Increased processing of SINE B2 ncRNAs unveils a novel type of transcriptome deregulation in amyloid beta neuropathology
Figures
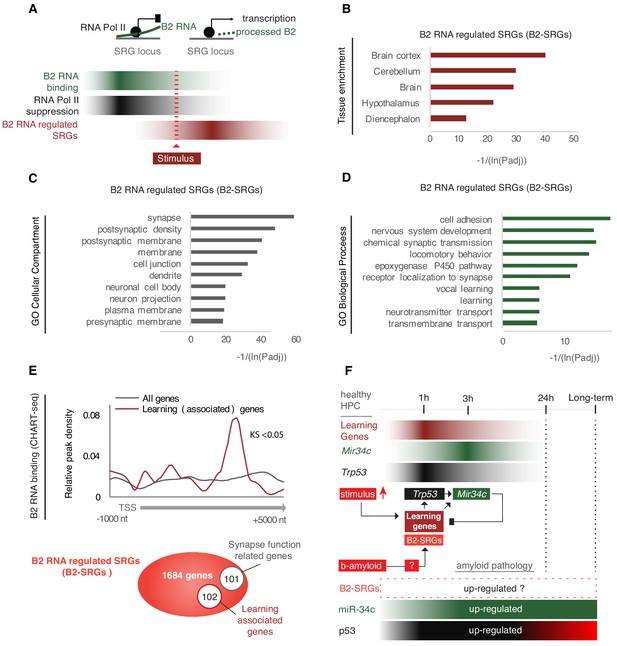
B2-SRGs are enriched in neural functions.
(A) Regulation mode of SRGs by B2 RNA processing based on previous works (Zovoilis et al., 2016; Yakovchuk et al., 2009; Ponicsan et al., 2010). Color intensity represents higher B2 RNA binding (green), Pol II suppression (black) or SRG expression (red). (B) Tissue enrichment analysis of B2-SRGs listed in Supplementary file 1 based on the DAVID functional annotation platform (Huang et al., 2009a; Huang et al., 2009b). The adjusted p-values of top five ranking terms are plotted as a function with higher scores representing lower adjusted p-values (higher statistical significance) ranging from padj 1.92E-04 for ‘Diencephalon’ to padj 8.91E-13 for ‘Brain Cortex’. A complete list of all terms that pass the EASE 0.05 score threshold is presented in Supplementary file 1. (C) Gene ontology (GO) analysis (DAVID) of B2 RNA regulated genes on the basis of cellular compartments. The adjusted p-values of top 10 ranking terms are plotted as a function with higher scores representing lower adjusted p-values ranging from 2.23E-06 for ‘Presynaptic membrane’ to 1.88E-18 for ‘Synapse’. A complete list of all terms that pass the EASE 0.05 score threshold is presented in Supplementary file 1. (D) GO analysis (DAVID) of B2 RNA regulated genes on the basis of biological processes. As above, adjusted p-values of top ranking terms are plotted as a function, with higher scores representing lower adjusted p-values ranging from 0.02 for ‘Transmembrane transport’ to 5.44E-06 for ‘Cell adhesion’. A complete list of all terms that pass the EASE 0.05 score threshold is presented in Supplementary file 1. (E) Upper panel: Metagene analysis of distribution of genomic B2 RNA-binding sites across the start of genes from Zovoilis et al., 2016, comparing learning-associated genes from Peleg et al., 2010 with all genes (Kolmogorov Smirnov test, KS <0.05). TSS represents the Transcription Start Site of these genes. Lower panel: Representation of the number of B2-SRGs that are (i) learning associated based on Peleg et al., 2010 (Supplementary file 1) and (ii) synapse function related based on GO term enrichment (Supplementary file 1, right panel). (F) Regulatory loop of learning genes-Trp53-Mir34c in mouse hippocampus based on Peleg et al., 2010; Zovoilis et al., 2011; Yamakuchi and Lowenstein, 2009. A potential role of B2-SRGs, that include many learning genes, in affecting this loop in amyloid pathology remained unknown after these studies (noted with a question mark in the figure).
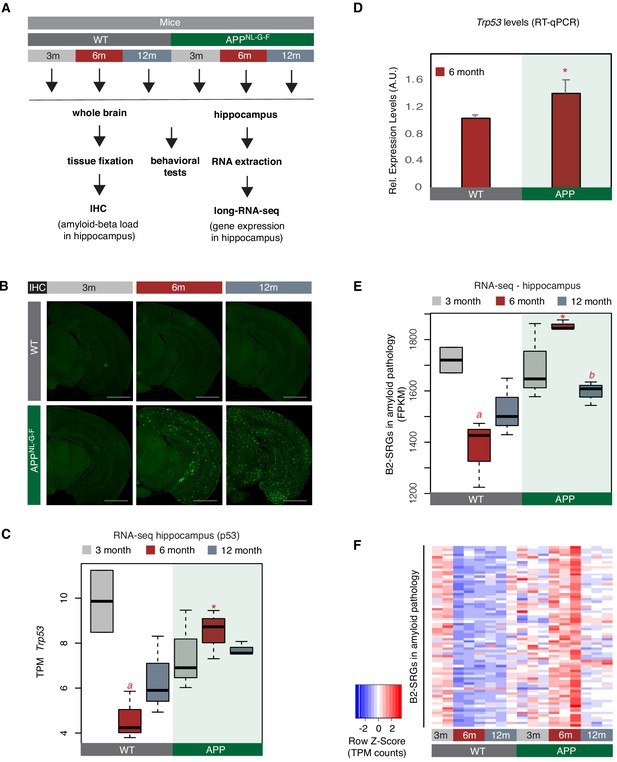
A number of B2-SRGs are hyper-activated in amyloid pathology.
(A) Experimental design for study of B2-SRGs in the hippocampus of the amyloid pathology mouse model (APP) and the respective wild type (WT) control. (B) Immunohistochemistry for identifying amyloid-beta load in the brain sections of mice from the same cohort as in (A) and our previous study (Mehla et al., 2019). Higher fluorescence intensity corresponds to higher amyloid load in APP mice from 6-month-old onwards. (C) Expression levels of the pro-apoptotic gene Trp53 (official symbol Trp53) as defined by long-RNA-seq. Trp53 transcripts per million reads (TPM) from the APP mice compared to WT, grouped by age. Boxplot depicts distribution of expression levels of Trp53 gene among different age groups of mice (WT: wild type, APP: mice with amyloid pathology). Black line denotes median. Statistical significance (p value threshold 0.05) for the comparison between 6-month-old APP and 6-month-old WT (p=0.01) (depicted as asterisk, unpaired, non-directional t-test, n = 3/group) and the comparison between 3-month-old WT (n = 2) and 6-month-old WT (n = 3) (p=0.04) (depicted as a). No significance for the rest of the comparisons between APP and WT in the other two age groups, or between other different ages within the same group (APP or WT). (D) Expression levels of the pro-apoptotic gene Trp53 (official symbol Trp53) in 6-month-old mice as defined by RT-qPCR. Statistical significance (p value threshold 0.05) for 6-month-old APP greater than 6-month-old WT(depicted as asterisk, unpaired directional t-test, n = 3/group, error bars represent standard deviation from the mean). (E) Expression levels as defined by long-RNA-seq of B2-SRGs that are upregulated in amyloid pathology (72 genes, Supplementary file 1). Boxplot depicts distribution of expression levels (in FPKM/Fragments per Kilobase per Million) of B2-SRGs among different age groups of wild type and APP mice. Statistical significance (p value threshold 0.05) for the comparison between 6-month-old APP and 6-month-old WT (p=0.02) (depicted as asterisk, unpaired, non-directional t-test, n = 3/group), for the comparison between 3-month-old WT (n = 2) and 6-month-old WT (n = 3) (p=0.03) (depicted as a) and for the comparison between 6-month-old APP and 12-month-old APP (p=0.005) (depicted as b). No significance for the rest of the comparisons between APP and WT in the other two age groups, or between other different ages within the same group (APP or WT). (F) Gene expression levels of B2-SRGs that are upregulated in amyloid pathology (72 genes, Supplementary file 1) show strong association with amyloid pathology status in the hippocampus of APP and WT mice of different ages. Heatmap depicts gene expression with rows representing B2 RNA regulated genes and columns representing mouse samples. Expression values are normalized per row and correspond to TPM values. Red color represents higher expression.
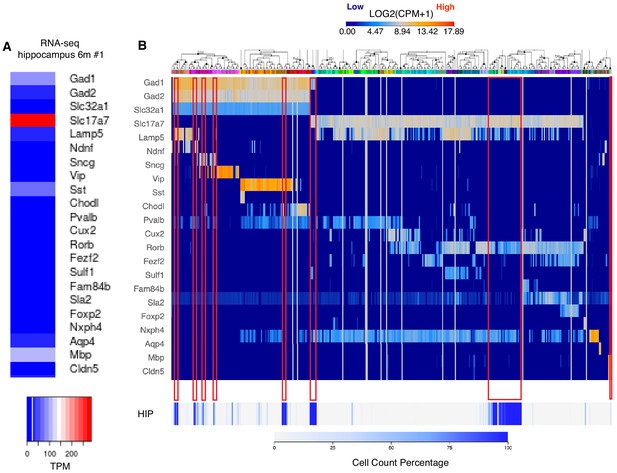
Expression of known hippocampal markers in our RNA-seq data of mice hippocampi.
(A–B) Expression levels comparison between RNA-seq data from WT 6-month-old mouse hippocampus used in our study (panel A) and RNA-seq data from the Allen Brain Atlas (Panel B). Comparison is done for the standard panel genes of reference genes used by Allen Brain Atlas. Every column in panel B represents a different cell type in brain with cell types belonging to hippocampus corresponding to increased density (cell count percentage) at the lower part of panel B labeled as HIP. Red rectangles mark the most prominent cell types that are enriched in Hippocampus.
© 2015, Allen Institute for Brain Science. Image credit: Panel B was taken from the Allen Mouse Brain Atlas, available at: https://portal.brain-map.org/atlases-and-data/rnaseq#Mouse_Cortex_and_Hip. It is not covered by the CC-BY 4.0 licence and further reproduction of this panel would need permission from the copyright holder.
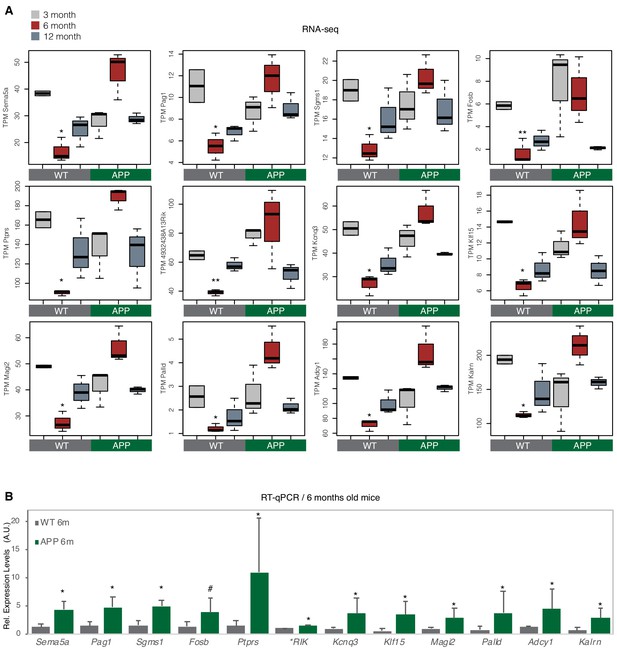
Validation of RNA-seq data in WT and APP mice by RT-qPCR.
(A) Expression levels calculated through RNA-seq for the selected genes to be tested through RT-qPCR in (B). These are 12 from the genes that were found to be upregulated in APP 6-month-old mice compared to WT mice of the same age (Supplementary file 1). The selected subset includes both B2-SRGs (first 10 boxplots) that are part of the 72 B2-SRGs upregulated in APP mice (Supplementary file 1) but also two non-B2-SRGs (last two plots; Adcy1 and Kalrn) that are used later as negative controls (see Figure 7C). Values are based on TPM counts of long-RNA-seq data. One asterisk marks statistical significance (p<0.05) for the comparison between 6-month-old APP and 6-month-old WT (n = 3/group, unpaired non-directional t-test), while two asterisks marks statistical significance only for 6-month-old APP greater than 6-month-old WT (n = 3/group, unpaired directional t-test). All the other comparisons between groups and different ages are non-significant. (B) Confirmation through RT-qPCR of the expression differences between 6-month-old WT and APP mice. Statistical significance (p value threshold 0.05), is depicted as asterisk, for expression in 6-month-old APP mice greater than the expression in 6-month-old WT mice (unpaired directional t-test, n numbers as in (A), error bars represent standard deviation from the mean). # represents a p=0.08 for the same comparison and test. * RIK stands for 4932438A13Rik.
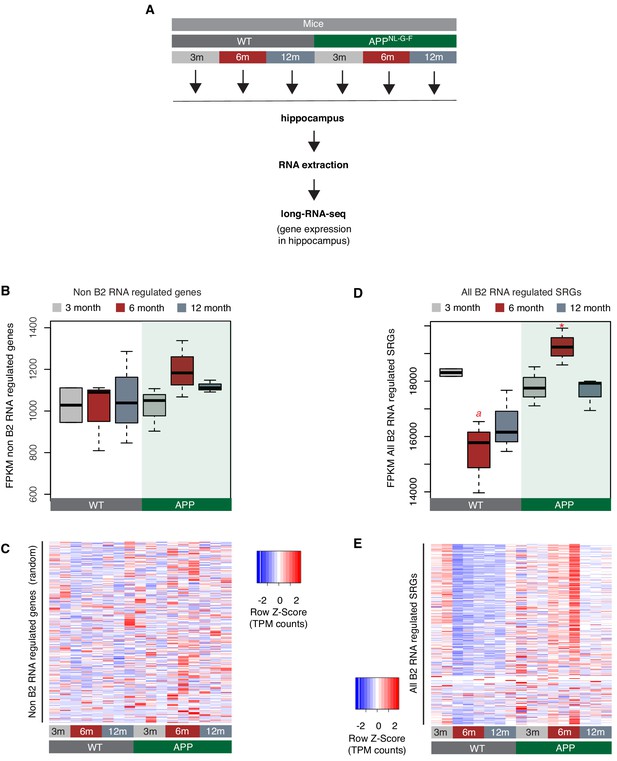
Expression levels of all B2-SRGs in amyloid pathology.
(A) Experimental design for study of B2-SRGs in the hippocampus of the amyloid pathology mouse model (APP) and the respective wild type (WT) control. (B) Expression levels of non-B2 RNA regulated genes as defined by long-RNA-seq. Boxplot depicts distribution of expression levels (in FPKM/Fragments per Kilobase per Million) among different age groups of wild type and APP mice. Not significant difference between 6-month-old APP and 6-month-old WT. (C) Gene expression levels (defined by long-RNA-seq) of a random set of genes not associated with B2 RNAs show weak or no association with amyloid pathology status in the hippocampus of APP and WT mice of different ages. Heatmap depicts gene expression with rows representing 1385 non-B2 RNA regulated genes (Supplementary file 1) and columns representing the different mouse samples. Expression values are normalized per row and correspond to TPM values. (D) Expression levels of B2-SRGs (1684 genes, Supplementary file 1) as defined by long-RNA-seq. Boxplot depicts distribution of expression levels (in FPKM/Fragments per Kilobase per Million) of B2 RNA regulated genes among different age groups of wild-type and APP mice. Statistical significance (p value threshold 0.05) for the comparison between 6-month-old APP and 6-month-old WT (p=0.02)(depicted as asterisk, unpaired, non-directional t-test, n = 3/group) and the comparison between 3-month-old WT and 6-month-old WT (p=0.05)(depicted as a). No significance for the rest of the comparisons between APP and WT in the other two age groups, or between other different ages within the same group (APP or WT). (E) Gene expression levels of B2-SRGs (1684 genes, Supplementary file 1) show strong association with amyloid pathology status in the hippocampus of APP and WT mice of different ages. Heatmap depicts gene expression with rows representing B2 RNA regulated genes and columns representing mouse samples. Expression values are normalized per row and correspond to TPM values. Red color represents higher expression. (Note) When expression dynamics for all B2-SRGs are tested, these are similar to the ones observed for genes in Figure 2, while this pattern in B2 RNA regulated SRG expression was not observed when a random set of genes is tested. Thus, it cannot be excluded that the expression pattern observed in genes of Figure 2 may extend to additional B2-SRGs described in our previous work that have been filtered out by DEseq. This could be attributed to our current sequencing depth and the large dynamic range of some of them as shown in panel E, but also due to genome wide mRNA expression level changes in our system that are not taken into account by DEseq’s conservative hypothesis that expression levels of most genes remain unchanged. This is relevant in our case given the ability of full-length B2 RNAs (which are elevated in our system) to suppress global gene expression and thus, it may have resulted in a number of genes being filtered out during DESeq correction for multiple testing.
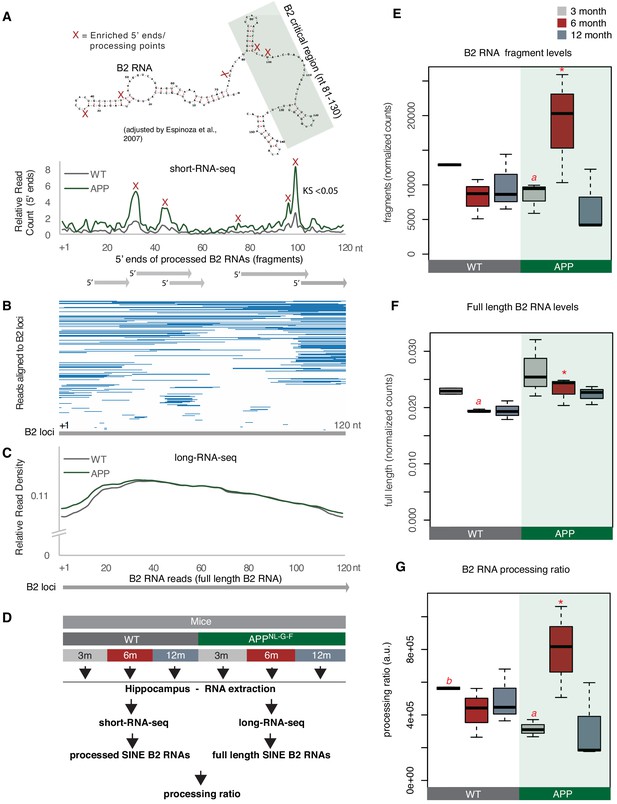
B2 RNA processing ratio is increased in 6-month-old APP mice.
(A) Plotting of the position of the first base (5′ end) of B2 RNA fragments across the B2 loci to depict increased levels of B2 RNA fragments in 6-month-old APP mice. Upper panel: Secondary structure and processing points of B2 RNA. Secondary structure of B2 RNA adapted from Espinosa and colleagues (Espinoza et al., 2007). As in our previous study (Zovoilis et al., 2016), we depict the B2 RNA processing points based on short RNA-seq data and mapping of the 5′ ends of B2 RNA fragments. X marks which cleavage sites of B2 RNA in the upper panel correspond to enriched processing points (the peaks of 5′ end fragments distribution) at the lower panel. The green rectangle depicts the critical region that binds and suppresses RNA Pol II (Yakovchuk et al., 2009; Espinoza et al., 2007; Ponicsan et al., 2010; Ponicsan et al., 2015) that may be affected by such processing points. Lower panel: Distribution of the 5’ends of B2 RNA fragments across the B2 RNA loci based on mapped short RNA-seq from the hippocampi of 6-month-old APP and WT mice. The x axis represents a metagene combining all B2 RNA loci aligned at the start site of their consensus sequence (position +1) and the y axis shows the relative 5′ end count for B2 RNA fragments aligning to any position downstream of position +1. Figure depicts the difference between B2 RNA fragment distribution of 6-month-old APP mice (higher peaks) vs 6-month-old WT mice (lower peaks) (Kolmogorov Smirnov test, KS <0.05). (B) Alignment of short-RNA seq reads mapped against the multiple B2 loci. Only B2 loci with at least one read are depicted. Short RNA-seq reads from one of the 6-month-old APP mice, whose 5′ ends are plotted at (A) lower panel are used here as an example of the length and position of fragments across B2 loci. (C) Metagene analysis of long-RNA-seq reads of mice across B2 RNA loci that is enriched in full-length B2 RNAs but not in B2 RNA fragments. Comparative analysis shows low difference in relative read density of full-length B2 RNAs between WT and 6-month-old APP. X axis as in A. Y axis corresponds to read coverage across the B2 loci. (D) Experimental design for estimation of B2 RNA processing ratio based on short and long-RNA seq data for all APP mice and controls. (E) Boxplot depicts distribution of levels of processed SINE B2 RNA fragments among different age groups of mice between wild type and APP. Statistical significance (p value threshold 0.05) only for 6-month-old APP greater than 3-month-old APP (p=0.04), 12-month-old APP (p=0.04) and 6-month-old WT (p=0.04) (depicted as asterisk, n = 3/group, unpaired directional t-test). All the other comparisons between groups and different ages non-significant except 3-month-old WT less than 3-month-old APP (p=0.03)(depicted as a). (F) Boxplot depicts distribution of levels of full-length SINE B2 RNAs among different age groups of mice between wild type and APP. Statistical significance (p value threshold 0.05) for the comparison between 6-month-old APP and 6-month-old WT (p=0.05; depicted as asterisk, n = 3/group, non-directional unpaired t-test) and for the comparison between 3-month-old WT (n = 2) and 6-month-old WT (n = 3) (p=0.004; depicted as a). All the other comparisons between groups and different ages are non-significant. (G) Boxplot depicts distribution of SINE B2 RNA processing ratio among different age groups of mice between wild type and APP. Statistical significance (p value threshold 0.05) for the comparison between 3-month APP and 6-month-old APP (p=0.04; depicted as a, n = 3/group, non-directional unpaired t-test), for the comparison between 3-month-old WT (n = 2) and 3-month-old APP (n = 3) (p=0.008; depicted as b), and for 6-month-old APP greater than 6-month-old WT (p=0.05; depicted as asterisk, n = 3/group). All the other comparisons between groups and different ages are non-significant.
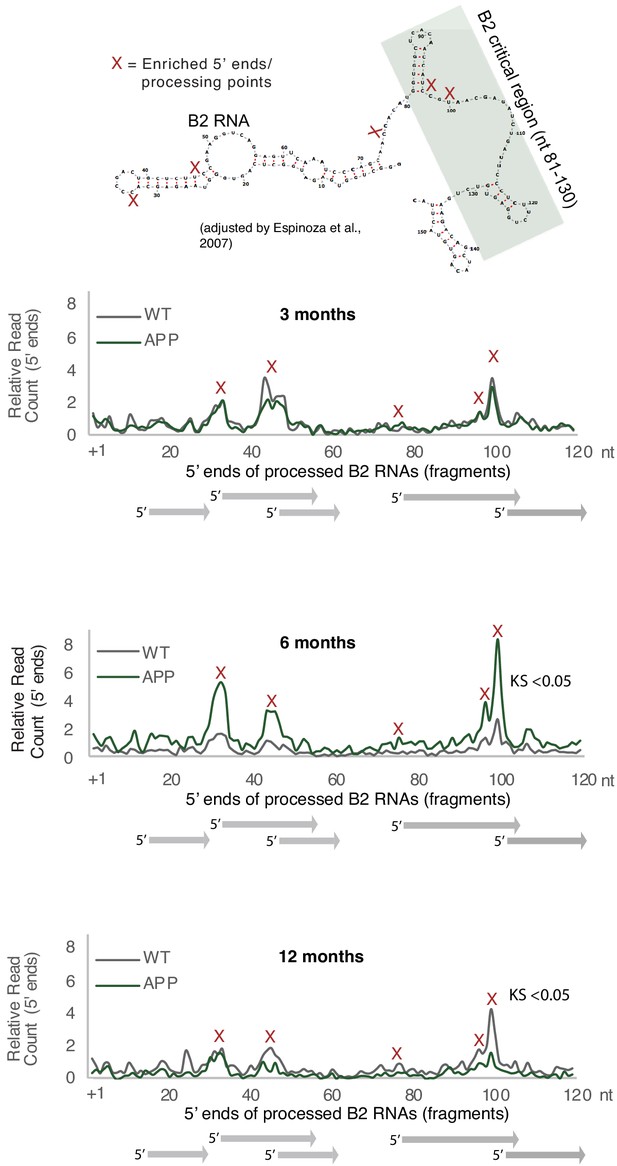
Plotting of the position of the first base (5′ end) of B2 RNA fragments across the B2 loci to compare levels of B2 RNA fragments between APP and WT mice in the three different age groups.
This figure relates to Figure 3A. Upper panel: As in Figure 3A. Secondary structure and processing points of B2 RNA. Secondary structure of B2 RNA adapted from Espinosa and colleagues (Espinoza et al., 2007). Lower panels: Distribution of the 5′ ends of B2 RNA fragments across the B2 RNA loci based on mapped short RNA-seq from the hippocampi of 3-month, 6-month, and 12-month-old APP and WT mice. The x axis represents a metagene combining all B2 RNA loci aligned at the start site of their consensus sequence (position +1) and the y axis shows the relative 5′ end count for B2 RNA fragments aligning to any position downstream of position +1. Figure depicts the difference between B2 RNA fragment distribution of 6-month-old APP mice (higher peaks) vs 6-month-old WT mice (lower peaks) (Kolmogorov Smirnov test, KS <0.05) and the reverse difference of 6-month-old APP mice (higher peaks) vs 6-month-old WT mice (lower peaks) (Kolmogorov Smirnov test, KS <0.05).
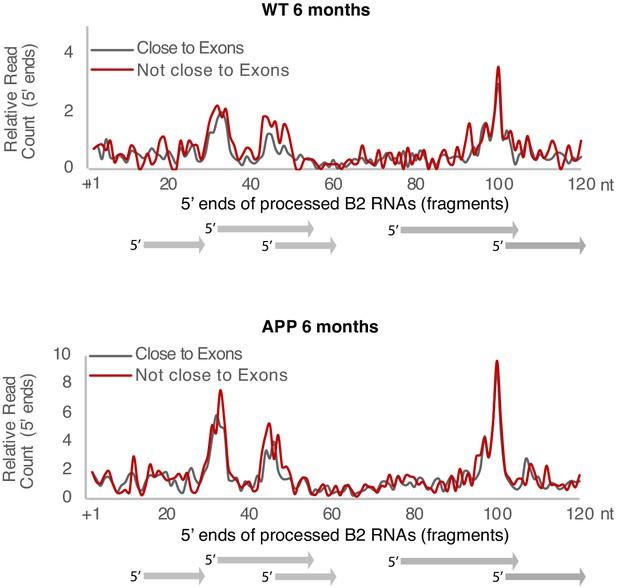
Plotting of the position of the first base (5′ end) of B2 RNA fragments across the B2 loci to compare levels of B2 RNA fragments in 6-month-old mice between B2 elements that overlap exonic/genic regions and those that do not.
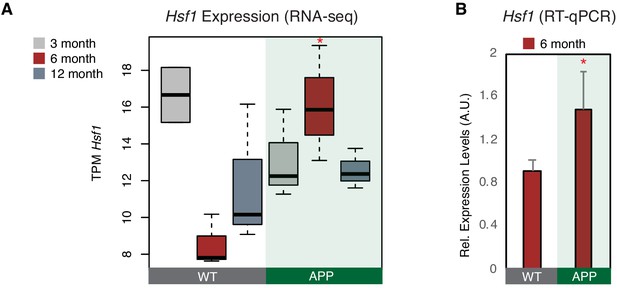
Hsf1 is upregulated in 6-month-old APP mice.
(A) Boxplot depicts distribution of expression levels of Hsf1 gene among different age groups of mice between wild type and APP. Values are based on TPM counts of long-RNA-seq data. Statistical significance (p value threshold 0.05) for the comparison between 6-month-old APP and 6-month-old WT (p=0.03) (n = 3/group, unpaired non-directional t-test). All the other comparisons between groups and different ages are non-significant. (B) Expression levels of Hsf1 in 6-month-old mice as defined by RT-qPCR. Statistical significance (p value threshold 0.05) for 6-month-old APP greater than 6-month-old WT (depicted as asterisk, unpaired directional t-test, n = 3/group, error bars represent standard deviation from the mean).
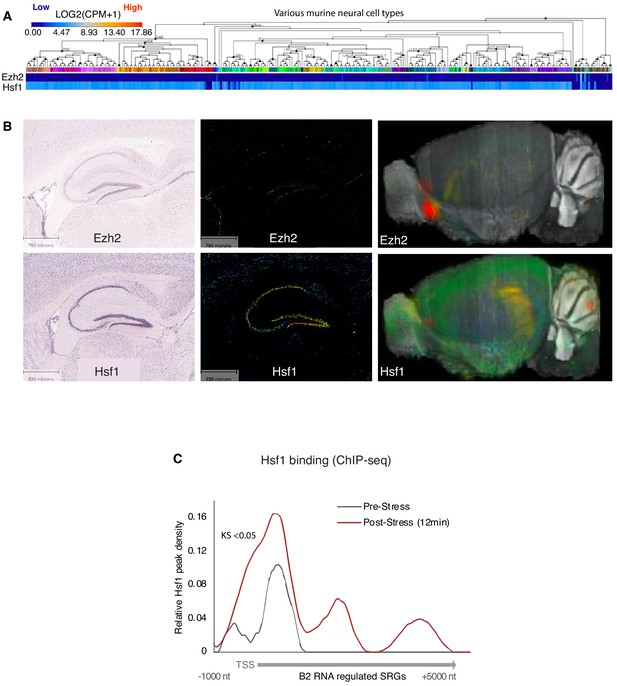
Expression of Hsf1 in neural tissues.
(A) Mouse cortex and hippocampus gene expression levels for Ezh2 and Hsf1 depicted in the Allen Brain Atlas Transcriptomics explorer showing limited expression of Ezh2 across multiple neural tissues compared to Hsf1. Levels are per cell type based on RNA-seq data. (B) In situ hybridization brain image data with cellular-level resolution for Ezh2 (upper panel) and Hsf1 (lower panel) depicting low expression levels for Ezh2. Left panel, ISH images, middle panel, gene expression images, right panel, expression in 3D reconstructed whole mouse brain. (C) Metagene analysis of increase in Hsf1 binding (ChIP-seq data) across the start of B2 RNA regulated genes (Supplementary file 1) between pre-stress and 12 min post-stress (KS <0.05). ChIP-deq data are from Mahat et al., 2016. TSS represents the Transcription Start Site of these genes. This figure provides the rational why we started investigating Hsf1 as a candidate with a potential relation to B2 Biology and a potential effect on B2 RNA processing.
© 2015, Allen Institute for Brain Science. Image credit: The image in panel A was taken from the Allen Mouse Brain Atlas, available at: https://portal.brain-map.org/atlases-and-data/rnaseq#Mouse_Cortex_and_Hip. The images for panel B were taken from the Allen Mouse Brain Atlas, available at https://mouse.brain-map.org/experiment/show/100142521 for Eh2 and https://mouse.brain-map.org/experiment/show/68196972 for Hsf1. These panels are not covered by the CC-BY 4.0 licence and further reproduction of this panel would need permission from the copyright holder.
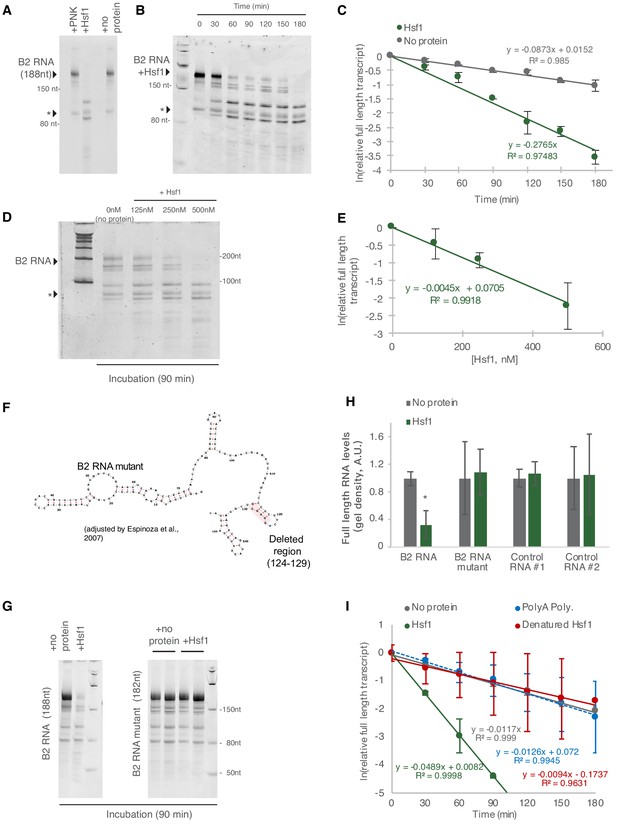
Hsf1 accelerates B2 RNA processing.
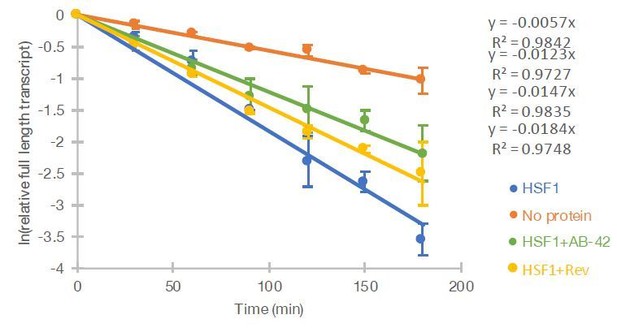
(A) In vitro incubation of B2 RNA. In vitro transcribed and folded B2 RNA at 200 nM incubated with PNK as a control (lane 1), 250 nM Hsf1 (lane 2) and without protein (lane 4). Incubations occurred for 6 hr at 37°C. (B) In vitro incubation of B2 RNA for different incubation periods. In vitro transcribed B2 RNA (100 nM) incubated at 37°C with 500 nM Hsf1 in the course of 180 min with time intervals of 30 min. (C) Relative full-length RNA remaining from (B) using ImageJ area under the curve software over time. The full gels are available as source files. (D) Titration of Hsf1 protein in incubation with 200 nM in vitro transcribed B2 RNA. Concentrations of Hsf1 range from 0 to 500 nM (0 means incubation was performed in the absence of protein and only in the presence of TAP buffer, for the same duration as the other three conditions). (E) Relative full-length RNA remaining from (D) using ImageJ area under the curve software over time. The trendline is a linear fit, displaying standard deviation on the data points. The full gels are available as source files. (F) Graphical representation of the deletion in the B2 RNA mutant on the secondary structure of B2 RNA as adapted from Espinoza et al., 2007. (G) In vitro incubation of the B2 RNA and the B2 RNA mutant for 90 min at 37°C with either Hsf1 or no protein (just TAP buffer, see Materials and methods). (H) Comparison among B2 RNA, the B2 RNA mutant (6nt deletion) and two control RNAs regarding the full-length RNA levels remaining after in vitro incubation for 90 min at 37°C with Hsf1. Sizes of control RNAs are control for RNA #1, 143nt and for control RNA #2, 163nt. or no protein (just TAP buffer, see Materials and methods). Incubation in the absence of Hsf1 but presence of the same buffer (TAP) was used as control to take into account any non-Hsf1-specific RNA destabilization due to non-specific degradation. Asterisk represents statistical significance p=0.005 (unpaired, non-directional t-test) for the comparison between Hsf1 and no protein incubation (three replicates for B2 RNA). The full gels are available as source files. (I) Comparison among Hsf1 (~60 KDa), denatured Hsf1, Poly A polymerase (~56 KDa) and no protein (just TAP buffer) with regard to B2 RNA processing (estimating relative full-length RNA remaining as in C). The full gels are available as source files. For Hsf1, after 90 min no detectable full-length B2 RNA could be measured anymore and no further data points were included in the linear model.
-
Figure 5—source data 1
Full gel images for Figure 5, part 1.
- https://cdn.elifesciences.org/articles/61265/elife-61265-fig5-data1-v2.ps
-
Figure 5—source data 2
Full gel images for Figure 5, part 2.
- https://cdn.elifesciences.org/articles/61265/elife-61265-fig5-data2-v2.ps
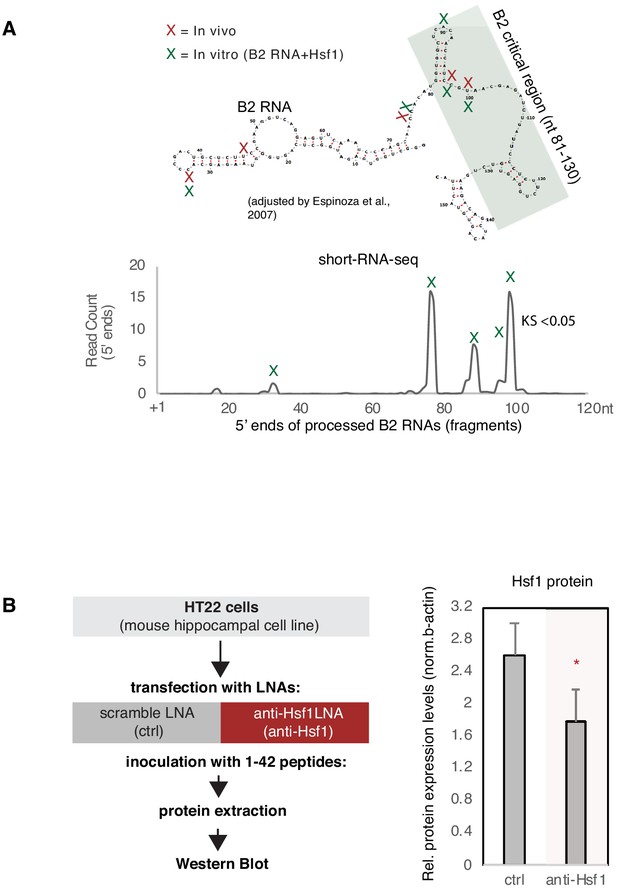
Position of B2 RNA fragments generated in vitro and downregulation of Hsf1 protein levels.
(A) Plotting of the position of the first base (5′ end) of B2 RNA fragments across the B2 loci produced by B2 RNA that has been processed in vitro in the presence of Hsf1. Mapping was done as in case of the in vivo samples (Figure 3) except the normalization step. (B) Estimation by western blotting of Hsf1 protein levels after treatment of HT22 cells with the anti-Hsf1 LNA. Left panel represents the experimental design. Asterisk depicts statistical significance p<0.05 for expression in anti-Hsf1 LNA-treated samples being less than that of samples treated with the scramble LNA (ctrl) (unpaired, directional t-test, n = 3/group).
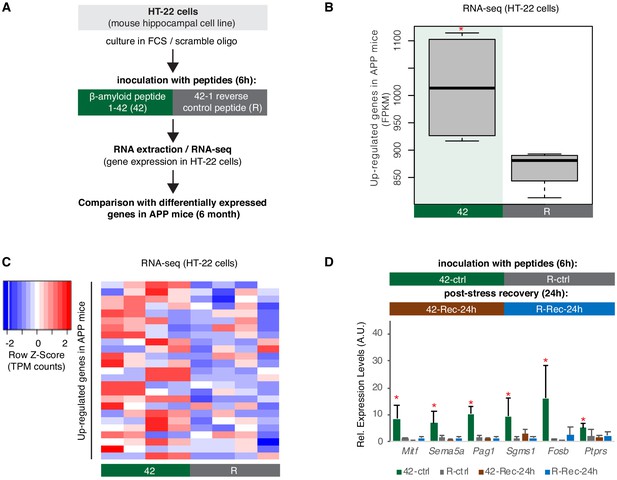
A hippocampal cell culture assay for tracking effects of amyloid beta toxicity on B2 RNA stability.
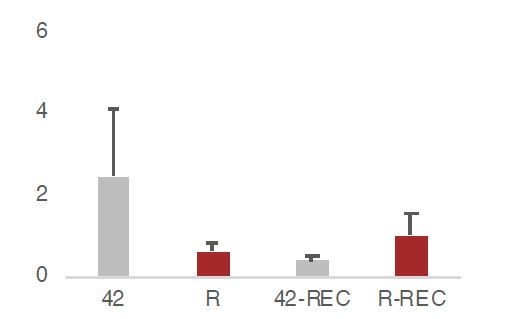
(A) Experimental design for the amyloid toxicity cell culture assay employing HT-22 cells. Cells culture media were supplemented with Fetal Calf Serum (FCS) and the scramble LNA described in Figure 8 to allow comparison with the LNA experiments. (B) Expression levels as defined by long-RNA-seq of genes upregulated in amyloid pathology (6-month-old APP mice) that are also upregulated in amyloid toxicity (25 genes, Supplementary file 1). Boxplot depicts distribution of expression levels (in FPKM/Fragments per Kilobase per Million) of these genes between the two conditions in HT-22 cells. p=0.05 for the comparison between HT22 samples transfected with 1–42 (Espinoza et al., 2007) vs 42–1 reverse control peptide (R) (depicted as asterisk, unpaired non-directional t-test, n = 4/group). (C) Heat map showing gene expression changes during incubation with amyloid peptides in HT22 cells for genes that are also upregulated in amyloid pathology (25 genes, Supplementary file 1) to test whether our cell culture system can simulate the transcriptome observed in amyloid pathology in vivo. Heatmap depicts increase in expression values calculated by long-RNA-seq (TPM values) in the HT22 samples transfected with 1–42 (Espinoza et al., 2007) vs 42–1 reverse control peptide (R) (heatmap columns), for these genes (Supplementary file 1) (heatmap rows). TPM values are normalized per row. Red color represents higher expression. (D) Upper panel: Experimental design for the amyloid toxicity assay and a control experiment testing recovery 24 hr after application of the stress stimulus to confirm return of gene expression levels to pre-stress levels. Lower panel: Confirmation through RT-qPCR of the expression differences of selected B2-SRGs identified in RNA-seq data to differ between HT22 cells transfected with 1–42 (Espinoza et al., 2007) and cells transfected with the 42–1 reverse control peptide (R) for 6 hr (Supplementary file 1). Statistical significance (p value threshold 0.05) is depicted as asterisk for expression in treated cells (Espinoza et al., 2007) greater than the expression in controls (R) (unpaired directional t-test, n = 4, error bars represent standard deviation from the mean). In the same graph, we include also expression levels of cells from the same experiment that were left to recover in standard medium without any peptides for 24 hr after the initial 6 hr application of the peptides (42-Rec and R-Rec). Expression levels in all genes tested returned to pre-stress levels. Asterisk depicts statistical significance as described above also for the comparison between 42-ctrl and 42-Rec-24h.
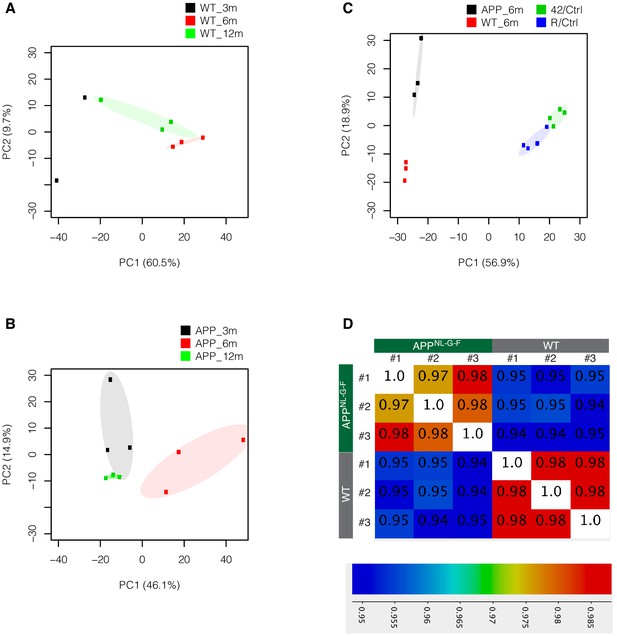
PCA plots and correlation matrix for sequenced samples in amyloid pathology and amyloid toxicity models.
Plots are based on amyloid pathology genes (Supplementary file 1). Correlation matrix was constructed using Semonk for the reads per million per gene length counts.
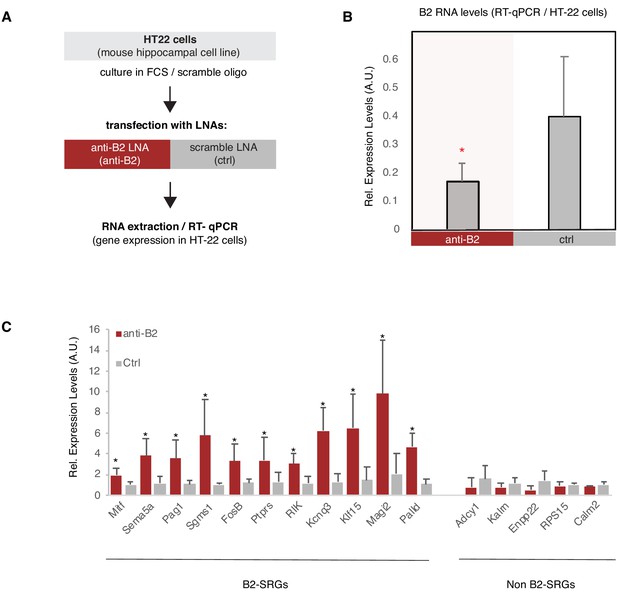
B2 NA destabilization leads to increase in expression of B2-SRGs.
(A) Experimental design for the B2 RNA knock-down cell culture assay employing HT-22 cells. (B) Expression levels of full-length B2 RNA (RT-qPCR) in the B2 RNA KD experiment. Statistical significance (p value threshold 0.05) for anti-B2 less than control (depicted as asterisk, n = 3, unpaired directional t-test, error bars represent standard deviation from the mean). (C) Expression levels (RT-qPCR)_of selected B2-SRGs (see Figure 2—figure supplement 2 and Figure 6D) and the respective negative controls during B2 RNA KD. Statistical significance (p value threshold 0.05) for anti-B2 greater than control (depicted as asterisk, n = 3/group, unpaired directional t-test, error bars represent standard deviation from the mean). Negative controls include five non-B2-SRGs that show no statistically significant difference between the two conditions.
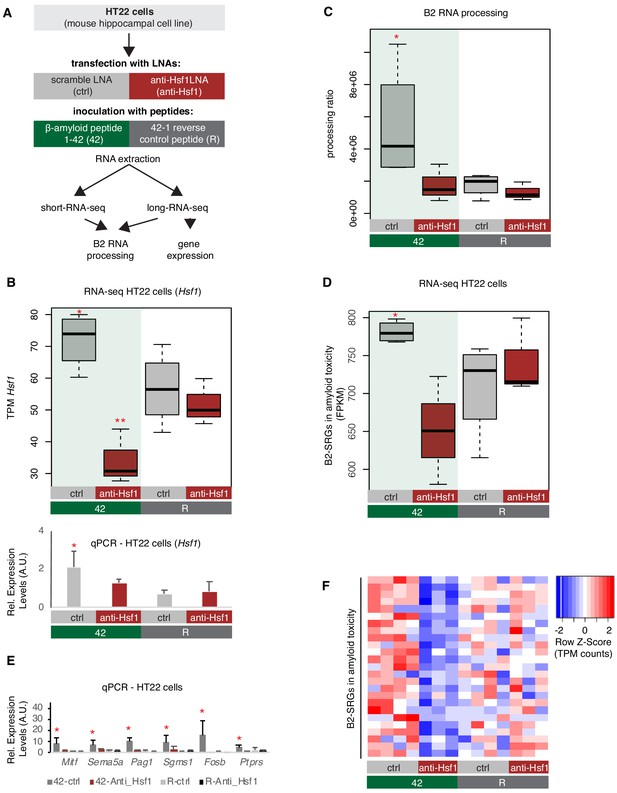
Hsf1 mediates B2 RNA processing in amyloid toxicity.
(A) Experimental design of the combined Hsf1 Knock Down – amyloid toxicity assay in HT22 cells followed by short and long RNA-seq. (B) Expression levels of Hsf1 as defined by long-RNA-seq (upper panel) and RT-qPCR (lower panel). Boxplots compare Hsf1 expression (TPM values) during incubation with the scramble LNA, and anti-Hsf1-specific LNA, incubated with either the 42 or R amyloid peptides. Statistical significance (p value threshold 0.05) for the comparison between 42/ctrl (n = 4) and 42/anti-Hsf1 (n = 3), unpaired non-directional t-test and for 42/ctrl (n = 4) greater than R/ctrl (n = 4) (both depicted as one asterisk), while two asterisks represent p<0.05 between 42/anti-Hsf1 and the other three groups. (C) B2 RNA processing ratio based on a combination of short and long-RNA-seq. Boxplot depicts distribution of total SINE B2 RNA processing ratio among different groups of HT22 cells between 42 and R. Statistical significance (p value threshold 0.05) for 42/ctrl greater than R/ctrl (p=0.04, n = 4/group, unpaired directional t-test, depicted as asterisk). No significant difference was observed between 42/anti-Hsf1 and R/anti-Hsf1 (n = 3/group) and between R/anti-Hsf1 and R/ctrl. (D) B2-SRGs expression levels in amyloid beta toxicity based on long-RNA-seq data (FPKM values) for genes of Supplementary file 1 (25 genes). Boxplot depicts distribution of expression levels in HT22 cells between 42 and R. p=0.05 for 42/ctrl greater than R/ctrl (n = 4/group, unpaired directional t-test, depicted as asterisk). No significant difference was observed between 42/anti-Hsf1 and R/anti-Hsf1 (NS, n = 3/group) and between R/anti-Hsf1 and R/ctrl. (E) Expression levels of selected B2-SRGs in the four conditions tested in our amyloid toxicity model though RT-qPCR. Statistical significance (p value threshold 0.05) for 42/ctrl greater than R/ctrl, unpaired directional t-test, n numbers as in subfigure B, with p<0.05 depicted as asterisk (error bars represent standard deviation from the mean). Samples and values depicted for non-Hsf1 LNA treated samples are the same as in Figure 6D and are used as controls as these samples were treated with a scramble LNA to allow comparison with the Hsf1 LNA treated samples. (F) Gene expression levels (long-RNA-seq) of B2-SRGs that are upregulated in amyloid toxicity (Supplementary file 1, 25 genes) show strong association with Hsf1 treatment during response to amyloid toxicity in HT22 cells. Heatmap depicts gene expression with rows as B2-SRGs in amyloid toxicity (Supplementary file 1) and columns as different HT22 cell treatments. TPM values are normalized per row. Red color represents higher expression.
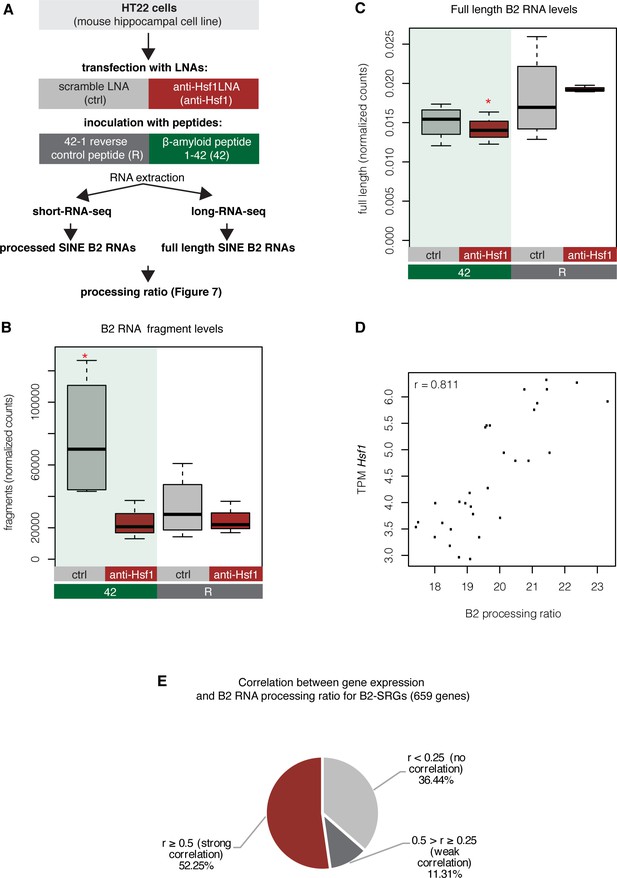
B2 RNA levels in HT22 cells and relationship with Hsf1 levels.
(A) Experimental design for estimation of B2 RNA processing ratio based on short and long-RNA seq data in our HT22 amyloid toxicity model. (B) Boxplot depicts distribution of levels of processed SINE B2 RNA fragments among different conditions. Statistical significance (p value threshold 0.05) for 42/ctrl greater than R/ctrl (p=0.03)(n = 4/group, unpaired directional t-test, depicted as asterisk). No significant difference was observed between 42/anti-Hsf1 and R/anti-Hsf1 (NS, n = 3/group) and between R/anti-Hsf1 and R/ctrl. (C) Boxplot depicts distribution of levels of full-length SINE B2 RNAs among different conditions. Statistical significance (p value threshold 0.05) for the comparison between 42-anti-Hsf1 and R-anti-Hsf1 (p=0.04)(depicted as asterisk, n = 3/group, unpaired non-directional t-test). No significant difference was observed for the other comparisons. (D) Correlation plot for the relationship between Hsf1 levels and processing ratio in all samples sequenced in the current study (Pearson correlation, r: correlation co-efficient, p<0.001). (E) Correlation between B2-SRGs expression and B2 RNA processing ratio calculated based on RNA-seq data and short-RNA-seq data from hippocampal samples in the current study. Only B2-SRGs, independently from amyloid beta status, that had read coverage across all samples were considered. For every gene a correlation coefficient was calculated (Pearson correlation) together with the respective p-value. Genes with a correlation p value less than 0.05 were considered in the subsequent analysis (659 genes) that classified them into three categories based on r: (i) no correlation (r < 0.25), (ii) weak correlation (r = 0.25 or r < 0.05), and (iii) strong correlation (r = 0.5 or r > 0.05). The pie diagram represents the percentage of each category. The exact r and p values of these genes are listed in Supplementary file 1.
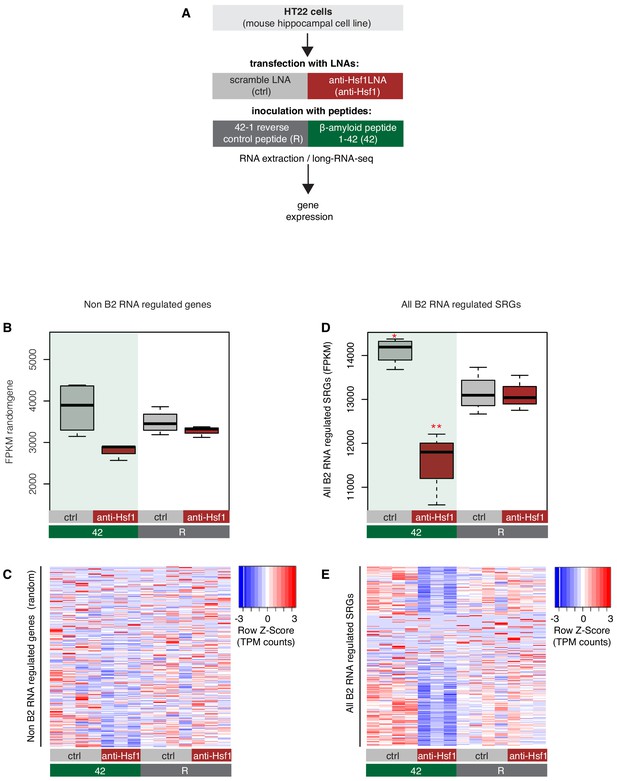
Expression levels of all B2-SRGs in amyloid beta toxicity.
(A) Experimental design of the combined Hsf1 Knock Down – amyloid toxicity assay in HT22 cells followed by short- and long-RNA-seq. (B) Expression levels of non-B2 RNA regulated genes as defined by long-RNA-seq. Boxplot depicts distribution of expression levels (in FPKM[Fragments per Kilobase per Million]) among different groups of HT22 cells. (C) Gene expression levels (defined by long-RNA-seq) of a random set of genes not associated with B2 RNAs (Supplementary file 1) show weak or no association with Hsf1 treatment during response to amyloid toxicity in HT22 cells. Heatmap depicts gene expression with rows as random genes (non-B2 RNA regulated) and columns corresponding to the different HT22 cell treatments. Expression values are normalized per row and correspond to TPM values, with red color represents higher expression. (D) B2 RNA regulated SRG expression levels based on long-RNA-seq data (FPKM values). Boxplot depicts distribution of expression levels of B2 RNA regulated genes among different groups of HT22 cells between 42 and R (1684 genes, Supplementary file 1). Statistical significance (p value threshold 0.05) for the comparison between 42/ctrl and R/ctrl (p=0.01)(n = 4/group, unpaired directional t-test, depicted as one asterisk) and for 42/anti-Hsf1 less than R/anti-Hsf1 (p=0.03)(n = 3/group, depicted as two asterisks, unpaired directional t-test). (E) Gene expression levels (long-RNA-seq) of B2-SRGs (Supplementary file 1) show strong association with Hsf1 treatment during response to amyloid toxicity in HT22 cells. Heatmap depicts gene expression with rows as B2 RNA regulated genes (1684 genes, Supplementary file 1) and columns as different HT22 cell treatments. TPM values are normalized per row. Red color represents higher expression.
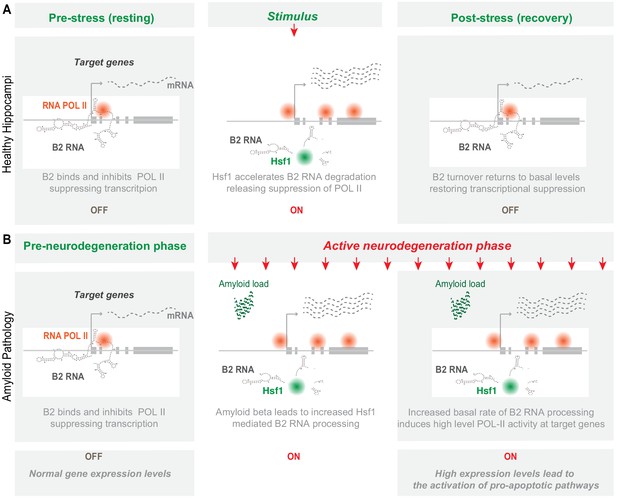
Representation of the role of B2 RNA processing in amyloid pathology.
Upon removal of the stress-generating stimulus, healthy cells restore the expression levels of Hsf1, specific B2 RNA regulated target genes and processing ratio of B2 RNAs returns to base levels. In contrast, in amyloid pathology, increased amyloid beta load acts as a continuous stimulus that causes the Hsf1 - B2 RNA – B2-SRG axis to ‘lock’ into an activated mode. ON/OFF represent active and suppressed SRG transcription, respectively.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (Mus musculus) | HT-22 | Millipore Sigma | Cat#SCC129, RRID:CVCL_0321 | |
| Sequence-based reagent | Amyloid Beta peptides ( 1-42) | Sigma-Aldrich | Custom synthesis | DAEFRHDSGYEVHHQKLVFFAEDVGSNKGAIIGLMVGGVVIA |
| Sequence-based reagent | Amyloid Beta peptides (Reverse 42–1) | Sigma-Aldrich | Custom synthesis | AIVVGGVMLGIIAGKNSGVDEAFFVLKQHHVEYGSDHRFEAD |
| Peptide, recombinant protein | Hsf1 protein | Enzo life sciences | ADI-SPP-902-F | Synthesized in insect, human sequence |
| Commercial assay, kit | NEBNext Small RNA Library Prep set | NEB | Cat# E7330 | |
| Commercial assay, kit | NEBNext Ultra II directional RNA library prep kit | NEB | Cat# E7760 | |
| Commercial assay, kit | Superscript III RT | Invitrogen | Cat# 18080093 | |
| Commercial assay, kit | Luna universal master mix | NEB | Cat# M3003 | |
| Antibody | Anti-Hsf1 (Rabbit, polyclonal) | Enzo | Cat# ADI-SPA-901 RRID:AB_10616511 | WB: 1:1000 |
DNA/RNA sequences.
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Sequence-based reagent | FosB Forward | IDT | Custom | 5-CGAGCTGCAAAAAGAGAAGG −3 |
| Sequence-based reagent | FosB Reverse | IDT | Custom | 5- TTACAGAGCAAGAAGGGAGG −3 |
| Sequence-based reagent | Pag1 Forward | IDT | Custom | 5-GAGCACAACTTCAAAGCTGG-3 |
| Sequence-based reagent | Pag1 Reverse | IDT | Custom | 5- TCATCAGGTTCTCATGGTCC −3 |
| Sequence-based reagent | Sema5a Forward | IDT | Custom | 5- ATGAGGCTGTGCAGTTCAGT-3 |
| Sequence-based reagent | Sema5a Reverse | IDT | Custom | 5-GTAACCAGGGGCCAATTTCT-3 |
| Sequence-based reagent | Sgms1 Forward | IDT | Custom | 5- ACCATAGACCACACAGGCTA-3 |
| Sequence-based reagent | Sgms1 Reverse | IDT | Custom | 5- TTTCTTCCGGTCTGAGCACT-3 |
| Sequence-based reagent | Hsf1 Forward | IDT | Custom | 5- TGACACCGAGTTCCAGCATC-3 |
| Sequence-based reagent | Hsf1 Reverse | IDT | Custom | 5- TGACACTGTCCTGGCGTATT-3 |
| Sequence-based reagent | Mitf Forward | IDT | Custom | 5- AAGCTCAGAGGCACCAGGTA-3 |
| Sequence-based reagent | Mitf Reverse | IDT | Custom | 5- CCTGCTCTGCTCCTCAAACT-3 |
| Sequence-based reagent | 7SK Forward | IDT | Custom | 5-GACATCTGTCACCCCATTGA-3 |
| Sequence-based reagent | 7SK Reverse | IDT | Custom | 5- GCCTCATTTGGATGTGTCTG-3 |
| Sequence-based reagent | Hprt Forward | IDT | Custom | 5- TCCTCCTCAGACCGCTTTT-3 |
| Sequence-based reagent | Hprt Reverse | IDT | Custom | 5- CCTGGTTCATCATCGCTAATC-3 |
| Sequence-based reagent | B2 Forward | IDT | Custom | 5- GGGGCTGGTGAGATG-3 |
| Sequence-based reagent | B2 Reverse | IDT | Custom | 5-AGCTGTCTTCAGACACTCC −3 |
| Sequence-based reagent | Adcy1 Forward | IDT | Custom | 5- GCATGACAATGTGAGCATCC −3 |
| Sequence-based reagent | Adcy1 Reverse | IDT | Custom | 5-TCAAGTCCCATCTCCACACA −3 |
| Sequence-based reagent | Kcnq3 Forward | IDT | Custom | 5- AGCACCGTCAGAAGCACTTT −3 |
| Sequence-based reagent | Kcnq3 Reverse | IDT | Custom | 5-TCCAAGAGACCCAGCTTTTG-3 |
| Sequence-based reagent | Klf15 Forward | IDT | Custom | 5-TCATGGAGGAGAGCCTCTGT-3 |
| Sequence-based reagent | Klf15 Reverse | IDT | Custom | 5-TCCAAGAGACCCAGCTTTTG-3 |
| Sequence-based reagent | Magi2 Forward | IDT | Custom | 5-CGGGATCACACTTTTCACCT-3 |
| Sequence-based reagent | Magi2 Reverse | IDT | Custom | 5-CGGGATCACACTTTTCACCT-3 |
| Sequence-based reagent | Palld Forward | IDT | Custom | 5-CAGTGGCTCAGACAGCACAT-3 |
| Sequence-based reagent | Palld Reverse | IDT | Custom | 5-CTCCTGTTTTCGGAGCTGAG-3 |
| Sequence-based reagent | Enpp2 Forward | IDT | Custom | 5-GACTGTCGGTGTGACAACCT-3 |
| Sequence-based reagent | Enpp2 Reverse | IDT | Custom | 5-CTTCTGAGCAGTGACAGGCA-3 |
| Sequence-based reagent | RPS15 Forward | IDT | Custom | 5-AACCAGAGATGATCGGCCAC-3 |
| Sequence-based reagent | RPS15 Reverse | IDT | Custom | 5-ATGAATCGGGAGGAGTGGGT-3 |
| Sequence-based reagent | Calm2 Forward | IDT | Custom | 5-GACTGAAGAGCAGATTGCAG-3 |
| Sequence-based reagent | Calm2 Reverse | IDT | Custom | 5-CAGTTCTGCTTCTGTGGGGT-3 |
| Sequence-based reagent | Kalrn Forward | IDT | Custom | 5-CCCTGAACTCCATCCACAGT-3 |
| Sequence-based reagent | Kalrn Reverse | IDT | Custom | 5-GAGGGGTGTGTGTGACTCTT-3 |
| Sequence-based reagent | B2 RNA | IDT G-block | Custom synthesis. Zovoilis et al., 2016 | 5′- taatacgactcactata GGGGCTGGTGAGATGGCTCAGTGGGTAAGAGCACCCGACTGCTCTTCCGAAGGTCCGGAGTTCAAATCCCAGCAACCACATGGTGGCTCACAACCATCCGTAACGAGATCTGACTCCCTCTTCTGGAGTGTCTGAAGACAGCTACAGTGTACTTACATATAATAAATAAATAAATCTTTAAAAAAAAA - 3 |
| Sequence-based reagent | B2mut4b | IDT G-block | Custom synthesis. | taatacgactcactataGGGCTGGTGAGATGGCTCAGTGGGTAAGAGCACCCGACTGCTCTTCCGAAGGTCCGGAGTTCAAATCCCAGCAACCACATGGTGGCTCACAACCATCCGTAACGAGATCTGACTCCCTCTTCTTCTGAAGACAGCTACAGTGTACTTACATATAATAAATAAATAAATCTTTAAAAAAAAA |
Additional files
-
Supplementary file 1
Supplementary tables.
Supplementary Table 1. List of B2 RNA regulated SRGs(B2-SRGs). Data are compiled from Zovoilis et al., 2016 and include those genes that are close to B2 CHART peaks (genome-binding sites) before but not after the application of stress stimulus. Supplementary Table 2. Complete lists of enriched terms in B2 RNA regulated SRGs(B2-SRGs)(see Suppl.Table 1) for Tissue Enrichemnt (left), Biological Process (middle) and Cellular Compartment (right). Supplementary Table 3. List of B2 RNA regulated SRGs (B2-SRGs) (see Suppl.Table 1) that are associated with learning based on Peleg et al., 2010. Supplementary Table 4. Upregulated genes in hippocampi of APP 6-month-old mice compared to 6-month WT mice. Values were calculated using DESeq (see Materials and methods) on long-RNA-seq data. Only genes with an FDR < 0.2 are depicted. Supplementary Table 5. List of B2 RNA regulated SRGs (B2-SRGs) (see Suppl.Table 1) that are upregulated in 6-month-old APP mice compared to WT (see Suppl.Table 4) Supplementary Table 6. List of B2 RNA regulated SRGs (B2-SRGs) (see Suppl.Table 1) that are upregulated in 6-month-old APP mice (see Suppl.Table 4) and are associated with learning based on Peleg et al., 2010. Supplementary Table 7. Complete lists of enriched terms in B2 RNA regulated SRGs (B2-SRGs) that are upregulated in 6-month-old APP mice compared to WT (see Suppl.Table 5) for Biological Process (left) and Cellular Compartment (right). Supplementary Table 8. Upregulated genes in HT22 cells treated with amyloid beta and Scr LNA compared to cells treated with the control peptide and scr LNA. Values were calculated using DESeq (see Materials and methods) on long-RNA-seq data. Only genes with an FDR < 0.2 are depicted. Supplementary Table 9. List of genes that are upregulated in HT22 cells treated with amyloid beta (see Suppl.Table 8) and in 6-month-old APP mice (see Suppl.Table 4) Supplementary Table 10. List of B2 RNA regulated SRGs (B2-SRGs) (see Suppl.Table 1) that are upregulated in HT22 cells treated with amyloid beta and Scr LNA compared with cells treated with the control peptide and scr LNA (see Suppl.Table 8 ) Supplementary Table 11. Complete lists of enriched terms in B2 RNA regulated SRGs (B2-SRGs) that are upregulated in HT22 cells treated with amyloid beta (see Suppl.Table 10) for Biological Process (left) and Cellular Compartment (right). Supplementary Table 12. Correlation co-efficients and p-values for genes of Figure 8—figure supplement 2. Includes genes for which there was readcoverage across all sample and the correlation p value was less than 0.05. Supplementary Table 13. List of non-B2 RNA regulated genes (random set) used throughout the study.
- https://cdn.elifesciences.org/articles/61265/elife-61265-supp1-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/61265/elife-61265-transrepform-v2.docx