In-silico analysis of myeloid cells across the animal kingdom reveals neutrophil evolution by colony-stimulating factors
Figures

Population comparison of blood myeloid cell subset distribution in chordates demonstrates predominance of the neutrophilic granulocyte in birds and mammals.
(A) Species tree of the animals within Phylum Chordata examined, sub-classified by animal order or class. (B) Meta-analysis of aggregated phylum Chordata complete blood cell counts (excluding all lymphoid cells). Data is visualised as a floating bar and the line represents the mean value and shows absolute counts per cell type per order (i) and composition of myeloid cells per cell type (ii). (N) = Neutrophils, (H) = Heterophils, (A) = Azurophils, (M) = Monocytes.
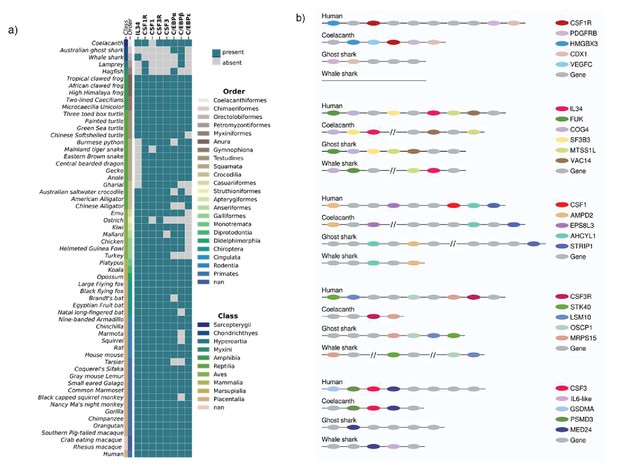
Analysis of chordate CSF1/CSF1R, CSF3/CSF3R, and neutrophil-related transcription factors reveals their loss in Jawed/Jawless fish lineages.
(A) Heat map of CSF1R/IL34/CSF1, CSF3R/CSF3, and neutrophil-related transcription factors in selected members of Phylum Chordata. (B) Syntenic maps of CSF1R/IL34/CSF1 and CSF3R/CSF3 in selected Jawed fish compared to human.
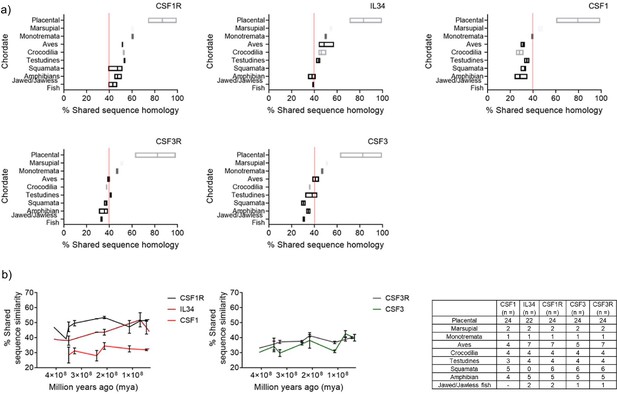
Shared sequence similarity analysis of Chordate CSFR/CSF protein homology further supports the ancestral pairing of CSF1R/IL34 and CSF3R/CSF3 in early lineages.
(A) Representative plots of % shared sequence similarity for CSF1R, CSF3R, IL34, CSF1, and CSF3 in the respective sub-groups of Phylum Chordata, data is visualised as a floating bar and the line represents the mean value. Baseline shared similarity is represented by the red line. (B) Graphical representation of % shared sequence similarity in sub-groups of Phylum Chordata versus time.
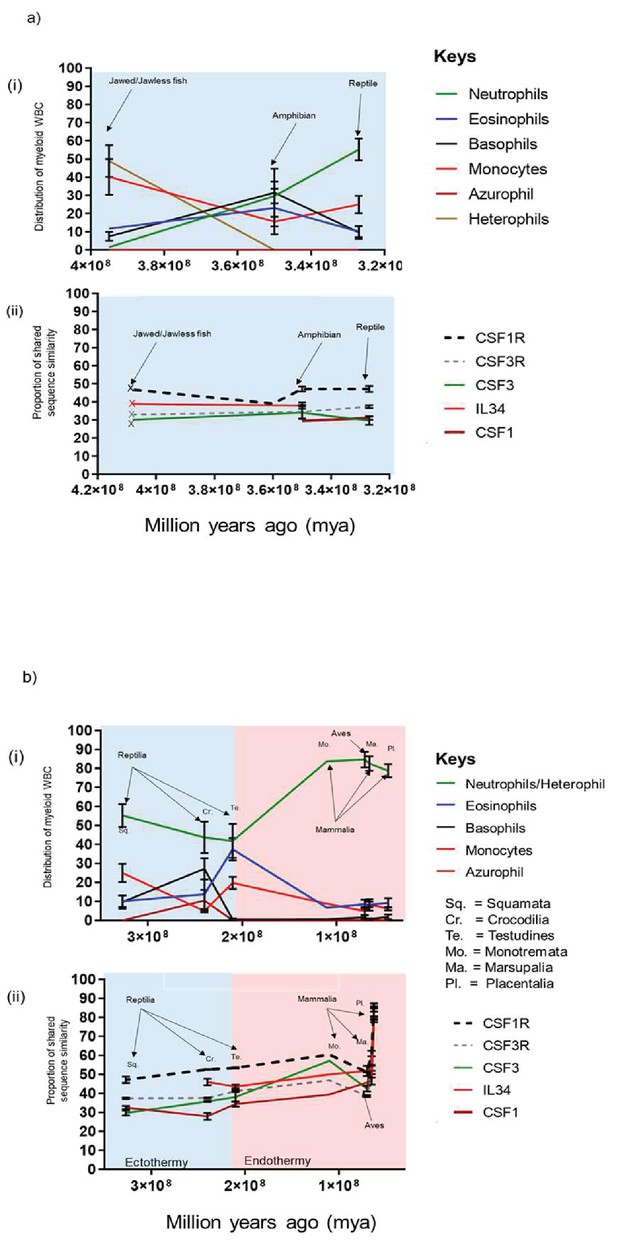
Changes in chordate blood granulocyte distribution from Jawed/Jawless lineages through to placental mammals are multi-factorial and likely driven by the emergence of CSF3 and onset of endothermy.
(A) Graphical representation of % population distribution of myeloid white blood cells versus time (i) and % shared sequence similarity of CSF1R and CSF3R protein families versus time for Jawed/Jawless fish, Amphibia and the reptilian order of Squamata (ii). (B) Graphical representation of % population distribution of myeloid white blood cells versus time (i) and % shared sequence similarity of CSF1R and CSF3R protein families versus time for the Reptilian orders of squamata, testudines and crocodilia, Aves, and the Mammalian orders of monotremata, marsupalia and Placental (ii). Error bars represent standard deviation.
Additional files
-
Supplementary file 1
Gene and related protein names for the CSF1R/IL34/CSF1 and CSF3R/CSF3 families.
- https://cdn.elifesciences.org/articles/60214/elife-60214-supp1-v2.docx
-
Supplementary file 2
Knock-out mouse models in cebpα, cebpβ and cebpε deficient mice.
- https://cdn.elifesciences.org/articles/60214/elife-60214-supp2-v2.docx
-
Supplementary file 3
References for Chordate haematological parameters.
- https://cdn.elifesciences.org/articles/60214/elife-60214-supp3-v2.docx
-
Supplementary file 4
Gene Identification numbers for species examined.
- https://cdn.elifesciences.org/articles/60214/elife-60214-supp4-v2.docx
-
Supplementary file 5
Protein accession numbers for species examined.
- https://cdn.elifesciences.org/articles/60214/elife-60214-supp5-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/60214/elife-60214-transrepform-v2.docx





