Identification of abscission checkpoint bodies as structures that regulate ESCRT factors to control abscission timing
Figures
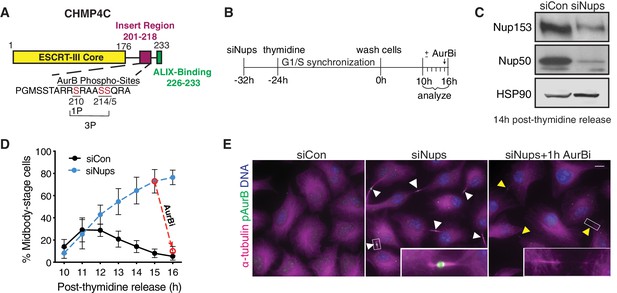
System for abscission checkpoint enrichment.
(A) Schematic of the CHMP4C protein. (B) Timeline for synchronizing cells with an active abscission checkpoint using siNup153/siNup50 (siNups), followed by thymidine treatment. (C) Western blot of lysates prepared from control (siCon) and checkpoint-active (siNups) HeLa cells harvested 14 hr post-thymidine release. (D) Quantification of % midbody-stage cells after treatment as in (B). Dashed line shows the loss of midbody-stage cells from the siNups condition at t = 16 hr following addition of AurBi at t = 15 hr. N = 1200–3300 (total) cells/timepoint from n = 11 independent biological replicates. n = 5 biological replicates for siNups + AurBi. (E) Images acquired 16 hr post-thymidine release after treatment as in (B). White arrowheads: midbodies. Yellow arrowheads: recently abscised midbodies. Scale bar, 10 µm. Insets have enhanced brightness. Throughout manuscript: DNA is detected with DAPI unless noted. Bar and line graphs represent mean ± standard deviation. Times refer to timeline in (B) unless noted. N = total measurements from all biological replicates (n).
-
Figure 1—source data 1
Source Data for Figure 1.
- https://cdn.elifesciences.org/articles/63743/elife-63743-fig1-data1-v2.xlsx
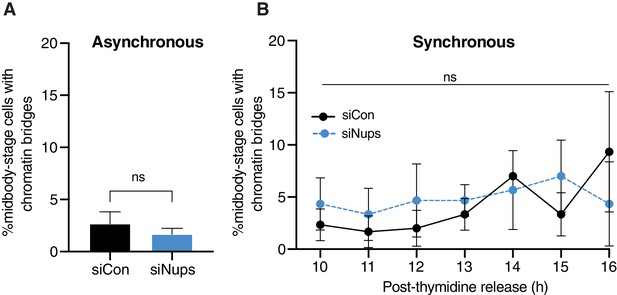
Nup depletion and cell synchronization delay abscission without increasing chromatin bridges.
(A) Quantification of % midbody-stage cells with chromatin bridges marked by Lap2ß under asynchronous conditions (48 hr after transfection with siNups or siControl). N = 300 midbodies, n = 3 biological replicates. (B) Timecourse quantification of % midbody-stage cells with chromatin bridges in control (siControl) and checkpoint-active (siNup153/50) cells. N = 300 midbodies per condition, n = 3 biological replicates. Throughout figure supplements: experimental timeline refers to Figure 1B unless otherwise noted. Line and bar graphs represent mean ± standard deviation unless noted. p<0.05:*, p≤0.01:**, p≤0.001***, p≤0.0001****. Exact p-values can be found in supplementary file 3. See Materials and methods for statistical tests used. N = total from all replicates.
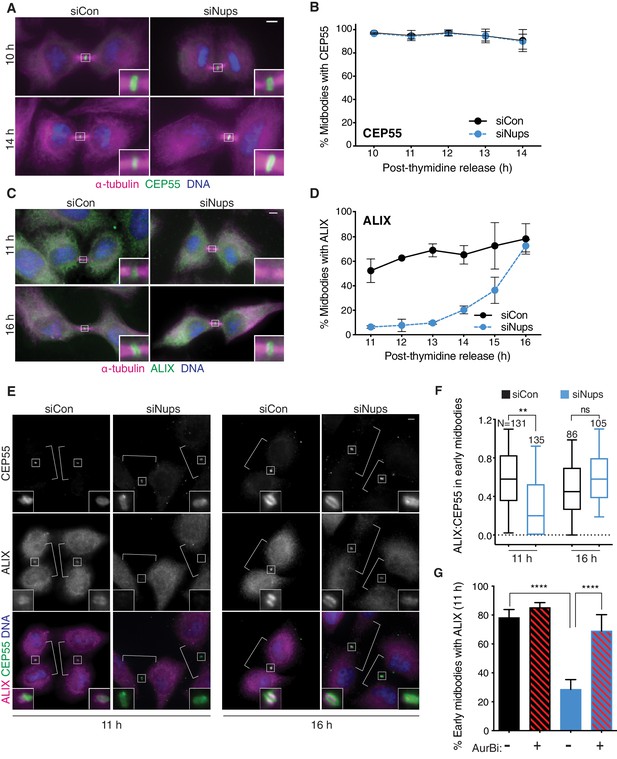
Abscission checkpoint activity delays ALIX recruitment to the midbody.
Immunofluorescence and time course quantifications of CEP55 (A, B) and ALIX (C, D) recruitment to early-stage midbodies in control and checkpoint-active cells. N = 300 midbodies/timepoint from n = 3 biological replicates. (E, F) Immunofluorescence and quantification of ALIX:CEP55 relative intensities from n = 2 biological replicates in individual control and checkpoint-active early-stage midbodies (see Figure 4B for example of early midbody stage). Only one midbody is shown in the siCon/16 hr condition because this sample had fewer midbody-stage cells than other conditions. DNA detected with Hoechst. (G) Quantification of ALIX recruitment to early-stage midbodies at 11 hr with/without checkpoint enrichment (blue/black bars) and with/without AurBi added at 10.5 hr (striped/solid bars). N = 500 midbodies/treatment from n = 5 biological replicates. Throughout manuscript: scale bars are 5 µm unless noted. White brackets mark midbody-stage cells, as detected by α-tubulin, not shown. Data points without visible error bars (as in 2D siCon 12 hr) have a SD too small to display outside the data point or box. Boxplots represent the 25th, median, and 75th percentile values. Whiskers represent the 10th and 90th percentiles. p<0.05: *, p≤0.01: **, p≤0.001: ***, p≤0.0001: ****. Exact p-values can be found in supplementary file 3. See Materials and methods for statistical tests used.
-
Figure 2—source data 1
Source Data for Figure 2.
- https://cdn.elifesciences.org/articles/63743/elife-63743-fig2-data1-v2.xlsx
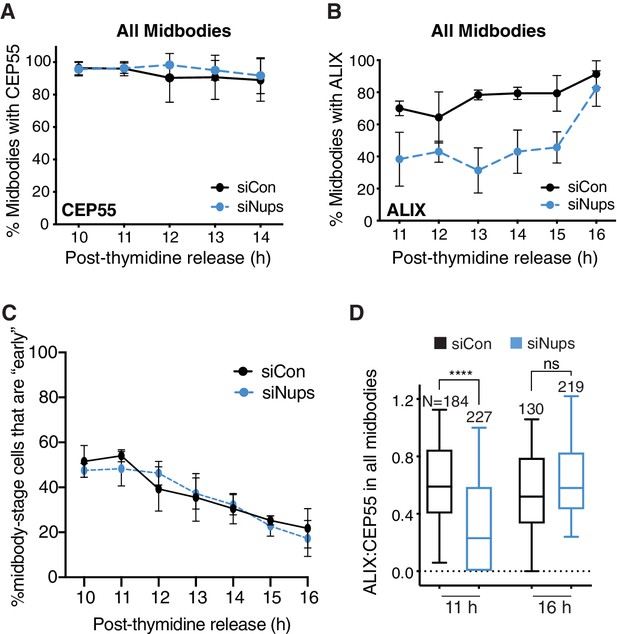
The abscission checkpoint delays ALIX recruitment in the total midbody population.
(A, B) Timecourse quantification of CEP55 (A) or ALIX (B) recruitment to midbodies in control (siControl) and checkpoint-active (siNup153/50) cells. N = 300 midbodies scored for each timepoint, n = 3 biological replicates. (C) Percent midbody-stage cells that are ‘early-stage’ throughout the abscission checkpoint timecourse. N = 400 midbody-stage cells per timepoint, n = 4 biological replicates. (D) Quantification of ALIX:CEP55 relative intensity at individual midbodies in control and siNups-treated cells from n = 2 biological replicates. Throughout figure supplements: Boxplots represent the 25th, median, and 75th percentile of values. Whiskers represent the 10th and 90th percentiles.
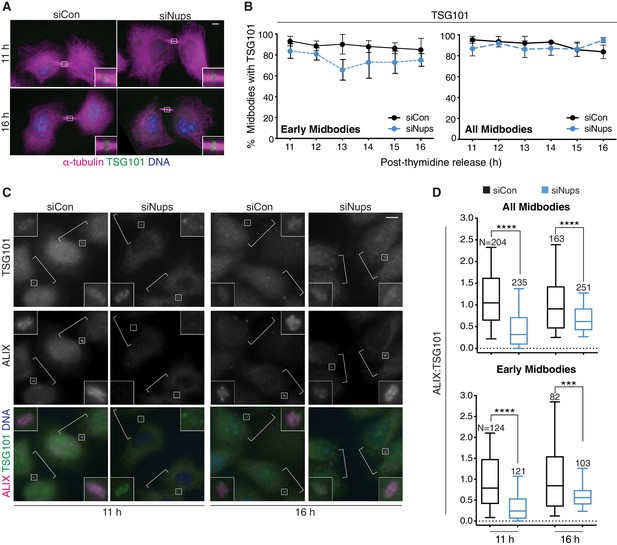
The abscission checkpoint does not delay TSG101/ESCRT-I recruitment to the midbody.
(A, B) Immunofluorescence and timecourse quantification of TSG101 recruitment to midbodies in control and checkpoint-active cells. N = 400 midbodies scored/timepoint from n = 4 biological replicates. (C, D) Immunofluorescence and quantification of ALIX:TSG101 relative intensity at individual midbodies in control and checkpoint-active cells from n = 2 biological replicates. DNA detected with Hoechst. Throughout figure supplements: White brackets mark cells confirmed to be midbody-stage by α-tubulin, staining not shown. Scale bars are 5 µm unless noted. Data points without visible error bars (as in B, “All Midbodies” siCon 14 hr) have SDs that do not extend outside the data point. DNA is detected with DAPI unless noted.
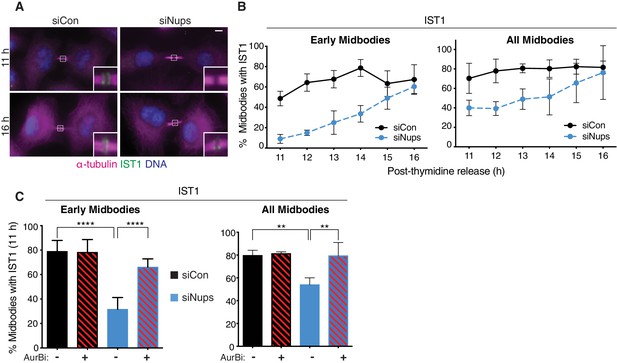
The abscission checkpoint delays IST1 recruitment to the midbody.
(A, B) Immunofluorescence and timecourse quantification of IST1 recruitment to midbodies in control and checkpoint-active midbodies. N = 300 midbodies scored/timepoint from n = 3 biological replicates. (C) Quantification of IST1 recruitment to early and total midbodies, 11 hr post-thymidine release, with/without siNups treatment (blue/black bars), and with/without 30 min AurBi added at 10.5 hr (striped/solid bars). All: N = 300 midbodies scored/condition from n = 3 biological replicates. Early: N = 500 midbodies scored/condition from n = 5 biological replicates.
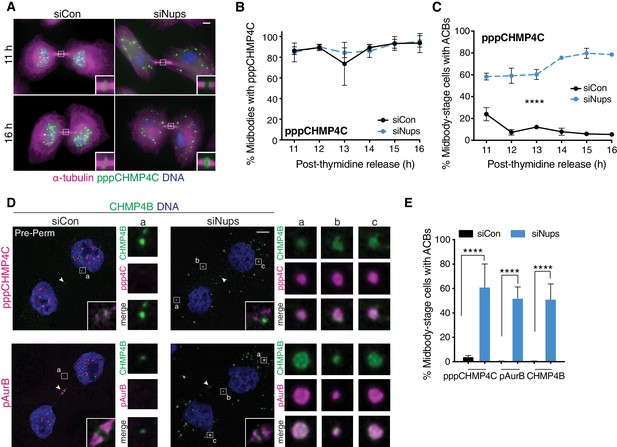
pppCHMP4C localizes to Abscission Checkpoint Bodies when the abscission checkpoint is active.
(A, B) Immunofluorescence and time course quantification of pppCHMP4C recruitment to midbodies in control and checkpoint-active cells, and (C) time course quantification of midbody-stage cells with ACBs present. N ≥ 300 midbodies/timepoint from n = 3 biological replicates. (D) Confocal z-projections of pre-permeabilized midbody-stage cells under asynchronous conditions (48 hr after transfection with siNups or siControl), stained as indicated. Note the appearance of ACB component substructure in some cases (siNups, CHMP4B). (E) Quantification of midbody-stage cells with ACBs present (treated as in D). N = 300 midbodies/condition from n = 3 biological replicates. Throughout manuscript: Components are brighter in ACBs than in midbodies and the images are optimized for ACBs, which reduces the appearance of midbody localization in the confocal images. Confocal midbody insets are 2.3 µm wide and have enhanced brightness. White arrowheads mark Flemming bodies (where detectable). ACB enlargements are 2 µm wide.
-
Figure 3—source data 1
Source Data for Figure 3.
- https://cdn.elifesciences.org/articles/63743/elife-63743-fig3-data1-v2.xlsx
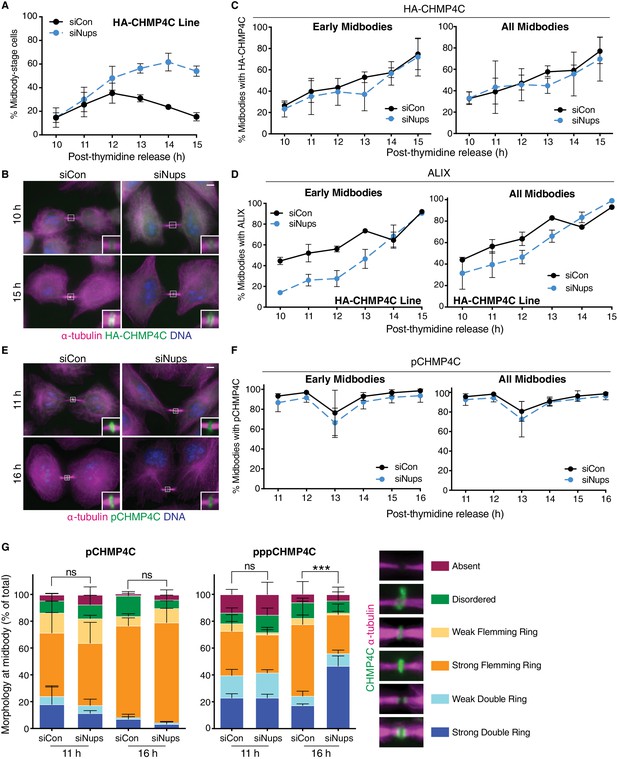
The abscission checkpoint does not delay CHMP4C recruitment to the midbody.
(A–C) Quantification of % midbody-stage cells (A), immunofluorescence (B), and timecourse quantification of HA-CHMP4C recruitment to midbodies (C) in control and checkpoint-active cells, using a HeLa cell line stably expressing HA-CHMP4C. (A) N = 950 cells scored/timepoint from n = 3 biological replicates. (C) N = 300 midbodies scored/timepoint from n = 3 biological replicates (n = 2 for 10 hr). (D) Timecourse quantification of ALIX recruitment to midbodies in control and checkpoint-active HA-CHMP4C-expressing cells. N = 200 midbodies scored/timepoint, n = 2 biological replicates. (E, F) Immunofluorescence and timecourse quantification of pCHMP4C recruitment to midbodies in control and checkpoint-active HeLa cells. N = 300 midbodies/timepoint, n = 3 biological replicates. (G) Quantification of p/pppCHMP4C morphology at the midbody in control and checkpoint-active cells 11 hr and 16 hr post-thymidine release. N = 300 midbodies/condition, n = 3 biological replicates. p-values compare combined weak and strong double rings.
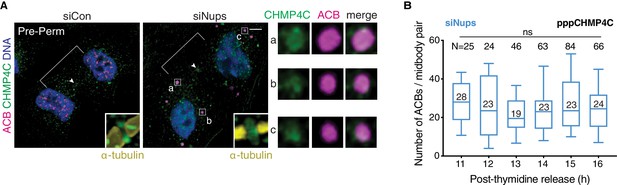
CHMP4C is detected in ACBs, which are maintained in stable numbers when the abscission checkpoint is active.
(A) Confocal z-projections of pre-permeabilized midbody-stage cells under asynchronous (48 hr) siControl and siNups conditions co-stained for CHMP4C and an ACB marker (detected by mAb SC35, see Figure 4). (B) Quantification of number of ACBs, tracked with α-pppCHMP4C, per midbody-stage cell after siNups treatment. n = 3 biological replicates; the median ACB number is indicated. (n = 2 biological replicates for 12 hr). Throughout figure supplements: White arrowhead marks Flemming body where detectable. Flemming body insets from confocal imaging are 2.3 µm wide and are enhanced for brightness. ACBs have a brighter signal than the Flemming body; thus, some Flemming bodies are seen only in enhanced inset. ACB enlargements are 2 µm wide.
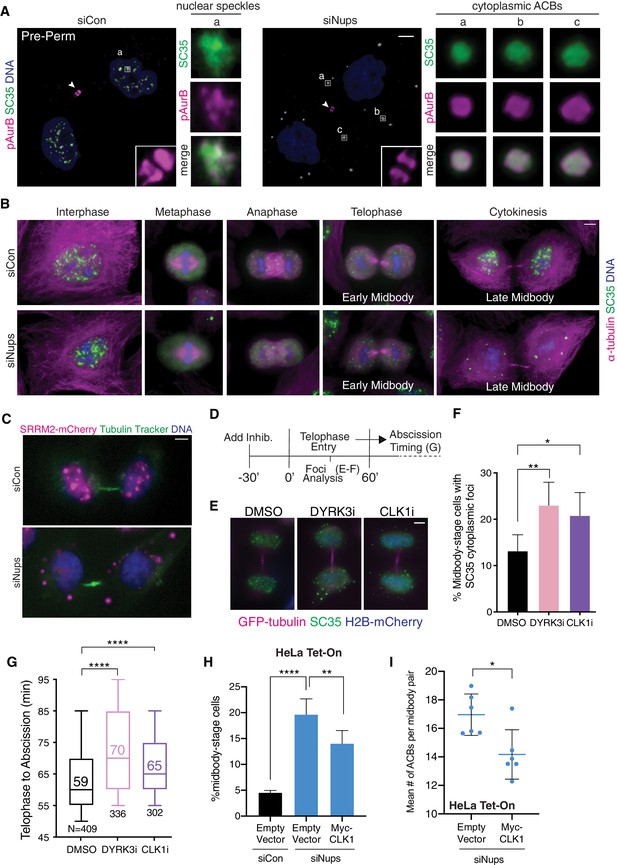
ACBs are related to MIGs and contribute to abscission delay.
(A) Confocal z-projections of pre-permeabilized (Pre-Perm) midbody-stage cells showing that SC35 antibody and antibody against pAurB co-stain ACBs (asynchronous cultures, 48 hr after transfection with siNups or siControl). (B) Immunofluorescence of nuclear speckles, MIGs, and ACBs in asynchronous conditions (as in A, but non-pre-permeabilized and at a variety of cell-cycle stages as designated). (C) Live-imaging of HeLa cells expressing SRRM2-mCherry after 48 hr treatment with siCon or siNups. Images were independently adjusted for brightness and contrast for optimal display. (D) Timeline of live-cell imaging of abscission timing following treatment with DMSO, 1 µM DYRK3i, or 1 µM CLK1i. (E, F) Immunofluorescence of midbody-stage cells fixed 60’ after addition of vehicle or inhibitors, with quantification of % midbody-stage cells with cytoplasmic foci marked by SC35 antibody. N = 300 midbodies/condition from n = 3 biological replicates. (n = 4 for DMSO) (G) Timing from telophase to abscission (cells treated as in D). n = 3 biological replicates. (H) Quantification of midbody-stage cells after 72 hr treatment with siCon or siNups and 48 hr expression of empty vector or CLK1 kinase as indicated (asynchronous). N = 4800 cells from n = 6 biological replicates. (I) Mean number of ACBs per midbody pair after treatment as in (H). N ≥ 663 midbody-stage cells per condition. n = 6 biological replicates.
-
Figure 4—source data 1
Source Data for Figure 4.
- https://cdn.elifesciences.org/articles/63743/elife-63743-fig4-data1-v2.xlsx
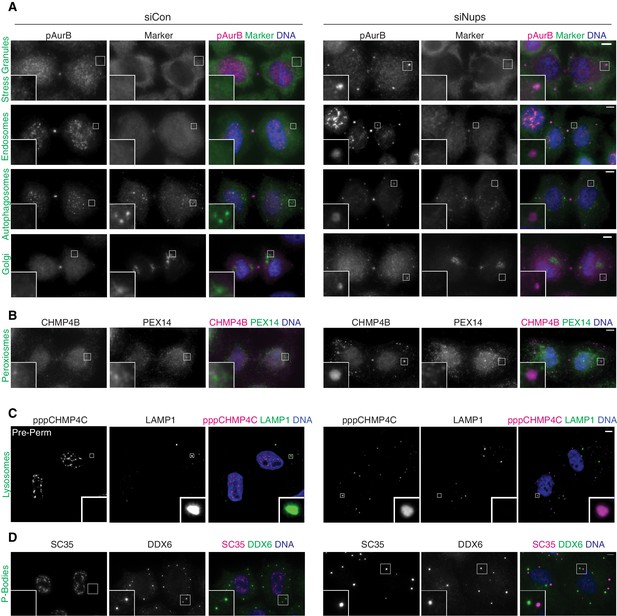
ACBs do not colocalize with a variety of subcellular organelles and structures.
(A, B) Immunofluorescence of asynchronous (48 hr) control and checkpoint-active (siNups) midbody-stage cells stained to detect ACBs (marked by (A) α-pAurB or (B) α-CHMP4B or (D) mAb-SC35), and specific organelle/substructure markers simultaneously. Antibodies used were as follows; stress granules:α-G3BP1, early endosomes:α-EEA1, autophagosomes:α-LC3ß, Golgi:α-giantin, peroxisomes:α-PEX14, P-bodies:α-DDX6. Typically ACBs and P-bodies are not present in the same cells, but to demonstrate their distinct identity a cell with both organelles present is shown. (C) Confocal z-projections of pre-permeabilized asynchronous (48 hr) control and checkpoint-activated midbody-stage cells co-stained for ACBs (marked by α-pppCHMP4C) and lysosomes (marked by α-LAMP1).
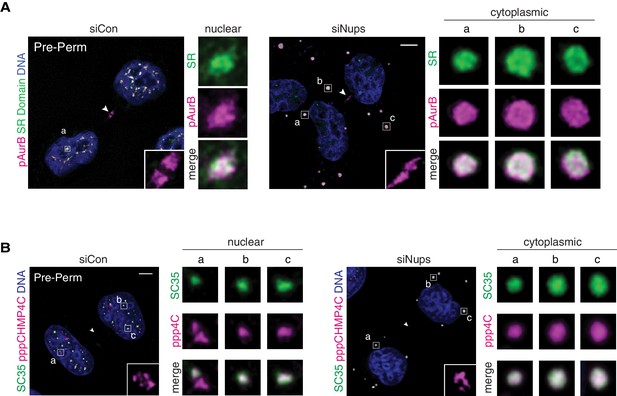
Colocalization of ACB components.
(A, B) Confocal z-projections of pre-permeabilized midbody-stage cells under asynchronous (48 hr) control and checkpoint-active conditions, co-stained for ACB components as indicated.
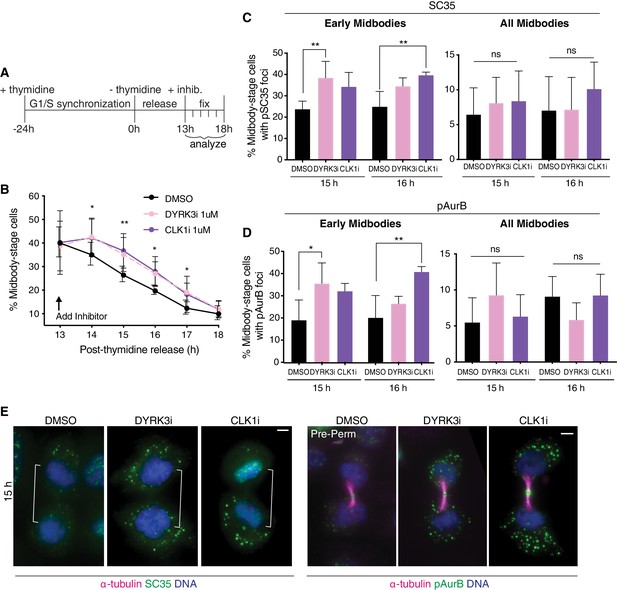
Interfering with timely resolution of MIGs delays cytokinetic abscission.
(A) Time schematic of fixed-imaging experiments after treatment with DMSO, 1 µM DYRK3i, or 1 µM CLK1i. Inhibitors were added 13 hr post-thymidine release, after most cells had completed metaphase, to avoid a confounding metaphase arrest. (B) Quantification of % midbody-stage cells after treatment as diagrammed in (A). N = 1200 cells scored/timepoint from n = 4 biological replicates. (C–E) Quantification and immunofluorescence of midbody-stage cells with cytoplasmic foci marked by mAb SC35 or α-pAurB present, treated as diagrammed in (A). In (E), cells were confirmed to be in equivalent midbody stages by tubulin staining (not shown for cells stained with mAb SC35). N = 400 midbody-stage cells scored/condition from n = 4 biological replicates.
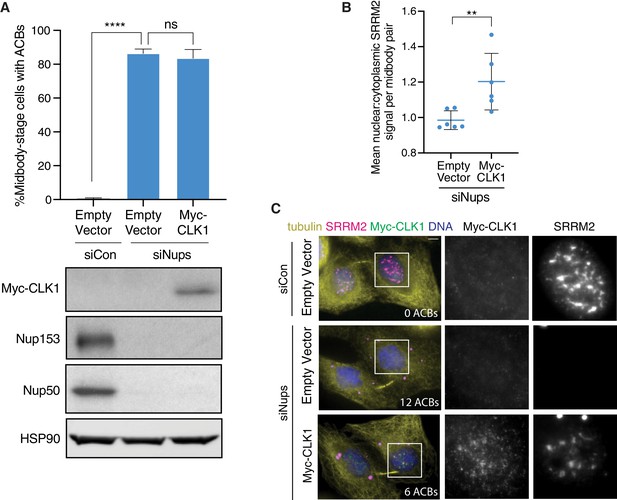
CLK1 expression partially dissolves ACBs to mitigate abscission arrest.
(A) Quantification of percent midbody-stage cells with ACBs present after treatment as in Figure 4H, with a representative western blot show below. N = 1200 midbodies from n = 6 biological replicates. (B) Relative intensity of SRRM2 nuclear signal versus its baseline cytoplasmic signal (a region without ACBs present) in midbody-stage cells with ACBs after treatment as in Figure 4H. N ≥ 663 midbody-stage cells from n = 6 biological replicates. (C) Immunofluorescence of cells treated with siCon or siNups and expressing empty vector or CLK1 as indicated. Cells were stained with α-Myc to mark exogenous CLK1 and α-SRRM2 to mark ACBs and NS.
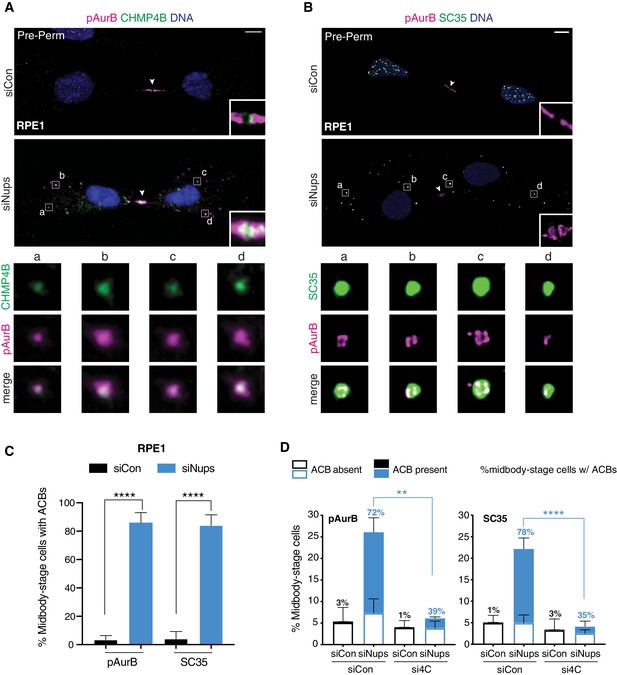
ACBs are conserved and dependent upon abscission checkpoint factor CHMP4C.
(A, B) Confocal z-projections of pre-permeabilized control and checkpoint-active RPE1 cells following 14 hr thymidine release, stained for ACB markers as indicated. (C) Quantification of control and checkpoint-active RPE1 midbody-stage cells with ACBs present 14 hr post-thymidine release, detected by α-pAurB or SC35 antibody. N = 300 midbody-stage cells scored/condition from n = 3 biological replicates. (D) Quantification of midbody-stage HeLa cells under asynchronous conditions (72 hr after transfection with indicated siRNAs) containing ACBs (marked by α-pAurB or SC35 antibody). The number above each bar represents the % midbody-stage cells with ACBs present. N = 300 midbody-stage cells/condition from n = 3 biological replicates. p-values compare % midbodies with ACBs present.
-
Figure 5—source data 1
Source Data for Figure 5.
- https://cdn.elifesciences.org/articles/63743/elife-63743-fig5-data1-v2.xlsx
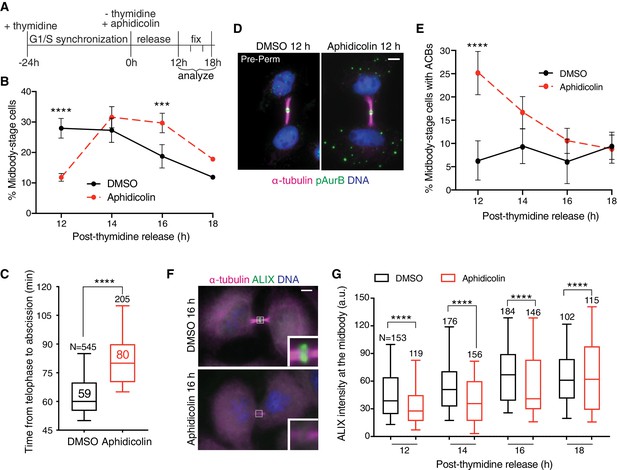
ACB formation and ALIX recruitment delay occur following replication stress.
(A, B) Timeline schematic (A) and quantification (B) of synchronized % midbody-stage cells after treatment with 0.4 µM aphidicolin or DMSO following thymidine release. N = 900 cells scored/timepoint from n = 3 biological replicates. (C) Timing from telophase to abscission after 48 hr treatment with DMSO or 0.4 µM aphidicolin. n = 3 biological replicates. (D–E) Immunofluorescence and quantification of % midbody-stage cells with ACBs present (treatment as in A). N = 300 midbody-stage cells scored/timepoint from n = 3 biological replicates. (F–G) Immunofluorescence and quantification of ALIX intensity at midbodies (treatment as in A). n = 3 biological replicates.
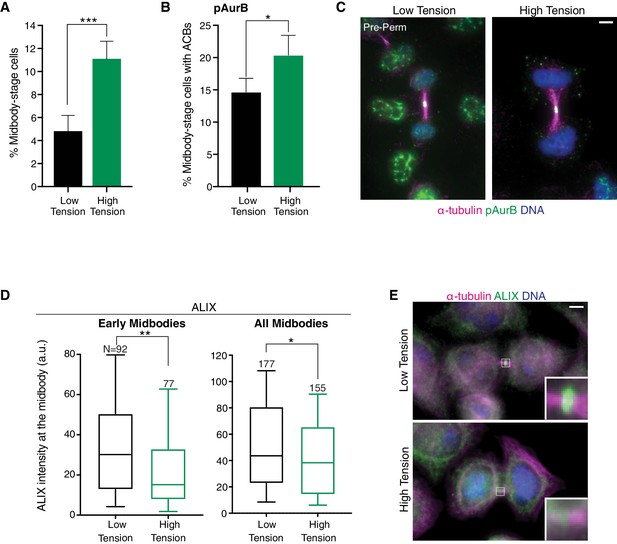
ACB formation and ALIX recruitment delay occur when intercellular tension is heightened.
(A) Quantification of % midbody-stage cells plated at high density (low tension) or low density (high tension). N = 1200 cells scored/condition from n = 4 biological replicates. (B, C) Quantification and immunofluorescence of midbody-stage cells with ACBs marked by α-pAurB present when plated at high or low density. N = 400 cells scored/condition from n = 4 biological replicates. (D, E) Quantification and immunofluorescence of ALIX intensity at midbodies in cells plated at high or low density. n = 4 biological replicates.
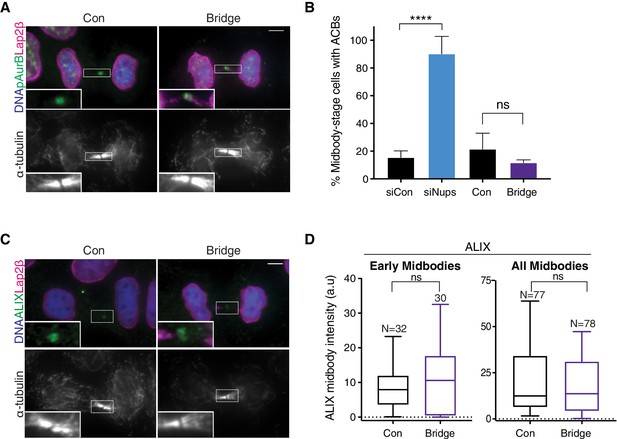
Chromatin bridges do not significantly promote ACB formation or ALIX recruitment delay.
(A) Immunofluorescence of representative cells with and without chromatin bridges. (B) Quantification of dividing cells with ACBs both with/without siNups treatment (for comparison) and with/without chromatin bridges. Rare, naturally occurring bridges were scored, accounting for the small sample size: N ≥ 142 midbodies scored/condition from n = 3 biological replicates. (C, D) Representative immunofluorescence and quantification of ALIX recruitment to midbodies in cells with and without chromatin bridges. n = 3 biological replicates. Insets in (A, C) were enhanced for brightness.
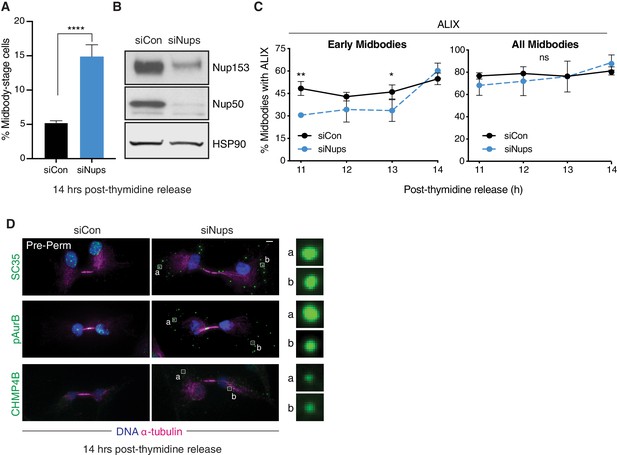
RPE1 cells display hallmarks of the abscission checkpoint, including ALIX recruitment delay and ACB formation.
(A, B) Quantification of % midbody-stage cells and western blot of lysates from RPE1 cells after siCon or siNups treatment 14 hr following thymidine release. N = 800 cells scored/condition from n = 3 biological replicates. (C) Timecourse quantification of ALIX recruitment to midbodies in control and checkpoint-active cells. N = 250 midbodies scored/timepoint from n = 3 biological replicates. (D) Immunofluorescence of pre-permeabilized control and checkpoint-active cells 14 hr following thymidine release, stained for ACB markers as indicated.
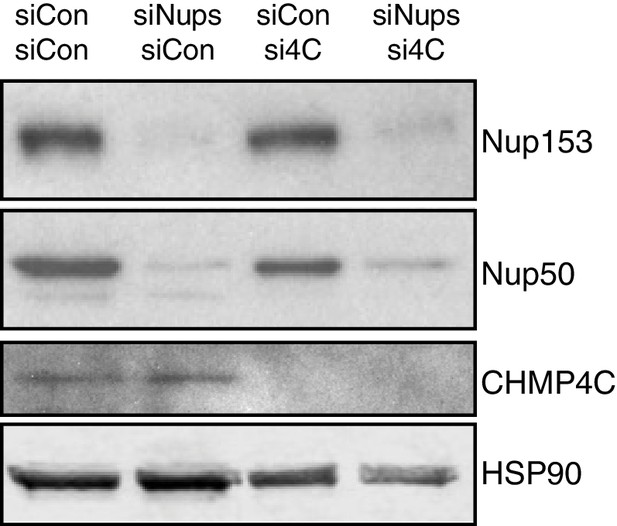
Confirmation of efficiency and specificity of protein depletion when CHMP4C and Nup153/50 are simultaneously targeted.
Western blots of the experiment shown in Figure 5D.
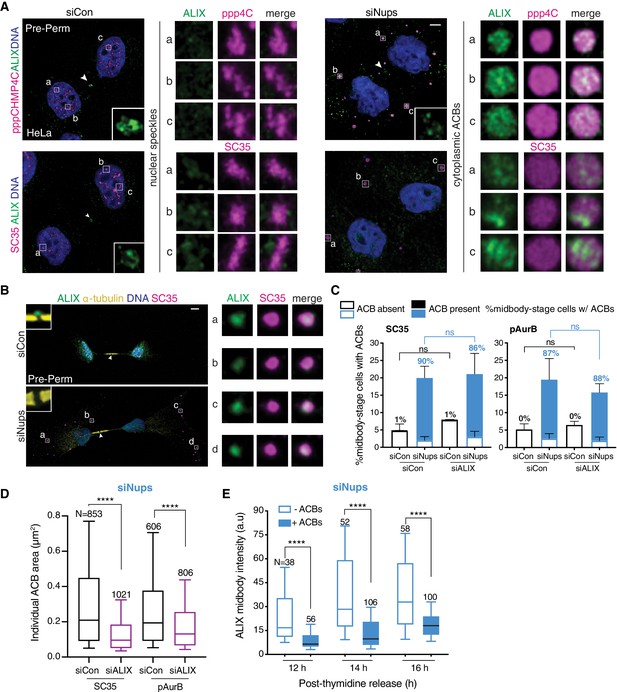
ACBs contain ALIX and cells with ACBs have delayed ALIX midbody recruitment.
(A) Confocal z-projections of pre-permeabilized midbody-stage HeLa cells under asynchronous conditions (48 hr after transfection with siNups or siControl), stained as indicated. (B) Confocal z-projections of pre-permeabilized control and checkpoint-active midbody-stage RPE1 cells (14 hr post-thymidine release) stained as indicated. (C, D) Quantification of (C) midbody-stage cells with/without ACBs detected with α-pAurB or SC35 antibody, (D) ACB size under asynchronous conditions (72 hr after transfection with indicated siRNAs). (C) N = 300 midbodies scored/condition from n = 3 biological replicates. The number above each bar represents the % midbody-stage cells with ACBs present. p-values compare % midbodies with ACBs. (D) n = 4 biological replicates. (E) Time course quantification of relative ALIX midbody intensity in checkpoint-active cells. Midbody-stage cells were binned into categories with or without ACBs (marked by α-pAurB) using images from n = 3 biological replicates.
-
Figure 6—source data 1
Source Data for Figure 6.
- https://cdn.elifesciences.org/articles/63743/elife-63743-fig6-data1-v2.xlsx
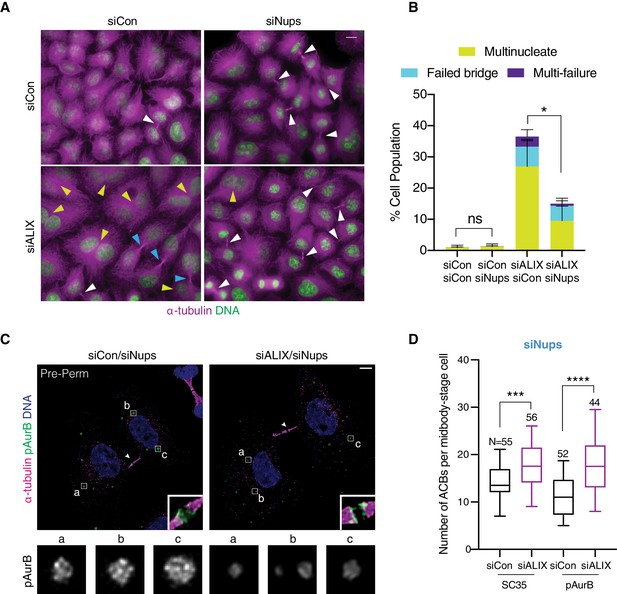
ACB size but not abscission checkpoint arrest is dependent on ALIX.
(A, B) Representative immunofluorescence and quantification of cells treated with indicated siRNAs for 72 hr. White arrowhead: midbody. Yellow arrowhead: multinucleate cell. Blue arrowhead: failed bridge. Scale bar, 10 µm. N = 900 cells from n = 3 biological replicates. p-values compare total failure events. (C) Confocal z-projections of pre-permeabilized cells with ACBs marked by pAurB. (D) Quantification of total number of ACBs per midbody-stage cell marked by α-pAurB or mAb SC35. n = 4 biological replicates.
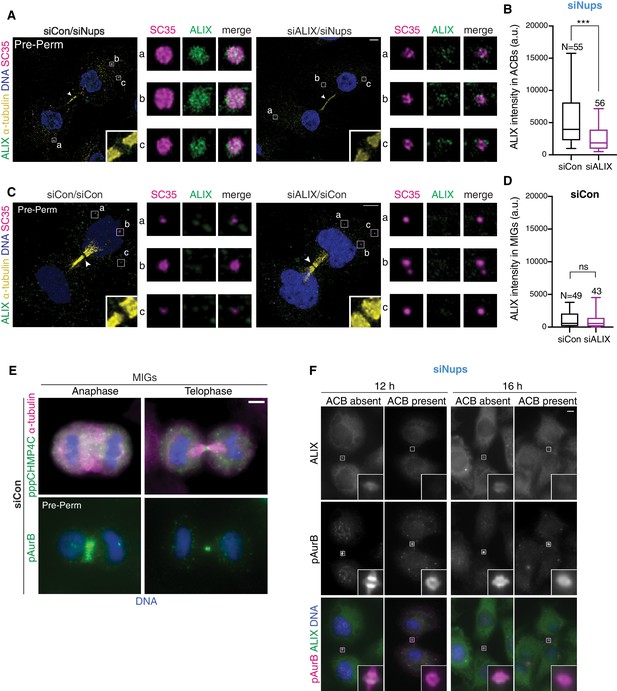
ALIX localizes specifically to ACBs.
(A–B) Confocal z-projections of pre-permeabilized cells (A) and quantification (B) of ALIX intensity per area in ACBs under asynchronous conditions (72 hr after transfection with siCon/siNups or siALIX/siNups). ACB area was defined by mAb SC35 staining. n = 4 biological replicates. (C, D) Confocal z-projections of pre-permeabilized cells (C) and quantification (D) of ALIX brightness per area in MIGs under asynchronous conditions (72 hr after transfection with siCon/siCon or siALIX/siCon). MIG area was defined by mAb SC35 staining. n = 4 biological replicates. (E) Representative images of MIGs marked by α-pppCHMP4C or α-pAurB. (F) Representative images from quantification in Figure 6E using α-pAurB to mark ACBs.

AurB inhibition decreases but does not completely dissolve ACBs.
(A) Quantification of midbody-stage cells (Intact) and recently-abscised midbody pairs (Cut) with ACBs marked by α-pAurB after treatment with siCon or siNups and thymidine synchronization. With and without 1 h treatment with AurBi added at 15 h, 1 h prior to harvest at 16 h. N=500 pairs per condition from n=5 biological replicates. (B) Number of ACBs per midbody pair (marked by α-pAurB) after treatment as in A (siNups only) n=3 biological replicates. (C) pAurB intensity per area in ACBs or at the midbody after treatment as in A (siNups only) n=4 biological replicates.
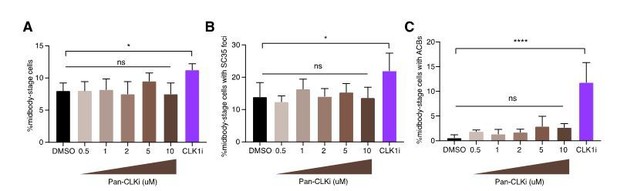
Specific CLK1/2 inhibition delays abscission timing.
(A) Quantification of midbody-stage cells after 5 h asynchronous treatment with DMSO, 0.5 – 10 µM Pan-CLK inhibitor (Pan-CLKi, TG003, Sigma), or 1 µM specific CLK1/2 inhibitor (CLK1i, 534350, Millipore). N=3200 cells from n=5 biological replicates. (B) Quantification of midbody-stage cells with foci marked by SC35 antibody after treatment as in A. N=800 midbody-stage cells from n=4 biological replicates. (C) Quantification of midbody-stage cells with ACBs (both larger and more numerous than MIGs, and present in later midbody-stage cells) marked by SC35 antibody after treatment as in A. N=800 midbody-stage cells from n=4 biological replicates.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (Homo sapiens) | HeLa-N | Maureen Powers Lab | HeLa cells selected for transfectability, ID confirmed by STR profiling | |
| Cell line (Homo sapiens) | RPE1 | Bruce Edgar Lab | RRID:CVCL_4388 | Non-transformed, ID confirmed by STR profiling |
| Transfected construct (Homo sapiens) | pLVX-CLK1 | This paper | Addgene Cat#:174088 | Lentiviral construct to inducibly express CLK1 |
| Antibody | anti-ALIX (Rabbit polyclonal) | Covance (This Lab) | RRID:AB_2892637 | IF (1:500), WB (1:500) |
| Antibody | anti-pAurB (Rabbit polyclonal) | Rockland | Cat#: 600-401-677 RRID:AB_206164 | IF (1:500) |
| Antibody | anti-pppCHMP4C (Rabbit polyclonal) | Pier Paolo D’Avino | N/A | IF (1:500) |
| Antibody | anti-SC35 (mouse monoclonal) | Abcam | Cat#: ab18826 RRID:AB_298608 | IF (1:5000) |
| Antibody | anti-SRRM2 (rabbit polyclonal) | Thermofisher | Cat#: PA5-66827 RRID:AB_2665182 | IF (1:1000) |
| Sequence-based reagent | siNup153 | Mackay et al., 2010 | siRNA | GGACUUGUUAGAUCUAGUU |
| Sequence-based reagent | siNup50 | Mackay et al., 2010 | siRNA | GGAGGACGCUUUUCUGGAU |
| Chemical compound, drug | CLK1/2 Inhibitor | Millipore | 534350 | 1 µM |
| Chemical compound, drug | DYRK3 Inhibitor | Tocris | GSK 626616 | 1 µM |
| Chemical compound, drug | AurB Inhibitor | Bio-Techne | ZM 447439 | 2 µM |
| Chemical compound, drug | Thymidine | CalBiochem | CAS 50-89-5 | 2 mM |
| Software, algorithm | Fiji | NIH | RRID:SCR_002285 |
Additional files
-
Supplementary file 1
Antibodies used for western blot and immunofluorescence in this study.
- https://cdn.elifesciences.org/articles/63743/elife-63743-supp1-v2.xlsx
-
Supplementary file 2
Plasmids and siRNAs used in this study.
- https://cdn.elifesciences.org/articles/63743/elife-63743-supp2-v2.xlsx
-
Supplementary file 3
Adjusted p-values calculated from this study.
- https://cdn.elifesciences.org/articles/63743/elife-63743-supp3-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/63743/elife-63743-transrepform-v2.docx





