Dot1l interacts with Zc3h10 to activate Ucp1 and other thermogenic genes
Figures
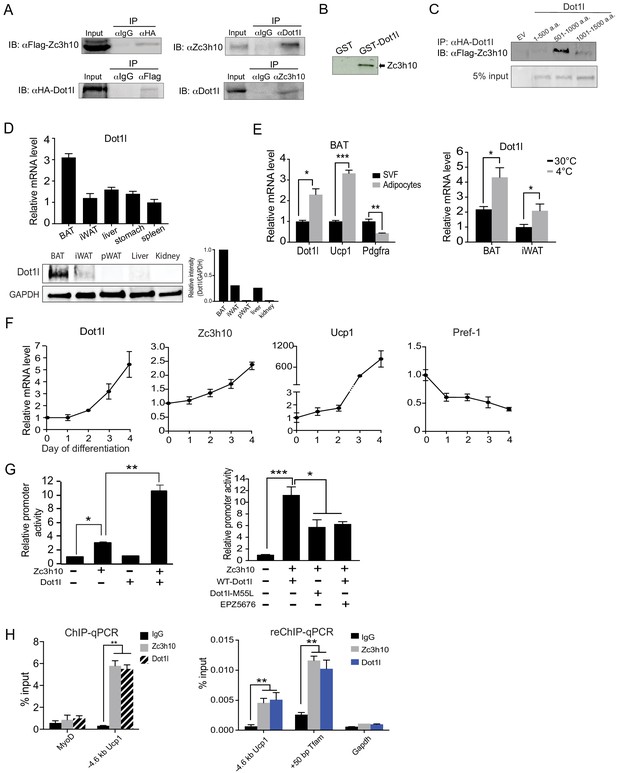
Dot1l directly interacts with Zc3h10 for its recruitment and activation of BAT gene program.
(A) (Left) CoIP using αFlag for Zc3h10 or αHA for Dot1l after immunoprecipitation with either αHA- or αFlag, respectively, using lysates from HEK293FT cells transfected with Flag-Zc3h10 and HA-Dot1l. (Right) CoIP of endogenous Zc3h10 and Dot1l protein using BAT tissue of C57BL/6 using αZc3h10 and αDot1l. (B) Autoradiograph of GST pull-down using GST-Dot1l and 35S-labeled in vitro transcribed/translated Zc3h10. (C) CoIP using αFlag for Zc3h10 after immunoprecipitation with αHA for Dot1l using lysates from HEK293FT cells transfected with Flag-Zc3h10 and various HA-Dot1l constructs. (D) RT-qPCR and western blot analysis of Dot1l in various tissues from 10-week-old C57BL/6 mice (n = 5). (Bottom right) Quantification of intensity of protein bands, calculated by Dot1l/Gapdh. (E) (Left) RT-qPCR for indicated genes in the adipocyte fraction and SVF from BAT. (Right) RT-qPCR for Dot1l mRNA of BAT and iWAT from mice housed at either 30°C or 4°C (n = 5). (F) RT-qPCR for indicated genes during the course of BAT cell differentiation. (G) (Left) HEK293 cells were cotransfected with the −5.5 kb Ucp1-Luc promoter with Zc3h10 or Dot1l either together or individually (n = 5). (Right) Luciferase assay with HEK293 cells transfected with M55L-Dot1l or WT-Dot1l with the −5.5 kb Ucp1-Luc promoter with Zc3h10. (H) (Left) ChIP-qPCR for Zc3h10 and Dot1l enrichment at the −4.6 kb region of Ucp1 promoter using Flag or HA antibodies. Differentiated BAT cells were transduced with either Flag-Zc3h10 or HA-Dot1l. (Right) ReChIP-qPCR for Dot1l and Zc3h10 co-occupancy using chromatin from BAT cells that were transduced with Zc3h10 and/or Dot1l. Data are expressed as means ± standard errors of the means (SEM). *p<0.05, **p<0.01, ***p<0.001.
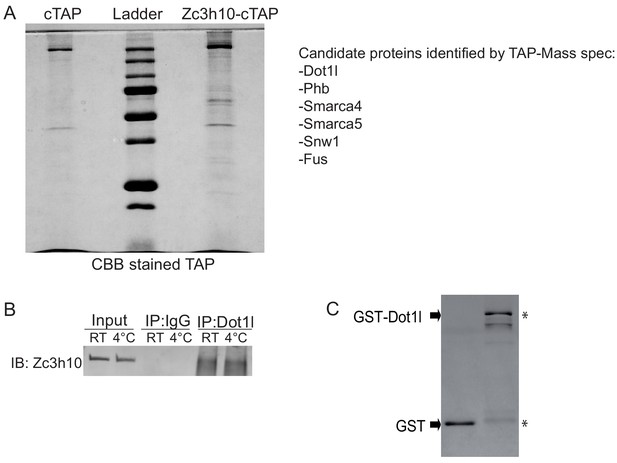
Dot1l directly interacts with Zc3h10 for UCP1 promoter activation.
(A) Gel stained with Coomassie Blue after cTAP purification assay and a list of potential Zc3h10 interacting proteins identified by Mass spec. (B) Immunoblotting for Zc3h10 after immonoprecipitating with αIgG or αDot1l, respectively, using lysates from BAT of WT mouse. (C) Coomassie staining for GST alone and GST-Dot1l fusion protein.
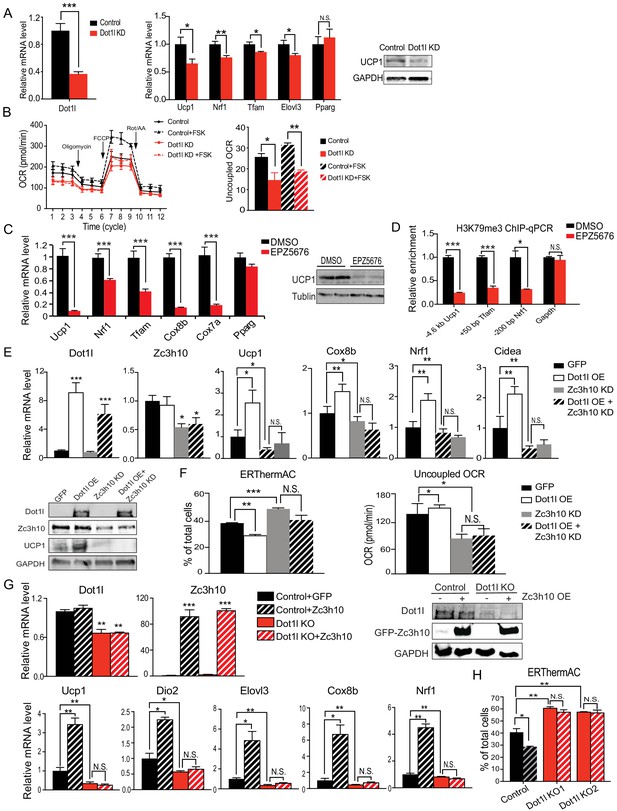
Dot1l is critical for the thermogenic gene program, and its action is dependent on Zc3h10.
(A) (Left) RT-qPCR for Dot1l and thermogenic genes in BAT cells infected either scrambled (Control) or adenovirus expressing short hairpin targeting Dot1l (Dot1l KD) after D2 of adipogenic differentiation (n = 6). (Right) Immunoblotting for Ucp1. (B) (Left) OCR measured in Dot1l KD cells using Seahorse XF24 analyzer (n = 5). (Right) Uncoupled OCR in BAT cells infected with control or shDot1l under oligomycin (0.5 uM). (C) (Left) RT-qPCR for BAT cells treated with Dot1l chemical inhibitor, EPZ5676 (5 nM). (Right) Western blotting analysis for Ucp1 protein. (D) ChIP-qPCR for H3K79me3 at indicating genes using chromatin from differentiated BAT cells overexpressing Dot1l and treated with EPZ5676 (n = 5). (E) (Top) RT-qPCR for indicated genes and (Bottom) immunoblotting for indicated proteins in differentiated BAT cells that were transduced with either AdGFP or AdDot1l or shZc3h10 individually or in combination for overexpression of Dot1l and knockdown of Zc3h10 (n = 6). The differentiated cells were treated with forskolin (10 uM) for 6 hr. (F) (Left) FACS analysis and quantification of ERthermAC, that inversely correlates with heat production. (Right) Uncoupled OCR measured in BAT cells by Seahorse assay (n = 5). (G) RT-qPCR for indicated genes and immunoblotting for indicated proteins in the control BAT cells or Dot1l-CRISPR KO pool, overexpressing either GFP or Zc3h10. (H) FACS analysis and quantification of ERthermAC in Zc3h10 overexpressing in the control or Dot1l-CRISPR KO pools treated with forskolin(10 µM). Data are expressed as means ± standard errors of the means (SEM). *p<0.05, **p<0.01, ***p<0.001.
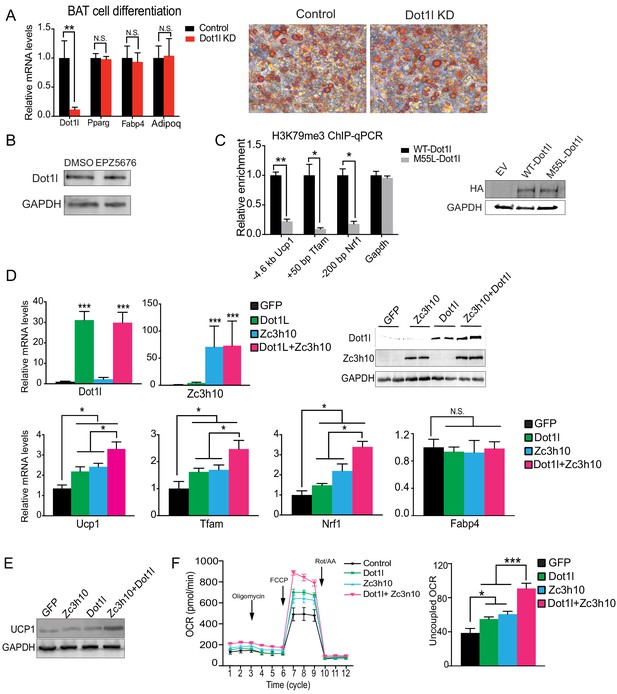
Dot1l promotes the thermogenic gene program in vitro.
(A) RT-qPCR for indiciated genes and Oil Red O staining for lipid accumulation in differentiated BAT cells that were tranduced with shDot1l before differentiation. (B) Immunoblotting for Dot1l in BAT cells treated with Dot1l inhibitor, EPZ5676, for testing Dot1l protein stability. (C) ChIP-qPCR for H3K79m3 in BAT cells transfected with WT-Dot1l and M55L-Dot1l mutant. (D) RT-qPCR for indicated genes and immunoblotting for indicated proteins in 3T3-L1 cells that were transduced with either AdGFP or AdZc3h10 or AdDot1l individually or in combination for overexpression (OE) of Zc3h10 and Dot1l (n = 6). The differentiated cells were treated with forskolin (10 uM) for 6 hr to induce beiging. (E) Immunoblotting for UCP1. (F) (Left) OCR measured in Zc3h10 OE and Dot1l OE, individually and in combination in differentiated 3T3-L1 adipocytes that were induced to beiging (n = 6). (Right) Uncoupled OCR under oligomycin (0.5 uM). Data are expressed as means ± standard errors of the means (SEM). *p<0.05, **p<0.01, ***p<0.001.
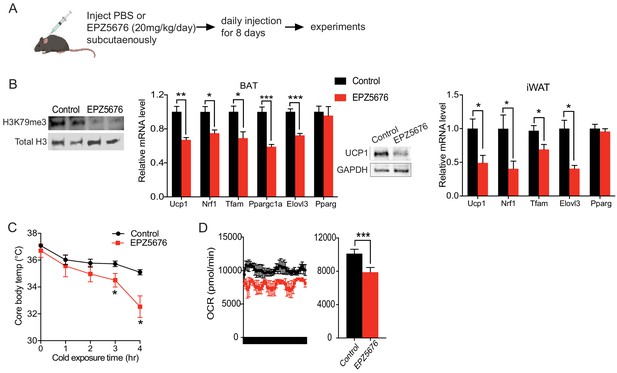
Inhibition of Dot1l activity impairs BAT gene program in vivo.
(A) Schematic diagram of the strategy used to inject either Saline for control or Dot1l chemical inhibitor, EPZ5676. (B) (Left) Immunoblotting for H3K79me3 and total H3. (Middle) RT-qPCR for indicated genes in BAT and iWAT from either PBS or EPZ5676 injected mice (n = 4 per group) and immunoblotting for Ucp1 protein. (C) Rectal temperature of 14-week-old mice maintained at 4°C at indicated time points (hr) (n = 4 per group). (D) VO2 assayed in mice that were housed at 4°C by indirect calorimetry using CLAMS. Data are expressed as means ± standard errors of the means (SEM). *p<0.05, **p<0.01, ***p<0.001.
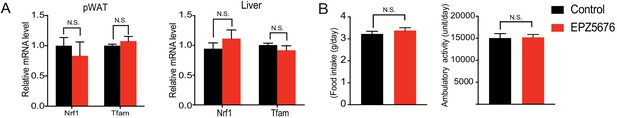
Inhibition of Dot1l activity impairs BAT gene program in vivo.
(A) RT-qPCR for indicated genes in pWAT and liver of control and EPZ5676 injected mice. (B) Food intake and locomotor activity of control and EPZ5676 injected mice. Data are expressed as means ± standard errors of the means (SEM). *p<0.05, **p<0.01, ***p<0.001.
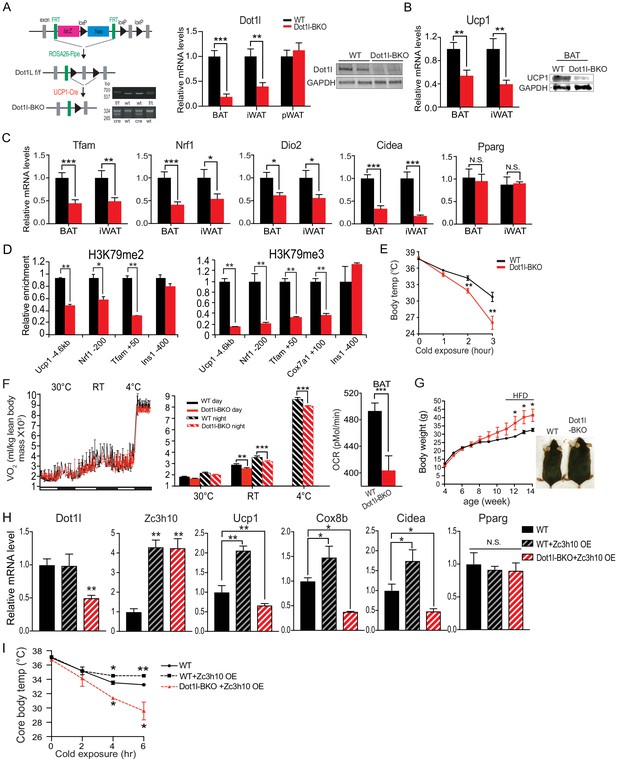
The requirement of Dot1l for cold-induced thermogenesis and the Dot1l-Zc3h10 function in mice.
(A) (Left) Schematic diagram of the strategy used to generate BAT-specific Dot1l conditional knockout mice. PCR genotyping of the mice: Top gel, Dot1l allele; bottom gel, Cre. (Middle) RT-qPCR for Dot1l in BAT, iWAT and pWAT from Dot1l f/f (WT) and Dot1l-BKO mice (n = 6 per group). (Right) Immunoblotting for Dot1l in BAT from Dot1l f/f (WT) and Dot1l-BKO mice. (B) RT-qPCR and immunoblotting for Ucp1 in BAT and iWAT from Dot1l f/f (WT) and Dot1l-BKO mice. (C) RT-qPCR for indicated genes in BAT and iWAT from Dot1l f/f (WT) and Dot1l-BKO mice. (D) ChIP-qPCR of H3K79me2 and H3K79me3 at Zc3h10-binding regions of BAT tissue from control (f/f) or Dot1l-BKO mice (n = 5 per group). (E) Rectal temperature measured in 13-week-old mice at 4°C at indicated time points (hours) (n = 6 mice per group). (F) (Left) whole body VO2 assayed in WT and Dot1l-BKO mice, housed at indicated ambient temperatures (n = 6 per group) by indirect calorimetry using CLAMS. (Right) OCR measured in BAT of WT and Dot1l-BKO mice using Seahorse XF24 Analyzer (n = 5 per group). (G) Representative photograph of 14-week-old control and body weight of WT and Dot1l-BKO mice fed HFD from wk 12. (H) RT-qPCR for indicated genes from WT injected with GFP or Zc3h10 adenovirus or Dot1l-BKO mice injected with Zc3h10 adenovirus for overexpression (n = 5 per group). (I) Rectal temperature measured at 4°C at indicated time points (hours) (n = 5 per group). Data are expressed as means ± standard errors of the means (SEM). *p<0.05, **p<0.01, ***p<0.001.
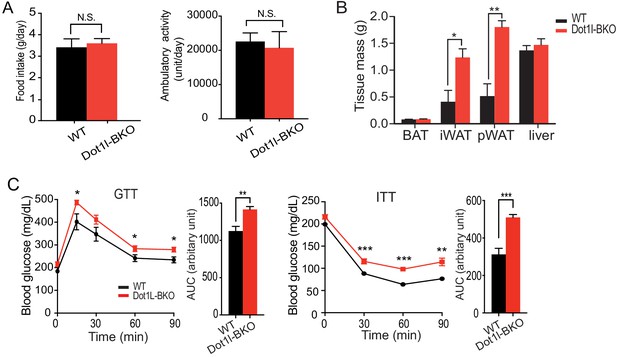
Dot1l is required for cold-induced thermogenesis in mice.
(A) Food intake and locomotor activity of WT and Dot1l-BKO mice. (B) Body weight and mass of various tissues taken from WT and Dot1l-BKO mice at 14 week of age. (C) GTT and ITT of Dot1l-BKO mice at 14 wk of age that were on HFD for 3 weeks (n = 5). Data are expressed as means ± standard errors of the means (SEM). *p<0.05, **p<0.01, ***p<0.001.
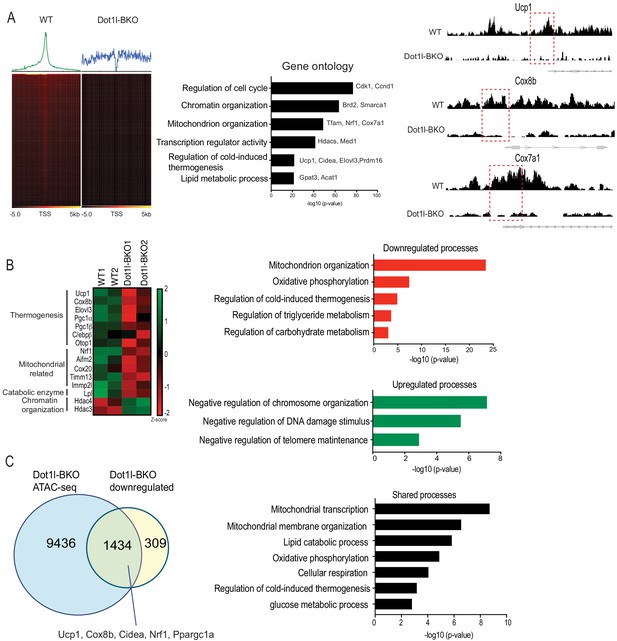
Genome wide analysis for Dot1l effect on chromatin accessibility for thermogenic gene program.
(A) ATAC-seq using BAT from WT and Dot1l-BKO mice after 4 hr cold exposure n = 2 per group. (Left) Heatmaps showing open chromatin regions focused at the transcription start site (TSS). (Middle) Gene Ontology analysis of pathways that are affected by Dot1l ablation in BAT. (Right) UCSC genome browser screenshot of representative peaks at a subset of promoter regions of thermogenic genes. (B) RNA-seq using BAT from WT and Dot1l-BKO mice, n = 2 pooled RNA samples per group. (Left) Heatmap showing changes in gene expression. Color scale shows changes in gene expression as determined by Z-score, green is –two and red is 2. (Middle) Representative top GO terms of upregulated and downregulated genes identified by differential expression analysis. (C) Venn diagrams showing number of unique or shared genes between ATAC- and RNA-seq datasets and charts for representative top gene ontology (GO) terms.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Mus musculus) | Dot1l | GenBank | ID: 208266 | |
| Strain, strain background (Escherichia coli) | BL21(DE3) | NEB | C2527H | |
| Strain, strain background (Mus musculus) | ES cell line Dot1ltm1a(KOMP) | KOMP Repository | MMRRC:054749-UCD | |
| Genetic reagent (Mus musculus) | Dot1l-G55L | This lab | ||
| Cell line (Homo sapien) | HEK293 | UCB Cell Culture Facility | RRID:CVCL_0045 | Authenticated using STR analysis |
| Cell line (Mus musculus) | 3T3-L1 | UCB Cell Culture Facility | RRID:CVCL_0123 | Authenticated using STR analysis |
| Cell line (Mus musculus) | BAT cells | Shingo Kajimura Ohno et al., 2013 | Harvard; authenticated using methods described in Aune et al., 2013 | |
| Transfected construct (Mus musculus) | Ad-m-Dot1l-shRNA | Vector Biolabs | shADV-257398 | |
| Transfected construct (Mus musculus) | Ad-Zc3h10-flag | Vector Biolabs | ADV-276549 | |
| Transfected construct (Mus musculus) | Ad-m-Dot1l | Vector Biolabs | ADV-257398 | |
| Antibody | Anti-Dot1l (rabbit monoclonal) | Cell signaling | Cat# 77087 RRID:AB_2799889 | 1:1000 |
| Antibody | Anti-GAPDH (rabbit monoclonal) | Cell signaling | Cat# 5174S RRID:AB_10622025 | 1:1000 |
| Antibody | Anti-UCP1 (rabbit polyclonal) | Abcam | Cat# 10983 RRID:AB_2241462 | 1:1000 |
| Antibody | Anti-H3K79me2 (rabbit polyclonal) | Abcam | Cat# ab3594 RRID:AB_303937 | ChIP: 5 µg (5 µl) |
| Antibody | Anti-H3K79me3 (rabbit monoclonal) | Abcam | Cat# ab2621 RRID:AB_303215 | ChIP: 5 µg (5 µl) |
| Antibody | Anti-Zc3h10 (rabbit polyclonal) | Thermo Fishcer | Cat# PA5-31814 RRID:AB_2549287 | 1:1000 |
| Recombinant DNA reagent | HA-Dot1l expression vector | Genecopoeia | EX-Mm14067-M07 | |
| Sequence-based reagent | Dot1l_F | This paper | RT-qPCR primer | CGA CTA ATG CTG CAC CTC CT |
| Sequence-based reagent | Dot1l_R | This paper | RT-qPCR primer | AGG AGT AGT GGT GTG GCT CA |
| Sequence-based reagent | Ucp1_F | This paper | RT-qPCR primer | ACT GCC ACA CCT CCA GTC ATT |
| Sequence-based reagent | Ucp1_R | This paper | RT-qPCR primer | CTT TGC CTC ACT CAG GAT TGG |
| Sequence-based reagent | Tfam_F | This paper | RT-qPCR primer | GTC CAT AGG CAC CGT ATT GC |
| Sequence-based reagent | Tfam_R | This paper | RT-qPCR primer | CCC ATG CTG GAA AAA CAC TT |
| Sequence-based reagent | Nrf1-F | This paper | RT-qPCR primer | GAC AAG ATC ATC AAC CTG CCT GTA G |
| Sequence-based reagent | Nrf1-R | This paper | RT-qPCR primer | GCT CAC TTC CTC CGG TCC TTT G |
| Sequence-based reagent | Zc3h10-F | This paper | RT-qPCR primer | CGA CTA ATG CTG CAC CTC CT |
| Sequence-based reagent | Zc3h10-R | This paper | RT-qPCR primer | AGG AGT AGT GGT GTG GCT CA |
| Commercial assay or kit | NE-PER Nuclear and Cytoplasmic Extraction Kit | Thermo Fisher | 78833 | |
| Commercial assay or kit | SimpleChIP Plus Sonication Chromatin IP Kit | Cell signaling | 56383S | |
| Commercial assay or kit | Dual-Luciferase Kit | Promega | E1980 | |
| Commercial assay or kit | Nextera DNA library Preparation Kit | Illumina | 15028212 | |
| Chemical compound, drug | EPZ5676 | MedChemExpress | HY-15593 | |
| Software, algorithm | Partek Flow Genomics Suite | Partek | RRID:SCR_011860 | |
| Other | Rectal probe | Physitemp | BAT-12 | For mice |





