SON and SRRM2 are essential for nuclear speckle formation
Figures
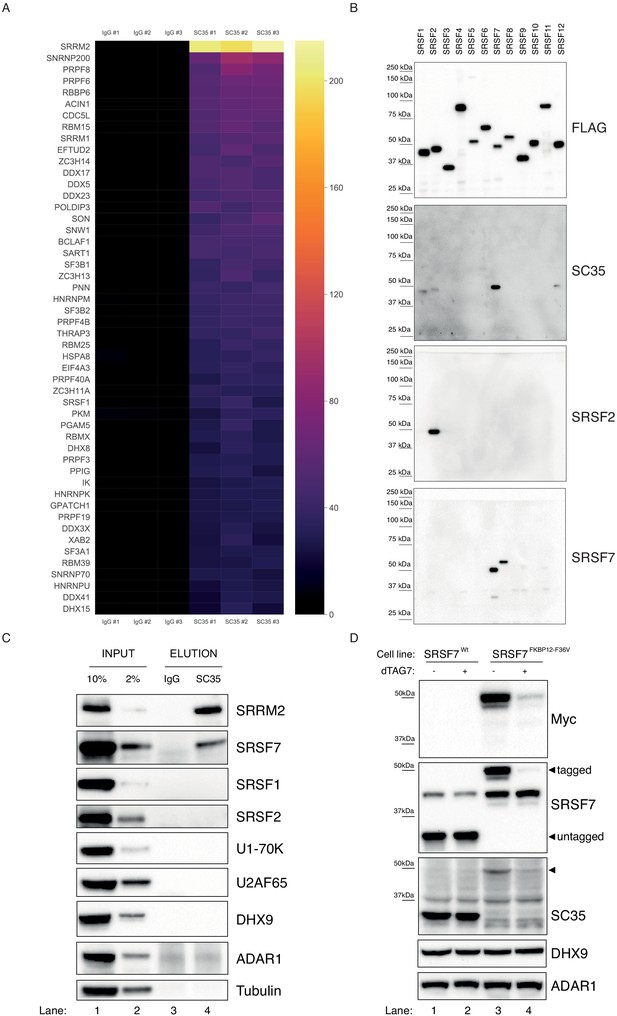
SC35 mAb immunoprecipitation followed by MS identifies SRRM2 as the top hit.
(A) The Top50 hits identified by the MS are depicted on a heatmap showing the number of unique peptides detected for each protein. Also see Figure 1—figure supplement 1A for an intensity vs MS/MS spectra plot and Supplementary file 2 for raw MaxQuant results (B) Streptavidin pull-down of biotin tagged ectopically expressed SRSF proteins 1 to 12 are blotted with FLAG antibody in order to show the amounts of loaded proteins on PAGE. Western blot using mAb SC35 detects SRSF7 with highest sensitivity in comparison to all other SRSF proteins, but also weakly reacts with SRSF1, 2, and 12. Specific antibodies against SRSF2 and SRSF7 are used to validate the authenticity of the purified proteins from stable cell lines in corresponding lanes and blots. (C) SC35-IP performed on lysates from wild-type HEK293 cells identifies SRRM2 as the most enriched protein with a weaker enrichment for SRSF7 but no enrichment for SRSF1 or SRSF2 using western blotting (compare Lane 4 across blots). (D) Homozygous knock-in of the 2xMyc-FKBP12F36V tag into SRSF7 gene locus shifts the SC35 band from 35 kDa to 50 kDa (compare Lanes 1 and 3 on the SC35 blot) and upon induction of degradation with dTAG7 the shifted band is lost (compare Lanes 3 and 4 in Myc, SRSF7, and SC35 blots). This blot validates that the 35 kDa band identified by mAb SC35 blots corresponds to SRSF7.
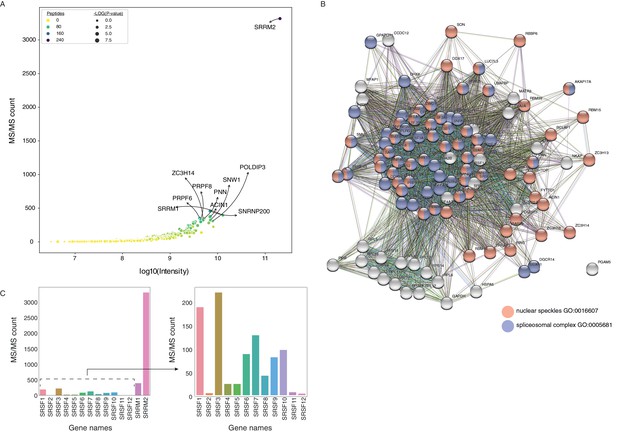
SC35 pull-down followed by MS identifies SRRM2 as the top hit and multiple spliceosomal components are co-purified together with SRRM2.
(A) The log of intensities is plotted against detected MS/MS spectra for significantly enriched proteins in the SC35 immunoprecipitations. SRRM2 together with nine most enriched proteins are annotated on the plot. (B) SC35 targets in the top quartile (108 proteins) are submitted into the STRING database and a network representation is shown. Red circles depict nuclear speckle GO-term category and blue circles depict spliceosomal complex GO-term category. (C) The MS/MS spectral counts for SRSF1 to 12 are plotted together with SRRM1 and SRRM2 (left) and without SRRM1 or SRRM2 (right) show a significant enrichment for SRRM2 amongst other SR-proteins.
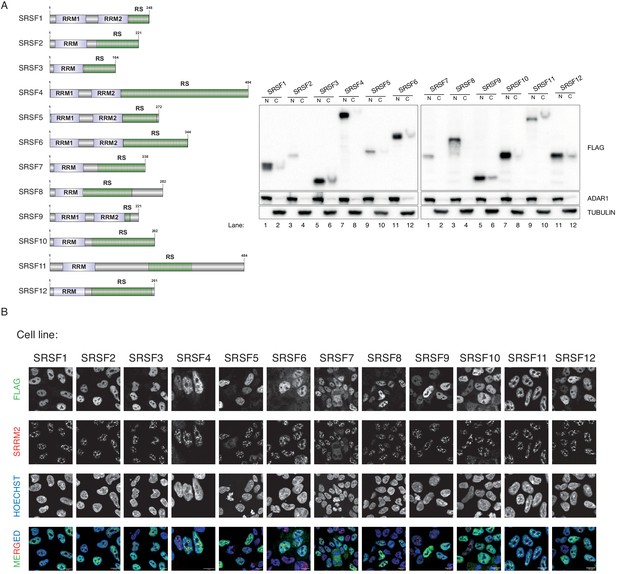
A validation for all stable cell lines with transgenic SRSF proteins show only weak localization to the NS.
(A) The domain and size representation of SRSF proteins are shown (left). The inducible stable cell lines are characterized using PAGE (right). The nuclear (N) and cytoplasmic (C) fractions are used to show the major localization to the nucleus is unaffected when the SRSF proteins are tagged and ectopically expressed. ADAR1 is used as a nuclear marker whereas TUBULIN is used as a cytoplasmic marker (B) The Immunofluorescence experiment is performed by using FLAG-M2 for the tagged SRSF protein and by using SRRM2 (pRb) antibody as an NS marker. Overall NS localization of SRSF proteins is noted, but in a more diffused pattern in comparison to SRRM2. Scale bars = 10 µm.
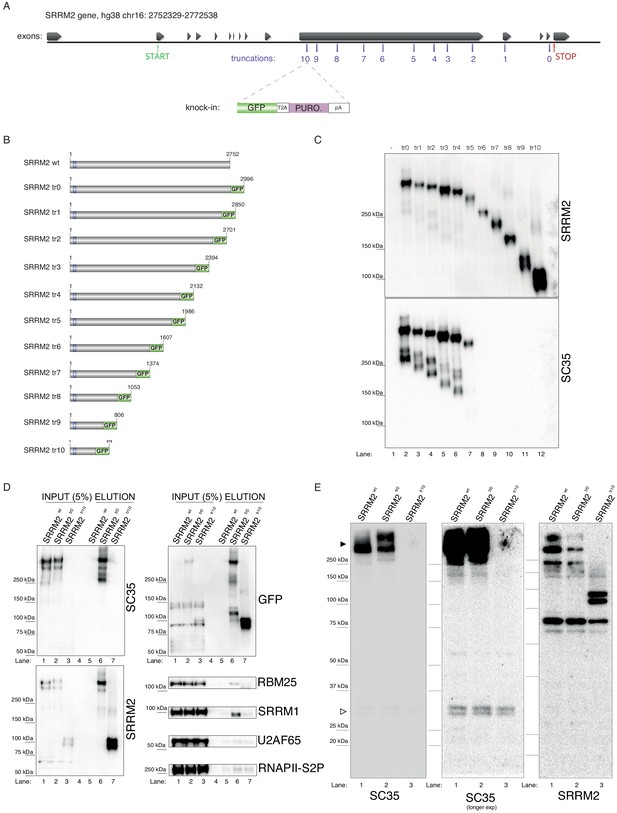
Endogenous truncating mutations of SRRM2 prove mAb SC35 as an SRRM2 antibody.
(A) The strategy for the CRISPaint generated endogenous truncating mutations (0-to-10) accompanied by the TagGFP2 (depicted as GFP for simplicity) fusion are shown. (B) The sizes of SRRM2 truncated GFP fusion proteins are displayed. (C) Protein purified using a GFP-trap pull-down from lysates of corresponding stable HAP1 cell lines carrying the truncated SRRM2 alleles are run on PAGE. Western blotting of SRRM2 using an antibody generated against the common N-terminus is used to show the amount of loaded protein on the gel. SC35 blot shows a significant reduction in signal intensity of SRRM2-tr5 and a complete loss of signal from SRRM2-tr6 to tr10. (D) GFP-trap pull-down performed on lysates from wild-type, tr0 and tr10 HAP1 cells enrich for SRRM2 in tr0 cells, indicating the GFP-tagged allele is specific to SRRM2 and is detected by also SC35 blot (Lanes 1 and 2 inputs compared to Lanes 5 and 6 on the upper and lower left-side blots). SRRM2 also co-purifies two other NS-associated proteins; SRRM1 and RBM25 (Lane 6 on lower right-side blots). SRRM2-tr10 is not detected by SC35 but the pull-down efficiency (Lane 7 on upper left-side blot) and loading is validated by SRRM2 (Lanes 3 and 7 on lower left-side blot) and GFP blots (Lanes 3 and 7 on upper left-side blot). (E) Total cell lysates from wild-type, tr0 and tr10 HAP1 cells are run on 4–12% polyacrylamide gel and blotted with SC35 reveal the high-molecular weight (~300 kDa) as the most intense band and the absence of signal in tr10 cell lines validates that this band represents SRRM2 (filled arrow head). Longer exposure of the blot reveals a weak cross-reactivity with a 35 kDa protein, most likely to be SRSF7, around 35 kDa (empty arrow head).
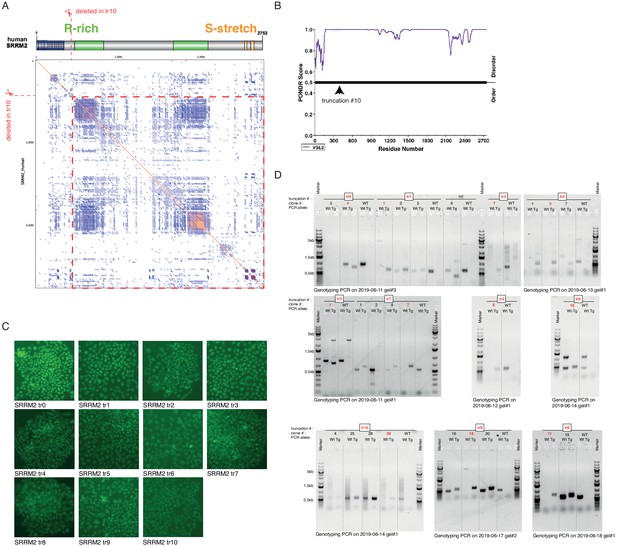
The strategy for making the truncating mutations of SRRM2.
(A) The position of the deepest truncation (tr10) is shown with respect to annotated domains of the protein as well as the dot matrix of the SRRM2 protein sequence. Repetitive regions appear as densed blue squares on the matrix. (B) The PONDR score for disordered regions of SRRM2 protein is displayed with the arrow head pointing to the position of truncation 10. (C) A summary of GFP fluorescence images for truncating mutations before the colony picking is shown. (D) The results of genotyping PCRs are summarized (Wt: PCR carried out using the oligos for wild type allele; Tg:PCR carried out using the oligos for wild type allele. Genotyping oligos and the expected PCR product sizes are listed in Supplementary file 1). Every clone used in this study is highlighted with red text color.
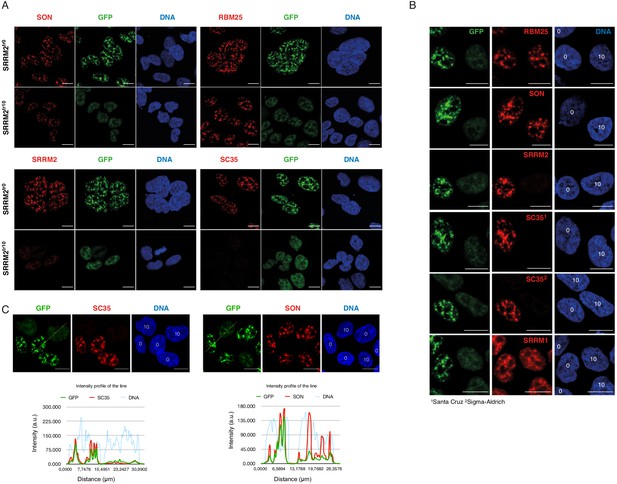
SRRM2 truncation#10 leads to loss of SC35 domains but not NS.
(A) SON and RBM25 antibodies are used as NS markers for IF analysis of both SRRM2tr0 and SRRM2tr10 HAP1 cells. No significant impact on the formation of NS in SRRM2tr10 cells in comparison to SRRM2tr0 cells is observed. Lack of signal for SC35 in SRRM2tr10 cells validates SC35 as an SRRM2 antibody. (B) The SRRM2tr0 and SRRM2tr10 cells are plated together before the IF protocol is performed and the GFP signal intensity as well as SC35 staining are used to distinguish SRRM2tr10 cells from SRRM2tr0 HAP1 cells. The DNA stain marking the nuclei are annotated with ‘0’ or ‘10’ on top to indicate the corresponding cell line. (C) The SRRM2tr0 and SRRM2tr10 HAP1 cells are imaged side-by-side and a line is drawn to quantify the signal intensity across two cell lines. The intensity profile of the lines shows dramatically reduced signal for SC35 between SRRM2tr0 and SRRM2tr10 cells, whereas similar signal intensities for SON and DNA in SRRM2tr0 cells is observed. Scale bars = 10 µm.
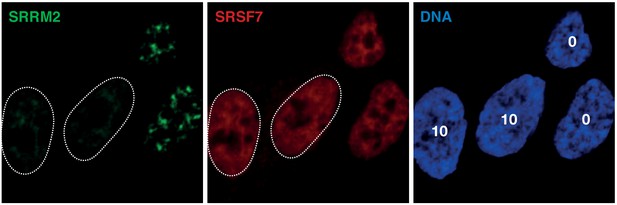
SRSF7 staining in SRRM2tr0 compared to SRRM2tr10 cells.
Immunofluorescence staining of SRSF7 in SRRM2tr0 and SRRM2tr10 cells that are imaged side-by-side show little to no difference in SRSF7 signal in cells that have a truncated SRRM2 that is not recognized by mAb SC35. Green: TagGFP2-SRRM2tr0 or tr10, Red: SRSF7, stained with pAb RN079PW (MBL), Blue: DNA.
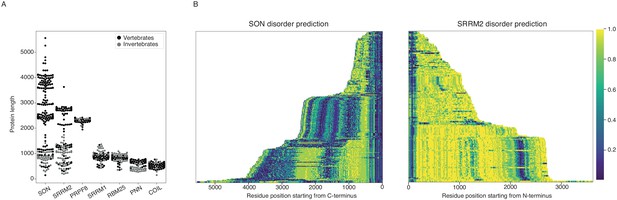
SON and SRRM2 are rapidly evolving and largely disordered proteins.
(A) The size distribution of SON and SRRM2 is highly variable across metazoan species with a mean length of 2227.9 a.a. and SD of 1149.5 for SON and a mean length of 1928.6 a.a. and SD of 919.3 for SRRM2. The lengths of other NS-associated proteins are less variable with a mean length of 895.1 a.a and SD of 104.2 for SRRM1; mean length of 835.4 a.a. and SD of 77.9 for RBM25; mean length of 652.8 a.a. and SD of 118.7 for PNN, mean length of 2332.8 a.a. SD of 40.8 for PRP8. (B) The disorder probability of SON and SRRM2 is predicted using the MobiDB-Lite algorithm, which shows an increase of disordered content with the increase of protein length for SRRM2, and to some extent, SON. The SON and SRRM2 graphs plotted side-by-side do not correspond to the same species, for a phylogeny resolved version of this graph see Figure 4—figure supplement 1 and for the alternative algorithm (IUPred2A) see Figure 4—figure supplement 2. The color is scaled from dark blue to yellow indicating a decrease in order as the value approaches 1.0 (yellow).
-
Figure 4—source data 1
Contains the numerical values of protein lengths shown in Figure 4A and disorder predictions shown in Figure 4B and Figure 4—figure supplements 1 and 2 (.csv) using two alternative algorithms (IUPred2A and MobiDB-Lite).
- https://cdn.elifesciences.org/articles/60579/elife-60579-fig4-data1-v2.zip
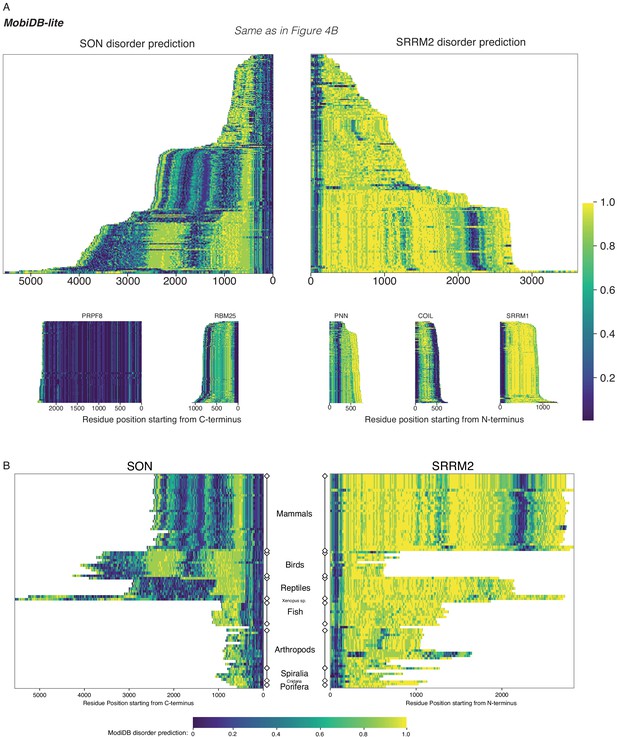
SON and SRRM2 are rapidly evolving and largely disordered proteins.
(A) The disorder probability of SON and SRRM2 is shown the same way as in Figure 4B calculated using MobiDB-Lite (up). The same analysis is also carried out on a spliceosomal core protein PRPF8, on other NS-associated proteins RBM25, PNN and SRRM1, and on a different nuclear body (Cajal bodies) scaffold protein COIL. The length for the highly disordered protein SRRM1 does not vary to a similar extent as SON or SRRM2, indicating the length changes are not a direct consequence of disorderedness. (B) The disorder probability of SON and SRRM2 is shown in phlogenetically matched order using MobiDB-Lite for disorder prediction. In mammals the length of both proteins seems to be fixed, whereas birds have shorter SRRM2 compared to mammals but longer SON. Strikingly, Xenopus tropicalis and laevis have both SON and SRRM2 increased in length.
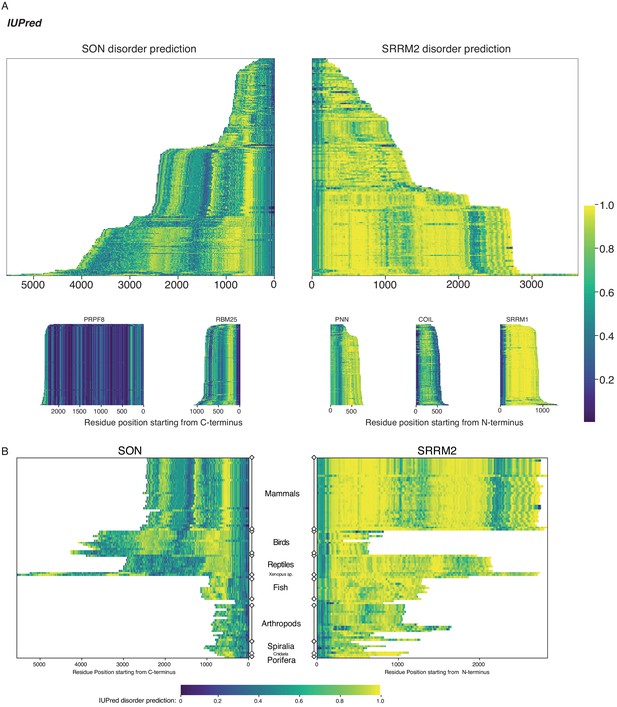
SON and SRRM2 are rapidly evolving and largely disordered proteins.
(A) The disorder probability of SON and SRRM2 is shown similar to Figure 4B and Figure 4—figure supplement 1A calculated using IUPred2A. Both disorder prediction tools resulted in similar plots. (B) The disorder probability of SON and SRRM2 is shown in phlogenetically matched order using IUPred2A for disorder prediction, similar to Figure 4—figure supplement 1B. Both disorder prediction tools resulted in similar plots.
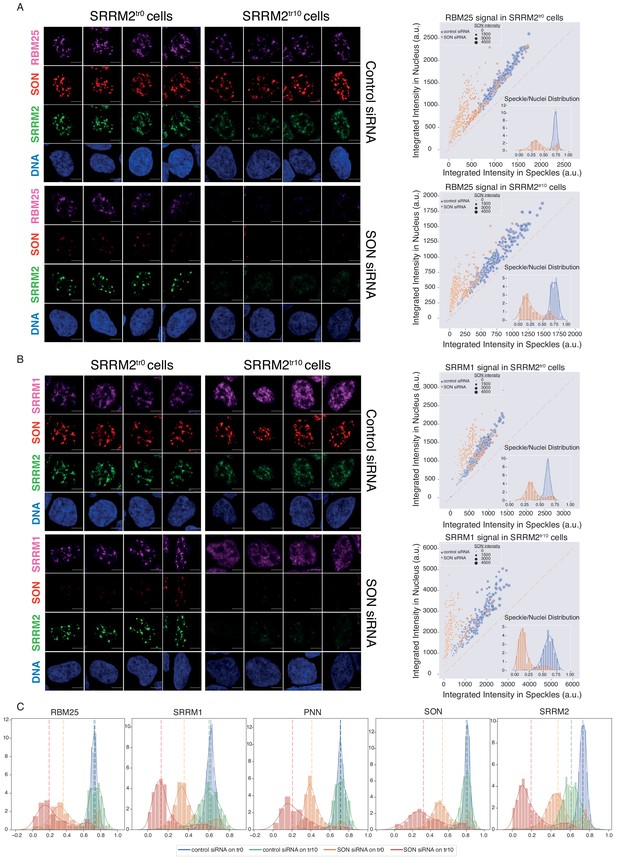
SON and SRRM2 form NS in human cells.
(A) RBM25 IF signal is shown for four individual cells in each siRNA treatment (control or SON siRNA) in SRRM2tr0 and SRRM2tr10 HAP1 cells. The NS localization of RBM25 is severely reduced upon SON knock-down in SRRM2tr0 cells, and completely lost upon SON knock-down in SRRM2tr10 cells. The quantification of the RBM25 signal within the nucleus is plotted against the RBM25 signal within NS (right panel) using ilastik to train detection of NS and CellProfiler for quantification on 10 imaged fields with a 63X objective (in SRRM2tr0 cells control n = 329, SON-KD n = 422; in SRRM2tr10 cells control n = 329, SON-KD n = 402). Each circle represents a cell and the size of the circles is proportionate to the signal intensity of SON. Inset shows the distribution of the ratio of signal detected in NS over signal detected in the nucleus of each cell. (B) SRRM1 IF signal is shown for four individual cells in each siRNA treatment (control or SON siRNA) in SRRM2tr0 and SRRM2tr10 HAP1 cells. The NS localization of SRRM1 is reduced in SON knock-down in SRRM2tr10 cells and lost upon SON knock-down in SRRM2tr10 cells. The quantification of the SRRM1 signal within the nucleus is plotted against the SRRM1 signal within NS (right panel) using ilastik to train detection of NS and CellProfiler for quantification on 10 imaged fields with 63X objective (in SRRM2tr0 cells control n = 494, SON-KD n = 229; in SRRM2tr10 cells control n = 225, SON-KD n = 247). Inset shows the distribution of the ratio of signal detected in NS over signal detected in the nucleus of each cell. (C) Distribution plots showing the ratio of signal detected in NS over signal detected in the nucleus of each cell, in each condition. The dashed line indicates the median ratio in each condition. See Figure 5—figure supplement 3A for a full version of this analysis for PNN. Scale bars = 5 µm.
-
Figure 5—source data 1
Contains two folders, ‘NS’ and ‘Cajal’.
Both folders contain ilastik models (.ilp) used to train for nuclear speckles and Cajal bodies and Cell Profiler pipelines (.cpproj) used to process the probability maps generated by ilastik. The outputs from Cell Profiler (.csv) files are used to generate the figures in the Jupyter Lab environment (.ipynb), shown in Figure 5 and Figure 5—figure supplement 3 respectively.
- https://cdn.elifesciences.org/articles/60579/elife-60579-fig5-data1-v2.zip
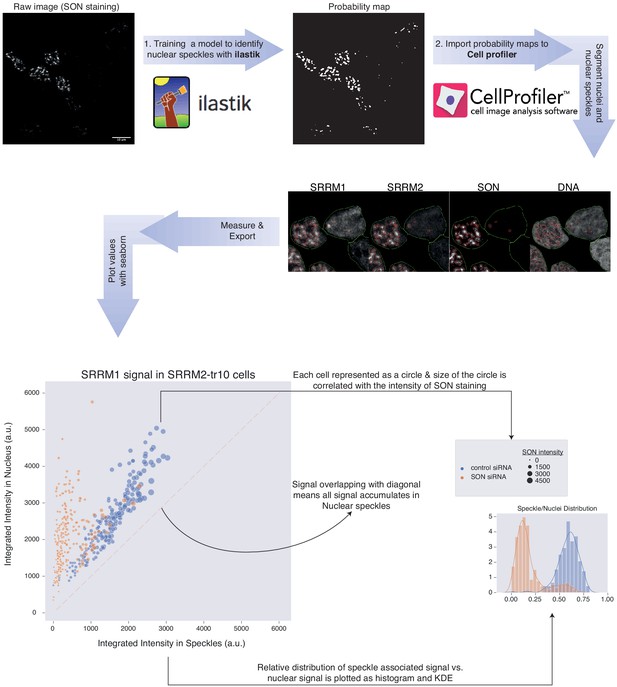
Training a machine learning method for detection of NS.
The pipeline used for the quantification of protein localization in NS under various conditions is depicted here. The probability maps generated with ilastik are imported into CellProfiler analysis software for segmentation and analysis. The numerical values obtained from CellProfiler are then used for the plots which are shown in Figure 5, Figure 5—figure supplements 3–4. The green line marks the nuclear boundaries and the red circles are detected as NS by the algorithm.
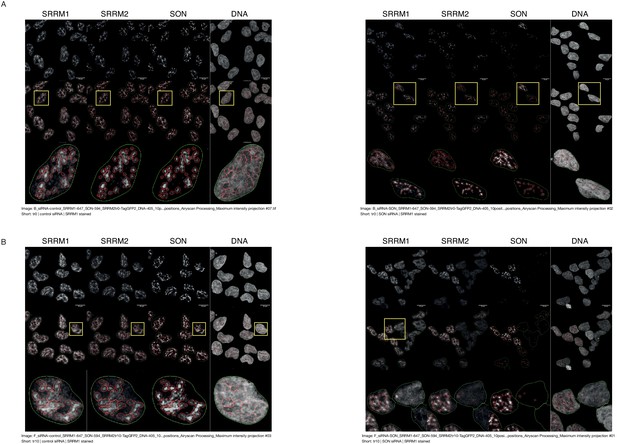
Examples of the trained module for detection of NS on different antibody stainings indicate the model predicts NS robustly for each stained protein.
(A) The outcome of the trained model on recognition of NS is shown for SRRM2tr0 HAP1 cells treated with control siRNA (left) or SON siRNA (right). (B) The outcome of the trained model on recognition of NS is shown for SRRM2tr10 HAP1 cells treated with control siRNA (left) or SON siRNA (right). The green line marks the nuclear boundaries and the red circles are detected as NS by the algorithm. Scale bars = 10 µm.
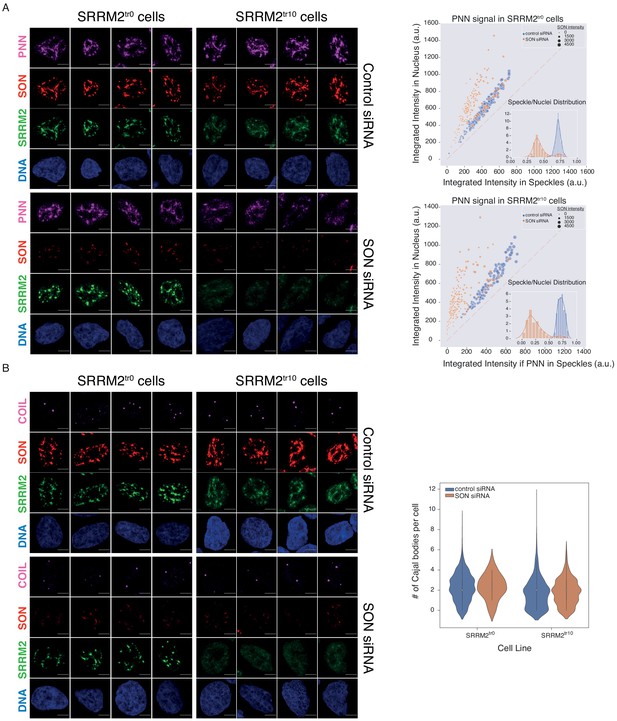
Depletion of SON in SRRM2tr10 cells leads to loss of NS but not of Cajal bodies.
(A) PNN IF signal is shown for four individual cells in each siRNA treatment (control or SON siRNA) in SRRM2tr0 and SRRM2tr10 HAP1 cells. The NS localization of PNN is lost upon SON knock-down in SRRM2tr10 cells. The quantification of the PNN signal within the nucleus is plotted against the PNN signal within NS (right panel) using ilastik to train detection of NS and CellProfiler for quantification on 10 imaged fields with 63X objective (in SRRM2tr0 cells control n = 346, SON-KD n = 172; in SRRM2tr10 cells control n = 138, SON-KD n = 199). Each circle represents a cell and the size of the circles is proportionate to the signal intensity of SON. Inset shows the distribution of the ratio of signal detected in NS over signal detected in the nucleus of each cell. (B) COIL IF signal is shown for four individual cells in each siRNA treatment (control or SON siRNA) in SRRM2tr0 and SRRM2tr10 HAP1 cells. There is no significant change in the localization or of the signal intensity of COIL upon SON knock-down. The quantification of the number of Cajal bodies (based on COIL signal) within the nucleus is shown in the right panel (in SRRM2tr0 cells control n = 337, SON-KD n = 63; in SRRM2tr10 cells control n = 309, SON-KD n = 244). Scale bars = 5 µm.
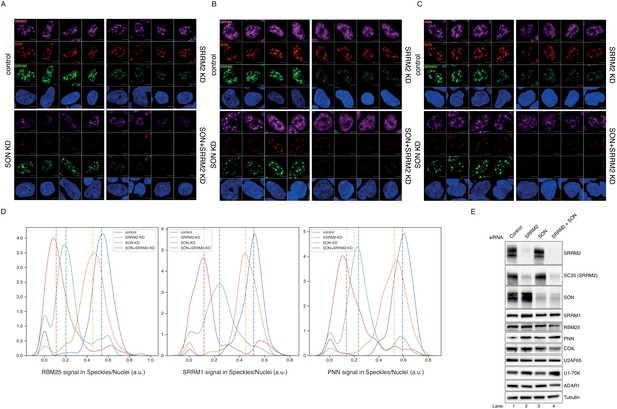
Co-depletion of SON and SRRM2 in SRRM2tr0+GFP HEK293 cells leads to loss of NS.
(A) RBM25 IF signal is shown for four individual cells in each siRNA treatment (control, SRRM2, SON or SRRM2, and SON siRNA) in SRRM2tr0+GFP HEK293 cells. RBM25 signal diffuses out of NS upon SON depletion but is completely lost only in the double knock-down cells. (B) SRRM1 IF signal is shown for four individual cells in each siRNA treatment (control, SRRM2, SON or SRRM2 and SON siRNA) in SRRM2tr0+GFP HEK293 cells. SRRM1 signal diffuses out of NS upon SRRM2 depletion and is mostly diffused in the nucleus in the double knock-down cells. (C) PNN IF signal is shown for four individual cells in each siRNA treatment (control, SRRM2, SON or SRRM2 and SON siRNA) in SRRM2tr0+GFP HEK293 cells. PNN signal diffuses out of NS upon SON depletion and is mostly diffused in the nucleus in the double knock-down cells. Scale bars = 5 µm (D) Distribution plots showing the ratio of signal detected in NS over signal detected in the nucleus of each cell, in each condition. The dashed line indicates the median ratio in each condition, similar to Figure 5C (for RBM25 staining in control siRNA n = 321, in SON-KD n = 187; in SRRM2-KD n = 249, in double-KD n = 202; for SRRM1 staining in control siRNA n = 155, in SON-KD n = 150; in SRRM2-KD n = 197, in double-KD n = 262; for PNN staining in control siRNA n = 204, in SON-KD n = 264; in SRRM2-KD n = 299, in double-KD n = 229). The double knock-down of SRRM2 and SON leads to the most significant loss of signal localised to the NS for RBM25, SRRM1 and PNN. (E) The diffusion of RBM25, SRRM1 or PNN out of the NS is not caused by the down-regulation of the protein levels as shown by western blot (see Lanes 1 and 4).
-
Figure 5—figure supplement 4—source data 1
Contains ilastik models (.ilp) used to train for nuclear speckles, and Cell Profiler pipelines (.cpproj) used to process the probability maps generated by ilastik.
The outputs from Cell Profiler (.csv) files are used to generate the figures in the Jupyter Lab environment (.ipynb), shown in Figure 5—figure supplement 4.
- https://cdn.elifesciences.org/articles/60579/elife-60579-fig5-figsupp4-data1-v2.zip
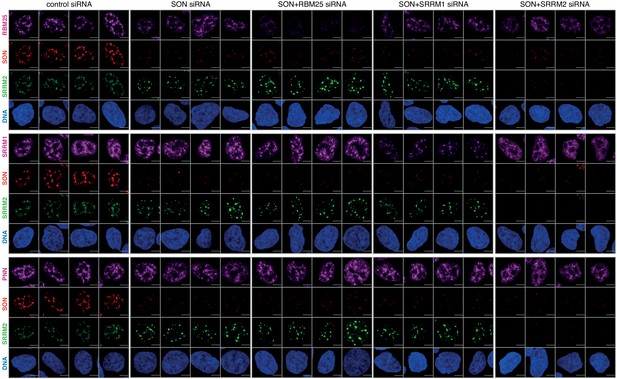
Co-depletion of SON with RBM25 or SRRM1 in SRRM2tr0 HAP1 cells does not lead to loss of spherical NS.
Knock-down experiments are performed similar to Figure 5—figure supplement 4, but in SRRM2tr0 HAP1 cells, however this time SON is also co-depleted together with RBM25 or SRRM1. Depletion of SON together with RBM25 or SRRM1 does not lead to loss of NS as can be seen in GFP signal of SRRM2, indicating the depletion of any NS-associated protein together with SON does not lead to dissolution of NS. The NS are lost only when SRRM2 and SON are co-depleted. Scale bars = 5 µm.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Homo sapiens) | SRRM2 | NCBI | Gene ID: 23524 | |
| Gene (Homo sapiens) | SRSF7 | NCBI | Gene ID: 6432 | |
| Gene (Homo sapiens) | SON | NCBI | Gene ID: 6651 | |
| Cell line (Homo sapiens) | HAP1 | Horizon | Cat. #: C631 | |
| Cell line (Homo sapiens) | Flp-In T-REx HEK293 | Thermo Fisher Scientific | Cat. #: R78007, RRID:CVCL_U427 | |
| Antibody | SC-35 (Mouse monoclonal) | Sigma-Aldrich (Merck) | Cat. #: S4045, RRID:AB_47751 | IF(1:200) WB(1:1000) |
| Antibody | SC-35 (Mouse monoclonal) | Santa Cruz Biotechnology | Cat. #: sc-53518, RRID:AB_671053 | IF(1:100) |
| Antibody | SRRM2 (Rabbit polyclonal) | Thermo Fisher Scientific | Cat. #: PA5-66827, RRID:AB_2665182 | IF(1:100) WB(1:1000) |
| Antibody | SON (Mouse monoclonal) | Santa Cruz Biotechnology | Cat. #: sc-398508 RRID:AB_2868584 | IF(1:100) |
| Antibody | SON (Rabbit polyclonal) | Sigma-Aldrich (Merck) | Cat. #: HPA023535, RRID:AB_1857362 | IF(1:200) WB(1:1000) |
| Antibody | RBM25 (Rabbit polyclonal) | Sigma-Aldrich (Merck) | Cat. #: HPA070713, RRID:AB_2686302 | IF(1:200) WB(1:1000) |
| Antibody | SRRM1 (Rabbit polyclonal) | abcam | Cat. #: ab221061, RRID:AB_2683778 | IF(1:600) WB(1:2000) |
| Antibody | PNN (Rabbit polyclonal) | abcam | Cat. #: ab244250, RRID:AB_2868585 | IF(1:200) WB(1:1000) |
| Antibody | coilin (Rabbit monoclonal) | Cell Signaling | Cat. #: 14168, RRID:AB_2798410 | IF(1:800) WB(1:2000) |
| Antibody | GFP (Rabbit polyclonal) | Chromotek | Cat. #: PAGB1, RRID:AB_2749857 | WB(1:1000) |
| Antibody | SRSF7 (Rabbit polyclonal) | MBL | Cat. #: RN079PW, RRID:AB_11161213 | IF(1:200) WB(1:1000) |
| Antibody | SRSF1 (Mouse monoclonal) | Santa Cruz Biotechnology | Cat. #: sc-33652, RRID:AB_628248 | WB(1:1000) |
| Antibody | SRSF2 (Rabbit monoclonal) | Abcam | Cat. #: ab28428, RRID:AB_777854 | WB(1:1000) |
| Antibody | U1-70K (Mouse monoclonal) | Santa Cruz Biotechnology | Cat. #: sc-390899, RRID:AB_2801569 | WB(1:1000) |
| Antibody | U2AF65 (Mouse monoclonal) | Santa Cruz Biotechnology | Cat. #: sc-53942, RRID:AB_831787 | WB(1:1000) |
| Antibody | DHX9 (Rabbit monoclonal) | abcam | Cat. #: ab183731, RRID:AB_2868586 | WB(1:1000) |
| Antibody | ADAR (Mouse monoclonal) | Santa Cruz Biotechnology | Cat. #: sc-73408, RRID:AB_2222767 | WB(1:1000) |
| Antibody | Tubulin (Mouse monoclonal) | Santa Cruz Biotechnology | Cat. #: sc-32293, RRID:AB_628412 | WB(1:2000) |
| Antibody | Myc (Rabbit monoclonal) | Cell Signaling | Cat. #: 2276, RRID:AB_331783 | WB(1:1000) |
| Antibody | FLAG (Mouse monoclonal) | Sigma-Aldrich (Merck) | Cat. #: F3165, RRID:AB_259529 | IF(1:200) WB(1:2000) |
| Antibody | RNAPII-S2P (Rabbit monoclonal) | Cell Signaling | Cat. #: 13499, RRID:AB_2798238 | WB(1:1000) |
| Recombinant DNA reagent | CRISPaint Gene Tagging Kit | Addgene | Cat. #: 1000000086, RRID:Addgene_1000000086 | |
| Sequence-based reagent | RBM25 siRNA | Thermo Fisher Scientific | Cat. #: s33912 | 10 nM |
| Sequence-based reagent | SRRM1 siRNA | Thermo Fisher Scientific | Cat. #: s20020 | 10 nM |
| Sequence-based reagent | SON siRNA | Thermo Fisher Scientific | Cat. #: s13278 | 10 nM |
| Sequence-based reagent | SRRM2 siRNA | Thermo Fisher Scientific | Cat. #: s24004 | 10 nM |
| Commercial assay or kit | Pierce MS- Compatible Magnetic IP Kit (Protein A/G) | Thermo Fisher Scientific | Cat. #: 90409 | |
| Commercial assay or kit | Lipofectamine RNAiMAX Reagent | Thermo Fisher Scientific | Cat. #: 13778075 | |
| Chemical compound, drug | dTAG-7 | Tocris | Cat. #: 6912 | 1 µM |
| Software, algorithm | ilastik | https://www.ilastik.org/ | RRID:SCR_015246 | |
| Software, algorithm | CellProfiler | https://cellprofiler.org/ | RRID:SCR_007358 | |
| Software, algorithm | Jupyter Lab | https://github.com/ jupyterlab/jupyterlab; Kluyver, 2016 | RRID:SCR_018315 |
Additional files
-
Supplementary file 1
The oligos used for: 1. the cloning of SRSF1-12 constructs for making stable-cell lines (shown in Figure 1 and Figure 1—figure supplement 1), 2. guide RNAs used to generate SRRM2 truncation HAP1 cell lines and 3. the genotyping oligos used for characterization of SRRM2 truncation HAP1 cell lines (shown in Figure 2 and Figure 2—figure supplement 1).
- https://cdn.elifesciences.org/articles/60579/elife-60579-supp1-v2.xlsx
-
Supplementary file 2
The MaxQuant processed output files showing peptide and protein identification, accession numbers, % sequence coverage of the protein, and q-values.
(shown in Figure 1 and Figure 1—figure supplement 1) are listed.
- https://cdn.elifesciences.org/articles/60579/elife-60579-supp2-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/60579/elife-60579-transrepform-v2.docx





