True S-cones are concentrated in the ventral mouse retina and wired for color detection in the upper visual field
Figures
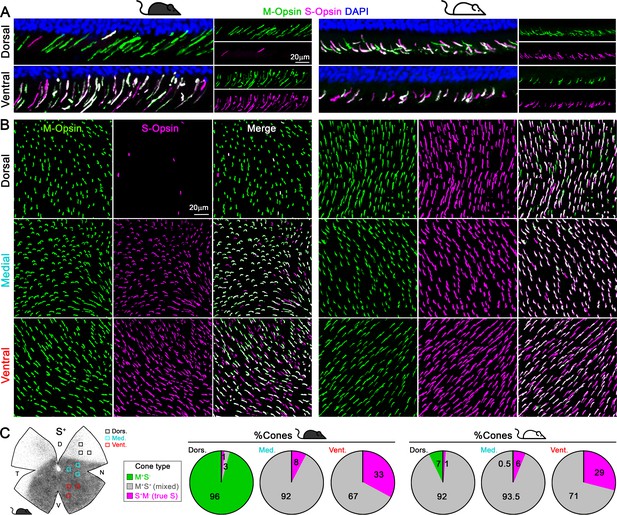
Cone outer segments across retinal areas.
Immunodetection of M and S wavelength-sensitive opsins in retinal sections (A) and flat-mount retinas (B) in two mouse strains (pigmented and albino mice, left and right columns respectively). (C) Retinal scheme of S-opsin expression used for image sampling to quantify and classify cones in three different retinal regions. Pie graphs showing the percentage of cones manually classified as M+S- (green), S+M- (true S, magenta) and M+S+ (mixed, gray) based on the opsin expression in different retinal areas from four retinas per strain. Black mouse: pigmented mouse strain (C57BL6), white mouse: albino mouse strain (CD1).
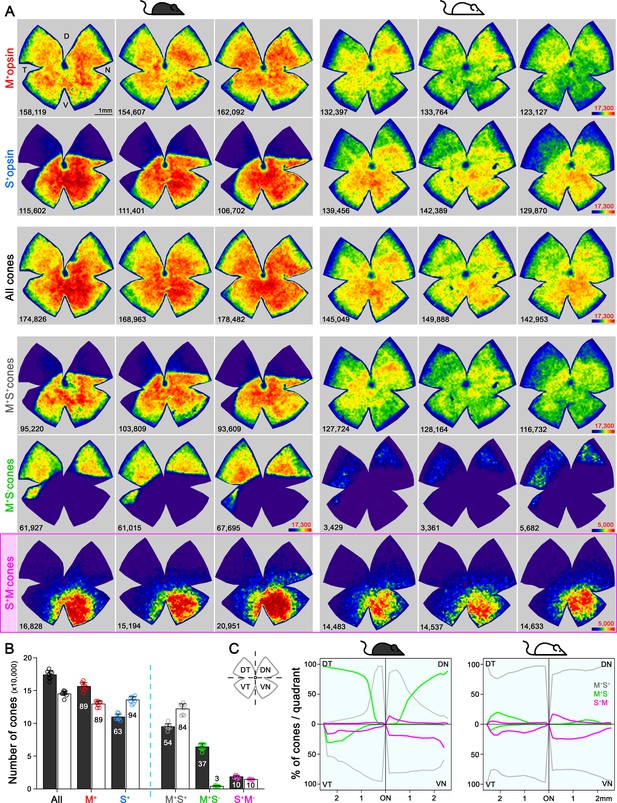
Topography and total number of different opsins (M+, S+) and cone-type populations in the whole mouse retina.
(A) Density maps depicting the distributions of different opsins expressing cones (M+ and S+) and different cone populations classified anatomically as: All, M+S+ (mixed), M+S-, S+M- (true S) cones in pigmented and albino mice (left and right side respectively). Each column shows different cone populations from the same retina and, at the bottom of each map is shown the number of quantified cones. Color scales are shown in the right panel of each row (from 0 [purple] to 17,300 [dark red] for all cone types except to 5000 cones/mm2 [dark red] for the true S-cones and M+S--cone in the albino strain). Retinal orientation depicted by D: dorsal, N: nasal, T: temporal, V: ventral. (B) Histogram showing the mean ± standard deviation of different cone subtypes for eight retinas per strain (Supplementary file 1B). The percentages of each cone subtype are indicated inside of each bar, where 100% indicates the total of the ‘all cones’ group. (C) Opsin expression profile across the different retinal quadrants (retinal scheme, DT: dorsotemporal, DN: dorsonasal, VT: ventrotemporal, VN: ventronasal). Line graphs show the spatial profile of relative opsins expression (mixed [gray], M+S- [green], true S-cones [magenta]), where the sum of these three cone populations at a given distance from the optic nerve (ON) head equals 100%. Black mouse: pigmented mouse strain, white mouse: albino mouse strain.
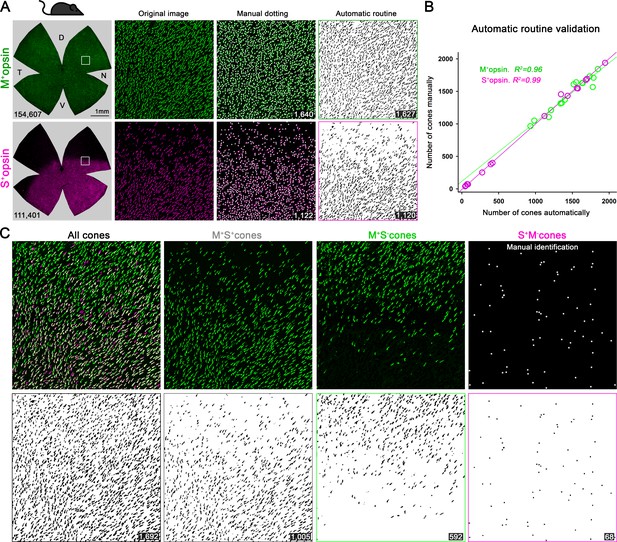
Validation of automatic routine for cone outer segment quantification.
(A) Retinal photomontages for M- and S-opsin signal in the same pigmented retina (correspond to second column in Figure 2A). The square depicts an area of interest selected (transition zone of S-opsin expression) to perform the automatic routine validation by comparing manual and automatic quantifications. The images processed by the automatic routine using ImageJ show the selection of positive objects from the corresponding original image. (B) X, Y graph showing the linear correlation (Pearson coefficient, (R2) between manual and automatic quantifications. 21,898 M+ and 13,705 S+cones were manually annotated while 21,689 M+ and 13,661 S+cones were automatically identified in 3 random images obtained from 5 retinal photomontages. (C) All, mixed, M+S-- and true S-cone populations are extracted from the original M- and S-cone images. All-cones were quantified after overlapping M- and S-signals. mixed (M+S+) cones were obtained by subtracting the background of the S-opsin image in the M-opsin one. M+S- cones for pigmented mice are obtained after subtracting the S-opsin signal to the M-opsin photomontage. Finally, M+S- cones for albino and true S-cones (S+M-), in both strains, are manually marked on the retinal photomontage (Adobe Photoshop CC). The B and W images show the processed image after quantifying automatically. At the bottom of each image is shown the number of quantified cones. Black mouse: pigmented mouse strain.
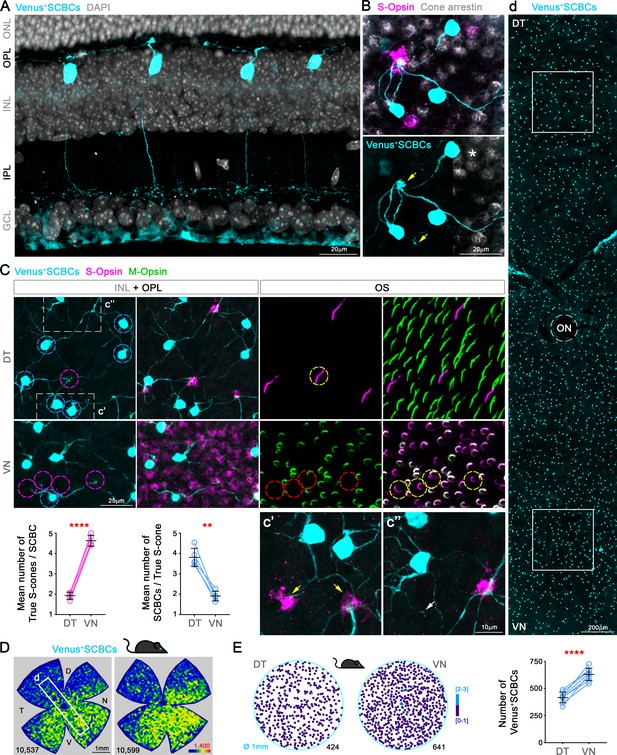
S-cone Bipolar cells (SCBCs) in Cpne9-Venus mouse retina.
(A) Retinal cross section showing the characteristic morphology of SCBCs (Behrens et al., 2016; Breuninger et al., 2011). (B) Detailed view of the selective connectivity between Venus+SCBCs and true S-cone terminals (yellow arrows). Note that SCBCs avoid contacts with cone terminals lacking S-opsin expression (M+S--cone pedicles, identified using cone arrestin), as well as a mixed cone pedicle, marked with an asterisk. In fact, on the contrary, the SCBCs prefer to develop multiple contacts to the same true S-cone pedicle. (C) Images from flat-mount retinas focused on the inner nuclear and outer plexiform layers (INL+OPL) or in the photoreceptor outer segment (OS) layer of the corresponding area. Magnifications showing divergent and convergent connectivity patterns from true S-cone pedicles in dorsal and ventral retinal domains, respectively. In the DT retina, six Venus+ SCBCs (cyan circles) contact a single true S-cone pedicle (magenta circle in DT); while one Venus+ SCBC contacts at least four true S-cone pedicles in the VN retina (magenta circles in VN), which belong to cones possessing S+M-OSs (yellow circles). Connectivity between true S-cones and SCBCs in DT and VN retina was assessed as the average number of true S-cone pedicles contacting a single SCBC per retina (magenta plot) or the average number of SCBCs contacting a single true S-cone pedicle per retina (cyan plot) (p<0.0001, p<0.01, respectively; n = 5). (c’) Detailed view of a secondary SCBC bifurcation contacting independently two true S-cone pedicles. (c”) Detailed view of a ‘blind’ SCBC process. (D) Density maps depicting the distributions of SCBCs in Cpne9-Venus mice. (d) Venus+ SCBCs along the DT-VN axis from a flat-mount retina (corresponding to the white frame in D) showing the gradual increase of SCBCs towards the VN retina where true S-cone density peaks (last row in Figure 2A). (E) Demonstration of Venus+ SCBC densities color-coded by the k-nearest neighbor algorithm according to the number of other Venus+ SCBCs found within an 18 μm radius in two circular areas of interest (DT and VN). Although, Venus+ SCBCs exhibit a sparse density without forming clusters (circular maps), they were significantly denser in VN retina (p<0.0001; n = 8).
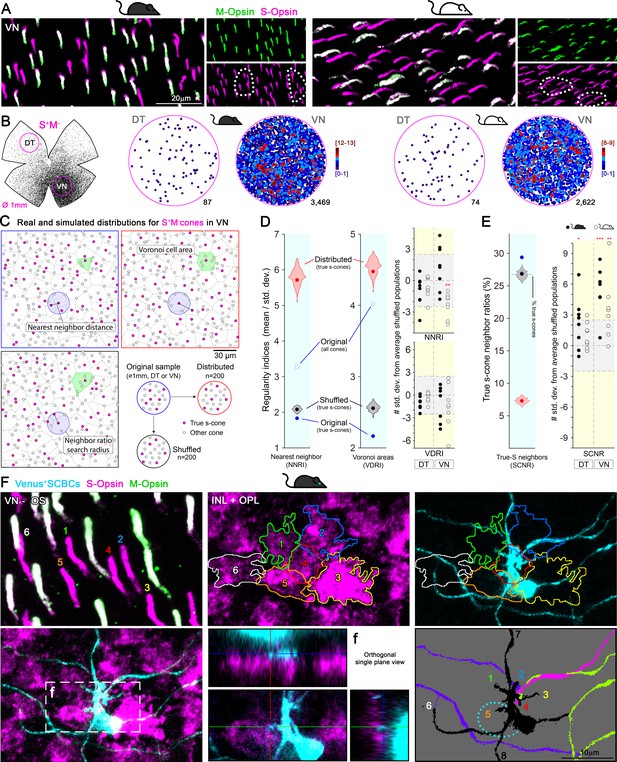
Clustering of true S-cones in the ventronasal (VN) retina.
(A) Retinal magnifications from flat-mount retinas demonstrating grouping of true S-cones in the VN area, where true S-cone density peaks. White dashed lines depict independent groups of true S-cones that are not commingled with mixed cones (M+S+, white outer segments in the merged image). (B) Retinal scheme of true S-cones used for selecting two circular areas of interest along the dorsotemporal-ventronasal (DT-VN) axis. Circular maps demonstrate true S-cone clustering in these regions. True S-cone locations are color-coded by the k-nearest neighbor algorithm according to the number of other true S-cones found within an 18 μm radius. (C–E) Analytical comparisons of DT and VN populations of true S-cones to their simulated alternatives. (C) Example real and simulated true S-cone populations and their quantification. Images depict true S-cone locations (magenta dots) and boundaries of their Voronoi cells (dashed lines) from original and example simulated (‘distributed’, ‘shuffled’) cone populations. Gray dots indicate the locations of other cone types. Observed cone locations were used for all simulated populations; only their cone identities were changed. The annotated features are examples of those measurements used in the calculations presented in D-E. (D) Comparison of sample regularity indices for one albino VN retinal sample to violin plots of those values observed for n = 200 simulated cone populations. Note that average regularity indices for true S-cones were lower than that of shuffled populations, whereas those values lay between shuffled and distributed populations when all cones were considered. Plots on the right show values for all actual retinal samples normalized using the mean and standard deviations of their simulated ‘shuffled’ counterparts. The y-axis range corresponding to ±2.5 standard deviations from the mean (i.e., that containing ~99% of shuffled samples) is highlighted in gray. (E) Comparison of the real average SCNR for the example in C-D to those values for its simulated counterparts. Note that the average SCNR for all cones in this sample was equal to that predicted by random chance (i.e., the ratio of true S-cones to all cones), which in turn was equal to the average for true S-cones for shuffled samples. In contrast, the real true S-cone SCNR was higher. Plot on the right shows true S-cone SCNR values for all samples, normalized as described for D. (F) Convergent connectivity from a true S-cone cluster to a single SCBC in the VN retina. Images of a true S-cone cluster, in a flat-mount retina, focused on the photoreceptor outer segment layer and the inner nuclear-outer plexiform layers (INL+OPL). The upper left panel show the numerical and colored identification of each true S-outer segment in the cluster (note that the number positions indicate the locations where outer segments contact the photoreceptor inner segment). Each true S-cone pedicle belonging to this cluster is outlined and color coded (middle upper panel) and are overlaid upon the SCBC dendritic profile (right upper panel). To identify synaptic contacts between the SCBC and the cone pedicles (maximum intensity projection -excluding the SCBC soma- shown in lower left panel), we acquired orthogonal single plane views zooming into putative dendritic tips. An example for the contact with cone #5 is shown in lower middle panel, corresponding to the box area in lower left panel (f). The lower right panel shows dendritic endings of this SBCB (black) contacting the marked cones (#1–6). It also contacts two additional cones outside of the field of view (#7,8). Dashed line depicts the soma of the SCBC. Dendrites from other SCBCs are color coded for differentiation.
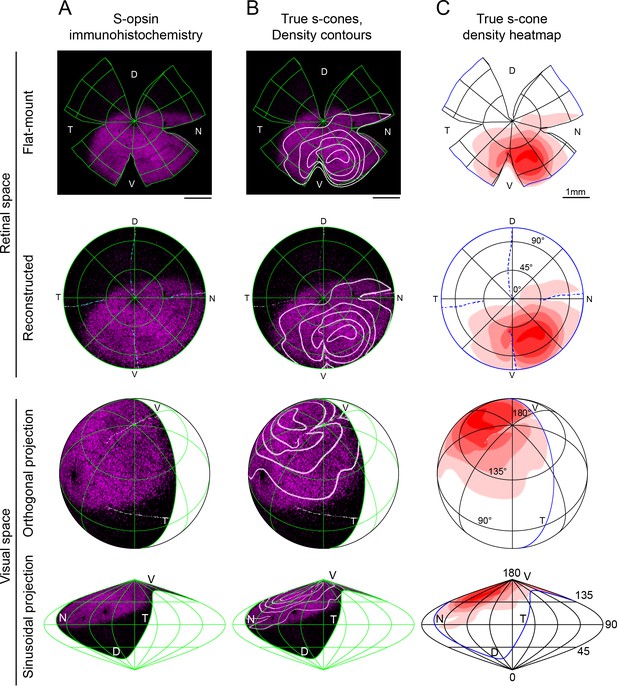
Reconstruction and mapping of true S-cone densities into visual space.
Representative left eye from a 3-month-old pigmented mouse (C57). (A) S-opsin antibody labeling; (B) true s-cone density contour lines separated by quintiles overlaid onto s-opsin labeling; (C) quintile heatmap contours of true s-cone density. The top two rows demonstrate the flat-mount retina with marks for edges and relaxing cuts, followed by its reconstruction into uncut retinal space with lines of latitude and longitude that have been projected onto the flat-mount. The bottom two rows show the reconstructed retina inverted into visual space using orthogonal and sinusoidal projections. For these views, eye orientation angles for elevation and azimuth of 22° and 64°, respectively, have been used as in Sterratt et al., 2013. For orthogonal projections, the globe has been rotated forward by 50° to emphasize the relationship of true S-cone densities to the upper pole of the visual field. S-opsin labeling is restricted to the upper visual field, but true S-cones are concentrated toward its lateral edges.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mus musculus, male) | C57BL/6J mouse strain | Jackson Laboratory | Cat#000664, RRID:IMSR_JAX:000664 | Pigmented mouse inbred strain |
| Strain, strain background (Mus musculus, male) | Crl:CD-1(ICR) mouse strain | Charles River | Cat#022, RRID:IMSR_CRL:022 | Albino mouse strain |
| Strain, strain background (Mus musculus, male) | Copine9-Venus mouse line | This paper | Material and methods section 8.3.1 | |
| Antibody | anti-OPN1SW (N-20) (Goat polyclonal) | Santa Cruz Biotechnology | Cat#sc-14363, RRID:AB_2158332 | IF (1:1200) |
| Antibody | anti-Opsin Red/Green (Rabbit polyclonal) | Millipore/Sigma | Cat#AB5405, RRID:AB_177456 | IF (1:1000) |
| Antibody | anti-Cone Arrestin (Rabbit polyclonal) | Millipore/Sigma | Cat#AB15282, RRID:AB_1163387 | IF (1:300) |
| Antibody | anti-GFP (Chicken polyclonal) | Millipore/Sigma | Cat#AB16901, RRID:AB_11212200 | IF (1:100) |
| Antibody | anti-Rabbit 488 (Donkey polyclonal) | Jackson Immunoresearch | Cat#711-547-003, RRID:AB_2340620 | IF (1:500) |
| Antibody | anti-Rabbit Cy3 (Donkey polyclonal) | Jackson Immunoresearch | Cat#711-165-152, RRID:AB_2307443 | IF (1:500) |
| Antibody | anti-Goat 647 (Donkey polyclonal) | Jackson Immunoresearch | Cat#705-605-147, RRID:AB_2340437 | IF (1:500) |
| Antibody | anti-Goat Cy3 (Donkey polyclonal) | Jackson Immunoresearch | Cat#705-166-147, RRID:AB_2340413 | IF (1:500) |
| Antibody | anti-Chicken 488 (Donkey polyclonal) | Jackson Immunoresearch | Cat#703-545-155, RRID:AB_2340375 | IF (1:500) |
| Sequence-based reagent | Copine9_gRNA_L(73/25) | This paper | 5’GAGACATGACTGGTCCAA3’ | |
| Sequence-based reagent | Copine9_gRNA_R(62/4.40), | This paper | 5’GCCTCGGAGCGTAGCGTCC3’ | |
| Software, algorithm | Zen | Zeiss | Zen lite Black edition 2.3 SP1 | |
| Software, algorithm | FIJI-ImageJ | NIH | v1.52r | https://imagej.nih.gov/ij/ |
| Software, algorithm | Sigma Plot | Systat Software | 13.0 | |
| Software, algorithm | GraphPad Prism | Graph Pad Software | 8.3.0 | |
| Software, algorithm | Photoshop | Adobe | CC 20.0.6 | |
| Software, algorithm | MATLAB | MathWorks | 2016 | |
| Software, algorithm | R | The R Project for Statistical Computing | 3.5.3 | https://www.r-project.org/ |
| Software, algorithm | Retina and Visual Space Retistruct Package | Sterratt DC et al., PLoS Comput Biol. | ||
| Software, algorithm | Zotero | Corporation for Digital Scholarship | 5.0 | https://www.zotero.org/ download/ |
| Other | DAPI | ThermoFisher Scientific | Cat# D3571, RRID:AB_2307445 | (1 ug/ml) |
Additional files
-
Source data 1
Raw quantitative data and statistics analysis.
- https://cdn.elifesciences.org/articles/56840/elife-56840-data1-v2.xlsx
-
Supplementary file 1
Quantifications of cone-type populations and S-cone bipolar cells.
(A) Cone numbers in different retinal areas along the dorsoventral axis in pigmented and albino mouse. Three images/area (dorsal, medial and ventral) from four retinas/strain. Different cone type quantifications are shown as average ± SD, corresponding to the percentages shown in Figure 1C. The total number of cones analyzed per location and strain are shown in the last column. Total number of cones (B) or S-cone Bipolar cells (SCBCs, C) in eight retinas/mouse strain or line (average ± SD, see also Figure 2B). Significant differences between strains p<0.05 (*), p<0.01 (**), p<0.001 (***), p<0.0001 (****).
- https://cdn.elifesciences.org/articles/56840/elife-56840-supp1-v2.docx
-
Supplementary file 2
True S-cone terminals and Cpne9-Venus+SCBCs connectivity in dorsotemporal (DT) and ventronasal retina (VN).
Quantitative data are shown as mean ± SD from the average of five DT and VN retinal areas (Figure 3C). Significant differences between retinal areas, p<0.01 (**), p<0.0001 (****).
- https://cdn.elifesciences.org/articles/56840/elife-56840-supp2-v2.docx
-
Supplementary file 3
True S-cones and S-cone bipolar cells in dorsotemporal and ventronasal retinas.
Numbers of true S-cones (A) and Cpne9-Venus+SCBCs (B) in dorsotemporal (DT) and ventronasal (VN) circular areas (1 mm diameter, Figures 3E and 4B). Quantitative data are shown as average ± SD from eight retinas/strain or line. The mean of true S-cones and Venus+SCBCs in these circular areas was used to calculate the DT:VN and true S-cone:SCBC (C) ratios. Significant differences between strains p<0.05 (*), p<0.001 (***). True S-cones and SCBCs were significant different between DT and VN retina (p<0.0001).
- https://cdn.elifesciences.org/articles/56840/elife-56840-supp3-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/56840/elife-56840-transrepform-v2.pdf





