRNA objects that reside in space occupied by the tessellated cell. These mRNAs would be assigned to that particular cell. Shown is one stripe of sfGFP-Sens expressing cells on one side of the DV boundary marked by pixel position 0.
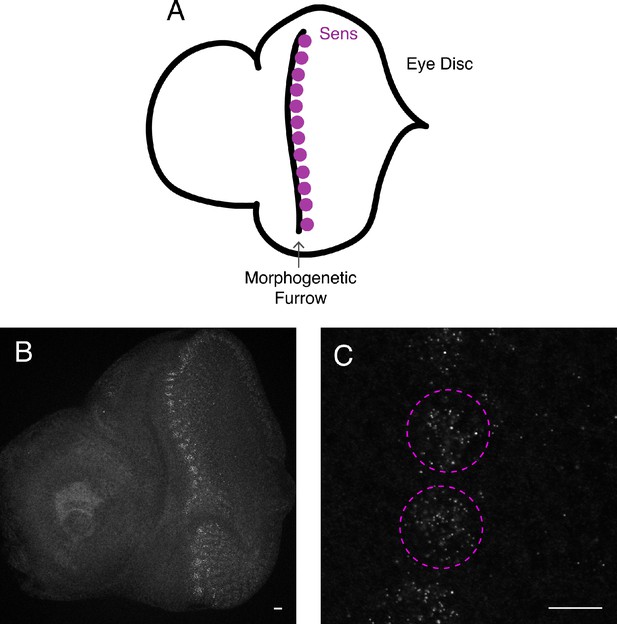
smFISH imaging of the eye imaginal disc.
(A) Schematic of the eye antennal disc complex showing the approximate location of cells that express the sens gene. Anterior is to the left. (B,C) Optical sections through a representative eye antennal disc complex probed for sfGFP-Sens mRNAs by smFISH. Anterior is to the left. (B) Low magnification shows a vertical stripe of positive fluorescence that oscillates between clusters of high and low mRNA abundance. This is the pattern that has been reported for cells in the morphogenetic furrow (Nolo et al., 2000). Scale bar = 5 μm. (C) Higher magnification of an optical section through the morphogenetic furrow showing two complete clusters of Sens-positive cells (dashed purple lines). Scale bar = 5 μm.
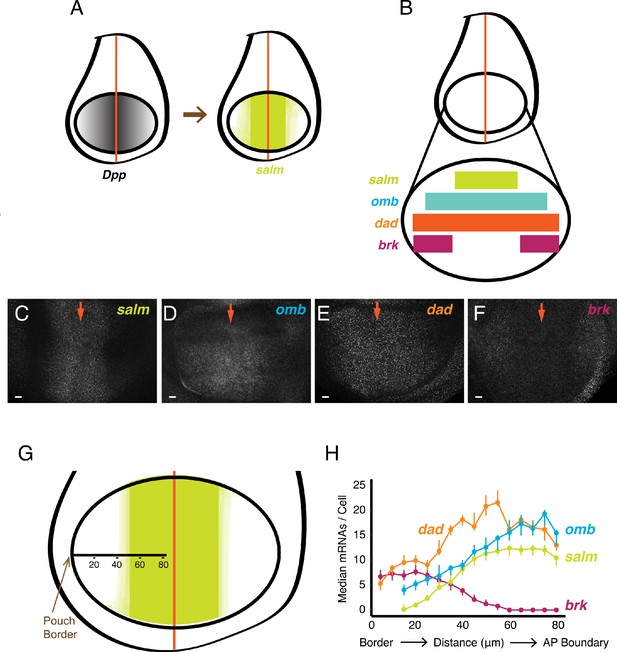
smFISH analysis of mRNA levels from Dpp-responsive genes.
(A) Schematic of wing discs highlighting the graded distribution of Dpp protein in the wing pouch, centered around the AP boundary, and the expression domain for salm, one of the targets of Dpp regulation. Not shown is Dpp localization in the notum domain of the disc. (B) Expression domains of four target genes of Dpp signaling. (C-F) Confocal sections of wing pouches probed for mRNAs synthesized from the salm (C), omb (D), dad (E), and brk (F) genes. Orange arrows mark the position of the AP boundary in each image. (G, H) mRNA number as a function of cell distance from the anterior-most border of the wing pouch. (G) A border-to-boundary axis, orthogonal to the AP boundary, is used to map cell position, along which distances are displayed in μm from the wing pouch border. (H) Cells were binned according to position along the border-to-boundary axis. Median mRNA number/cell for each bin is plotted with 95% bootstrapped confidence intervals.
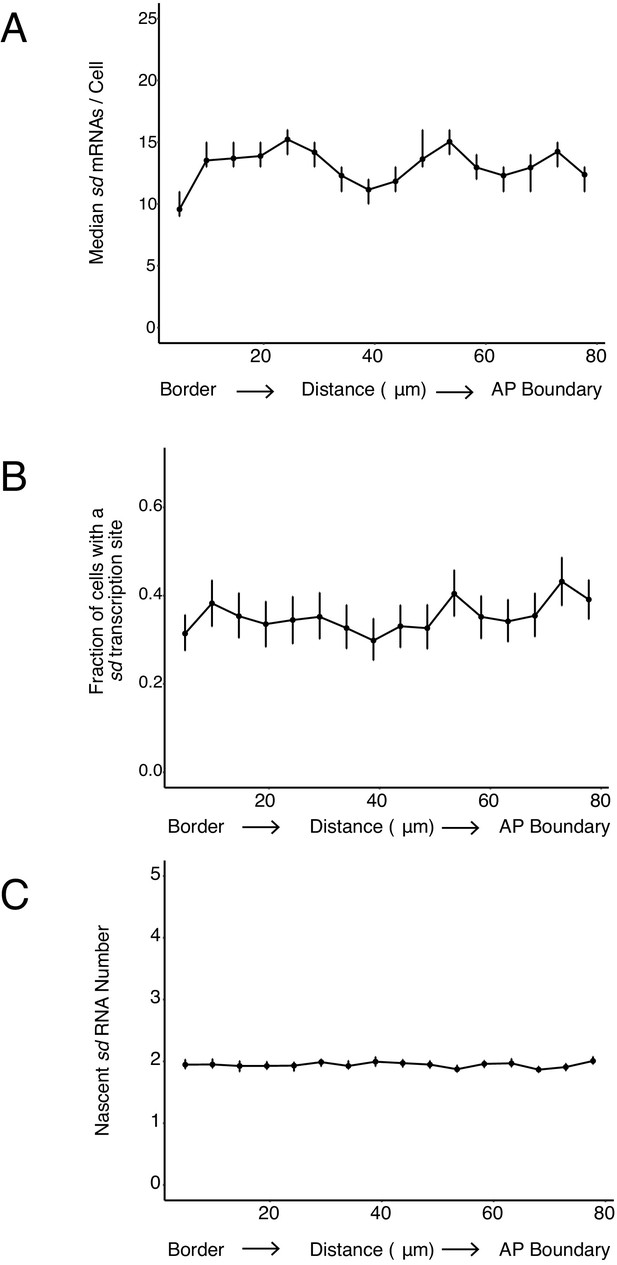
Detection of RNAs corresponding to the sd gene.
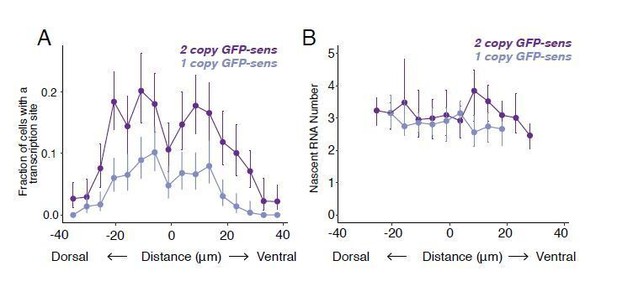
(A) sd mRNA number as a function of cell distance from the anterior-most border of the wing pouch. An axis orthogonal to the AP boundary is used to map cell position. Numbers refer to distance in μm from the wing pouch border. (B) The probability of detecting a cell with a sd transcription site does not vary with the cell's location relative to the source of morphogens. Error bars are 95% bootstrapped confidence intervals. Cells are binned according to their distance from the pouch border, and the fraction of cells in each bin with a transcription site is shown. (C) The average number of nascent RNAs in a sd transcription site does not vary with the cell's location. Error bars are bootstrapped 95% confidence intervals. Cells are binned according to their distance from the pouch border, and the average number of nascent RNAs per site in each bin is shown.
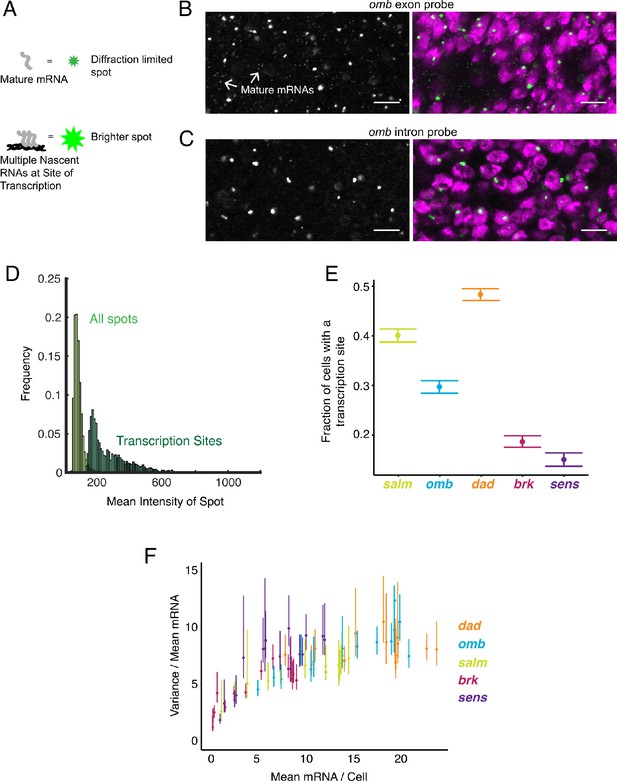
Sites of nascent transcription are detected by smFISH.
(A) Sites of nascent transcription can fluoresce more brightly than single mRNA molecules due to multiple nascent transcripts localized to one gene locus. (B) Probes recognizing an omb exon generate many small dim spots and a few large bright spots. Right image shows the merge of probe and DAPI fluorescence. The bright spots are associated with nuclei whereas most dim spots are not. (C) Probes recognizing an omb intron only generate large bright spots that are associated with nuclei. Scale bars = 5 μm. (D) Frequency distribution of intensity for all spots identified in a wing disc probed for sens RNAs. Using a threshold of twice the median spot intensity, all single mRNA spots were filtered out, leaving only spots that are associated with transcription sites. The frequency distribution for this class of spots is shown. (E) Transcription sites are assigned to cells. For each cell that contains one or more mRNA molecules, it is scored for whether it also has one or more transcription sites. The average fraction of all such cells with a transcription site is shown for each gene. Error bars represent 95% confidence intervals. (F) The ratio of the variance of mRNAs/cell to its mean, as a function of the mean, for all genes. This ratio is larger than one, irrespective of the mRNA number for binned sub-populations of cells and the gene. Error bars represent 95% confidence intervals.
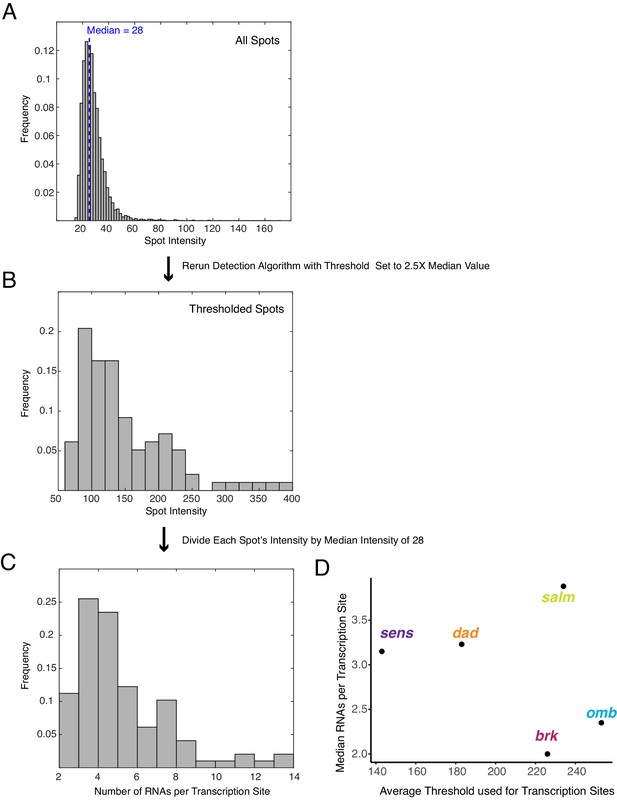
Detection of transcription sites and their quantification.
(A) A representative frequency distribution of fluorescence intensity for 3D spots identified in one wing disc expressing sfGFP-sens. The median intensity is 28 units. (B) The same wing disc was reanalyzed for 3D spots but using a threshold of 70 units as a cutoff, below which spots are not counted. (C) The fluorescence intensity of each 3D spot in B is divided by the median intensity of 28 units to provide a normalized number of RNAs that are localized to that 3D spot. This is not an actual number of RNA molecules but the output from partially transcribed RNAs annealing to a variable number of probes depending on the composition of binding sites in the RNA composite. (D) The mean threshold used for transcription site identification for each data set plotted against the median normalized RNA molecules per transcription site for all transcription sites in that data set.
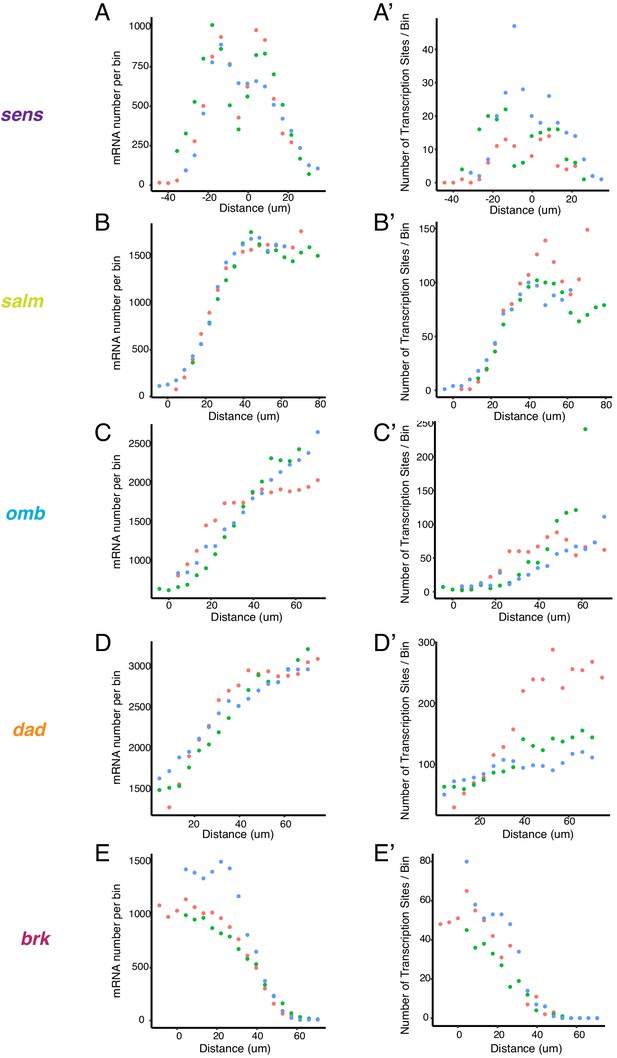
Transcription sites and mRNA patterns in unsegmented images.
(A,B,C,D,E) Three discs were analyzed independently (green, blue, orange dots) for spots that corresponded to the mRNAs from sens (A), salm (B), omb (C), dad (D) and brk (E). Spots were binned according to their positions along the AP or DV axes, and total mRNAs per bin were plotted. Note the strong concordance of independent discs for all genes. (A',B',C',D',E') The same three discs were analyzed independently (green, blue, orange dots) for spots that corresponded to transcription sites from sens (A), salm (B), omb (C), dad (D) and brk (E). Spots were binned according to their positions along the AP or DV axes, and total transcription sites per bin were plotted. Note the strong concordance of independent discs for all genes.
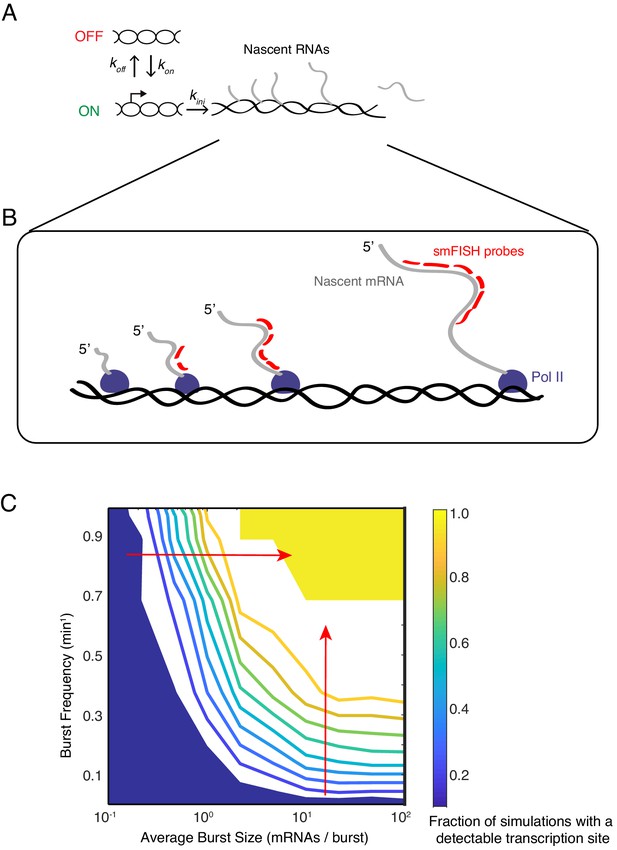
Modeling transcription sites using bursting dynamics.
(A) Model framework showing the three rate parameters affecting transcription initiation. Two parameters affect the promoter state, while the third parameter only affects how many initiation events occur when the promoter is ON. (B) Pol II molecules in elongation mode are distributed along the transcription unit. If Pol II is upstream of the probe binding sites, the nascent transcript will not be detected. If Pol II is downstream, the nascent transcript will be detected as 100% signal. If Pol II is transcribing within the binding sites, the nascent transcript will be detected as a partial signal. These three different scenarios are explicitly accounted for in our mathematical model. For example in the simulation result shown here, four Pol II's are situated such that a total of 12 virtual probe-binding sites are present. Since each mRNA has six binding sites, it means that this simulated transcription site has 12/6 or 2 units of normalized signal. Applying our filter cutoff for identifying a transcription site as two or more units, this simulated site would be scored as a positive. (C) The phase diagram of transcription site detection as a function of burst size and frequency in the model. Both burst size and frequency impact the likelihood of detecting a transcription site. When burst size increases at low burst frequency, the likelihood of detecting a transcription site remains fairly constant. When burst size increases at high burst frequency (horizontal red arrow), the likelihood of detecting a transcription site is ultrasensitive to burst size. Likewise, when burst frequency increases at low burst size, the likelihood of detecting a transcription site remains fairly constant. When burst frequency increases at high burst size (vertical red arrow), the likelihood of detecting a transcription site is ultrasensitive to burst size. The phase diagram makes manifest that a range of combinations of burst frequency and size could explain observed transcription site frequency data.
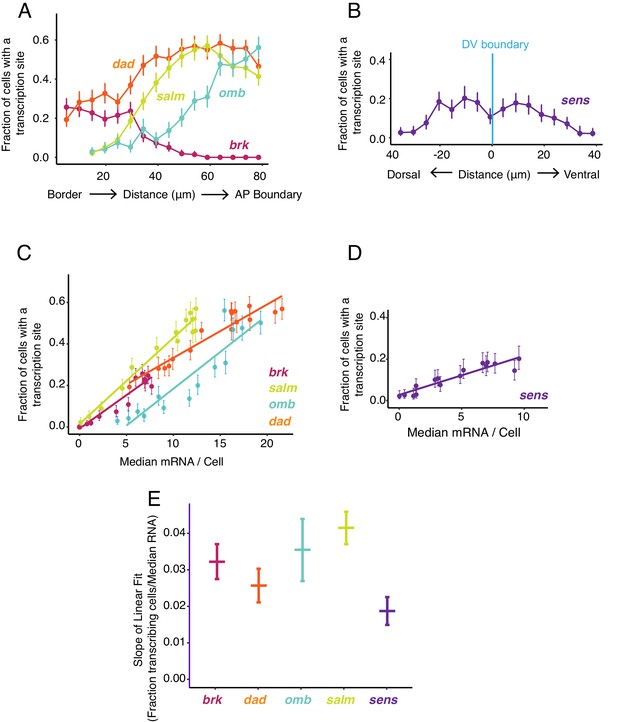
Transcription site detection correlates with mRNA number.
(A,B) The probability of detecting a cell with a transcription site varies with the cell's location relative to the source of morphogen. Error bars are 95% bootstrapped confidence intervals. (A) Cells are binned according to their distance from the pouch border, and the fraction of cells in each bin with a transcription site are shown for each Dpp-responsive gene. (B) Cells are binned according to their distance from the DV boundary, and the fraction of cells in each bin with a transcription site is shown for the sens gene. (C,D) The probability of detecting a cell with a transcription site varies linearly with the number of mRNA molecules in the cell. Fitted lines are from linear regression. Error bars are 95% confidence intervals. (C) Cells are binned according to the number of mRNAs they contain, and the fraction of cells in each bin with a transcription site are shown for each Dpp-responsive gene. (D) Cells are binned according to the number of mRNAs they contain, and the fraction of cells in each bin with a transcription site is shown for the sens gene. (E) Linear regression analysis was performed on samples from C and D, shown is the slope with a parametric 95% confidence interval.
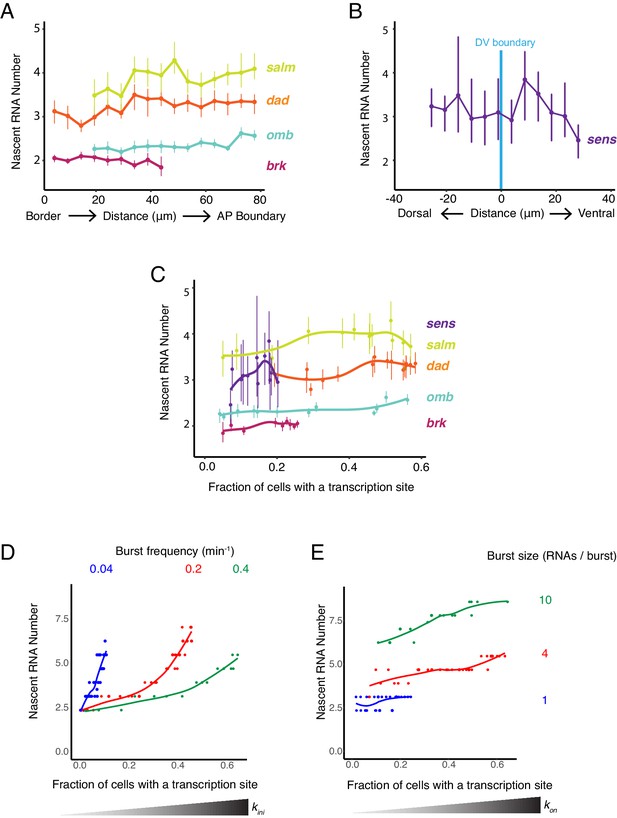
Burst frequency is regulated by Dpp and Wg.
(A,B) The average number of nascent RNAs in a transcription site does not vary with the cell's location relative to the source of morphogen. Error bars are bootstrapped 95% confidence intervals. (A) Cells are binned according to their distance from the pouch border, and the average number of nascent RNAs per site in each bin are shown for each Dpp-responsive gene. (B) Cells are binned according to their distance from the DV boundary, and the average number of nascent RNAs per site in each bin is shown for the sens gene. (C) The average number of nascent RNAs in a transcription site does not vary with the probability of detecting a cell with a transcription site. Error bars are 95% confidence intervals. (D,E) Model predictions of the relationship between average number of nascent RNAs in a transcription site and the probability of detecting a site for the dad gene. (D) Simulations are performed where the rate parameter kini has been systematically varied so as to modulate burst size alone. Resulting values for nascent RNA number and fraction of cells with a site are shown. Each datapoint is the average of 1000 simulations. Simulations are repeated for three different values of kon to specifically set the burst frequency to 0.04, 0.2 and 0.4 min−1. (E) Simulations are performed where the rate parameter kon has been systematically varied so that burst frequency alone is variable. Resulting values for nascent RNA number and fraction of cells with a site are shown. Each datapoint is the average of 1000 simulations. Simulations are repeated for three different values of kini to specifically set the burst size to 1, 4 and 20.
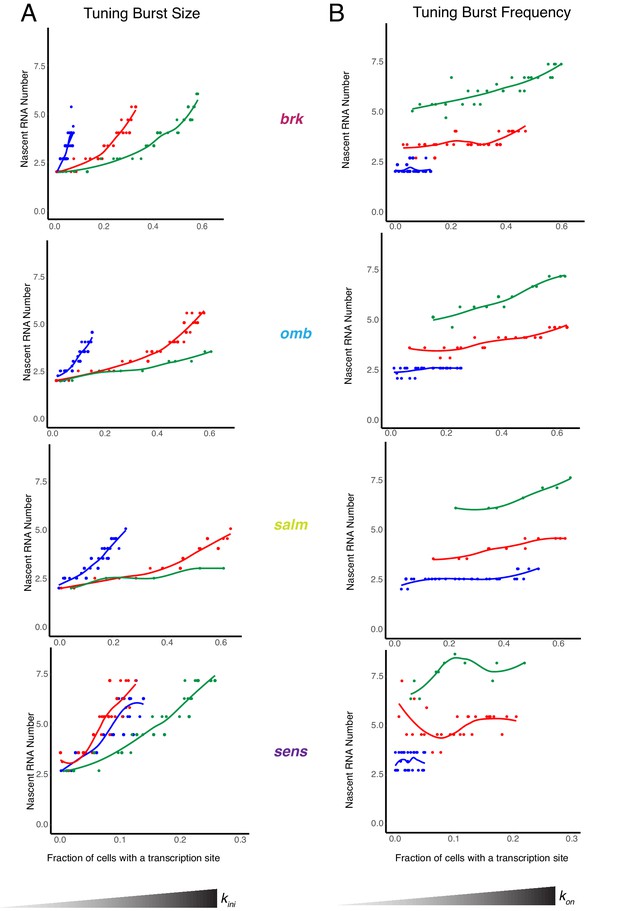
Modeling the relationship between average number of nascent RNAs in a transcription site and the probability of detecting a site for the brk, omb, salm, and sens genes.
(A) Simulations are performed where the rate parameter kini has been systematically varied so that burst size alone is variable. Resulting values for nascent RNA number and fraction of cells with a site are shown. Each datapoint is the average of 1000 simulations. Simulations are repeated for three different values of kon to specifically set the burst frequency to 0.04, 0.2 and 0.4 min-1. (B) Simulations are performed where the rate parameter kon has been systematically varied so that burst frequency alone is variable. Resulting values for nascent RNA number and fraction of cells with a site are shown. Each datapoint is the average of 1000 simulations. Simulations are repeated for three different values of kini to specifically set the burst size to 1, 4 and 10.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Drosophila melanogaster) | white1118 | BloomingtonDrosophilaStock Center | BDSC: 3605 Flybase: FBst0003605 RRID:BDSC_3605 | |
| Gene (Drosophila melanogaster) | sensE1 | Nolo et al., 2000 | Flybase: FBal0098024 | From Hugo Bellen |
| Genetic reagent (Drosophila melanogaster) | sfGFP-sens [VK37] | Venken et al., 2006. From Hugo Bellen | Pacman construct containing sens gene with N-terminal 3xFlag-TEV-StrepII-sfGFP-FlAsH fusion tag inserted at 22A3 (VK37) | |
| Genetic reagent (Drosophila melanogaster) | dad-GFP [VK37] | BloomingtonDrosophilaStock Center | BDSC: 81273 Flybase: FBti0150281 RRID:BDSC_81273 | y w; PBac{y[+mDint2] w[+mC]=Dad GFP.FLAG} inserted at 22A3 (VK37) |
| Genetic reagent (Drosophila melanogaster) | brk-GFP [VK33] | BloomingtonDrosophilaStock Center | BDSC: 38629 Flybase: FBti0147730 RRID:BDSC_38629 | w1118; PBac{y[+mDint2] w[+mC]=brk GFP.FPTB} inserted at 65B2 (VK33) |
| Sequence-based reagent | GFP hybridization oligo probes | Biosearch Technologies | Custom probe set | Set of oligos with 3' modification mdC(TEG-Amino). Sequence of all oligos is in Supplementary file 1 |
| Sequence-based reagent | sens hybridization oligo probes | Biosearch Technologies | Custom probe set | Set of oligos with 3' modification mdC(TEG-Amino). Sequence of all oligos is in Supplementary file 1 |
| Sequence-based reagent | salm hybridization oligo probes | IDT | Custom probe set | Set of oligos. Sequence of all oligos is in Supplementary file 1 |
| Sequence-based reagent | omb hybridization oligo probes | IDT | Custom probe set | Set of oligos. Sequence of all oligos is in Supplementary file 1 |
| Sequence-based reagent | sd hybridization oligo probes | IDT | Custom probe set | Set of oligos. Sequence of all oligos is in Supplementary file 1 |
| Sequence-based reagent | omb intron hybridization oligo probes | IDT | Custom probe set | Set of oligos. Sequence of all oligos is in Supplementary file 1 |
| Sequence-based reagent | omb 5' exon hybridization oligo probes | IDT | Custom probe set | Set of oligos. Sequence of all oligos is in Supplementary file 1 |
| Chemical compound, drug | NHS-ester ATTO 633 dye | Sigma | #01464 | |
| Chemical compound, drug | NHS-ester ATTO 565 dye | Sigma | #72464 | |
| Chemical compound, drug | amino-11-ddUTP | Lumiprobe | A5040 | |
| Chemical compound, drug | Paraformaldehyde (powder) | Polysciences | 00380–1 | |
| Chemical compound, drug | Triton X-100 | Sigma Aldrich | T9284-500ML | |
| Chemical compound, drug | VectaShield | Vector Labs | H-1000 | |
| Chemical compound, drug | 4′,6-diamidino-2-phenylindole (DAPI) | Life Technologies | D1306 | |
| Chemical compound, drug | salmon sperm single stranded DNA | Invitrogen | #15632 | |
| Chemical compound, drug | vanadyl ribonucleoside | New England Biolabs | #S14025 | |
| Software, algorithm | MATLAB pipeline to process raw smFISH images with no prior preprocessing | This paper | https://github.com/elifesciences-publications/smfish_pipeline | |
| Other | Graces’ Insect Medium | Sigma | #69771 | Growth medium for organ culture |
Additional files
-
Supplementary file 1
Excel file containing the sequences of all oligonucleotide probes used for smFISH experiments in this paper.
Each worksheet lists the oligos specific for a gene, as indicated. Sequences are ordered 5' - to - 3'.
- https://cdn.elifesciences.org/articles/56076/elife-56076-supp1-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/56076/elife-56076-transrepform-v2.pdf