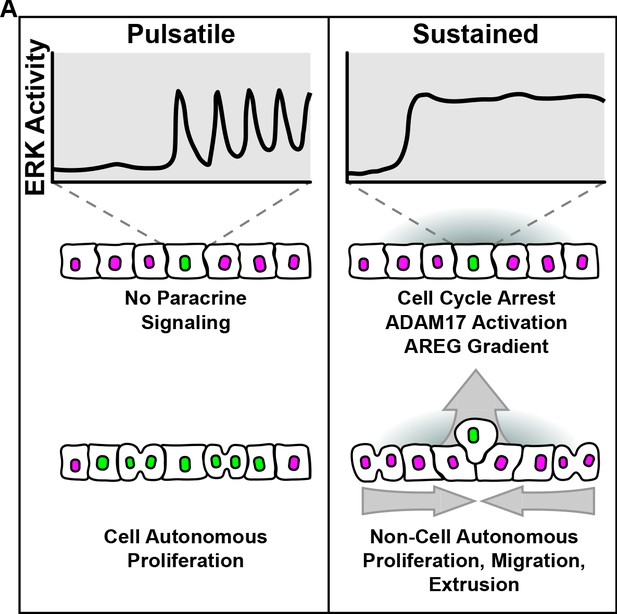
MAPK activity dynamics regulate non-cell autonomous effects of oncogene expression
Figures
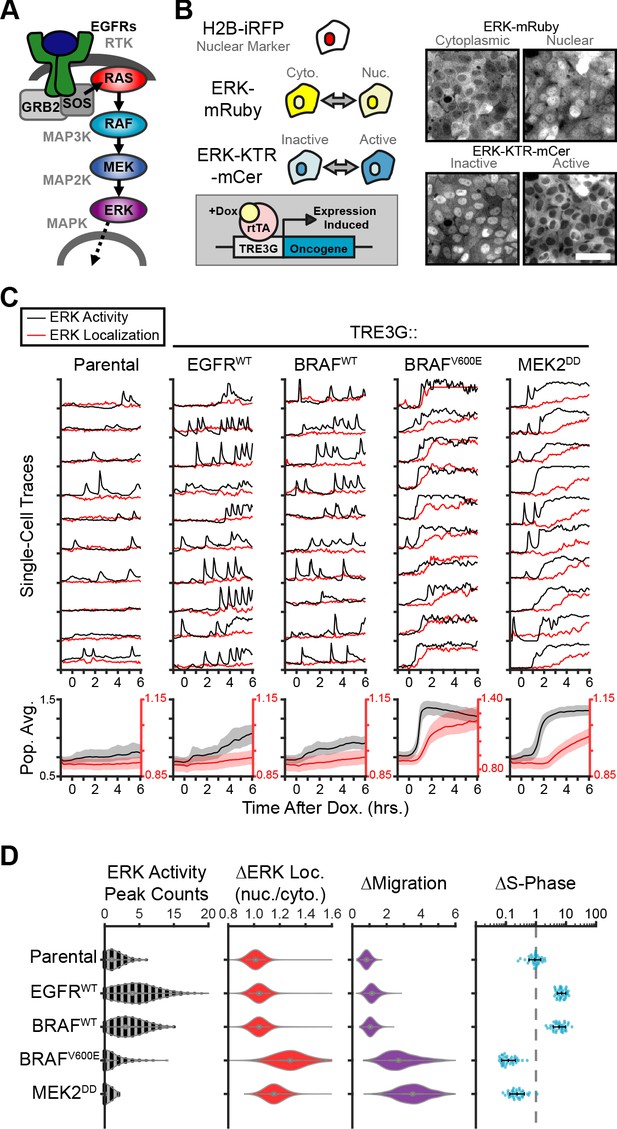
Oncogenic ERK signaling dynamics promote qualitatively different cell behaviors.
(A) Schematic representation of the RTK/RAS/ERK signaling pathway. (B) MCF10A cells were transduced with lentiviral vectors expressing ERK KTR-mCerulean3 and ERK-mRuby2. The doxycycline inducible system (rTtA and TRE3G) was used to drive the expression of oncogenes during live imaging. Representative images of cytoplasmic and nuclear ERK-mRuby2 (top) and inactive or active ERK as reported by ERK KTR-mCerulean3 (bottom). Scale bar = 50 µm. (C) Cells described in B with indicated inducible oncogenes were imaged every 5 min for 6 hr upon doxycycline induction (2 µg/ml) at t = 0. Single cells were analyzed as described in methods. Population averages represent more than 1000 cells per condition. Shaded regions indicate the 25th-75th percentiles. (D) Quantification of data obtained in C. Single-cell counts of ERK activity peaks after induction (6–12 hr), ERK kinase localization fold change (final N/C ratio over basal N/C ratio per cell), and cell migration (final over basal distance traveled per cell) were extracted as described in methods. For proliferation analysis the fraction of S phase cells was measured using Edu incorporation and the change over the no dox control was calculated and normalized to the mean of parental cells (dashed line) (see Materials and methods). Data represents 36 independent observations.
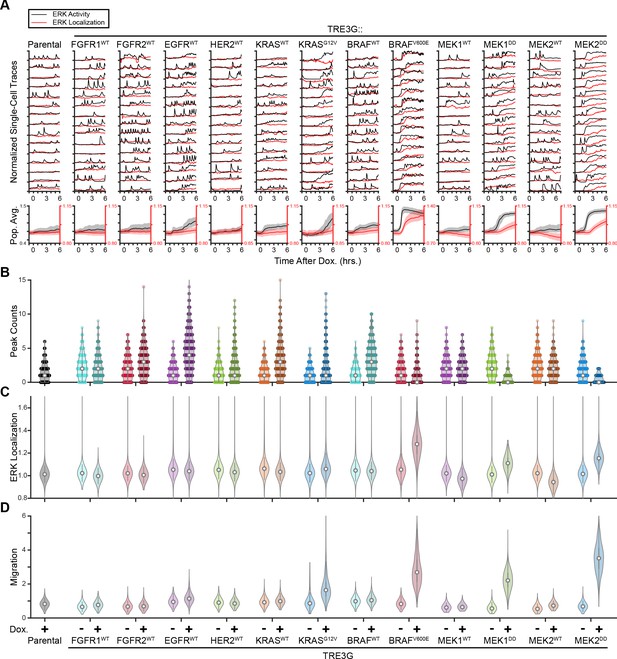
A screen for oncogenic effects on ERK dynamics and cell behavior.
(A) Inducible cells expressing indicated genes treated and plotted as in 1C. Single traces and population data are reproduced for oncogenes appearing in Figure 1. (B–D) Single-cell peaks, ERK nuclear localization, and migration are quantified directly from imaging data as in 1D for indicated inducible cells in the presence or absence of doxycycline (2 µg/ml). Data appearing in Figure 1 is reproduced.

Relative expression of inducible genes.
(A) Parental, BRAFWT, and BRAFV600E inducible cells were grown in the presence or absence of Dox (2 µg/ml) for 24 hr before sample collection and western blot for BRAF and HSC70 (see Materials and methods).

Oncogene-induced cell behaviors are distinct from epithelial-to-mesenchymal transition (EMT).
(A) Indicated parental or inducible MCF10As were grown in the presence or absence of doxycycline (+Dox., 2 µg/ml) for 24 hr before fixing for immunofluorescence using primary antibodies for E-Cadherin (E-Cad) or N-Cadherin (N-Cad), see Materials and methods. Staining is compared to parental cells treated with TGFβ continuously for several passages over 10 days to induce EMT (5 ng/ml or 50 ng/ml). Nuclear marker (H2B-iRFP, magenta), IF staining (green), an overlay of nuclear marker with IF staining, and ERK activity (ERK-KTR, cyan) are shown. Scale bar = 100 µm.
Different ERK dynamics following oncogene induction.
Dual Sensor MCF10A cell lines containing H2B-iRFP nuclear marker (red), ERK-mRuby2 localization reporter (yellow), and an ERK KTR-mCerulean3 representing kinase activity (cyan), and indicated inducible oncogene constructs were treated with doxycycline (2 µg/ml) at time 0 (see Figure 1). Images were collected every 5 min for an hour basal period and 12 hr after induction. Scale bar = 100 µm.
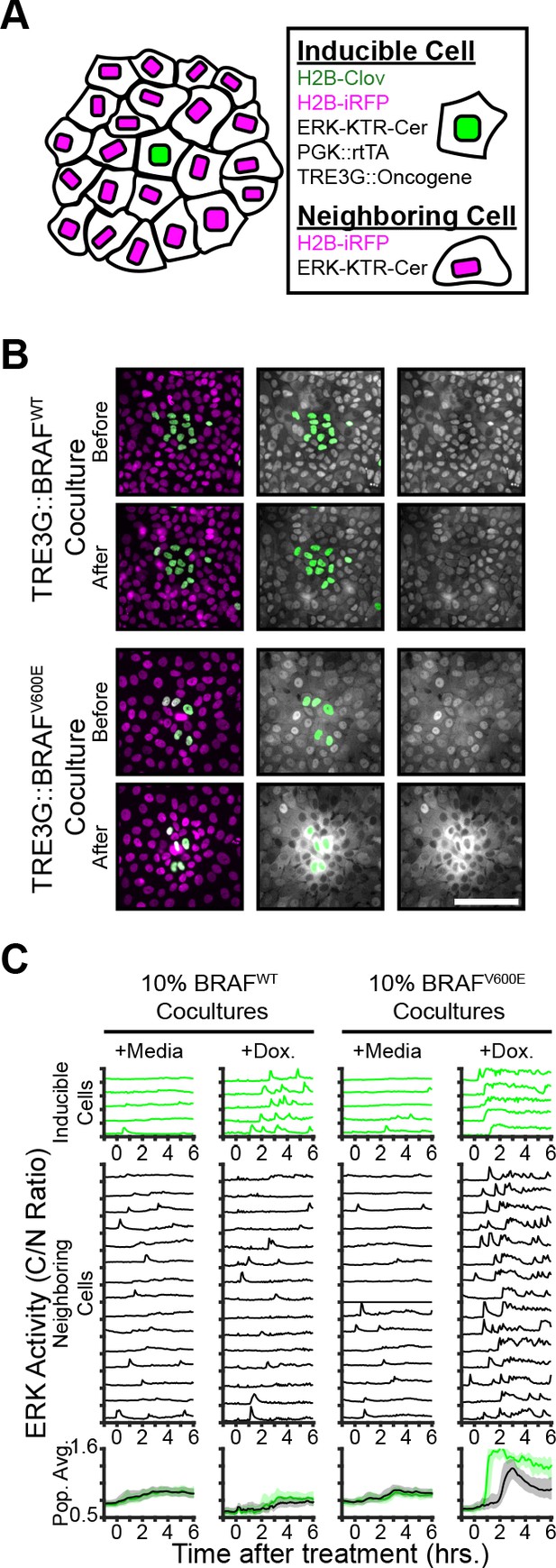
Oncogene induction results in dynamics-dependent paracrine ERK signaling.
( A) Schematic representation of coculture assay. H2B-iRFP (magenta) and ERK KTR are expressed in all cells for segmentation and quantification. H2B-mClover (green) was used to label inducible cells. (B) BRAFWT or BRAFV600E inducible cells were cocultured at 10% with ERK KTR cells and treated with doxycycline (2 µg/ml). Representative images are shown. Scale bar = 100 µm. (C) BRAFWT or BRAFV600E cocultures, as in B, were treated with vehicle (+Media) or with doxycycline (+Dox, 2 µg/ml). Single cells were quantified as described in methods. ERK activity traces in inducible (top, green) and neighboring cells (bottom, black) are shown. Population averages and 25th-75th percentiles (shaded) are shown for n > 450 cells per coculture condition.
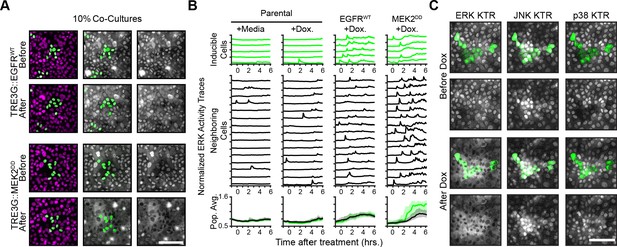
ERK dynamics-dependency of paracrine ERK activation.
(A) Images from cocultures of inducible EGFRWT and MEK2DD cells presented as in 2B. Scale bar = 100 µm. (B) ERK activity traces from cocultures of indicated cells treated as in 2C. Parental cells have an H2B-mClover nuclear marker without any inducible gene system. Population averages and 25th-75th percentiles (shaded region) shown for n > 1000 cells. (C) The TRE3G-BRAFV600E construct was expressed in a cell line containing ERK KTR-mCerulean3, p38 KTR-mClover, and JNK KTR-mRuby2. These cells were incubated with CellTracker Deep Red dye (green, Invitrogen) and cocultured with unlabeled neighboring cells. Images display the activity of ERK, p38, and JNK before and 6 hr after addition of doxycycline (2 µg/ml). Scale bar = 100 µm.
Oncogene-induced ERK signaling waves.
Inducible BRAFV600E cells with a H2B-mClover nuclear marker (green) were cocultured in monolayers with neighboring cells (H2B-iRFP, magenta) (see Figure 2). Images were collected every 5 min for an hour basal period and 12 hr following addition of doxycycline (2 µg/ml) at time 0. ERK KTR-mCer3 (grey) reports ERK activity in all cells. Three examples are shown for both WT (left) and ADAM17KO (right) inducible cells. Scale bar = 100 µm.
Spontaneous cell-death induces ERK signaling waves.
Nine examples are shown for spontaneous cell death events in serum-starved MCF10A monolayers imaged every 5 min.
The timing of cell death in each example is aligned. ERK KTR-mCer3 represents ERK activity (grey). Scale bar = 100 µm.
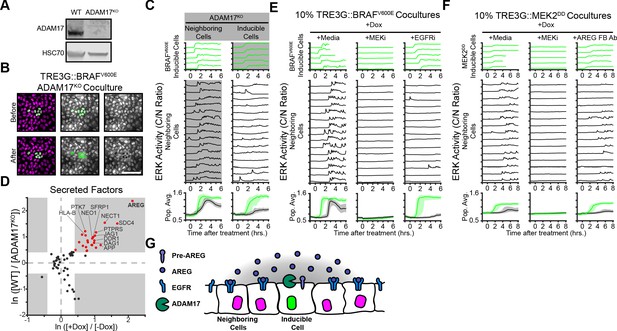
ERK activity waves require ADAM17 release of AREG and neighboring cell EGFRs.
(A) Immunoblot against ADAM17 and HSC70 in WT and ADAM17KO cells generated by CRISPR-Cas9 editing (see Materials and methods for details). (B) Representative images of ADAM17KO BRAFV600E inducible cells cocultured and treated as in Figure 2B. (C) ADAM17KO cells (gray boxed traces) were used as inducible cells (right) or neighboring cells (left) in cocultures. Data for n > 1100 cells is presented as in Figure 2C. (D) ADAM17 substrates profiled by TMT mass spectrometry. Supernatants from ADAM17KO or WT cells expressing (+Dox) or not expressing (-Dox) BRAFV600E were collected and analyzed by Tandem-Mass-Tag (TMT) mass spectrometry as described in methods. Scatter plots show the natural log of fold change values of all statistically significant (p<0.05) proteins in both WT vs. ADAM17KO and +Dox vs. -Dox comparisons. Grey boxes indicate >1.5 fold change. (E) BRAFV600E co-cultured monolayers were plated as in Figure 2C and pretreated with indicated inhibitors (MEKi, 5 µM PD0325901; EGFRi, 5 µM Gefitinib) for one hour before induction with doxycycline (2 µg/ml). Representative single cell traces and population averages for n > 1000 cells are shown as in 2C. (F) MEK2DD co-cultured monolayers were plated as in Figure 2C and pretreated with indicated inhibitors (MEKi, 5 µM PD0325901; AREG FB Ab, 50 µg/ml function-blocking antibody) for one hour before induction with doxycycline (2 µg/ml). Representative single cell traces and population averages for n > 1000 cells are shown as in Figure 2C. (G) Schematic representation of ADAM17-AREG-EGFR paracrine signaling.
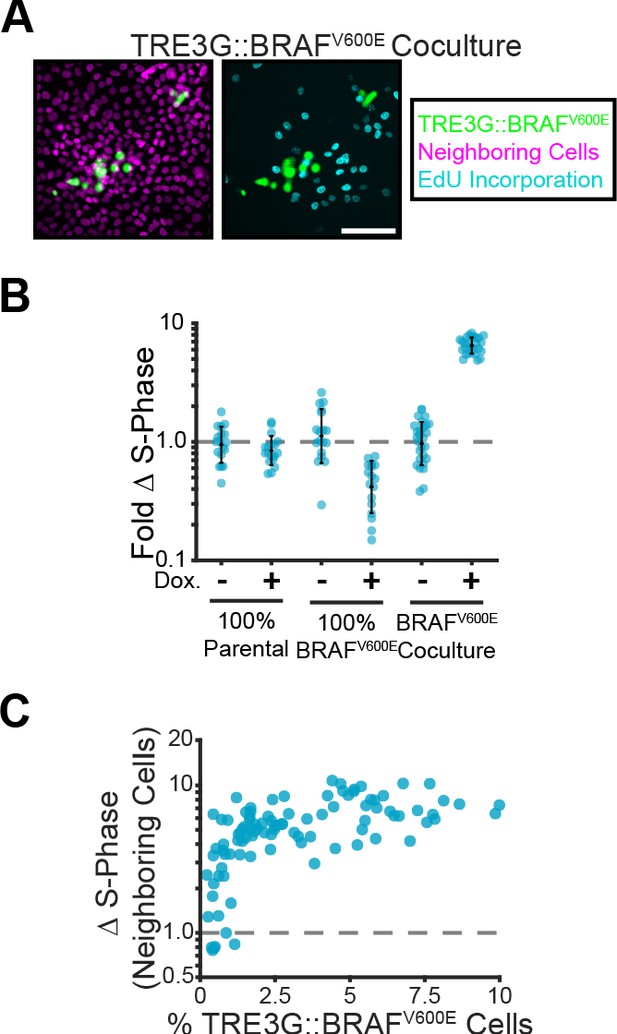
Paracrine ERK signaling leads to non-cell autonomous proliferation.
(A) Representative images of BRAFV600E cocultures treated with doxycycline and EdU as described in methods. Inducible cell nuclei (H2B-mClover, green), all nuclei (H2B-iRFP, magenta) and EdU staining (cyan) are shown. Scale bar = 100 µm. (B) Indicated monolayers were cultured and incubated with or without doxycycline for 24 hr. The change in S-phase cell fractions was determined by EdU incorporation as described in methods and normalized to parental mean (dashed line). Bar represents mean and standard deviation for n ≥ 16 observations. (C) Inducible BRAFV600E cocultures were plated at different proportions and labelled with EdU as in A. The fold-change in S-phase cell fractions is plotted against the percent of BRAFV600E-expressing cells for each position. 98 total observations shown.
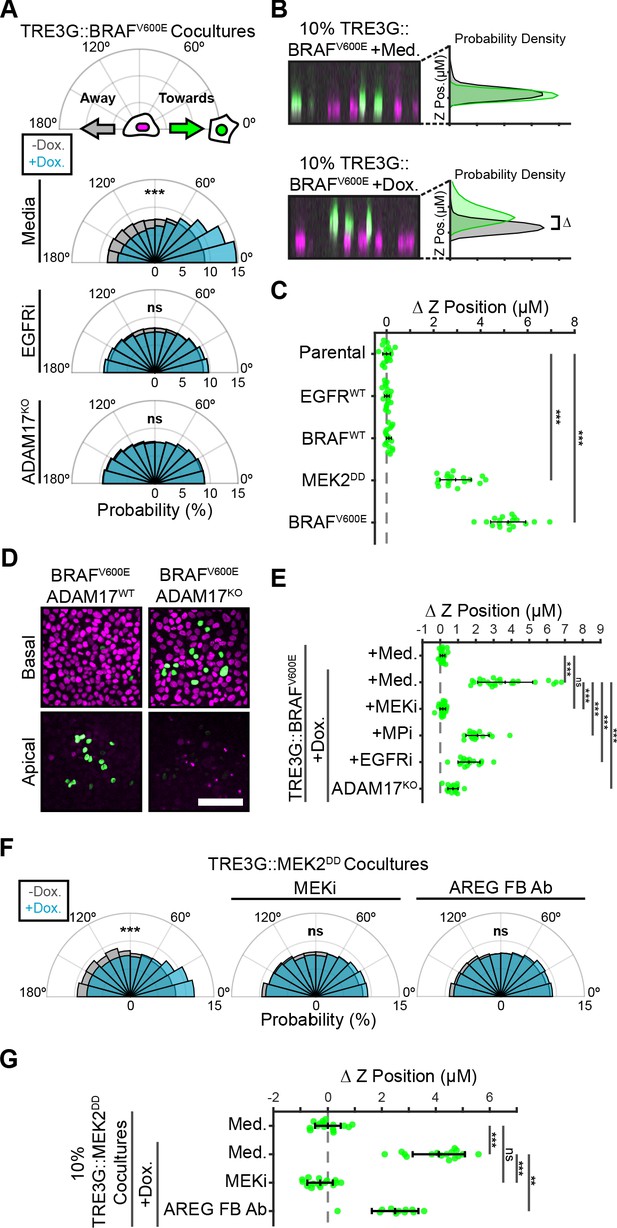
Paracrine ERK activation coordinates extrusion of aberrantly signaling cells through directed migration of the neighboring epithelium.
(A) Inducible BRAFV600E cells (WT or ADAM17KO) were plated in 1% cocultures and treated with doxycycline (2 µg/ml) in the presence or absence of EGFR inhibitor gefitinib (5 µM) as indicated. Radial histograms represent migration angle distributions of neighboring cells before (grey) and 2–6 hr after (cyan) induction (see Materials and methods). Data represents angles from n > 1000 cells from 10 independent observations per condition. Data was assessed using subsampling and a two-sample KS test with ‘ns’ not significant, ***p<0.001 (see Materials and methods). (B) 10% BRAFV600E cocultured monolayers were seeded as described in methods. After 24 hr with doxycycline (2 µg/ml), monolayers were imaged by spinning disk confocal. Representative orthogonal Z projections and probability densities for nuclear height of inducible (green) and neighboring (grey) cells are shown (see methods). Extrusion (ΔZ) is calculated as the height difference between gaussian-fitted maxima of the green and black distributions. (C) 10% cocultures of indicated parental or inducible cells were treated with 24 hr doxycycline (2 μg/ml), imaged, and analyzed as in B. Data represents difference in nuclear height (ΔZ) for n = 18 observations normalized to the mean height of parental cells (dashed line), with mean and +/- standard deviation (black bars). Significance was calculated by two-sample t-test with ‘ns’ indicating no significance, ***p<0.001. (D) Representative basal and apical images (+6 µm) of WT or ADAM17KO, BRAFV600E inducible cells (green) in WT monolayers (red) after 24 hr of doxycycline treatment. (E) 10% BRAFV600E cocultures were pretreated with inhibitors (MEKi, 5 μM PD0325901, MPi, 5 μM Batimastat, EGFRi, 5 μM Gefitinib) and 24 hr doxycycline (2 μg/ml) or media, imaged and analyzed as in B. Data represents difference in nuclear height (ΔZ) for n ≥ 16 independent observations presented as in C. (F) Inducible MEK2DD cells were plated in 1% cocultures and treated with doxycycline (2 µg/ml) in the presence of MEK inhibitor (MEKi, 5 µM PD0325901) or amphiregulin function-blocking antibody (AREG FB Ab, 50 ng/ml) as indicated. Radial histograms are presented as in A for angles of n > 100 cells from two to three independent observations per condition. Data was assessed using subsampling and a two-sample KS test with ‘ns’ not significant, ***p<0.001 (see methods). (G) 10% MEK2DD cocultures were pretreated with MEK inhibitor (MEKi, 5 µM PD0325901) or Amphiregulin function-blocking antibody (AREG FB Ab, 50 ng/ml) and 24 hr doxycycline (2 µg/ml) or media, as indicated, then imaged and analyzed as in B-C. Data represents difference in nuclear height (ΔZ) for n ≥ 11 independent observations normalized to the mean height of media-treated MEK2DD cells (dashed line), with mean and +/- standard deviation (black bars). Significance was calculated by two-sample t-test with ‘ns’ indicating no significance, **p<0.01, and ***p<0.001.
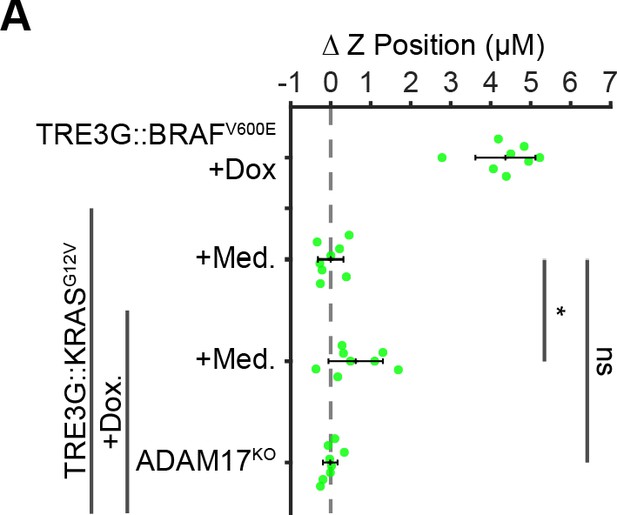
KRASG12V expressing cells do not extrude at 24 hr.
(A) 10% BRAFV600E or KRASG12V (WT or ADAM17KO) cocultures were treated with 24 hr doxycycline (2 µg/ml) or media, imaged and analyzed as in Figure 5B. Data represents difference in nuclear height (ΔZ) for n = 8 independent observations normalized to the mean height of un-induced KRASG12V cells (dashed line), with mean and +/- standard deviation (black bars). Significance was calculated by two-sample t-test with ‘ns’ indicating no significance, *p<0.05.

Actin dynamics and sphingosine kinase requirements during oncogenic cell extrusion.
(A) Representative images of immunofluorescence for E-cadherin (cyan) from 1% BRAFV600E cocultures treated for 24 hr with doxycycline (2 µg/ml) or media imaged by confocal. TRE3G::BRAFV600E cells carry H2B-mClover nuclear markers (green), while neighboring cells have only H2B-iRFP (magenta). A basal and an apical plane are shown for each position, while the bottom shows an XZ orthogonal slice from the indicated position (grey line) in the above images. Orthogonal view shows positions of basal (hollow triangle) and apical planes (solid triangle). Scale bar = 50 µm. (B) Manual quantification of fully extruded TRE3G::BRAFV600E cells from cocultures induced for indicated times (0, 4, 8, 24 hr) with doxycycline (2 µg/ml) after membrane immunofluorescence staining of E-Cadherin. In order to be considered fully extruded rather than elongated, cells must have no evidence of basal attachment. Data represents n ≥ 35 cells. (C) Time-course of actin dynamics and basal protrusions of neighboring cells in 1% BRAFV600E coculture imaged with Utrophin-261-EGFP (cyan, top, see Figure 5—Video 2). TRE3G::BRAFV600E cell is indicated by bold asterisk (*), and H2B-iRFP (bottom, magenta) from cells in the plane of the monolayer are shown to observe timing of extrusion. Scale bar = 50 µm. (D) Quantification of actin enrichment from BRAFV600E cocultures. Median actin intensities (AUs) from randomly selected neighbor-neighbor cell borders are compared to the leading edges of all neighboring cells adjacent to TRE3G::BRAFV600E cells that were treated with doxycycline alone (2 µg/ml, from images in C) or with doxycycline plus EGFR inhibitor (Gefitinib, 5 µM). See Materials and methods for details. (E) MEK2DD cocultures were plated as in 2B, and treated with doxycycline (+Dox, 2 µg/ml) in the presence of media or inhibitors (MEKi, 5 µM PD0325901; SphKi, 10 µM SKII). Single-cell ERK activity traces are presented as in 2C with population averages and 25th-75th percentiles (shaded) shown for n > 1000 cells per coculture condition. (F) Neighboring cell migration angles from 1% Inducible MEK2DD cells cocultures treated with doxycycline (2 µg/ml) in the presence or absence of Sphingosine Kinase inhibitor (5 µM SKII). Data represents angles from n > 100 cells from two independent observations per condition presented as in 5A, and Dox condition reproduced from Figure 5F for comparison. Data was assessed using subsampling and a two-sample KS test with *p<0.05, ***p<0.001 (see Materials and mmethods). (G) Extrusion from 10% MEK2DD cocultures pretreated with Sphinosine-1-Phosphate pathway inhibitors (SphKi, 10 µM SKII; S1PR2i, 10 µM JTE-013) and 24 hr doxycycline (2 µg/ml) or media, imaged and analyzed as in 5B-D (green). Some conditions reproduced from Figure 5 for comparison (grey). Data represents difference in nuclear height (ΔZ) for n ≥ 15 independent observations normalized to the mean height of parental cells (dashed line), with mean and +/- standard deviation (black bars). Significance was calculated by two-sample t-test with ‘ns’ indicating no significance, *p<0.05.
Extrusion of BRAFV600E-Expressing Cells.
Cocultured monolayers of inducible BRAFV600E cells were imaged every 10 min for 24 hr following doxycycline (2 µg/ml) treatment. The first frame shows the starting location of inducible cells (H2B-mClover, green) and neighboring cells (H2B-iRFP, magenta). The following video indicates cells in a basal position within the monolayer in magenta, and cells in an apical z-plane (+8 µm, cyan). Scale bar = 100 µm.
Live actin dynamics during extrusion.
Inducible BRAFV600E cells were cocultured with neighboring cells stably expressing Utrophin-261-EGFP, an actin-binding protein (cyan, see Figure 5—figure supplement 2). Actin dynamics and basal protrusions of neighboring cells (cyan) and the nuclei of all cells (H2B-iRFP, magenta), were imaged within the basal plane of the monolayer every 20 min from hours 1–7 after addition of doxycycline (2 µg/ml). Scale Bar = 50 µm.

ERK activity in neighboring cells is required for coordinating extrusion.
(A) Representative images showing WT or ADAM17KO cells with inducible MKK3DD (green), cocultured at 10% with neighboring ERK-KTR cells (grey). Cocultures were treated with doxycycline (2 µg/m) in the presence of media, p38 inhibitor (5 µM BIRB-796), EGFR inhibitor (5 µM Gefitinib), or MEK inhibitor (5 µM PD 0325901). Scale bar = 100 µm. (B) ERK activity traces of neighboring cells in coculture with MKK3DD-inducible cells (WT or ADAM17KO) plated at 10%, pretreated with inhibitors (p38i, 5 µM BIRB 796; EGFRi, 5 µM Gefitinib; MEKi, 5 µM PD 0325901) and doxycycline (2 µg/ml) or media, and imaged as in Figure 2C. 15 representative neighboring cell ERK activity traces are shown for each condition. (C) Inducible MKK3DD cells (WT or ADAM17KO) were plated in 1% cocultures and treated with doxycycline (2 µg/ml) in the presence or absence of inhibitors. Radial histograms of migration angles before (grey) and 6–9 hr after (cyan) induction presented as in Figure 5A. Data represents angles from n > 900 cells from ≥6 observations per condition assessed using subsampling and a two-sample KS test with ‘ns’ not significant, *p<0.05, ***p<0.001 (see Materials and methods). (D) 10% MKK3DD cocultures were pretreated with inhibitors (p38i, 5 µM BIRB 796; EGFRi, 5 µM Gefitinib; MEKi, 5 µM PD 0325901) and 24 hr doxycycline (2 µg/ml) or media, imaged and analyzed as in Figure 5B–D. Data represents difference in nuclear height (ΔZ) for n ≥ 16 observations normalized to the mean height of parental cells (dashed line), with mean and +/- standard deviation (black bars). Significance was calculated by two-sample t-test with ‘ns’ indicating no significance, ***p<0.001.
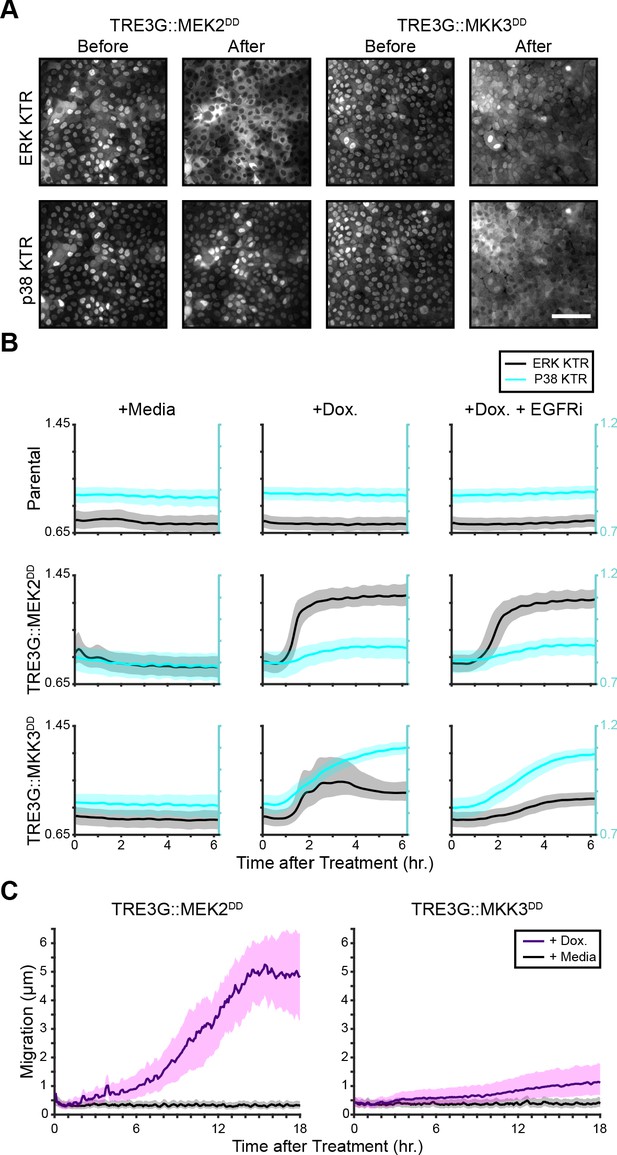
MAPK specificity of MEK2DD and MKK3DD.
(A) Representative images from inducible MEK2DD and MKK3DD cells expressing H2B-iRFP, ERK KTR-mCer3 and p38 KTR-mClov before and after induction with doxycycline (2 µg/ml). Scale bar = 100 µm. (B) Population average traces of ERK (black) and p38 (cyan) activity for indicated parental and inducible ERK/p38 KTR cells from A. Cocultures were treated with media, doxycycline, or doxycycline with EGFR inhibitor (5 µM gefitinib) to limit paracrine ERK activation. Population averages and 25th-75th percentiles (shaded regions) are shown for n > 2000 cells. (C) Migration of inducible cells from B plotted as µm change for every 5 min timepoint.

Mosaic p38 activation leads to ADAM17-EGFR -ependent proliferation of neighboring cells.
(A) EdU of 10% MKK3DD inducible cell cocultures treated for 24 hr with vehicle (+Media) or doxycycline (2 µg/ml) in the presence of MEK inhibitor (MEKi, 5 µM PD 0325901), p38 inhibitiors (5 µM BIRB 796 or 25 µM SB 203580) or in ADAM17KO cells. Data presented as in 4B for n ≥ 16 observations. See Materials and methods for details about EdU incorporation.
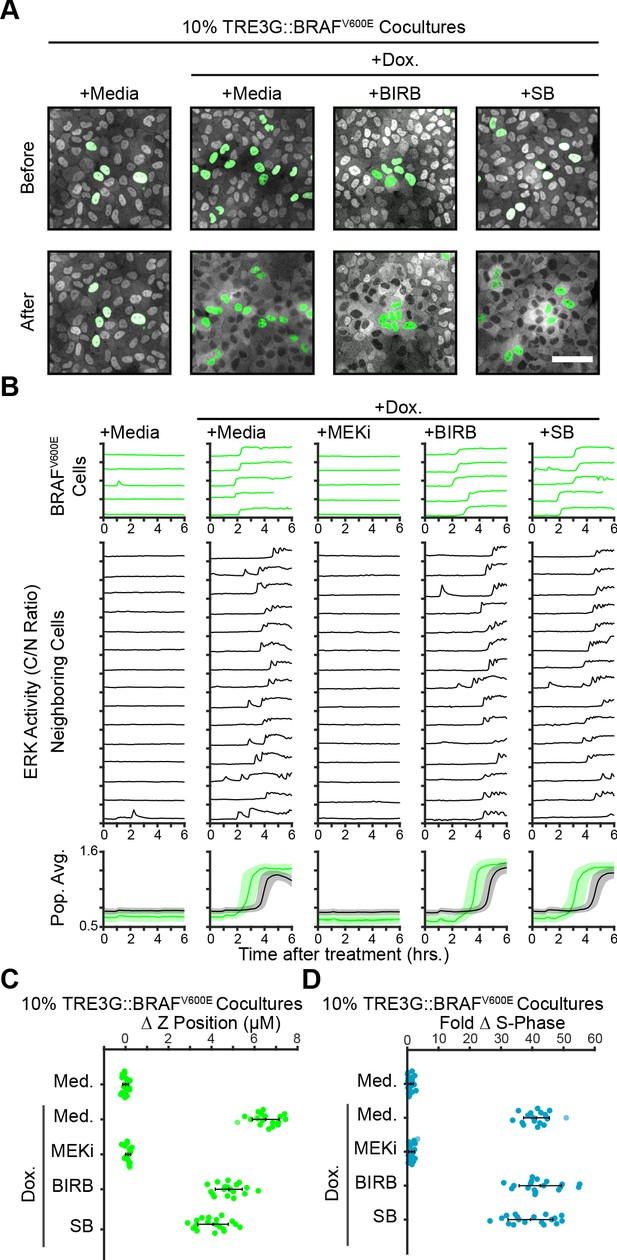
Oncogene-induced paracrine ERK activity and resulting cell behaviors are p38-independent.
(A) Representative images from 10% TRE3G::BRAFV600E cocultures plated as in 2B before and after treatment with media or doxycycline (2 µg/ml) alone or with p38 inhibitors (5 µM BIRB 796 or 25 µM SB 203580). Scale bar = 100 µm. (B) ERK activity traces, averages and 25th-75th percentiles from inducible BRAFV600E cocultures from a represented as in 2C for n > 1600 cells. (C) Extrusion of BRAFV600E inducible cells from 10% cocultures treated as indicated and presented as in 5C for n = 18 observations. (D) EdU incorporation in 10% BRAFV600E cocultures treated as indicated and presented as in Figure 4B for n = 18 observations (see Materials and methods).
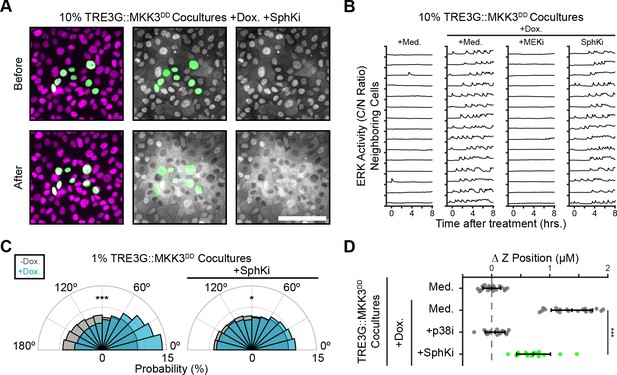
Partial involvement of S1P signaling in extrusion of p38-active cells.
(A) Representative images from MKK3DD cocultures pretreated with sphingosine kinase inhibitor (SphKi, 10 µM SKII) before induction with doxycycline (2 µg/ml). Scale bar = 100 µm. (B) ERK activity traces of neighboring cells in coculture with MKK3DD inducible cells plated at 10%, pretreated with inhibitors (MEKi, 5 µM PD 0325901; SphKi, 10 µM SKII) and doxycycline (2 µg/ml) or media, imaged as in 2C and presented as in 6B. 15 representative neighboring cell ERK activity traces are shown for each condition. (C) Inducible MKK3DD cells were plated in 1% cocultures and treated with doxycycline (2 µg/ml) in the presence of Sphingosine Kinase inhibitor (10 µM SKII). Radial histograms of migration angles before (grey) and after (cyan) induction presented as in Figure 6C, and the +Dox alone condition (left) is reproduced from Figure 6C for comparison. Data represents angles from n > 900 cells from ≥6 observations per condition assessed using subsampling and a two-sample KS test with *p<0.05, ***p<0.001 (see methods). (D) 10% MKK3DD cocultures were pretreated with inhibitors (p38i, 5 µM BIRB 796; SphKi, 10 µM SKII) and 24 hr doxycycline (2 µg/ml) or media, imaged and analyzed as in 5B-D. Data represents difference in nuclear height (ΔZ) for n ≥ 16 observations normalized to the mean height of parental cells (dashed line), with mean and +/- standard deviation (black bars). Some conditions are reproduced from Figure 6D for comparison (grey). Significance was calculated by two-sample t-test with ***p<0.001.
p38-induced ERK signaling waves.
Inducible MKK3DD cells with a H2B-mClover nuclear marker (green) were cocultured in monolayers with neighboring cells (H2B-iRFP, magenta) (see Figure 4). Images were collected every 5 min for an hour basal period and 24 hr following addition of doxycycline (2 µg/ml) at time 0. ERK KTR-mCer3 (grey) reports ERK activity in all cells. Three examples are shown for both WT (left) and ADAM17KO (right) inducible cells. Scale bar = 100 µm.
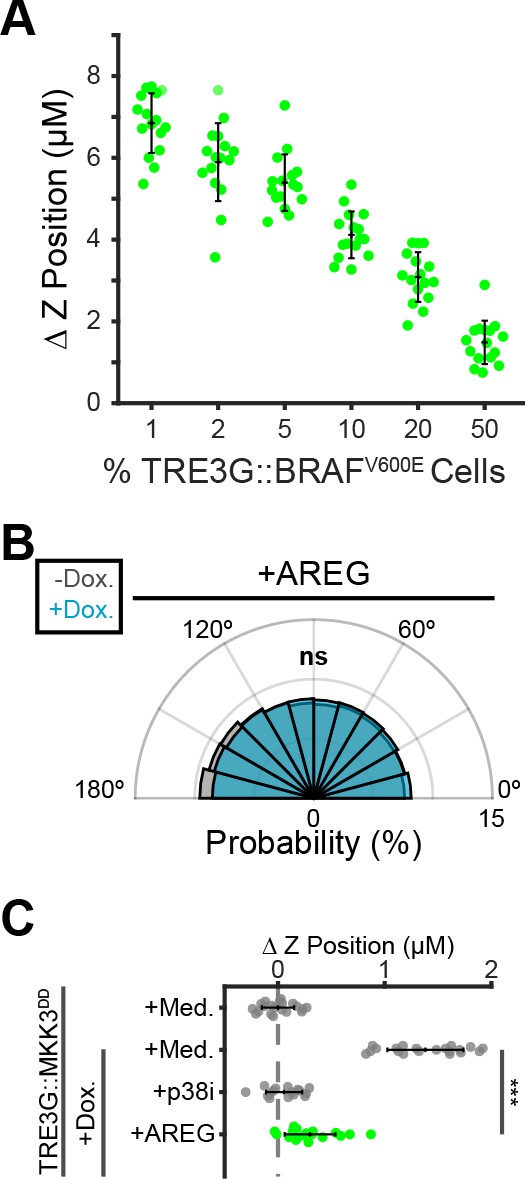
Localized paracrine signals coordinate directed migration and extrusion.
(A) Inverse relationship between fraction of oncogenic cells in coculture and extrusion efficiency. Inducible BRAFV600E cells were cocultured at indicated proportions, treated 24 hr with doxycycline (2 µg/ml), imaged, and analyzed as in Figure 5B. Data represents difference in nuclear height (ΔZ) for n ≥ 15 observations presented as in 5C. (B) Inducible MKK3DD cells were plated in 1% cocultures and treated with doxycycline (2 µg/ml) in the presence or absence of Amphiregulin (20 ng/ml). Radial histograms of migration angles before (grey) and after (cyan) induction presented as in Figure 6C. Data represents angles of n > 900 cells from ≥6 observations assessed using subsampling and a two-sample KS test with ‘ns’ not significant, ***p<0.001 (see Materials and methods). (C) 10% MKK3DD cocultures were pretreated with Amphiregulin (20 ng/ml, green) and 24 hr doxycycline (2 µg/ml) or media, imaged and analyzed as in Figure 5B–D, and compared to selected conditions reproduced from Figure 6D (grey). Data represents difference in nuclear height (ΔZ) for n ≥ 16 observations normalized to the mean height of parental cells (dashed line), with mean and +/- standard deviation (black bars). Significance was calculated by two-sample t-test with ‘ns’ indicating no significance, ***p<0.001.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (Human) | MCF10A | ATCC | ||
| Cell line (Human) | HEK293FT | Thermo-Fisher | ||
| Recombinant DNA reagent | pLenti H2B-iRFP | PMID:24949979 | H2B-iRFP; 'Nuclear marker' | pSR1881 |
| Recombinant DNA reagent | pLenti PGK-ERK1-mRuby2 | This paper | ERK Localization Sensor | pSR1214, Regot Lab |
| Recombinant DNA reagent | pLenti PGK-ERK-KTR-mCerulean3 | Addgene #90229 PMID:24949979 | ERK-KTR; ERK Kinase Translocation Reporter | pTA30, Regot Lab |
| Recombinant DNA reagent | pLenti H2B-mClover | This paper | H2B-Clover | pTA54, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G::FGFR1WT | This paper | TRE3G::FGFR1WT | pTA46, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G::FGFR2WT | This paper | TRE3G::FGFR2WT | pHC127, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G::EGFR1WT | This paper | TRE3G::EGFR1WT | pHC132, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G::HER2WT | This paper | TRE3G::HER2WT | pHC123, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G::KRASWT | This paper | TRE3G::KRASWT | pHC131, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G::KRASG12V | This paper | TRE3G::KRASG12V | pHC136, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G::BRAFWT | This paper | TRE3G::BRAFWT | pHC142, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G::BRAFV600E | This paper | TRE3G::BRAFV600E | pHC125, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G::MEK1WT | This paper | TRE3G::MEK1WT | pHC134, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G:: MEK1DD | This paper | TRE3G:: MEK1DD | pAP53, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G:: MEK2WT | This paper | TRE3G:: MEK2WT | pHC126, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G:: MEK2DD | This paper | TRE3G:: MEK2DD | pHC141, Regot Lab |
| Recombinant DNA reagent | pLenti PGK::rtTA, TRE3G:: MKK3DD | This paper | TRE3G:: MKK3DD | pAP55, Regot Lab |
| Recombinant DNA reagent | CMV Puro DEST | Addgene #17452 PMID:19657394 | ||
| Recombinant DNA reagent | PGK Puro DEST | Addgene #19068 PMID:19657394 | ||
| Recombinant DNA reagent | pEGFP-C1 Utr261-EGFP | Addgene #58471 PMID:26317264 | ||
| Recombinant DNA reagent | pLenti PGK-Utr261-EGFP puro | This paper | pTA152, Regot Lab | |
| Recombinant DNA reagent | pLenti PGK-P38-KTR-mClover | This paper | pAP50, Regot Lab | |
| Recombinant DNA reagent | pLenti PGK-JNK-KTR-mRuby2 | Addgene #59154 PMID:24949979 | pSR1846 | |
| recombinant DNA reagent | lentiCRISPR_V2_Neo | Gift from Dr. Andrew Holland | lentiCRISPR_V2_Puro on addgene as #52961 | |
| sequence-based reagent | ADAM17KO guide | This paper, from IDT | 5’-CTACAGATACATGGGCAGAG-3’ (targets R241 of exon 6) | |
| Recombinant DNA reagent | pLenti CRISPR ADAM17KO Neo | This paper | ADAM17KO | pTA70, Regot Lab |
| Chemical compound, drug | PD-0325901 | Selleck Chemicals #S1036 | MEKi; MEK inhibitor | 5µM |
| Chemical compound, drug | Batimastat | Selleck Chemicals #S7155 | MPi; MMP/ADAM inhibitor | 5µM |
| Chemical compound, drug | Gefitinib | Selleck Chemicals #S1025 | EGFRi; EGFR inhibitor | 5µM |
| Chemical compound, drug | BIRB-796 | Selleck Chemicals # S1574 | P38i; p38 inhibitor; BIRB | 5µM |
| Chemical compound, drug | SB-203580 | Sigma # S8307 | P38i; p38 inhibitor; SB | 25µM |
| Chemical compound, drug | SKII | Selleck Chemicals #S7176 | SKi; Sphingosine Kinase inhibitor | 10µM |
| Chemical compound, drug | JTE-013 | Selleck Chemicals # S128 | S1PR2i; S1PR2 inhibitor | 10µM |
| Peptide, recombinant protein | Doxycycline | Sigma #D9891 | Dox | 2μg/ml |
| Peptide, recombinant protein | Amphiregulin | Peprotech #100-55B | AREG | 20ng/ml |
| Peptide, recombinant protein | TGFβ | R&D Systems #7754-BH | TGFβ | 5ng/ml or 50ng/ml |
| Peptide, recombinant protein | EGF | Peprotech #AF-100-15 | EGF | MCF10A culture |
| Peptide, recombinant protein | Insulin | Sigma #I0516 | MCF10A culture | |
| Peptide, recombinant protein | Penicillin/ Streptomycin | Sigma #P0781 | MCF10A culture | |
| Peptide, recombinant protein | Cholera Toxin | Sigma # C-8052 | MCF10A culture | |
| Peptide, recombinant protein | Hydrocortisone | Sigma #H-0888 | MCF10A culture | |
| Other | Horse Serum | Gibco #16050-122 | MCF10A culture | |
| Other | DMEM/F12 | Gibco #11030-032 | MCF10A culture | |
| Other | 0.25% Trypsin-EDTA | Gibco #25300-054 | MCF10A culture | |
| Other | Puromycin | Sigma #P8833 | 1 μg/ml | |
| Other | Blasticidin | Corning #30-100-RB | 3μg/ml | |
| Other | Hygromycin | Gibco #10687010 | 10 μg/ml | |
| Other | Neomycin | Sigma #N6386 | 500μg/ml | |
| Other | Lipofectamine 2000 | Thermo Fisher #11668-027 | For lentiviral production | |
| Other | Polybrene | EMD/Millipore #TR-1000-G | 10 μg/ml, for lentiviral infection | |
| Other | Human Plasma Fibronectin | EMD/Millipore #FC010 | ||
| Chemical compound, drug | EdU | Thermo Fisher # A10044 | EdU | 10μM |
| Chemical compound, drug | Alexa-Fluor Azide 488 click labelling | Thermo Fisher # A10266 | ||
| Chemical compound, drug | DAPI | Thermo Fisher # D3571 | ||
| Antibody | Anti-Amphiregulin Antibody (mouse monoclonal) | R & D Systems #MAB262 | AREG FB-Ab | 50ng/ml |
| Antibody | Anti-ADAM17 Antibody (rabbit polyclonal) | CST #3976S | α-ADAM17 | 1:1,000 |
| Antibody | Anti-BRAF Antibody (rabbit monoclonal) | CST #14814S | α-BRAF | 1:1,000 |
| Antibody | Anti-E-Cadherin Antibody (rabbit monoclonal) | CST #3195S | α-ECad | 1:500 |
| Antibody | Anti-N-Cadherin Antibody (rabbit monoclonal) | CST #13116S | α-NCad | 1:200 |
| Antibody | Anti-HSC70 Antibody (mouse monoclonal) | Santa Cruz Biotechnology | α-HSC70 | 1:1,000 |
| Antibody | IRDye anti-rabbit 800 (donkey polyclonal) | Licor #925-32212 | 1:10,000 | |
| Antibody | IRDye anti-mouse 680 (goat polyclonal) | Licor #925-68070 | 1:10,000 | |
| Antibody | anti-Rabbit IgG Alexa Fluor 405 (donkey polyclonal) | Abcam #175649 | 1:400 | |
| Software, algorithm | CellProfiler | https://cellprofiler.org/ | ||
| Software, algorithm | findPeaks matlab script | T. C. O’Haver, 2014; Mathworks.com | modified to detect rate of change between gaussian-fitted minima and maxima from single cell traces | |
| Software, algorithm | Proteome Discoverer | Thermo Fisher, v2.3 | ||
| Software, algorithm | Mascot | Matrix Science, v2.6.2 | ||
| Software, algorithm | preprocessImagesCaller.py | Aikin T., Peterson A., Pokrass M., Clark H., Regot S., PreprocessImagesCaller. GitHub. https://github.com/tjaikin/Regot-Lab/blob/Aikin_2020/preprocessImagesCaller.py. dc08aeb. | ||
| Software, algorithm | preprocessImages.py | Aikin T., Peterson A., Pokrass M., Clark H., Regot S., PreprocessImages. GitHub. https://github.com/tjaikin/Regot-Lab/blob/Aikin_2020/preprocessImages.py. dc08aeb. | ||
| Software, algorithm | flatfielding.py | Aikin T., Peterson A., Pokrass M., Clark H., Regot S., Flatfielding. GitHub. https://github.com/tjaikin/Regot-Lab/blob/Aikin_2020/flatfielding.py.dc08aeb. | ||
| Software, algorithm | registerAndCrop.py | Aikin T., Peterson A., Pokrass M., Clark H., Regot S., registerAndCrop. GitHub. https://github.com/tjaikin/Regot-Lab/blob/Aikin_2020/registerAndCrop.py.dc08aeb. | ||
| Software, algorithm | trackOrganizeCpDataCaller.py | Aikin T., Peterson A., Pokrass M., Clark H., Regot S., trackOrganizeCpDataCaller. GitHub. https://github.com/tjaikin/Regot-Lab/blob/Aikin_2020/trackOrganizeCpDataCaller.py.dc08aeb. | ||
| Software, algorithm | trackOrganizeCpData.py | Aikin T., Peterson A., Pokrass M., Clark H., Regot S., PreprocessImages. GitHub. https://github.com/tjaikin/Regot-Lab/blob/Aikin_2020/trackOrganizeCpData.py.dc08aeb. |
Additional files
-
Supplementary file 1
Table of secreted factors identified by mass spectrometry.
Supernatants from ADAM17KO or WT cells expressing (+Dox) or not expressing (-Dox) BRAFV600E were collected and analyzed by Tandem-Mass-Tag (TMT) mass spectrometry as described in methods. Table shows fold change and p-values for the 24 identified factors presented in Figure 3D in red.
- https://cdn.elifesciences.org/articles/60541/elife-60541-supp1-v1.pdf
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/60541/elife-60541-transrepform-v1.pdf