Microtubules originate asymmetrically at the somatic golgi and are guided via Kinesin2 to maintain polarity within neurons
Figures
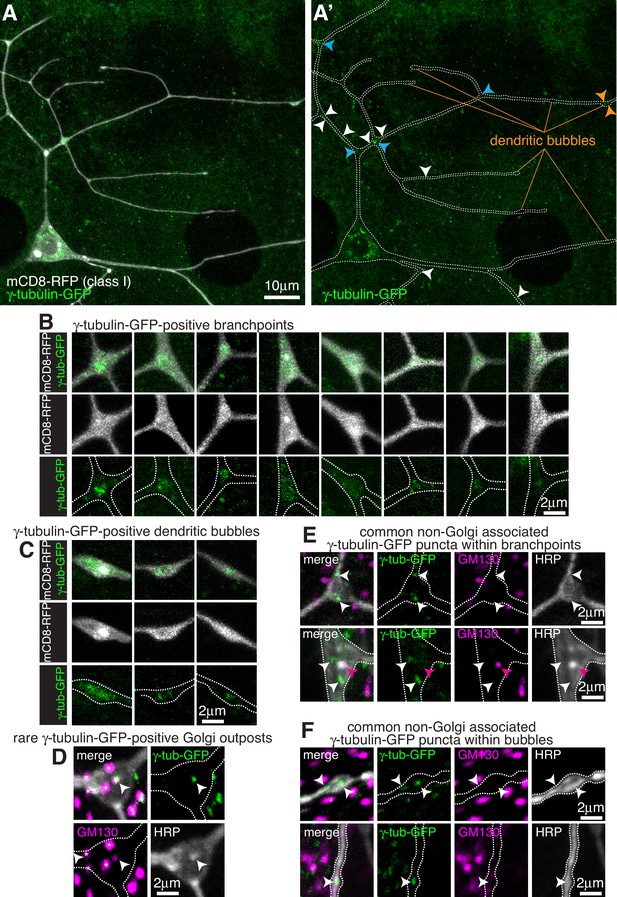
Endogenously-tagged γ-tubulin-GFP localises to a fraction of branchpoints and dendritic bubbles within class I da neurons.
(A) Fluorescent confocal images of the proximal region of a class I da neuron expressing mCD8-RFP (greyscale) within a living 3rd instar larva expressing endogenously-tagged γ-tubulin-GFP (green). Left panel (A) shows an overlay of the GFP and RFP signals, right panel (A’) shows only the GFP signal with the outline of the neuron drawn for clarity; white, blue and orange arrowheads indicate γ-tubulin-GFP puncta/accumulations within dendritic stretches, branchpoints, and dendritic bubbles, respectively. (B,C) Selected images of γ-tubulin-GFP-positive branchpoints (B) or dendritic bubbles (C) from living neurons as in (A). Individual mCD8-RFP channel images (greyscale) have been included for clarity. (D–F) Confocal images show branchpoints (D,E) or dendritic bubbles (F) from 3rd instar larvae expressing endogenous γ-tubulin-GFP fixed and immunostained for GFP (green), GM130 (magenta) and HRP (greyscale). γ-tubulin-GFP signal was rarely observed co-localising with GM130 and HRP signal (D), and frequently observed independent of the Golgi markers at both branchpoints (E) and dendritic bubbles (F).
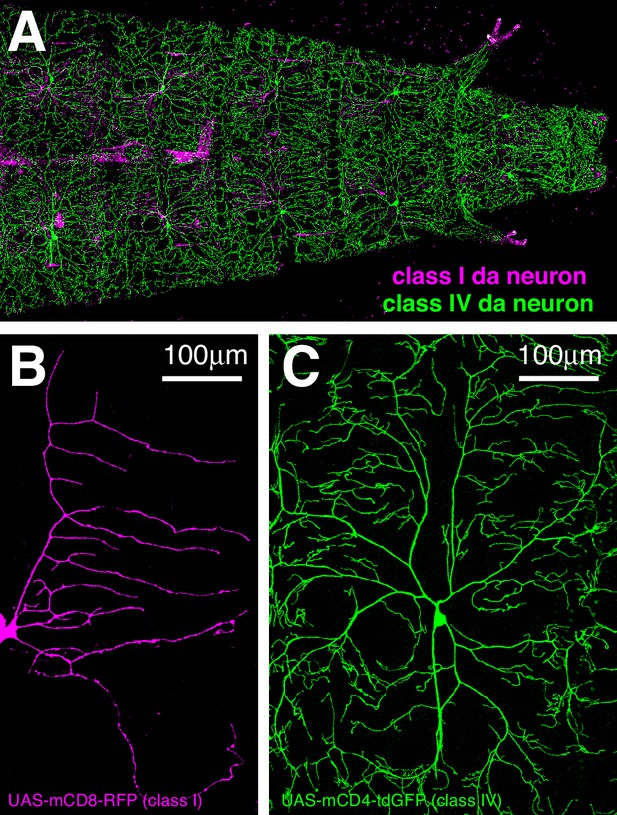
Class I and class IV da neuron morphology.
(A) A fluorescent confocal image of the anterior region of a living 3rd instar larva where mCD8-RFP and mCD4-tdGFP are expressed in class I (magenta) and class IV (green) neurons, respectively. (B,C) Enlarged images of a class I (B) and a class IV (C) da neuron.
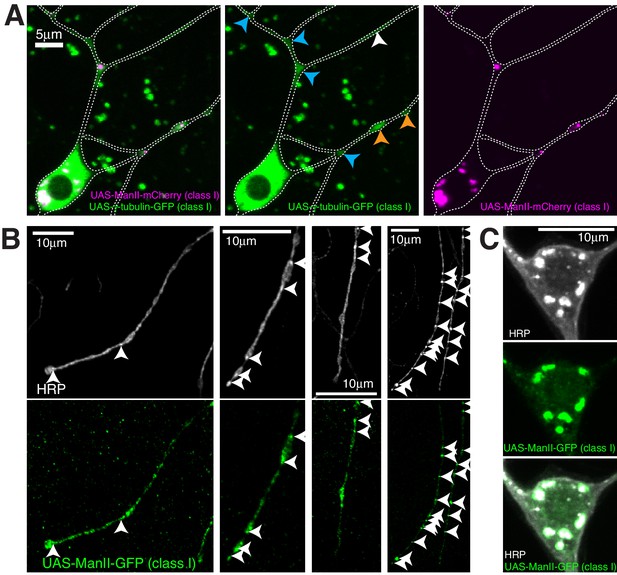
Ectopically expressed γ-tubulin-GFP is strongly enriched in branchpoints and dendritic bubbles and ectopically expressed ManII is not a reliable marker of Golgi outposts in class I da neurons.
(A) Fluorescent confocal images of the proximal region of a class I da neuron expressing 221-gal4 > UAS-γ-tubulin-GFP (green) and 221-gal4 > UAS ManII-mCherry (magenta) within a living 3rd instar larva. Left panel shows an overlay of the GFP and mCherry channels, middle and right panels show the GFP and mCherry channels, respectively. Blue and orange arrowheads indicate enrichments of ectopically expressed γ-tubulin-GFP within branchpoints and dendritic bubbles, respectively. (B,C) Fluorescent confocal images of distal dendrites (B) or a soma (C) from different class I da neurons expressing 221-gal4 > UAS ManII-GFP fixed and immunostained with antibodies against HRP (greyscale) and GFP (green). Arrowheads in (B) indicate HRP puncta that represent internal membrane, at least some of which could be Golgi outposts. Note that UAS-ManII-GFP spreads into regions of the class I dendrites, including dendritic bubbles, that do not contain HRP puncta, suggesting that over-expressing ManII in class I neurons leads to ‘leaking’ of the protein outside of Golgi outposts within dendrites. In contrast, the UAS-ManII-GFP signal always colocalises with the HRP signal within soma (C).
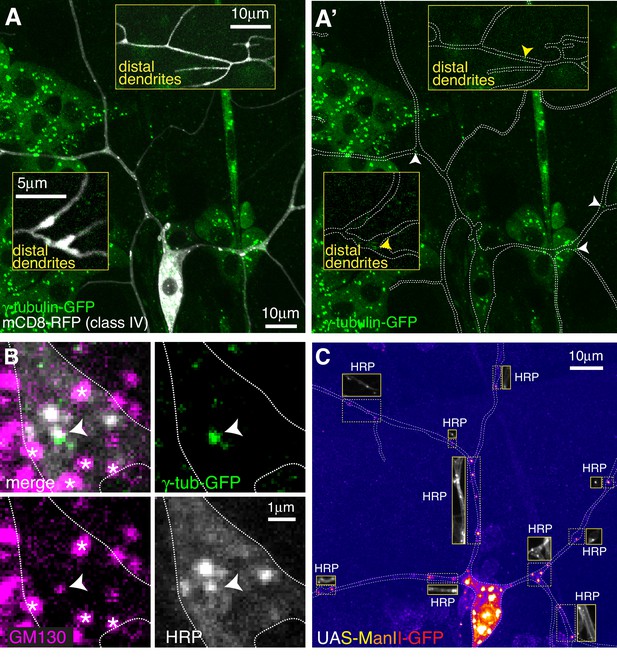
Endogenously-tagged γ-tubulin-GFP localises to a small fraction of proximal Golgi outposts within class IV da neurons.
(A) Fluorescent confocal images of the proximal region of a class IV da neuron expressing UAS-mCD8-RFP (greyscale) within a living 3rd instar larva expressing endogenously-tagged γ-tubulin-GFP (green). Left panel (A) shows an overlay of the GFP and RFP signals, right panel (A’) shows only the GFP signal with the outline of the neuron drawn for clarity; insets show examples of distal dendrites located >200 μm from the soma; white and yellow arrowheads indicate bright or weak γ-tubulin-GFP puncta, respectively, within proximal or distal dendrites. (B) Images show a γ-tubulin-GFP-positive proximal branchpoint from a 3rd instar larva expressing endogenous γ-tubulin-GFP fixed and immunostained for GFP (green), GM130 (magenta) and HRP (greyscale). Asterisks indicate GM130 foci from overlapping epidermal cells. Arrowhead indicates an example of a γ-tubulin-GFP-positive Golgi outpost observed within proximal class IV branchpoints. (C) Confocal images show the proximal region of a class IV dendritic arbor within a 3rd instar larva expressing ManII-GFP (to mark medial Golgi) fixed and immunostained for GFP (fire) and HRP (greyscale within insets). Insets show HRP staining alone.
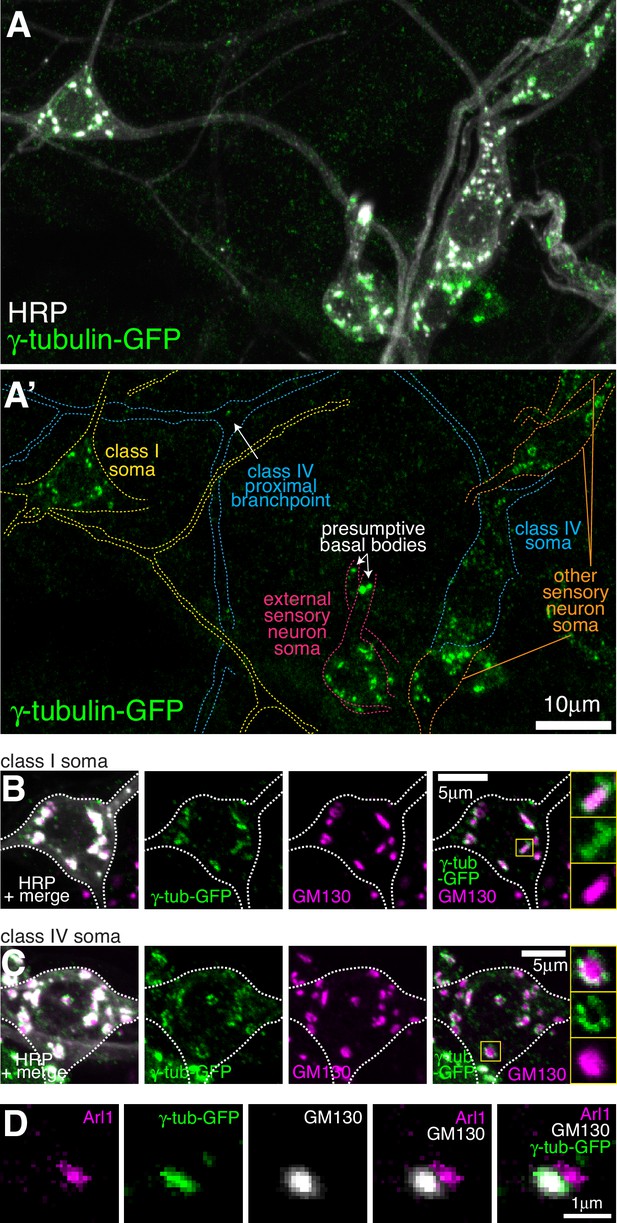
Endogenously-tagged γ-tubulin-GFP localises to the somatic Golgi of sensory neurons.
(A) Confocal images show the somas and some proximal dendrites of sensory neurons within the dorsal cluster: class I (yellow), class IV (blue), as (pink) and other (orange) from a 3rd instar larva expressing endogenously-tagged γ-tubulin-GFP and immunostained for GFP (green) and HRP (marking Golgi stacks, greyscale). Upper panel (A) shows an overlay of the GFP and HRP signals, lower panel (A’) shows only the GFP signal with coloured neuronal outlines drawn for clarity. (B, C) Confocal images show the somas of class I (B) or class IV (C) da neurons from a 3rd instar larva expressing endogenously-tagged γ-tubulin-GFP fixed and immunostained for GFP (green), GM130 (magenta) and HRP (greyscale). Enlarged boxes in (B) and (C) show side-on and face-on stacks, respectively. (D) Confocal images show an example of a single Golgi stack within a da neuron from a 3rd instar larva expressing endogenously-tagged γ-tubulin-GFP fixed and immunostained for GFP (green), Arl1 (trans-Golgi, magenta) and GM130 (cis-Golgi, greyscale). The larva was also stained with HRP antibodies to identify neurons, but this channel has been omitted for clarity.
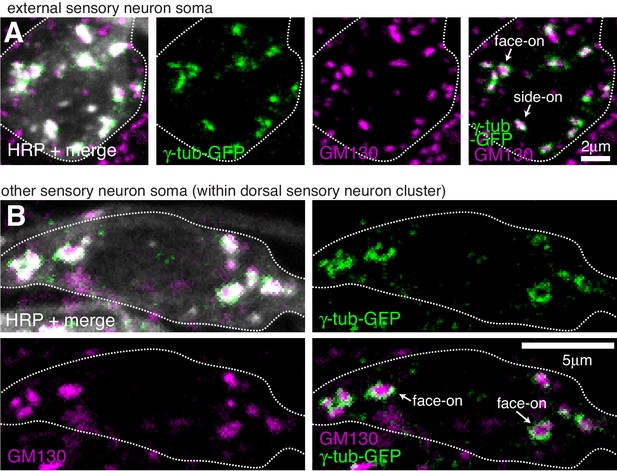
Endogenously-tagged γ-tubulin-GFP localises to the somatic Golgi of sensory neurons.
(A,B) Confocal images show the soma of an es neuron (A) or another sensory neuron within the dorsal cluster (B) from 3rd instar larva expressing two copies of endogenous γ-tubulin-GFP fixed and immunostained with antibodies against GFP (green), GM130 (magenta) and HRP (greyscale). γ-tubulin-GFP and GM130 signals associate at presumptive side-on stacks (extended GM130 signal) and face-on stacks (round GM130 signal) in a way that suggests γ-TuRCs are recruited to the rims of the cis-Golgi.
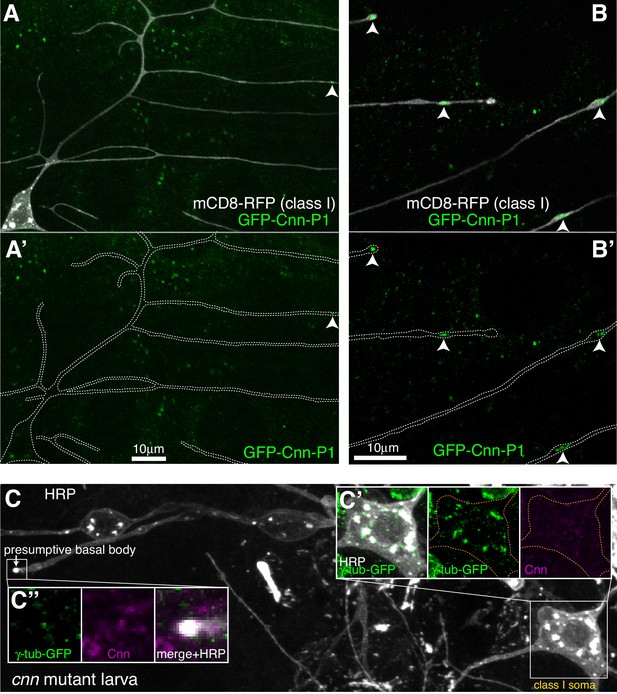
Cnn-P1 is dispensable for γ-tubulin-GFP recruitment to the somatic Golgi.
(A,B) Fluorescent confocal images of proximal (A) and distal (B) regions of class I da neurons expressing mCD8-RFP (greyscale) within a living 3rd instar larva expressing endogenously-tagged GFP-Cnn-P1 (green). Upper panels show an overlay of the GFP and RFP channels, lower panels show only the GFP channel with the outline of the neurons drawn for clarity. Arrowhead in (A) indicates a rare dendritic GFP-Cnn-P1 puncta within a proximal dendritic bubble, while arrowheads in (B) indicate more frequent GFP-Cnn-P1 accumulations within distal dendritic bubbles. (C) Confocal images show the somas and some proximal dendrites of sensory neurons within the dorsal cluster from a 3rd instar cnn mutant larva expressing endogenously-tagged γ-tubulin-GFP and immunostained for GFP (green), HRP (greyscale) and Cnn (magenta). Images in (C’) and (C’’) show the separate channels for the class I soma and the presumptive basal body of an es neuron, respectively.
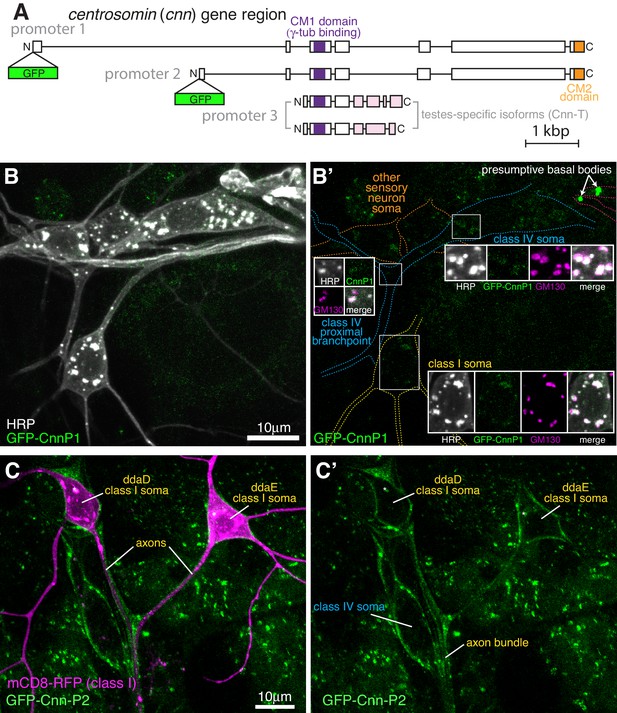
Cnn-P1 is not strongly associated with the somatic Golgi or Golgi outposts of sensory neurons, while Cnn-P2 is expressed in ensheathing glia.
(A) A gene diagram of cnn to indicate the different Cnn isoforms. Boxed regions are exons, lines are introns. CRISPR combined with homologous recombination was used to insert GFP just after the start codon of either the promoter 1 (P1) or promoter 2 (P2) isoform. Promotor three isoforms are expressed only in testes and so were not analysed in this study. (B) Confocal images show the somas and some proximal dendrites of sensory neurons within the dorsal cluster from a 3rd instar larva expressing endogenously-tagged GFP-Cnn-P1 and immunostained with nanobodies against GFP covalently coupled to ATTO-488 (green) and antibodies against HRP (greyscale) and GM130 (magenta). Left panel (B) shows an overlay of the GFP and anti-HRP signals, while the right panel (B’) shows the GFP signal with coloured outlines of the different neurons drawn for clarity, with insets showing all channels, including an overlay, for the class I neuron soma, regions of the class IV neuron soma and a class IV proximal branchpoint that contains Golgi outposts. The GFP-Cnn-P1 signal appears very weakly, if at all, at the somatic Golgi and Golgi outposts marked by HRP and GM130 staining. (C) Confocal images show class I da neurons from a living 3rd instar larva expressing 221-Gal4 > UAS-mCD8-RFP (magenta) and two copies of endogenously tagged GFP-Cnn-P2. Left panel (C) shows an overlay of the GFP and RFP channels, while the right panel (C’) shows just the GFP channel. Note that the GFP-Cnn-P2 surrounds the axons and somas of the neurons, suggestive of localisation within ensheathing glia. The * star indicates a region of very bright RFP signal that has bled through into the GFP channel.
Endogenously-tagged GFP-Cnn-P1 localises as expected to centrosomes within Drosophila syncytial embryos.
The Movie was taken using multi-Z-stack time-lapse epifluorescence microscopy and shows Z-projected images through time of a syncytial embryo expressing endogenously-tagged GFP-Cnn-P1. The cycle starts in M-phase and the centrosomes separate when the cycle transitions to the following S-phase. GFP-Cnn-P1 localises to centrosomes during both M-phase, where the signal is rounded, and S-phase, where the signal displays typical Cnn ‘flares’. Images were collected every 20 s and the Movie plays at 20 frames/second.
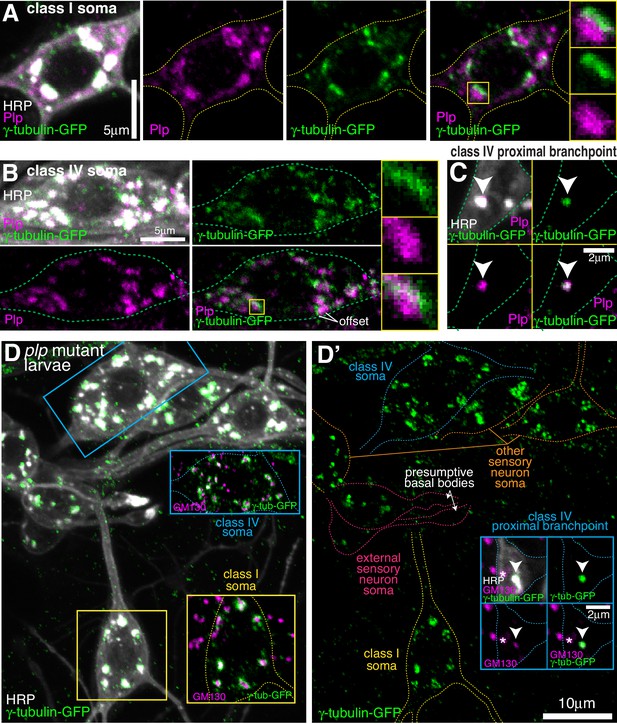
Endogenous Plp localises to Golgi but is dispensable for γ-tubulin-GFP recruitment.
(A–C) Confocal images show the somas of a class I (A) or a class IV (B) neuron and a proximal branchpoint of a class IV neuron (C) from 3rd instar larvae expressing γ-tubulin-GFP fixed and immunostained for GFP (green), Plp (magenta) and HRP (greyscale). Arrowhead in (C) indicates a γ-tubulin-GFP-positive Golgi outpost that contains Plp. (D) Confocal images show the somas and some proximal dendrites of sensory neurons within the dorsal cluster: class I (yellow), class IV (blue), es (pink) and other (orange), from a plp mutant 3rd instar larva expressing endogenously-tagged γ-tubulin-GFP and immunostained for GFP (green), HRP (greyscale) and GM130 (magenta). Left panel (D) shows an overlay of the GFP and HRP signal, with insets showing overlays for different neurons, as indicated; right panel (D’) shows the GFP channel with coloured outlines of the different neurons drawn for clarity, with insets showing separate channels for a proximal branchpoint.
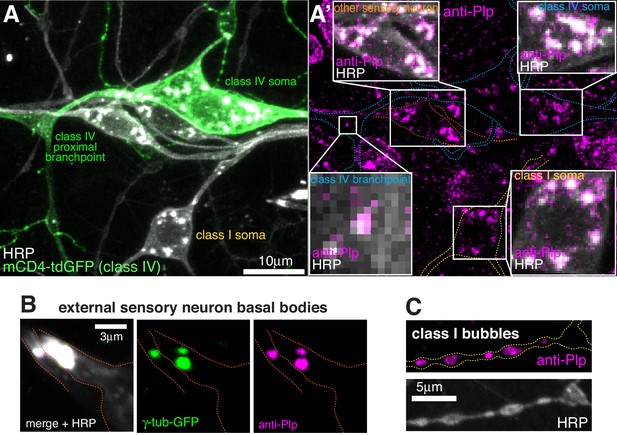
Plp localises to the somatic Golgi, Golgi outposts, basal bodies and class I -specific dendritic bubbles.
(A) Confocal images show the somas and some proximal dendrites of sensory neurons within the dorsal cluster from a 3rd instar larva expressing ppk-mCD4-tdGFP and immunostained with antibodies against GFP (green), HRP (greyscale) and Plp (magenta). Left panel (A) shows the GFP and anti-HRP channels, while the right panel (A’) shows the anti-Plp channel; insets show an overlay of the anti-Plp and anti-HRP channels. Plp signal colocalises with the HRP staining that marks the multiple well-distributed Golgi stacks within the neuronal soma of each neuron. Plp also localises to some HRP puncta (presumably Golgi outposts) within the proximal branchpoints of class IV neurons. (B) Confocal images show parts of the es neurons containing the basal bodies from a 3rd instar larva expressing two copies of endogenously tagged γ-tubulin-GFP and immunostained with antibodies against GFP (green), HRP (greyscale) and Plp (magenta). (C) Confocal images show dendritic bubbles of a class I da neuron from a 3rd instar larva immunostained with antibodies against HRP (greyscale) and Plp (magenta).
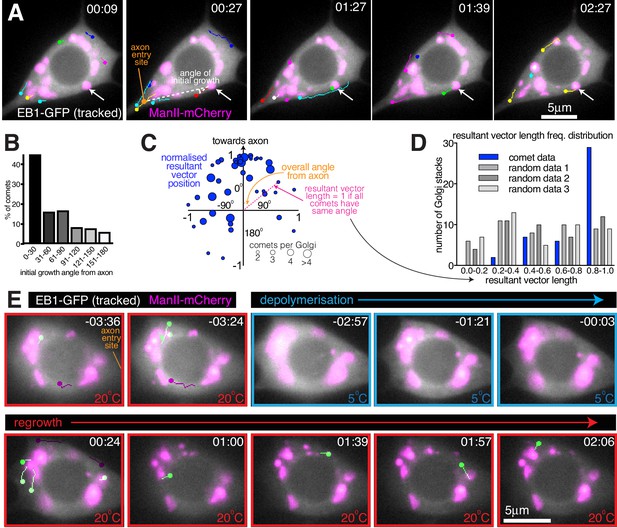
Somatic Golgi stacks nucleate microtubules.
(A) Widefield fluorescent images from a movie showing the soma of a class I da neuron expressing EB1-GFP (greyscale) and ManII-mCherry (magenta). Manually assigned multi-colour tracks of EB1-GFP comets were drawn over each image (filled circle = last location of comet). Arrow indicates a Golgi stack from which sequential EB1-GFP comets emerge; dotted lines show an example of an angle measurement of a comet’s initial growth relative to the axon entry point (orange circle). Time in min:s from the start of the movie is shown. (B) Graph shows a frequency distribution of the initial growth angle (before turning) relative to the axon entry site (as indicated in (A)) of EB1-GFP comets that emerged from Golgi stacks. Negative angles were made positive so as not to distinguish between comets growing to the left or right of the axon. 163 comets from 59 Golgi stacks from seven neurons were analysed. (C) Scatter graph shows the position of the normalised resultant comet vectors (see methods) for each of 44 Golgi stacks from seven neurons. Circle position is representative of the average angle from the axon and how similar the angles were (as indicated). Circle size reflects the number of comets analysed per Golgi stack (as indicated below). (D) Graph shows a frequency distribution of the resultant vector lengths for the comet data (blue) and for 3 sets of data produced using randomly generated angles (grey shades). (E) Widefield fluorescent images from a movie showing the soma of class I da neuron expressing UAS-EB1-GFP (greyscale) and UAS-ManII-mCherry (magenta). The sample was subjected to a warm-cool-warm temperature cycle to induce depolymerisation (5°C; blue outline) and then repolymerisation (20 °C; red outline) of dynamic microtubules. Tracks were manually colour coded: green = comets emerging from Golgi stacks; purple = comets emerging from elsewhere. Time in min:s from the 5°C to 20°C transition is shown.
-
Figure 6—source data 1
Calculating the angle of microtubule growth from Golgi stacks.
- https://cdn.elifesciences.org/articles/58943/elife-58943-fig6-data1-v3.xlsx
EB1-comets emerge from somatic Golgi stacks and initially grow preferentially towards the axon.
The Movie was taken using single-Z time-lapse epifluorescence microscopy and shows a class I da neuron expressing 221-Gal4 > UAS-EB1-GFP and 221-Gal4 > UAS ManII-mCherry. The left panel shows an overlay of the GFP and mCherry channels, while the right panel shows the same overlay with the ImageJ-generated EB1-GFP comet tracks drawn onto the images. The axon (A) and dendrites (D) are labelled in the right panel. The tracks are multi-coloured, as is default for ImageJ, and the colours have no reference to comet type. Comets can be seen emerging from Golgi stacks and this can occur repeatedly from the same stack (see bottom right stack). Images were collected every 3 s and the Movie plays at 10 frames/second.
The somatic Golgi stacks nucleate microtubules.
The Movie was taken using single-Z time-lapse epifluorescence microscopy and shows a class I da neuron expressing 221-Gal4 > UAS-EB1-GFP and 221-Gal4 > UAS ManII-mCherry. The sample was subjected to a warming-cooling-warming (20 °C-5°C-20°C) cycle, as indicated by the red (20 °C) and blue (5 °C) borders. The left panel shows an overlay of the GFP and mCherry channels, while the right panel shows the same overlay with the ImageJ-generated EB1-GFP comet tracks drawn onto the images. The axon (A) and dendrites (D) are labelled in the right panel. The tracks have been manually post-coloured such that green represent comets emerging from Golgi stacks and purple represent comets emerging from elsewhere within the soma. Comets can be seen emerging from Golgi stacks before cooling, and then all comets stop during the 3 min cooling period. After warming, comets can be seen emerging from the Golgi stacks, including comets that emerge >1 min after warming. Images were collected every 3 s and the Movie plays at 10 frames/second.
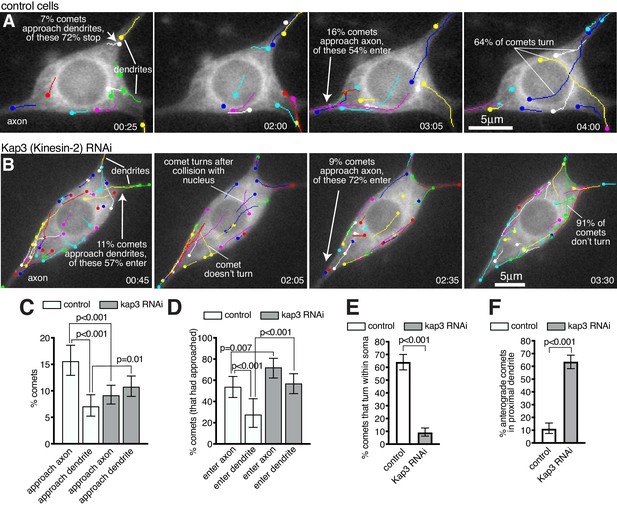
Microtubules within the soma grow preferentially towards the axon and are excluded from entering dendrites in a Kinesin-2-dependent manner.
(A,B) Widefield fluorescent images from movies showing the somas of class I da neurons expressing UAS-EB1-GFP and either γ-tubulin-37C RNAi (control) (A) or UAS-Kap3 RNAi (B). Manually assigned multi-colour tracks of EB1-GFP comets are drawn over each image (filled circle = last location of comet). Time in min:s from the start of the movie is shown. (C) Graph shows the % of comets that approach either the axon (green) or the dendrites (magenta) in either control (666 comets analysed across 13 movies) or Kinesin-2 RNAi (1058 comets analysed across nine movies) class I da neuron somas, as indicated. (D) Graph shows the % of comets that enter the axon (green) or the dendrites (magenta) as a proportion of those that had approached the axon in either control or Kinesin-2 RNAi class I da neuron somas, as indicated. (E) Graph shows the % of applicable comets that display turning events within either control (n = 257 comets across 13 movies) or Kinesin-2 RNAi (n = 386 comets across nine movies) class I da neuron somas, as indicated. (F) Graph shows the % of comets that are anterograde in the proximal primary dendrite (before any branches) in either control (n = 252 comets across 13 movies) or Kinesin-2 RNAi (n = 338 comets across nine movies) class I da neurons, as indicated. Error bars in (C–F) show the 95% confidence intervals.
-
Figure 7—source data 1
Calculating percentages of comets aproaching and entering axons and dendrites, turning events, and proximal dendrite polarity in control neurons.
- https://cdn.elifesciences.org/articles/58943/elife-58943-fig7-data1-v3.xlsx
-
Figure 7—source data 2
Calculating percentages of comets aproaching and entering axons and dendrites, turning events, and proximal dendrite polarity in Kap3 RNAi neurons.
- https://cdn.elifesciences.org/articles/58943/elife-58943-fig7-data2-v3.xlsx
-
Figure 7—source data 3
Calculating percentages of comets entering dendrites in Klp64D RNAi neurons.
- https://cdn.elifesciences.org/articles/58943/elife-58943-fig7-data3-v3.xlsx
Microtubules turn towards the axon and are excluded from entering dendrites.
The Movie was taken using single-Z time-lapse epifluorescence microscopy and shows a control class I da neuron expressing 221-Gal4 > UAS-EB1-GFP and 221-Gal4 > UAS-γ-tubulin-37c-RNAi. The left panel shows the GFP channel, while the right panel shows the GFP channel with the ImageJ-generated EB1-GFP comet tracks drawn onto the images. The axon (A) and dendrites (D) are labelled in the right panel. The tracks are multi-coloured, as is default for ImageJ, and the colours have no reference to comet type. Comets display turning events, indicating that the growing plus ends of microtubules can be guided along pre-existing microtubules. Comets that approach the axon frequently enter and grow down the axon; In contrast, comets that approach a dendrite entry site do not enter (quantified in Figure 7). Images were collected every 5 s and the Movie plays at 10 frames/second.
Kinesin-2 is required for growing microtubules to turn within the soma and to be excluded from entering dendrites.
The Movie was taken using single-Z time-lapse epifluorescence microscopy and shows a Kinesin-2 RNAi class I da neuron expressing 221-Gal4 > UAS-EB1-GFP and 221-Gal4 > UAS-Kap3-RNAi. The left panel shows the GFP channel, while the right panel shows the GFP channel with the ImageJ-generated EB1-GFP comet tracks drawn onto the images. The axon (A) and dendrites (D) are labelled in the right panel. The tracks are multi-coloured, as is default for ImageJ, and the colours have no reference to comet type. Comets can be seen emerging from multiple locations within the soma but unlike in control neurons they do not turn unless they encounter the nuclear envelope or cell cortex, suggesting that Kinesin-2 is required to guide growing microtubules towards the axon. Comets that approach the axon still frequently enter and grow down the axon, and, in contrast to control neurons, comets that approach dendrites also readily enter (quantified in Figure 7). This shows that Kinesin-2 is also required for excluding growing microtubules from entering dendrites. Images were collected every 5 s and the Movie plays at 10 frames/second.
Knockdown of Klp64D supports the conclusion that Kinesin2 is required for growing microtubules to be excluded from entering dendrites.
The Movie was taken using single-Z time-lapse epifluorescence microscopy and shows a Kinesin-2 RNAi class I da neuron expressing 221-Gal4 > UAS-EB1-GFP and 221-Gal4 > UAS-Klp64D RNAi. Images were collected every 5 s and the Movie plays at 10 frames/second.
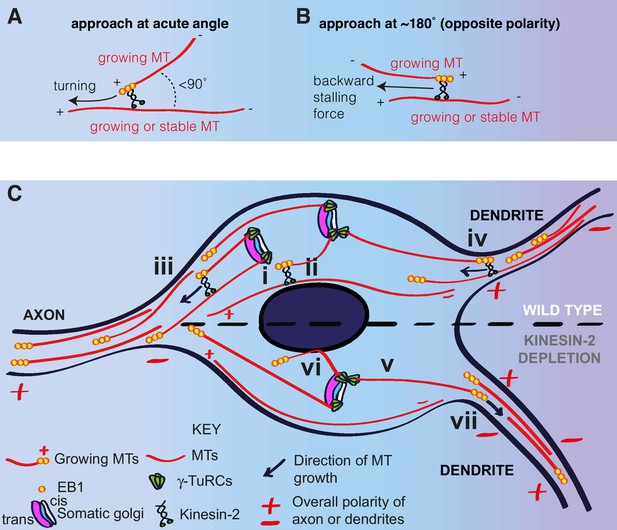
A model for microtubule nucleation and organisation within the soma of da neurons.
(A,B) Diagrams showing how plus-end-associated Kinesin-2 is predicted to induce turning of a growing microtubule along another microtubule when it approaches the other microtubule at an acute angle (A), or plus-end stalling when the growing microtubule approaches the other microtubules at an angle of ~180° (B). (C) A model depicting microtubule nucleation and growth within the soma of wild type (upper section) or Kinesin-2 depleted (lower section) da neurons. γ-TuRCs localise to the cis-Golgi (white) and nucleate microtubules preferentially towards the axon (i), creating a polarised microtubule network. In wild type neurons, plus-end-associated Kinesin-2 guides growing microtubules along this network towards the axon (ii). Growing microtubules readily enter axons (iii) but are excluded from entering dendrites (iv), because plus-end-associated Kinesin-2 engages with microtubules of opposite polarity, exerting a backward stalling force on the growing microtubule. In Kinesin-2 depleted neurons, microtubules nucleated from the somatic Golgi grow in straight lines (v) until contact with the nuclear envelope (vi) or the cell cortex (not depicted). Microtubules that approach dendrites can now readily enter (vii) resulting in a loss of overall minus-end-out microtubule polarity.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Escherichia coli) | 5-alpha competent E. coli | NEB | C2987I | High Efficiency chemically competent |
| Genetic reagent (Drosophila melanogaster) | cnnf04547 | Exelixis at Harvard Medical School | Exelixis:f04547; Flybase:FBst1019233 | FlyBase symbol: cnnf04547 ; Stock discontinued at the Exelixis and available from Conduit lab upon request. |
| Genetic reagent (D. melanogaster) | plp5 | Bloomington Drosophila Stock Center | BDSC:9567; FLYB:FBst0009567; RRID:BDSC_9567 | FlyBase symbol: ru1 Plp5 st1/TM6C, Sb1 Tb1 |
| Genetic reagent (D. melanogaster) | plps2172 | Bloomington Drosophila Stock Center | BDSC:12089; Flybase:FBst0012089; RRID:BDSC_12089 | FlyBase symbol: ‘w[1118]; P{w[+mC]=lacW}Plp[s2172]/TM3, Sb[1]’ |
| Genetic reagent (D. melanogaster) | UAS-mCD8-RFP | Bloomington Drosophila Stock Center | BDSC:27392; Flybase:FBti0115769; RRID:BDSC_27392 | FlyBase symbol: ‘w[*]; P{w[+mC]=UAS-mCD8.ChRFP}3’ |
| Genetic reagent (D. melanogaster) | UAS-EB1-GFP | Bloomington Drosophila Stock Center | BDSC:35512; Flybase:FBti0141213; RRID:BDSC_35512 | FlyBase symbol: ‘w[*]; P{w[+mC]=UAS-EB1-GFP}3’ |
| Genetic reagent (D. melanogaster) | UAS-ManII-GFP | Bloomington Drosophila Stock Center | BDSC:65248; Flybase:FBti018367; RRID:BDSC_65248 | FlyBase symbol: ‘w[1118]; P{w[+mC]=UAS-ManII-EGFP}2; TM2/TM6B, Tb[1]’ |
| Genetic reagent (D. melanogaster) | UAS-ManII-mCherry | Ori-McKenney et al., 2012 (doi:10.1016/j.neuron.2012.10.008) | Fly genotype: ‘w[1118]; P{w[+mC]=UAS-ManII-mCherry}/CyO’; gift from Yuh-Nung Jan | |
| Genetic reagent (D. melanogaster) | ppk-Gal4 | Bloomington Drosophila Stock Center | BDSC:32078; Flybase:FBti0127690; RRID:BDSC_32078 | FlyBase symbol: ‘w[*]; P{w[+mC]=ppk-GAL4.G}2’ |
| Genetic reagent (D. melanogaster) | ppk-Gal4 | Bloomington Drosophila Stock Center | BDSC:32079; Flybase:FBti013120; RRID:BDSC_32079 | FlyBase symbol: ‘w[*]; P{w[+mC]=ppk-GAL4.G}3’ |
| Genetic reagent (D. melanogaster) | 221-Gal4 | Bloomington Drosophila Stock Center | BDSC:26259; Flybase:FBti011433; RRID:BDSC_26259 | FlyBase symbol: ‘w[*]; Pin[1]/CyO; P{GawB}221w-’ |
| Genetic reagent (D. melanogaster) | ppkCD4-tdGFP | Bloomington Drosophila Stock Center | BDSC:35842; Flybase:FBti014342; RRID:BDSC_35842 | FlyBase symbol: ‘w[1118]; P{w[+mC]=ppk-CD4-tdGFP}1b’ |
| Genetic reagent (D. melanogaster) | UAS-Dicer 2 | Bloomington Drosophila Stock Center | BDSC:24646; Flybase:FBti010143; RRID:BDSC_24646 | FlyBase symbol: P{w[+mC]=UAS-Dcr2.D}1,w[1118] |
| Genetic reagent (D. melanogaster) | γ-tubulin37C RNAi; UAS-γ-tubulin37C RNAi | Bloomington Drosophila Stock Center | BDSC:32513; Flybase:FBti0132207; RRID:BDSC_32513 | FlyBase symbol: ‘y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]=TRiP.HMS00517}attP2’ |
| Genetic reagent (D. melanogaster) | Kap3 RNAi; UAS-Kap3 RNAi | ViennaDrosophila Resource Center | FBst0466097; VDRC ID:45400 | FlyBase symbol: ‘w1118;P{GD7377}v45400’ |
| Genetic reagent (D. melanogaster) | Klp64D RNAi | ViennaDrosophila Resource Center | FBst0466080; VDRC ID:45373 | FlyBase symbol: ‘w1118;P{GD7048}v45373’ |
| Genetic reagent (D. melanogaster) | γ-tubulin23c-sfGFP | Tovey and Conduit, 2018 (https://doi.org/10.1016/j.cub.2018.05.044) | insertion of sfGFP with a 4X GlyGlySer linker at the C-terminus | |
| Genetic reagent (D. melanogaster) | γ-tubulin23c-eGFP | This paper | insertion of eGFP with a 4X Ser linker at the C-terminus. Generated by inDroso | |
| Genetic reagent (D. melanogaster) | sfGFP-Cnn-P1 | This paper | insertion of sfGFP with a 4X GlyGlySer linker at the start of exon 1a | |
| Genetic reagent (D. melanogaster) | sfGFP-Cnn-P2 | This paper | insertion of sfGFP with a 4X GlyGlySer linker at the start of exon 1b | |
| Antibody | anti-GFP (mouse monoclonal) | Roche | Cat# 11814460001; RRID:AB_390913 | IF(1:250) |
| Antibody | anti-Cnn (rabbit polyclonal) | Lucas and Raff, 2007 (https://doi.org/10.1083/jcb.200704081) | IF(1:1000); gift from Jordan Raff | |
| Antibody | anti-Plp (rabbit polyclonal) | Martinez-Campos et al., 2004 (https://doi.org/10.1083/jcb.200402130) | IF(1:500); gift from Jordan Raff | |
| Antibody | anti-Arl1 (chicken polyclonal) | Torres et al., 2014 (doi:10.1242/jcs.122028) | IF(1:200); gift from Sean Munroe | |
| Antibody | anti-GM130 (rabbit polyclonal) | Abcam | Cat# ab30637; RRID:AB_732675 | IF(1:300) |
| Antibody | Alexa-647 conjugated Hrp (goat polyclonal) | Jackson | Jackson ImmunoResearch 123-605-021; RRID:AB_2338967 | IF(1:500) |
| Antibody | ATTO-488 GFP-booster (Camelid single domain ‘nanobody’) | Chromotek | gb2AF488-10; RRID:AB_2827573 | IF(1:200) |
| Antibody | anti-Mouse IgG H and L Alexa Fluor 488 (goat polyclonal) | Abcam | Cat# ab150117, RRID:AB_2688012 | IF(1:500) |
| Antibody | anti-Rabbit IgG H and L Alexa Fluor 568 (goat polyclonal) | Thermo Fisher Scientific | Cat# A-11036, RRID:AB_10563566 | IF(1:500) |
| Recombinant DNA reagent | pCDF3 (plasmid) | Addgene | RRID:Addgene_49410 | pCFD3-dU6:3gRNA; gift from Simon Bullock |
| Recombinant DNA reagent | pBluescript SK+ (plasmid) | Simon Bullock lab | Standard cloning vector; gift from Simon Bullock | |
| Recombinant DNA reagent | pBS-KS-attB1-2-PT-SA-SD-2-sfGFP-FIAsh-StrepII-TEV-3x-FLAG (plasmid) | Drosophila Genomics Resource Center | Stock number:1314 | Vector used to amplify sf GFP sequence |
| Sequence-based reagent | CnnNt_P1T2_f | This paper | Guide oligo | GTCGTGTTTAGACTGGTCCATGGG To generate Bbs1 cleavable fragment to clone into Bbs1 cut pCFD3 for Cnn promoter one tagging. Guide sequence underlined. |
| Sequence-based reagent | CnnNt_P1T2_b | This paper | Guide oligo | AAACCCCATGGACCAGTCTAAACA; To generate Bbs1 cleavable fragment to clone into Bbs1 cut pCFD3 for Cnn promoter one tagging. Guide sequence underlined. |
| Sequence-based reagent | CnnNt_P2T2_f | This paper | Guide oligo | GTCGTTAAATGAAACATAGAATA; To generate Bbs1 cleavable fragment to clone into Bbs1 cut pCFD3 for Cnn promoter two tagging. Guide sequence underlined. |
| Sequence-based reagent | CnnNt_P2T2_b | This paper | Guide oligo | AAACTATTCTATGTTTCATTTAA; To generate Bbs1 cleavable fragment to clone into Bbs1 cut pCFD3 for Cnn promoter two tagging. Guide sequence underlined. |
| Sequence-based reagent | cnn_UHA_P1_GFP_f | This paper | PCR primers | AAAGTTAACTATTTGAGGACCTCCCATGGTGTCCAAGGGCGAGGAG; used to amplify sfGFP with overhangs for HiFi cloning into pBS containing homology arms for for Cnn promoter 1 |
| Sequence-based reagent | cnn_DHA_P1_GFP_b | This paper | PCR primers | CCCGCAAAACCTGTTTAGACTGGTCGGATCCGCCGCTACCTCCGCTTCCACCGGAACCTCCCTTGTACAGCTCATCCATGCC; used to amplify sfGFP with overhangs for HiFi cloning into pBS containing homology arms for for Cnn promoter 1. Also contains sequence for 4X GlyGlySer linker |
| Sequence-based reagent | cnn_UHA_P2_GFP_f | This paper | PCR primers | GCAAATGTTAAATGAAAGATACAATATGGTGTCCAAGGGCGAGGAG; used to amplify sfGFP with overhangs for HiFi cloning into pBS containing homology arms for for Cnn promoter 2 |
| Sequence-based reagent | cnn_DHA_P2_GFP_b | This paper | PCR primers | GTGAGGTAGATCGAAAGATACCCGCGGATCCGCCGCTACCTCCGCTTCCACCGGAACCTCCCTTGTACAGCTCATCCATGCC; used to amplify sfGFP with overhangs for HiFi cloning into pBS containing homology arms for for Cnn promoter 2. Also contains sequence for 4X GlyGlySer linker |
| Sequence-based reagent | cnn_UHA_P1_f | This paper | PCR primers | GAAAGCTAAGCAGATTCTTCAGC; used to amplify upstream homology arm for promoter 1 |
| Sequence-based reagent | cnn_UHA_P1_b | This paper | PCR primers | GGGAGGTCCTCAAATAGTTAAC; used to amplify upstream homology arm for promoter 1 |
| Sequence-based reagent | cnn_DHA_P1_f | This paper | PCR primers | GACCAGTCTAAACAGGTTTTGC; used to amplify downstream homology arm for promoter 1 |
| Sequence-based reagent | cnn_DHA_P1_b | This paper | PCR primers | GCTCTCTGCACGTCCAAATAAAC; used to amplify downstream homology arm for promoter 1 |
| Sequence-based reagent | cnn_UHA_P2_f | This paper | PCR primers | CACTCGCTATAGTCTCGATCC; used to amplify upstream homology arm for promoter 2 |
| Sequence-based reagent | cnn_UHA_P2_b | This paper | PCR primers | ATTGTATCTTTCATTTAACATTTGCGCTC; used to amplify upstream homology arm for promoter 2 |
| Sequence-based reagent | cnn_DHA_P2_f | This paper | PCR primers | GCGGGTATCTTTCGATCTACC; used to amplify downstream homology arm for promoter 2 |
| Sequence-based reagent | cnn_DHA_P2_b | This paper | PCR primers | TCCCGTGGAACAGGAACCAC; used to amplify downstream homology arm for promoter 2 |
| Sequence-based reagent | cnn_DHA_P1_pBS_f | This paper | PCR primers | CGGTTTATTTGGACGTGCAGAGAGCGGGGGATCCACTAGTTCTAGAG; used to amplify pBS backbone for promoter 1. Contains overhangs for HiFi cloning. |
| Sequence-based reagent | cnn_UHA_P1_pBS_b | This paper | PCR primers | AAGCTGAAGAATCTGCTTAGCTTTCGGGCTGCAGGAATTCGATATC; used to amplify pBS backbone for promoter 1. Contains overhangs for HiFi cloning. |
| Sequence-based reagent | cnn_DHA_P2_pBS_f | This paper | PCR primers | TTGCTGTGGTTCCTGTTCCACGGGAGGGGGATCCACTAGTTCTAGAG; used to amplify pBS backbone for promoter 2. Contains overhangs for HiFi cloning. |
| Sequence-based reagent | cnn_UHA_P2_pBS_b | This paper | PCR primers | AAGTGGATCGAGACTATAGCGAGTGGGGCTGCAGGAATTCGATATC; used to amplify pBS backbone for promoter 2. Contains overhangs for HiFi cloning. |
| Other | Live imaging solution | Thermo Fisher Scientific | A14291DJ | Larvae imaging |





