Live imaging of hair bundle polarity acquisition demonstrates a critical timeline for transcription factor Emx2
Figures
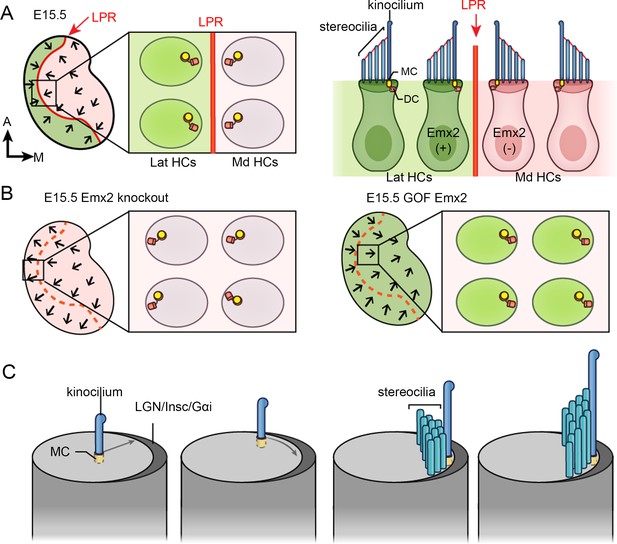
Hair bundle orientation establishment in the developing mouse utricle.
(A) In E15.5 utricle, hair bundles are pointing toward each other (arrows) across the line of polarity reversal (LPR, red). The kinocilium is located asymmetrically at the lateral region of the apical hair cell (HC) surface. The mother centriole (MC) (yellow), which forms the basal body of the kinocilium, is located more centrally relatively to the daughter centriole (DC) (orange). Emx2-positive domain is in green. (B) Schematics showing hair bundle orientation in Emx2 knockout and gain-of-Function utricles. (C) Model showing asymmetric hair bundle establishment requires the LGN/Inscuteable/Gαi complex. Orientations: A, anterior; M, medial.
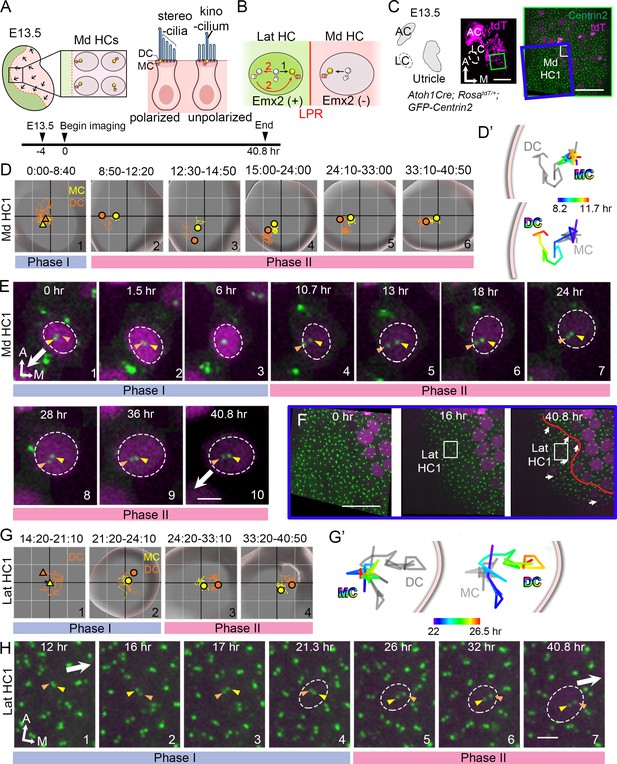
Live imaging of hair bundle establishment in medial and lateral hair cells (HCs) based on centriole movements.
(A) Schematic of E13.5 utricle. The line of polarity reversal (LPR) is not apparent (dotted line) since HCs are mostly absent in the Emx2-positive lateral region (green) at this age. In the medial utricle, while some HCs (Md HCs) are polarized showing the kinocilium located asymmetrically at the lateral region, others are immature and unpolarized with the kinocilium located at the center. (B) The mother centriole (MC)/kinocilium in a lateral HC (Lat HC) could migrate directly towards the medial side (black arrow, #1) or it could first migrate toward the lateral side before reversing its direction to the medial side (red arrows, #2). (C) Schematic drawing and images of an Atoh1Cre; RosatdT/+; CAG:GFP-Centrin2 utricular explant at E13.5 showing the location of Md HC1 (small white rectangle). (D–E) Centriole trajectories (D, D’) and selected time frames of apical views (E) of MC (yellow) and daughter centriole (DC) (orange) in Md HC1 from time-lapse recording (Figure 2—source data 1). (D) Yellow and orange triangles represent the beginning positions of respective MC and DC in each time period, and the circled dots represent the final positions in each time period. The yellow and orange lines represent trajectories of the respective MC and DC and are plotted relative to the center of the HC, which is represented by the centroid of the graph. Each small grid is 1.25 × 1.25 μm. The apical surface of the HC is shown in light grey and the white rim marks the width of the HC body (data extracted using Imaris software). All subsequent live-imaging graphs are organized in a similar manner. Initially, the DC is moving vigorously around the MC (Phase I), then the DC starts to move towards the periphery, which is followed by the MC (Phase II). (D’) Selected temporal trajectories (color-coded) of centrioles in Md HC1 from the end of Phase I to the beginning of Phase II (8:20-11:40 hr), showing the initial movements of DC toward the periphery ahead of the MC. The pink line indicates the edge of the apical HC surface. (E) In Phase I (blue bar), the basal body/MC (yellow arrowhead) is located at the center of the HC, whereas the DC (orange arrowhead) moves around the MC. In Phase II (pink bar), the DC starts to migrate toward the lateral periphery of the HC (#4-#5). This migration is followed by the MC (#6-#7). Then, both centrioles move towards the center as a pair (#8-#10). The white arrow represents the direction where the hair bundles should be pointing in this region of the utricle. (F–H) Lat HC1. (F) Time frames of the blue rectangle area in (C) at 0 hr, 16 hr and 40.8 hr of the recording showing the gradual appearance of tdTomato signals in the lateral utricle and the location of the Lat HC1. Arrows represent hair bundle orientation. (G–H) Total (G) and selected temporal (G’) trajectories as well as selected time frames of the MC and DC (H) in the Lat HC1 (Figure 2—source data 1). TdTomato signal in Lat HC1 was not detectable until 21 hr into the recording (F, H#1–3), which made it difficult to identify the center of the HC. Therefore, the position of the MC (yellow triangle) was used as a proxy for the center of the HC (asterisk) for #1 in (G) until the center of HC can be determined in #2–4 in (G). Lat HC1 shows similar centriole movements as Md HC1 with DC precedes MC to the periphery (G’). AC, anterior crista; LC, lateral crista; Md HC, medial utricular HC; Lat HC, lateral utricular HC. Scale bars: 100 μm (low magnification) and 30 μm (high magnification) in (C), 30 μm in (F), 3 μm in (E) and (H).
-
Figure 2—source data 1
Coordinates of centriole positions relative to the center of the HC or the MC in Md HC1 and Lat HC1.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig2-data1-v2.xlsx
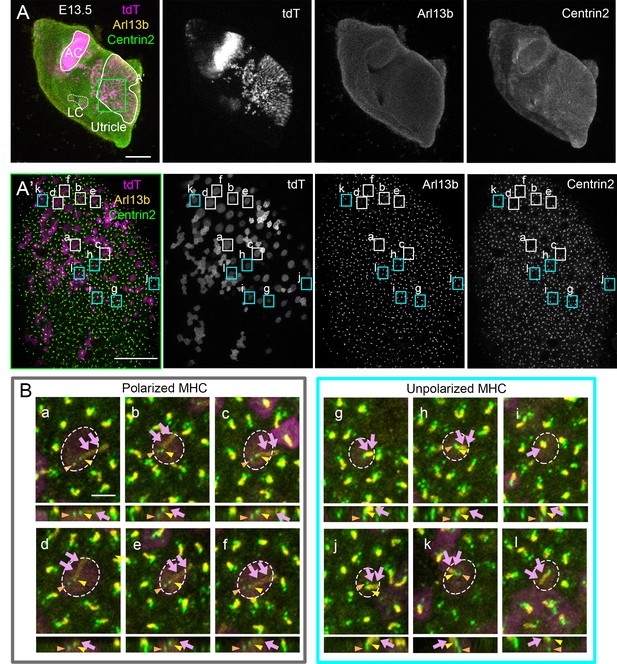
Identification of mother and daughter centrioles in polarized and unpolarized medial utricular hair cells (Md HCs).
(A, A’) A low (A) and high (A’) magnification of Atoh1Cre; RosatdT/+; CAG:GFP-Centrin2 utricular explant that was immuno-stained with anti-Arl13b antibodies (yellow) at E13.5. (B) Representative apical and side views of polarized (a–f) and unpolarized (g–l) HCs in (A’). From apical views (upper panels), the mother centriole (MC) (yellow arrowhead) is associated with the Arl13b-positive kinocilium (magenta arrows) and not the daughter centriole (DC) (orange arrowhead) in both polarized and unpolarized HCs. From side views (lower panels), the MC is positioned closer to the apical surface of the HC than the DC. Notably, the DC position is consistently more lateral than the MC in polarized HCs (a–f) but its position relative to the MC is variable in unpolarized HCs (g–l). Scale bars: 100 μm in (A), 30 μm in (A’) and 3 μm in (a), which applies to (b–l).
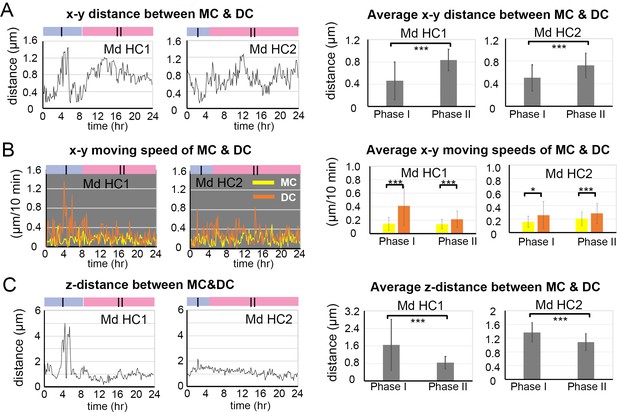
Traveled distance and speed between the mother centriole (MC) and daughter centriole (DC) in medial utricular hair cells (Md HCs).
(A) The x-y distance between MC and DC over time and their average distances in the two phases. Md HC1: number of time points measured (n) equal 53 and 92 for Phase I and II, Md HC2: n = 30 and 115 for Phase I and II. (B) The x-y moving speed of MC and DC in each 10 min time frame and their averages in each phase. Md HC1: number of time points measured (n) equal 52 and 91 for Phase I and II, Md HC2: n = 29 and 114 for Phase I and II. (C) The z-distance traveled between MC and DC over time and their average distances in the two phases. Md HC1: number of time points measured (n) equal 53 and 92 for Phase I and II, Md HC2: n = 30 and 115 for Phase I and II. Error bars represent standard deviation, SD. *p<0.05, ***p<0.001.
-
Figure 2—figure supplement 2—source data 1
The x-y moving speeds of centrioles in Md HC1 and Md HC2 during centriole migration.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig2-figsupp2-data1-v2.xlsx
-
Figure 2—figure supplement 2—source data 2
The x-y distance between centrioles in Md HC1 and Md HC2 during centriole migration.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig2-figsupp2-data2-v2.xlsx
-
Figure 2—figure supplement 2—source data 3
The z-distance between centrioles in Md HC1 and Md HC2 during centriole migration.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig2-figsupp2-data3-v2.xlsx
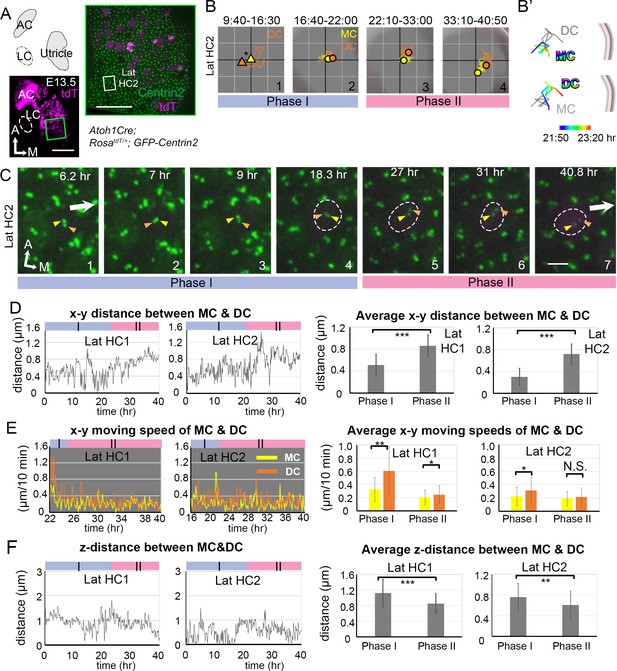
Live imaging of hair bundle establishment in lateral utricular hair cells (Lat HCs) based on centriole movements.
(A) Schematic drawing and images of an Atoh1Cre; RosatdT/+; CAG:GFP-Centrin2 utricular explant at E13.5. (B) The trajectory of the mother centriole (MC) and daughter centriole (DC) in Lat HC2. The position of the MC (yellow triangle) is used as a proxy for the center of the HC (asterisk) for #1, because of the weak tdTomato signal. For #2–4, the center of the graph represents the apical center of the HC, as described in Figure 2. The DC (orange) initially shows rapid movements around the MC in Phase I (#1–2). Then, the DC (orange) starts to move medially, which is followed by the MC in Phase II (#3–4). (B’) The temporal trajectories (color coded) of centrioles in Lat HC2 from the end of Phase I to the beginning of Phase II (8:20-11:40 hr). During this time, the DC moves from the center towards the periphery, while the MC stays at relatively similar positions in the center. (C) Selected frames from a time-lapse recording of centriole movements showing apical views of Lat HC2. In Phase I (#1–4), the DC moves around the MC, when tdTomato signal is not apparent. At the end of Phase I (#4), the tdTomato signal is detectable and it continues to increase in Phase II. During Phase II (#5–7), the DC starts to migrate towards the medial periphery of the HC, which is followed by the MC. (D) The x-y distance between MC and DC over time and their average distances in each phase. Lat HC1: number of time points measured (n) equal 30 and 100 for Phase I and II, Lat HC2: n = 30 and 113 for Phase I and II. (E) The moving speed of MC and DC in each 10 min time frame and their averages in each phase. Lat HC1: number of speeds measured (n) equal 18 and 99 for Phase I and II, Lat HC2: n = 34 and 112 for Phase I and II. (F) The z-distance between MC and DC over time and their average distances in each phase. Lat HC1: number of time points measured (n) equal 30 and 100 for Phase I and II, Lat HC2: n = 30 and 113 for Phase I and II. Error bars represent SD in (D), (E) and (F). *p<0.05, **p<0.01, ***p<0.001.
-
Figure 2—figure supplement 3—source data 1
Coordinates of centriole positions relative to the center of the hair cells (HC) or the mother centriole (MC) in Lat HC2.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig2-figsupp3-data1-v2.xlsx
-
Figure 2—figure supplement 3—source data 2
The x-y distance between centrioles in Lat HC1 and Lat HC2.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig2-figsupp3-data2-v2.xlsx
-
Figure 2—figure supplement 3—source data 3
The z distance between centrioles during migration in Lat HC1 and Lat HC2.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig2-figsupp3-data3-v2.xlsx
-
Figure 2—figure supplement 3—source data 4
The x-y moving speeds of centrioles during migration in Lat HC1 and Lat HC2.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig2-figsupp3-data4-v2.xlsx
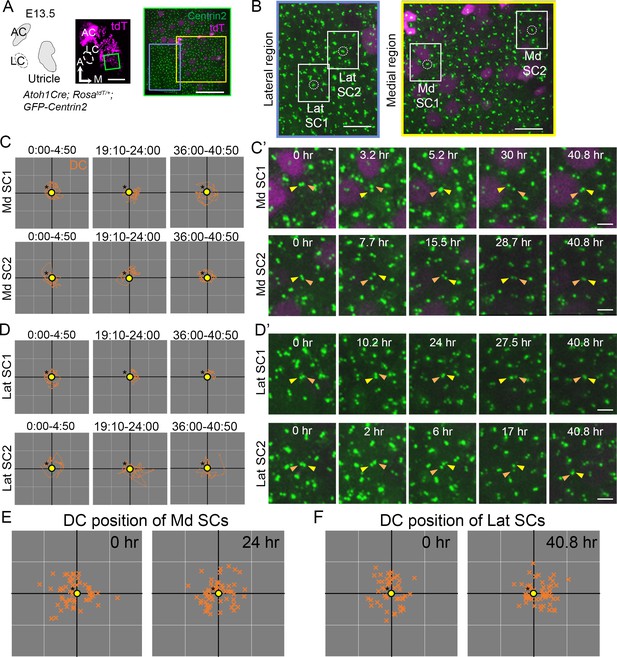
Absence of polarizing centriole trajectories in medial and lateral supporting cells.
(A) Schematic drawing and images of an Atoh1Cre; RosatdT/+; CAG:GFP-Centrin2 utricular explant at E13.5. (B) Magnified images of yellow (Medial) and light blue (Lateral) rectangles in A. (C, C’) Selected trajectories (C) and frames (C’) from a time-lapse recording of centriole movements in Md SC1 and Md SC2, which are tdTomato-negative. The daughter centriole (DC) moves around the mother centriole (MC) throughout the 40 hr of recording without a directional trajectory. (D, D’) Selected trajectories (D) and frames (D’) from a time-lapse recording of centriole movements in Lat SC1 and Lat SC2. (C) and (D) show trajectories of the DC relative to the MC (yellow circle with asterisk) without a specific directional advancement. (E, F) Relative positions of each DC compared to the MC at 0 hr and 24 hr of the recording in Md SCs (E, n = 55 at 0 hr, n = 55 at 24 hr) and at 0 and 40.8 hr of recording in Lat SCs (n = 46 at 0 hr, n = 55 at 40.8 hr). The position of the DC in relationship to the MC is inconsistent among SCs. Scale bars: 100 μm (low magnification) and 30 μm (high magnification) in (A), 10 μm in (B), 3 μm in (C’) and (D’).
-
Figure 2—figure supplement 4—source data 1
Coordinates of daughter centriole (DC) positions relative to the mother centriole (MC) in Md and Lat supporting cells.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig2-figsupp4-data1-v2.xlsx
-
Figure 2—figure supplement 4—source data 2
Relative positions of the daughter centriole (DC) compared to the mother centriole (MC) in Md and Lat supporting cells at different time points.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig2-figsupp4-data2-v2.xlsx
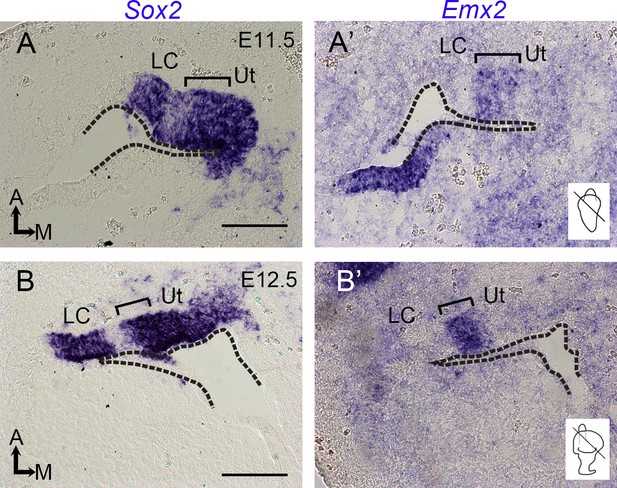
Expression of Emx2 in the developing utricle.
(A, A’, B, B’) In situ hybridization of adjacent sections at the levels of the utricle (Ut) and lateral crista (LC) at E11.5 (A, A’) and E12.5 (B, B’). The levels of sections are indicated in the inner ear schematics. Sensory tissues of the lateral crista and utricle are Sox2-positive, and the bracket indicates the Emx2-positive domain in the utricle, which includes the lateral sensory region. Dotted lines indicate the apical margin of the otic epithelium. Scale bars: 50 μm.
Md HC1 and Md HC2.
Time-lapse videos of the apical view (left panel) and the reconstructed image (right panel) of Md HC1 and Md HC2 in Atoh1Cre; RosatdT/+; CAG:GFP-Centrin2 utricle showing the daughter centriole (DC) first moves sporadically around the mother centriole (MC). Then, the DC moves toward the lateral side (white arrow). This trajectory is followed by the MC. Yellow and red spheres indicate mother and daughter centrioles, respectively. Magenta indicates tdTomato signal within the HC. Scale bar: 3 μm. Play back speed: 7 frames/s.
Lat HC1 and Lat HC2.
Time-lapse videos of the apical view (left panel) and the reconstructed image (right panel) of Lat HC1 and Lat HC2 in Atoh1Cre; RosatdT/+; CAG:GFP-Centrin2 utricle, showing the daughter centriole (DC) first moves sporadically around the mother centriole (MC). Then, the DC moves toward the medial side (white arrow), which is followed by the MC. Yellow and red spheres indicate mother and daughter centrioles, respectively. Magenta indicates tdTomato signal within the HC. Scale bar: 3 μm. Play back speed: 7 frames/s.
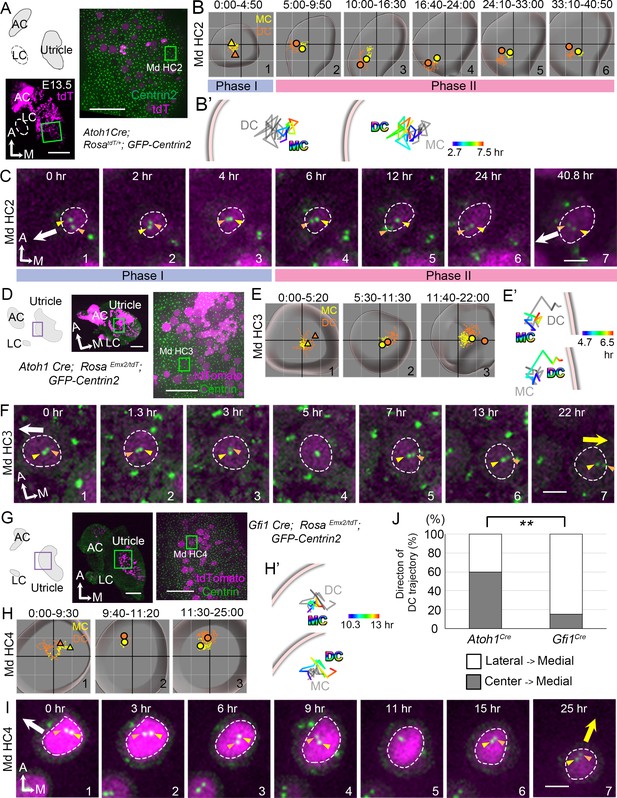
Trajectories of centriole movements in Emx2 gain-of-function Md HCs.
(A–C) Schematic drawing and images of Md HC2 in Atoh1Cre; RosatdT/+; CAG:GFP-Centrin2 control utricle at E13.5 (A). (B–C) Total (B) and temporal (B’) trajectories as well as selected frames (C) from a recording of mother centriole (MC) (yellow) and daughter centriole (DC) (orange) in Md HC2 (Figure 3—source data 1). Trajectory is similar to Md HC1 in Figure 2. Briefly, the DC is moving vigorously around the MC (Phase I), then the DC starts to move toward the periphery, where hair bundles should be pointing in this region of the utricle (white arrow in C). This trajectory is followed by the MC (Phase II). (D–F) Schematic drawing and low- and high-magnification images of Md HC3 in Atoh1Cre; RosaEmx2/tdT; CAG:GFP-Centrin2 utricle at E13.5 (D). Total (E) and selected temporal (E’) trajectories as well as selected apical views (F) of MC (yellow) and DC (orange) in Md HC3 over-expressing Emx2 (Figure 3—source data 1). (F) The DC (orange arrowhead) moves around the MC (yellow arrowhead) in the center of the apical HC surface (#1-4). Then, the DC starts to move toward the medial side (yellow arrow), which is followed by the MC (#5-7). (G–I) Schematic drawing, low- and high-magnification images of Md HC4 in Gfi1Cre; RosaEmx2/tdT; CAG:GFP-Centrin2 utricular explant (G). Total (H) and selected temporal (H’) trajectories as well as selected apical views (I) of the MC and DC in Md HC4 over-expressing Emx2 (Figure 3—source data 1). (H) Between 0:00 and 9:30 hr (#1), the centrioles are migrating toward the lateral side of the HC with the DC more lateral than the MC. In #2 (9:40-11:20 hr), the DC starts to change its position to the medial side of the MC, which becomes more apparent in #3 (11:30-25:00 hr). (H’) Temporal trajectories of centrioles in Md HC4 from 10:20 to 13:00 hr showing the DC in the periphery moving medial to the MC toward the center of the HC. (I) In panels 1–4, the DC (orange arrowhead) is heading toward the lateral side of the utricle (white arrow), then it changes course and moves to the medial side of MC (panels 6–7, yellow arrow). (J) Percentages of DC with two different trajectories in HCs ectopically expressing Emx2 using either Atoh1cre or Gfi1cre. Total number of HCs analyzed: Atoh1cre, n = 15; Gfi1cre, n = 39 (Figure 3—source data 2). Scale bars: 100 μm (low mag) and 30 μm (high mag) in (A), (D) and (G), 3 μm in (C), (F) and (I). **p<0.01.
-
Figure 3—source data 1
Coordinates of centriole positions relative to the center of the cell in Md HC2, Md HC3, and Md HC4.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig3-data1-v2.xlsx
-
Figure 3—source data 2
Quantification of daughter centriole trajectories in Atoh1Cre; RosaEmx2/tdT; CAG:GFP-Centrin2 and Gfi1Cre; RosaEmx2/tdT; CAG:GFP-Centrin2 utricles.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig3-data2-v2.xlsx
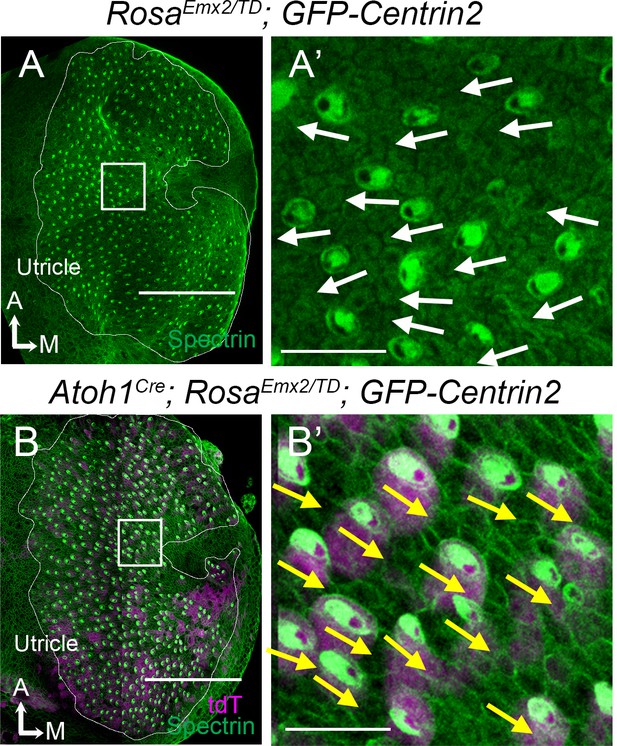
Gain-of-function of Emx2 reverses hair bundle orientation in Md HCs of Atoh1Cre; RosaEmx2/tdT; CAG:GFP-Centrin2 utricles.
(A) Low-magnification image of a RosaEmx2/tdT; CAG:GFP-Centrin2 utricle at E15.5. (A’) High-magnification image of the rectangular area in (A), showing normal lateral-oriented hair bundles in the medial utricle (white arrows) based on β2-spectrin immunostaining (green color). The void of β2-spectrin staining indicates the position of the kinocilium and thus the orientation of the hair bundle. (B) Low-magnification image of an Atoh1Cre; RosaEmx2/tdT; CAG:GFP-Centrin2 utricle at E15.5, in which hair cells (HCs) are tdTomato-positive (magenta) and stained with anti-β2-spectrin antibodies (green). (B’) High magnification of the rectangular area in (B) showing that hair bundles in the medial utricle all point towards the medial (yellow arrows) instead of the lateral direction as in controls (A’). The utricle is fan-shaped. Therefore, depending on selected regions, hair bundle orientation is not necessarily consistent among regions but always consistent within a region. Scale bars: 100 μm in (A, B) and 10 μm in (A’, B’).
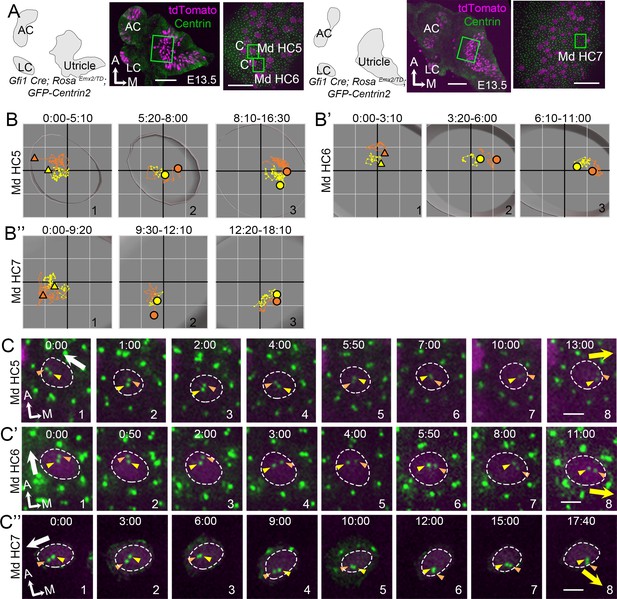
Centriole trajectories in Emx2 gain-of-function medial hair cells (Md HCs) using Gfi1Cre.
(A) Schematic drawings, low- and high-magnification images of two Gfi1Cre; RosaEmx2/tdT; CAG:GFP-Centrin2 utricular explants. (B–B’’) The trajectory of mother centriole (MC) and daughter centriole (DC) in Md HC5 (B), Md HC6 (B’), and Md HC7 (B”). At first, both centrioles are positioned in the lateral side of the HC (B-B", #1). Then, the DC starts to reverse its direction and moves toward the medial side of the HC (B-B", #2), followed by the MC (B-B", #3). Yellow and orange triangles show the initial MC and DC positions in #1, whereas yellow and orange circled dots represent the final positions in Panels #2-#3. Each small grid is 1.25 × 1.25 μm. (C–C’’) Selected apical views of a live-imaging recording of Md HC5 (C), Md HC6 (C’), and Md HC7 (C”) that overexpress Emx2. Initially, the DC (orange arrowhead) is located by the lateral side (white arrow, Md HC5 #1-4, Md HC6 #1–3, Md HC7 #1–4), then it changes course and moves to the medial side of the MC (yellow arrow, Md HC5, #7–8, Md HC6, #4–8, MD HC7 #6–8). Scale bars: 100 μm (low magnification) and 30 μm (high magnification) in (A), and 3 μm in (C), (C’) and (C’’).
-
Figure 3—figure supplement 2—source data 1
Coordinates of centriole positions relative to the center of the HC in Md HC5, Md HC6, and Md HC7.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig3-figsupp2-data1-v2.xlsx
Md HC3 and Md HC4.
Time-lapse videos of the apical view (left panel) and the reconstructed image (right panel) of Md HC3 (Atoh1Cre; RosaEmx2/tdT; CAG:GFP-Centrin2) and Md HC4 (Gfi1Cre; RosaEmx2/tdT; CAG:GFP-Centrin2). The daughter centriole (DC) in MHC3 first moves sporadically around the mother centriole (MC), similar to other MHCs (Figure 2—video 1). Then, the DC moves toward the medial side (yellow arrow) in an opposite direction from normal Md HCs (white arrow). This movement is followed by the MC. The DC in Md HC4 first leads the MC moving toward the lateral side (white arrow) and then it changes course and moves medial to the MC toward the medial side (yellow arrow). Yellow and red spheres indicate MC and DC, respectively. Magenta indicates tdTomato signal within the HC. Scale bar: 3 μm. Play back speed: 7 frames/s.
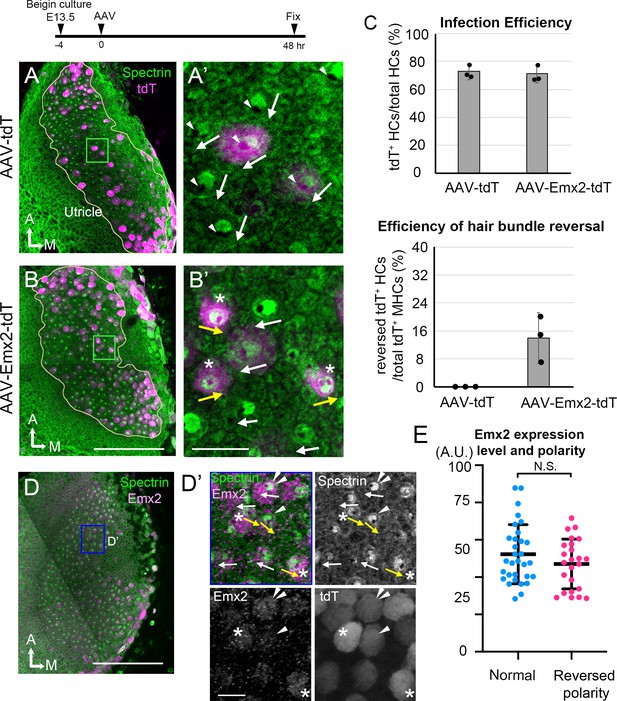
AAV-Emx2 infection alters hair bundle orientation in medial hair cells (Md HCs).
(A, A’) Low- (A) and high-magnification (A’) images of the rectangular area of a control explant infected with AAV-tdT (tdT: magenta) and stained with anti-β2-spectrin antibodies (green), in which the absence of staining indicates the kinocilium location (arrowhead) and HC orientation (arrow). The two AAV-tdT infected HCs (magenta) show hair bundle orientation similar to non-infected HCs. (B,B’) Low- (B) and high-magnification (B’) images of an utricular explant infected with AAV-Emx2-tdT (tdT: magenta), stained with anti-β2-spectrin antibodies (green). (B’) Some of the AAV-Emx2-tdT infected Md HCs show opposite hair bundle orientation (yellow arrow and asterisk) from the rest of the non-infected HCs (white arrows). (C) Efficiency of viral infections and efficiency of hair bundle reversal among infected HCs (n = 3 experiments for each condition). (D–D’) Low (D) and high magnification (D’) images of an utricular explant infected with AAV-Emx2-tdT and stained with anti-Emx2 (magenta) and β2-spectrin antibodies (green). (D’) Four HCs with normal bundle orientation (white arrows) showing variable Emx2 staining including, one with strong Emx2 expression (white arrow with double arrowheads). The three HCs with abnormal bundle orientation (yellow arrows pointing towards the medial) show one with weak (arrowhead) and two with strong (asterisks) Emx2 expression. (E) Quantification of Emx2 immunoreactivity among infected HCs with normal (n = 30) or reversed bundle orientation (n = 23). There is no significant difference in the levels of Emx2 expression between the two groups of HCs. Error bars represent SD. Scale bars: 100 μm in (B) and applies to (A) and 10 μm in (B’) and applies to (A’). 100 μm in (D) and 10 μm in (D’).
-
Figure 4—source data 1
Efficiency of infections and hair bundle reversal phenotype mediated by control (AAV-tdT) and AAV-Emx2-tdT viruses.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig4-data1-v2.xlsx
-
Figure 4—source data 2
Correlation of Emx2 expression levels with hair bundle orientation.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig4-data2-v2.xlsx
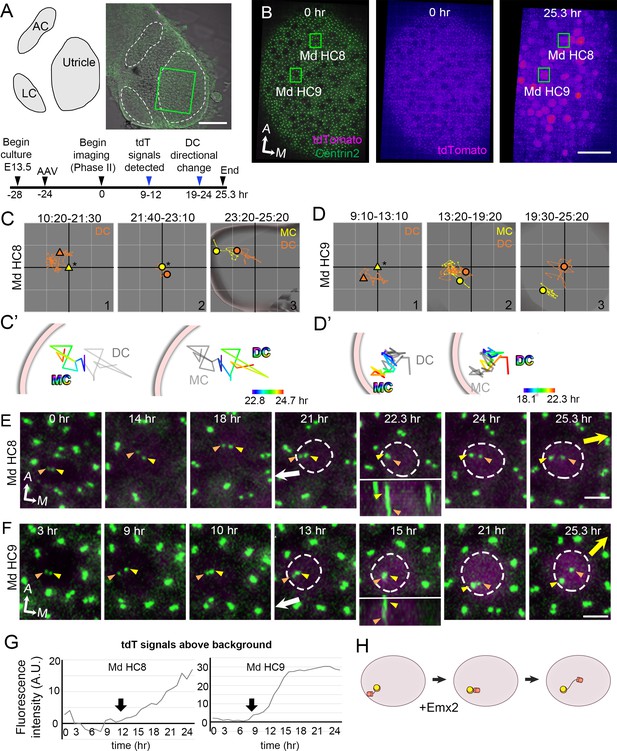
Altered daughter centriole (DC) trajectory in medial hair cells (Md HCs) infected with AAV-Emx2.
(A) Schematic and low-magnification image of the CAG:GFP-Centrin2 utricular explant infected with AAV-Emx2-tdT at E13.5. The timeline of experimental treatments (black arrowheads) and observations (blue arrowheads) are shown. (B) An utricular explant at the beginning (0 hr) and end (25.3 hr) of a time-lapse recording showing clear tdTomato-positive cells by the end of recording. (C, C’, D, D’) Total (C, D) and selected temporal (C’, D’) trajectories of the mother centriole (MC) and DC in Md HC8 and Md HC9 (Figure 5—source data 1). Initially, the DC (orange triangle, #1) is positioned lateral to the MC (yellow triangle) at the center (asterisk). Then, the DC (orange dot) moves sporadically around MC (yellow dot, #2), followed by DC moving medial to the MC in #3. (E, F) Selected frames of the recording of Md HC8 and Md HC9. At the beginning, the DC is located by the lateral side (white arrow) of each HC. Then, the DC overlaps with the MC briefly at 22.3 hr for Md HC8 and 15 hr for Md HC9 during recording (insets showing side views), followed by the DC moving medial to the MC toward the medial direction (yellow arrow). (G) tdTomato expression compared to the background level, indicating tdTomato signals exceeded background after 12 (Md HC8) and 9 hr (Md HC9) of recordings (arrows). (H) Schematic of centriole movements in the presence of Emx2. Scale bars: 100 μm in (A), 30 μm in (B) and 3 μm in (E) and (F).
-
Figure 5—source data 1
Coordinates of centriole positions relative to the center of the hair cell (HC) or the mother centrioles (MCs) in Md HC8 and Md HC9.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig5-data1-v2.xlsx
Md HC8 and Md HC9.
Time-lapse videos of the apical view (left panel) and the reconstructed image (right panel) of Md HC8 and Md HC9 in CAG:GFP-Centrin2 utricular explant infected with AAV2.7m8-CAG-Emx2-P2A-tdTomato. In both Md HC8 and Md HC9, the daughter centriole (DC) starts out on the lateral side of the mother centriole (MC) toward the lateral side of the utricle (white arrow). Then, it changes course to be on the medial side of the MC toward the medial utricle (yellow arrow). Yellow and red spheres indicate MC and DC, respectively. Magenta indicates tdTomato signal in the infected HC cytoplasm. Scale bar: 3 μm. Play back speed: 7 frames/s.
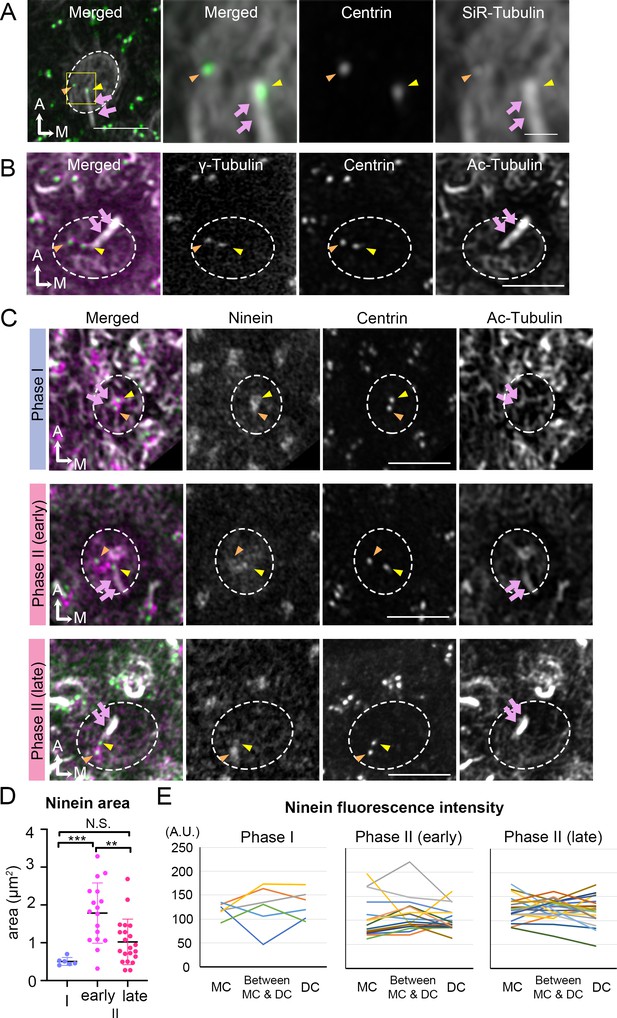
Broaden ninein localization during centriole migration.
(A) SiR-tubulin (white) labeling of a medial hair cell (Md HC) at Phase II from an Atoh1Cre; RosatdT/+; CAG:GFP-Centrin2 utricule at E13.5. The magnified views of the yellow rectangular area in the left panel are shown in the three right panels, illustrating that both centrioles (green in the merged picture) are associated with the microtubule network. MC, yellow arrowhead; DC, orange arrowhead; kinocilium, pink arrows. Dotted white lines indicate apical surface of the HC. (B) Anti-γ-tubulin staining of a Md HC at Phase II. Daughter (DC) and mother centrioles (MC) show similar expression of γ-tubulin (magenta in the merged picture). Anti-acetylated tubulin antibody labels mature microtubules and the kinocilium. (C) Immunostaining of ninein in Md HCs in Phase I, early and late Phase II. Phase I and late Phase II HCs show centrosomal ninein (magenta on merged picture) staining. At the beginning of Phase II, ninein staining is diffuse and broader than the centrioles. (D) Distribution of the ninein-positive area (μm2) during Phase I, early and late Phase II HCs. (E) Ninein fluorescence intensity associated with the MC, DC and the region in-between the two centrioles during Phase I, early and late Phase II HCs (see Materials and methods). No consistency of ninein staining associated with a specific centriole or region was observed among various samples (numbers of HCs for Phase I, Phase II early and late are 6, 17, and 21, respectively). Scale bars: bar in left panel of (A) equals 3 μm, bar in the right panel of (A) equals 1 μm and applies to the two adjacent panels on its left, bars on the fourth and third panels of respective (B) and (C) equal 3 μm and apply to other panels in (B–C). **p<0.01, ***p<0.001.
-
Figure 6—source data 1
Quantification of ninein area in hair cells (HCs) during each phase of centriole migration.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig6-data1-v2.xlsx
-
Figure 6—source data 2
Anti-ninein staining intensity associated with the mother centriole (MC), daughter centriole (DC) and area in between the centrioles.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig6-data2-v2.xlsx
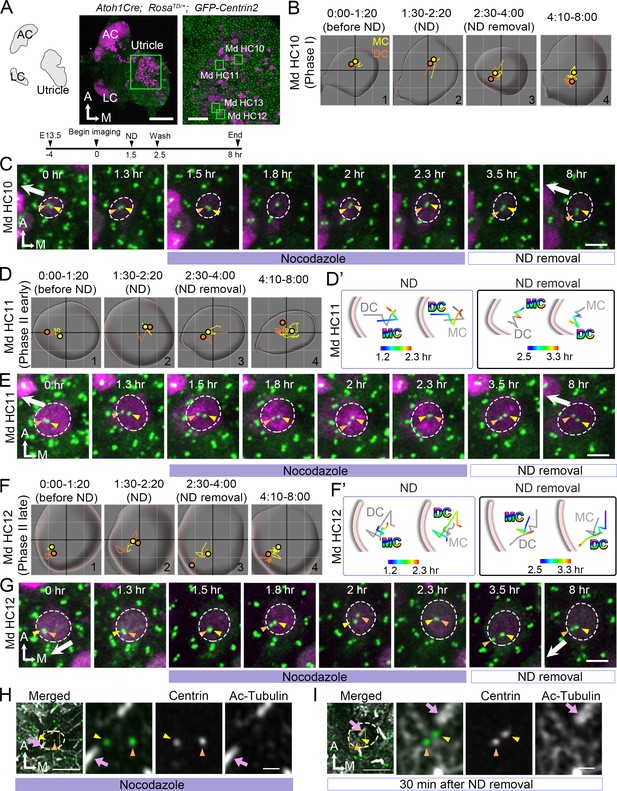
Nocodazole treatments affect centriole migration and positions.
(A) Schematic, low and high magnifications of an Atoh1Cre; RosatdT/+; CAG:GFP-Centrin2 utricular explant at E13.5. The timeline of the experiment is shown. (B, C) Md HC10 at Phase I when centrioles are at the center of the hair cell (HC). (B) Total trajectories as well as (C) selected frames of mother centriole (MC) (yellow) and daughter centriole (DC) (orange) in Md HC10 (Figure 7—source data 1). MC and DC remained in the center during nocodazole treatment (B#2, C). After nocodazole removal, the DC started to move to the periphery (B#3,4, C, n = 3 out of 6 Phase I HCs). (D–E) Md HC11 at the beginning of Phase II, in which the DC is located peripherally whereas the MC is at the center. (D) Total, selected temporal (D’) trajectories as well as selected frames (E) of MC and DC in Md HC11 (Figure 7—source data 1). During nocodazole treatment, both DC and MC relocated to the center of the HC (D#2, D’). After drug removal, the DC moved toward the periphery within 1 hr (D’, E, 3.5 hr, n = 11 out of 12 early Phase II HCs). (F–G) Md HC12 at late Phase II in which both centrioles are peripherally positioned. (F) Total, selected temporal (F’) trajectories as well as selected frames (G) of MC and DC in Md HC12 (Figure 7—source data 1). During nocodazole treatment, both centrioles move back to the center of the HC (F#2, F’, G). After removal of nocodazole, centrioles return to the periphery with the DC moving ahead of the MC (F’, n = 17 out of 18 late Phase II HCs). (H) The nocodazole-treated Md HC shows mispositioned centrioles (MC, yellow arrowhead; DC, orange arrowhead) that are no longer asymmetrically located in the periphery. The three panels on the right are merged, single centrin and acetylated tubulin images of the rectangular area in the left panel. Tubulin arrays are absent in the cytoplasm of HC. (I) A Md HC after removal of nocodazole for 30 min shows centrioles returning to their peripheral location. The three panels on the right are magnifications of the rectangular area in the left panel. Tubulin arrays (white) are radiating from the centrioles to the periphery Scale bars: 100 μm (low magnification) and 30 μm (high magnification) in (A), 3 μm in (C), (E), (G), and the first panel in (H) and (I), and 1 μm in high magnification of (H) and (I).
-
Figure 7—source data 1
Coordinates of centriole positions relative to the center of the cell for Md HC10, MD HC11 and Md HC12.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig7-data1-v2.xlsx
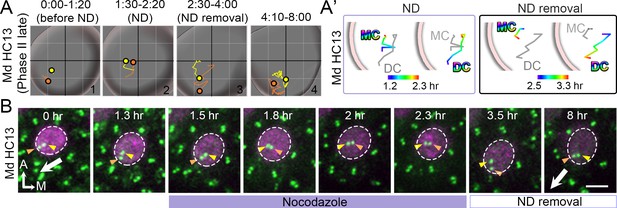
A late Phase II HC treated with nocodazole.
(A, A’, B) Total (A) and selected temporal (A’) trajectories as well as selected frames (B) of MC and DC in Md HC13 at late Phase II (Figure 7—figure supplement 1—source data 1). Before the introduction of nocodazole (ND, 0:00-1:20 hr), the DC is located more peripherally than the MC, towards the lateral edge of HC (A#1, B). During nocodazole treatment, both DC and MC moved back to the center of the HC (A’). After nocodazole removal, centrioles return to the periphery with the DC moving ahead of the MC (A’, B). Scale bar: 3 μm in (B).
-
Figure 7—figure supplement 1—source data 1
Coordinates of centriole positions relative to the center of Md HC13.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig7-figsupp1-data1-v2.xlsx
MD HC12 and MD HC13 - nocodazole treatment and removal.
Time-lapse videos of the apical view (left panel) and the reconstructed image (right panel) of Md HC12 and Md HC13 in Atoh1Cre; RosatdT/+; CAG:GFP-Centrin2 utricle, showing that the introduction of nocodazole causes the lateral-positioned centrioles (white arrow) to spring back to the center of the hair cell (HC) and then they return to the peripheral position after nocodazole removal. Yellow and red spheres indicate mother and daughter centrioles, respectively. Magenta indicates tdTomato signal in the HC cytoplasm. Scale bar: 3 μm. Play back speed: 3 frames/s.
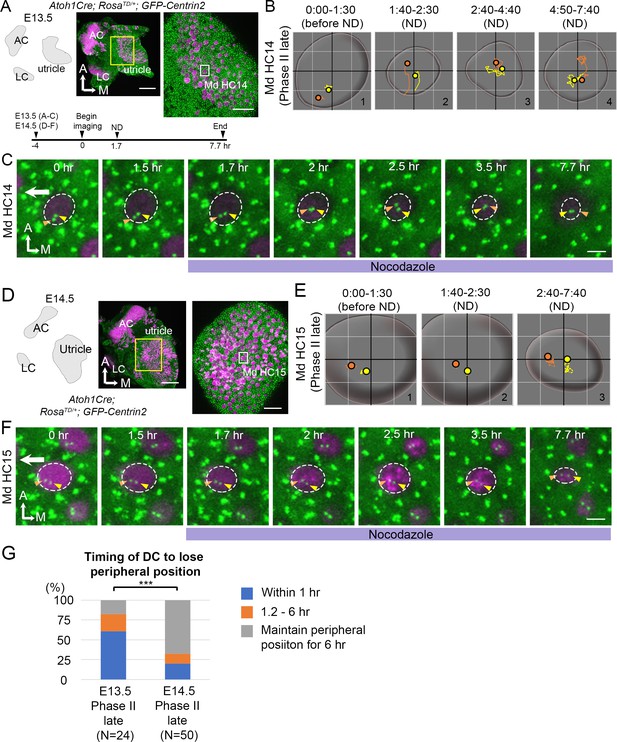
Mature HCs are less sensitive to the microtubule disruption.
(A) Schematic, low and high magnifications of an Atoh1Cre; RosatdT/+; CAG:GFP-Centrin2 utricular explant at E13.5. The timeline of the experiment is shown. (B) Total trajectories as well as (C) selected frames of MC (yellow) and DC (orange) in Md HC14 at late Phase II (Figure 8—source data 1). During nocodazole treatment, the centrioles move back to the center of the HC within 30 min (B#2, C 2 hr). Then, both centrioles moved around in the center of the HC in variable relative positions from each other till the end of the experiment (B#2–4, C 2.5–7.7 hr). (D) Schematic, low and high magnifications of an Atoh1Cre; RosatdT/+; CAG:GFP-Centrin2 utricular explant at E14.5. (E–F) Total trajectories (E) as well as selected frames (F) of MC and DC in Md HC15 at late Phase II (Figure 8—source data 1). The peripheral location of the centrioles and the relative relationship of DC lateral to the MC were maintained until the end of recording (E#2-#3, F 1.5–7.7 hr). The surface of the HC appeared reduced in size by the end of the experiment (E and F, 7.7 hr). (G) Timing of DC to lose the peripheral position. At E13.5, 60.7% (17 out of 28 HCs) of late Phase II HCs lose their peripheral positions of DCs within 1 hr-treatment of ND, whereas, at E14.5, it reduces to 20% (10 out of 50, p=0.000089, Figure 8—source data 2). Scale bars: 100 μm (low magnification) and 30 μm (high magnification) in (A) and (D), 3 μm in (C) and (F). ***p<0.001.
-
Figure 8—source data 1
Coordinates of centriole positions relative to the center of the cell for Md HC14 and MD HC15.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig8-data1-v2.xlsx
-
Figure 8—source data 2
Timing of the daughter centriole (DC) to lose its peripheral position under nocodazole treatments.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig8-data2-v2.xlsx
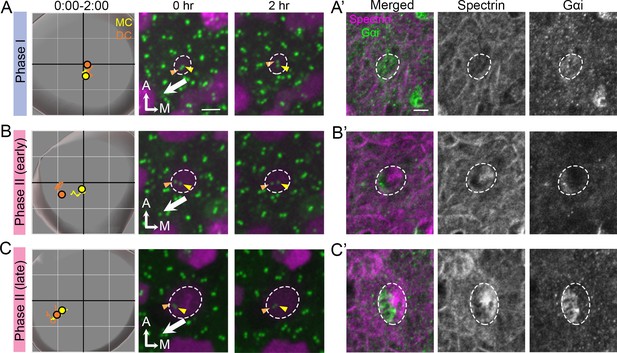
Correlation of centriole trajectory with Gαi and β2-spectrin accumulation in nascent HCs.
Trajectories and selected frames of typical Phase I (A), early (B) and late Phase II (C) Md HCs during 2 hr live imaging (Figure 9—source data 1). (A’, B’, C’) Immunostaining of β2-spectrin and Gαi in the same HCs as shown in A, B and C, after live imaging. (A’) At phase I, β2-spectrin signal is not detectable, and Gαi is diffusely expressed in the apical HC surface. (B’) At early Phase II, Gαi starts to accumulate in the lateral periphery and β2-spectrin signal is located in the medial side. (C’) At late Phase II, both Gαi and β2-spectrin signals become more apparent and complementary to each other. Scale bar of (A) equals 3 μm and apply to the other panels in (A–C). Scale bar on the first panel of (A’) equals 3 μm and apply to the other panels in (A’–C’).
-
Figure 9—source data 1
Coordinates of centriole positions during the 2 hr of live imaging.
- https://cdn.elifesciences.org/articles/59282/elife-59282-fig9-data1-v2.xlsx
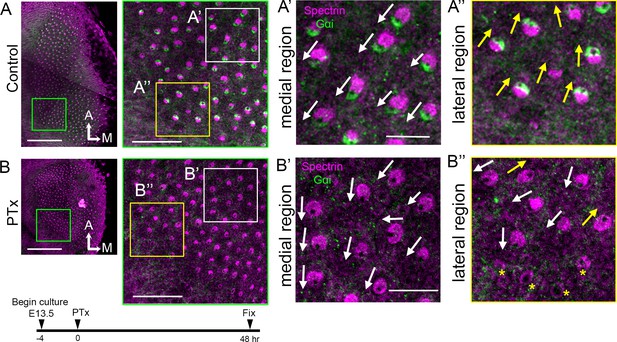
Pertussis toxin treatments affect hair bundle orientation in the lateral but not medial utricular hair cells.
(A,B) E13.5 utricular cultures treated with media (A, control) or media containing pertussis toxin (B, PTx) for 2 days. (A’, A’’) Magnified images of the medial (A’) and lateral (A’’) region of the control in A, showing the organized hair bundle orientation. White and yellow arrows indicate hair bundles pointing toward the lateral and medial direction, respectively. Gαi staining is associated with the kinocilium location, which is devoid of β2-spectrin staining. (B) Lower and higher magnifications of the utricle treated with PTx at 8.4 nM. (B’, B’’) Magnified images of a medial (B’) and lateral (B’’) region of B. (B’, B”) Gαi staining is lost with PTx treatment, but hair bundle polarity in the medial region is maintained (B’), but many HCs in the lateral utricle show abnormal hair bundle orientation (B", white arrows and yellow asterisks). Scale bars: 100 μm (low magnification) and 30 μm (high magnification) in (A) and (B), 10 μm in (A’) and (B’) that apply to (A’’) and (B’’).
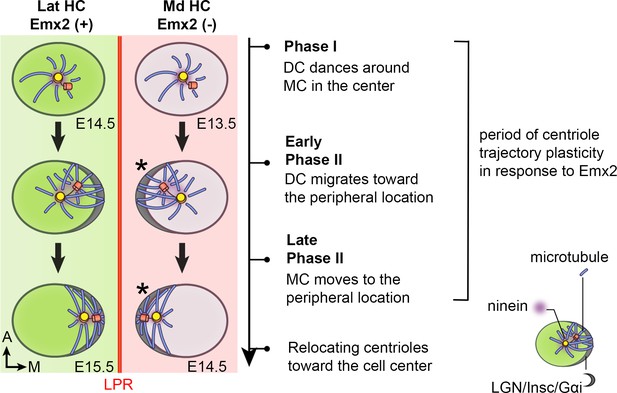
Summary of centriole trajectory during hair bundle establishment in nascent utricular HCs.
In Emx2-negative Md HCs, Phase I is represented by MC located in the apical center of HCs with the DC dancing around the MC. Ninein is associated with the centrioles, which serves as the nucleation center for microtubules. In early Phase II, when DC migrates toward the peripheral side, the MC starts to follow the direction of the DC and the peripheral crescent LGN/Insc/Gαi complex starts to be established (Ezan et al., 2013; Tarchini et al., 2013). The broad distribution of ninein surrounding both centrioles is likely to anchor the minus end of microtubules and facilitates centriole migration to the periphery. By the end of Phase II, both MC and DC are located in the periphery and ninein becomes restricted to the centrioles again. Then, centrioles are relocalized towards the cell center by the bare zone (Tarchini et al., 2013). Cuticular plate starts to form at the beginning of Phase II based on β2-spectrin staining (not drawn). Centriole trajectory in both Phase I and early Phase II are reversible in the presence of Emx2 but responsiveness to Emx2 decreases over time (Jiang et al., 2017). Emx2-positive Lat HCs (green) show similar but opposite trajectory pattern of centriolar migration. Nevertheless, LGN/Insc/Gαi may be dispensable for hair bundle establishment in Md HCs since blocking Gαi with pertussis toxin does not appear to affect bundle orientation (asterisk).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (M. musculus) | CAG:GFP-Centrin2 | PMID:21752934 | RRID:MGI:3793421 | Xiaowei Lu (University of Virginia) |
| Genetic reagent (M. musculus) | Atoh1-Cre | PMID:19609565 | RRID:MGI:3775845 | Bernd Fritzsch (University of Iowa) |
| Genetic reagent (M. musculus) | Gfi1-Cre | PMID:20533399 | RRID:MGI:4430258 | Lin Gan (Augusta University) |
| Genetic reagent (M. musculus) | Rosa26REmx2 | PMID:28266911 | ||
| Genetic reagent (AAV-Virus) | AAV2.7m8-CAG-Emx2-P2A-tdTomato | Vector Biolabs | 4.0 × 1010 GC in 100 µl culture medium | |
| Genetic reagent (AAV-Virus) | AAV2.7m8-CAG-tdTomato | Vector Biolabs | 4.0 × 1010 GC in 100 µl culture medium | |
| Chemical compound, drug | nocodazole | Sigma-Aldrich | M1404 | 5 μM |
| Chemical compound, drug | pertussis toxin | Millipore Sigma | 516560–50 UG | 8.4 nM |
| Chemical compound, drug | SiR-tubulin | Spirochrome | SC002 | 1 μM |
| Antibody | Mouse anti-βII spectrin | BD Biosciences | Cat# 612562, RRID:AB_399853 | IHC (1:500) |
| Antibody | Rabbit anti-Arl13b | Proteintech | Cat# 17711–1-AP, RRID:AB_2060867 | IHC (1:500) |
| Antibody | Mouse anti-acetylated tubulin | Sigma-Aldrich | Cat# T7451, RRID:AB_609894 | IHC (1:500) |
| Antibody | Rabbit anti-ninein | Abcam | Cat# Ab231181 | IHC (1:500) |
| Antibody | Rabbit anti-γ-tubulin | Sigma Aldrich | Cat# ab4447, RRID:AB_304460 | IHC (1:500) |
| Antibody | Rabbit anti-Emx2 | Trans Genic | Cat# KO609 | IHC (1:250) |
| Antibody | Rabbit anti-Gαi | Provided by B. Nurnberg | IHC (1:1000) | |
| Antibody | Alexa Fluor 488 donkey anti-mouse IgG | Thermo Fisher Scientific | Cat# A-21202 RRID:AB_2535788 | IHC (1:500) |
| Antibody | Alexa Fluor 647 donkey anti-mouse IgG | Thermo Fisher Scientific | Cat# A-31571 RRID:AB_162542 | IHC (1:500) |
| Antibody | Alexa Fluor 405 Donkey anti rabbit IgG | Abcam | ab175651 | IHC (1:500) |
| Antibody | Alexa Fluor donkey anti-rabbit 647 | Thermo Fisher Scientific | Cat# A-31573 RRID:AB_2536183 | IHC (1:500) |
| Software, algorithm | ImageJ | RRID:SCR_003070 | ||
| Software, algorithm | Imaris 9.5.0 | Bitplane | RRID:SCR_007370 |





