Redox controls RecA protein activity via reversible oxidation of its methionine residues
Figures
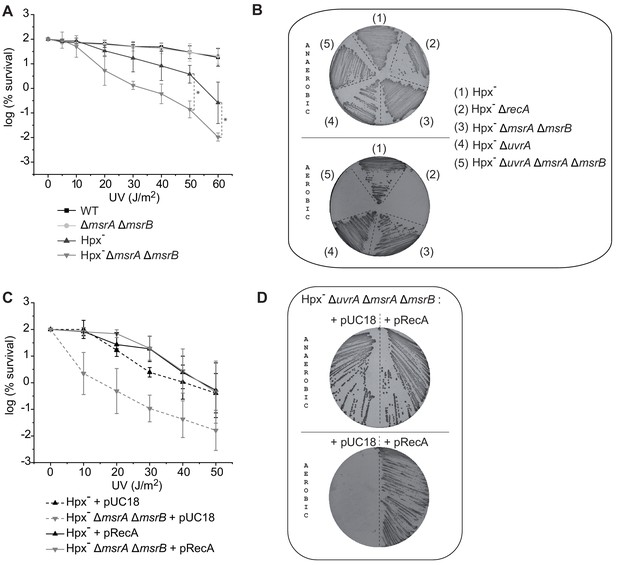
MsrA/B essential for maintaining RecA functional.
(A) UV sensitivity of wild-type (WT) and mutant strains (BE152 [ΔmsrA ΔmsrB]; BE007 [Hpx-]; BE080 [Hpx-ΔmsrA ΔmsrB]) was tested by monitoring colony-forming units at different UV doses. Curves represent the mean value of biological triplicates, and error bar ± represents the s.d. (B) Mutated strains (BE007 [Hpx-]; LL1609 [Hpx-ΔrecA]; BE080 [Hpx-ΔmsrA ΔmsrB]; BE032 [Hpx-ΔuvrA]; BE033 [Hpx-ΔmsrA ΔmsrB ΔuvrA]) on Lysogeny Broth (LB) plates incubated in the presence or absence of oxygen as indicated on the pictures. (C) Suppression assay for UV sensitivity with Hpx-ΔmsrA ΔmsrB carrying pRecA. (D) Suppression assay for oxygen sensitivity of BE033 (Hpx-ΔmsrA ΔmsrB ΔuvrA) mutant carrying the plasmid pRecA that overexpress RecA. Asterisks indicate a statistically significant difference between Hpx- and Hpx-ΔmsrA ΔmsrB, *p≤0.05 (Mann–Whitney U test).
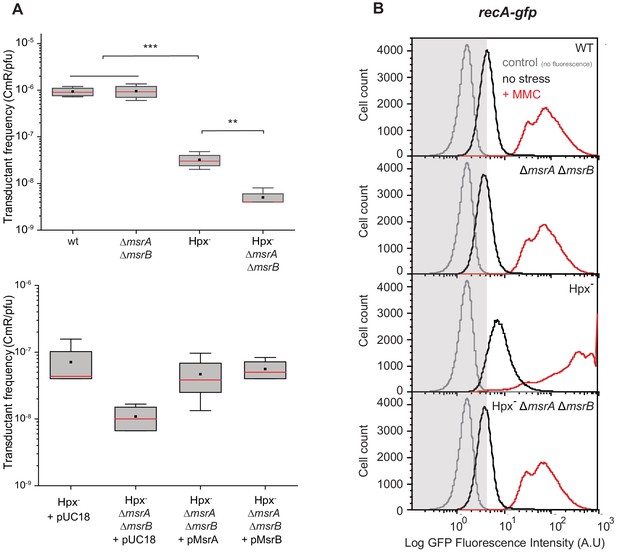
Physiological importance of RecA repair by MsrA and MsrB.
(A) Transductant frequency of a lacZ-cat marker in different genetics backgrounds. Top panel shows the transductant frequency of the BE080 (Hpx-ΔmsrA ΔmsrB) strain as recipient compared with the BE007 (Hpx-) strain, n = 6 and p=0.01. Bottom panel shows the Hpx- and Hpx-ΔmsrA ΔmsrB recipient strains carrying an empty vector pUC18, pMsrA, or pMsrB, n = 3–6, p=0.05. (B) SOS gene expression at the single cell level, monitored by flow cytometry. Green Fluorescent Protein (GFP) intensity of a transcriptional recA-gfp fusion in wild type (WT), ∆msrA ∆msrB, Hpx-, and Hpx- ∆msrA ∆msrB backgrounds cultivated in the absence (black line) or presence of mitomycin C (MMC) (0.25 μg/mL) (red line). The fluorescence intensity is proportional to the recA gene expression, which is used as an indicator of SOS response induction. In total, 100,000 events were analyzed for each sample.
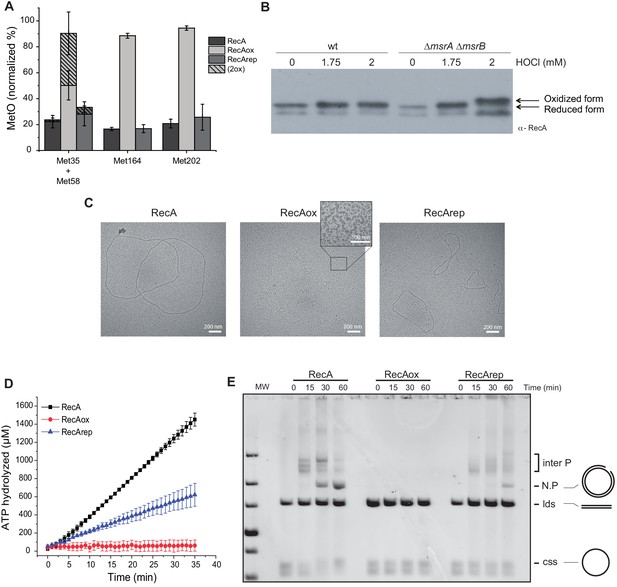
Reversible inactivation of RecA by MsrA and MsrB.
(A) RecA was treated with H2O2 to induce Met-O formation. Subsequently, oxidized RecA was incubated with MsrA and MsrB proteins in the presence of the reducing agent 1,4-dithiothreitol (DTT). The relative percentage of Met-O in various forms of RecA (native, oxidized [RecAox], and repaired [RecArep]) was determined by mass spectrometry analysis. Error ± means s.d., n = 3. (B) Wild type (WT) and BE152 (∆msrA ∆msrB) were cultivated in LB, and HOCl (1.75 and 2 mM) was added. Immunoblot analysis reveals RecA mobility on SDS-polyacrylamide gel electrophoresis. (C) Electron microscopy of RecA filaments. Filament formation was studied using different forms of RecA (native, RecAox, and RecArep) as indicated on the top of the pictures. (D) ATP hydrolysis rates by RecA, RecAox, and RecArep. (E) DNA strand exchange promoted by the different RecA species. The intermediate joint molecules (Inter P) product of exchange between circular single strand (css) and linear double strand (lds) DNA contains a three-stranded branch point that migrates along the molecule until nicked, forming a nicked circular duplex DNA (N.P.).

Quantification analysis of data from Figure 3C, E.
(A) Electron microscopy (EM) quantitation analysis, and DNA molecules observed by EM from Figure 3C were manually counted. For each condition, 2–4 squares of three or four grids of the replicates were fully counted. RecA nucleofilament/DNA molecules were categorized into six groups: fully filamented circles, medium filamented circles, small filamented circles, very small filamented circles, linear (broken circles), and single-strand binding protein (SSB)/DNA (circular single-strand DNA SSB coated). The proportion of each categories expressed in percentage is represented. Asterisks indicate a statistically significant difference, ****p≤0.0001 (Mann–Whitney U test). The numbers of molecules counted were respectively n:712 (RecA), n:829 (RecAox), and n:797 (RecArep). (B) Quantification analysis from Figure 3E. This analysis was carried out on four independents experiments. *p≤0.05 (Mann–Whitney U test).
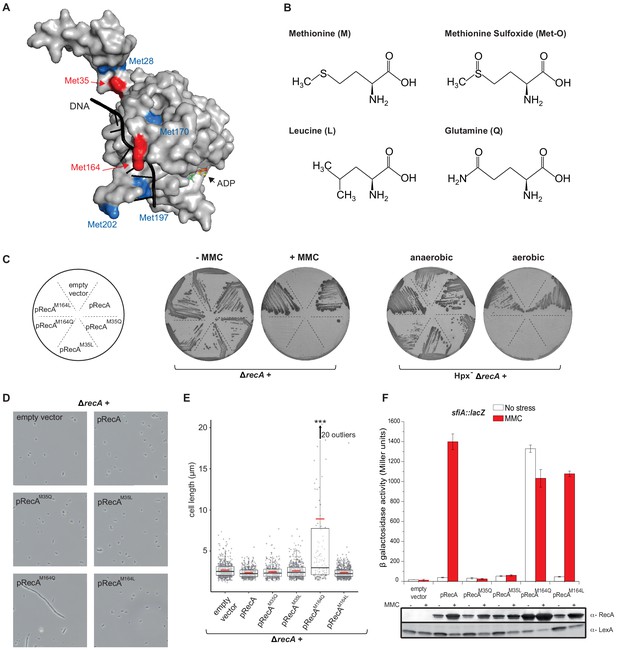
Mimicking Met oxidation yields to diverse functional consequences.
(A) Structure of the E. coli RecA-DNA complex adapted from PDB: 3CMW (Chen et al., 2008). The cartoon shows the structure of a RecA monomer (surface representation) in the presence of ssDNA in black and the ATP analogue adenosine diphosphate (ADP)-AlF4 binding site (for simplicity, only the structure of ADP is represented). Exposed methionine (Met) residues are colored in blue (Met28, Met170, Met197, Met202) or red (Met35, Met164). (B) Diagram of the structure of methionine (M), methionine sulfoxide (Met-O), leucine (L), and glutamine (Q). (C) Left panel shows the growth of LL1594 (ΔrecA) carrying plasmids with the different recA alleles under mitomycin C (MMC) stress. Right panel shows the growth of LL1609 (Hpx-ΔrecA) carrying the different recA alleles under aerobic condition. (D) LL1594 (ΔrecA) cells expressing in trans the different RecA variants were imaged in stationary phase by phase-contrast microscopy. While ΔrecA expressing RecA, RecAM35L, RecAM35Q, or RecAM164L exhibit a normal cell shape, long filaments were observed in the presence of RecAM164Q. (E) Quantitation of the cell length from the experiment shown in (D). This analysis was carried out on a large number of cells from at least 10 fields. The range of the number of cells is 151–554. Parametric statistical tests (t test) yield high significant statistic (p<10−5) between the ∆recA + pRecAM164Q group and any of the other groups. Among all the other groups, the differences are not significant. (F) SOS induction after 2 hr exposure to MMC in a ΔrecA strain expressing in trans the different recA alleles was monitored by sfiA::lacZ expression (top panel) and western blot anti-RecA and anti-LexA analyses (bottom panel). In the wild-type (WT) situation, MMC exposure leads to LexA cleavage, RecA production, and sfiA expression. Histogram graph of the mean expression-level value of the biological triplicates (top panel), error bar ± represents the s.d.
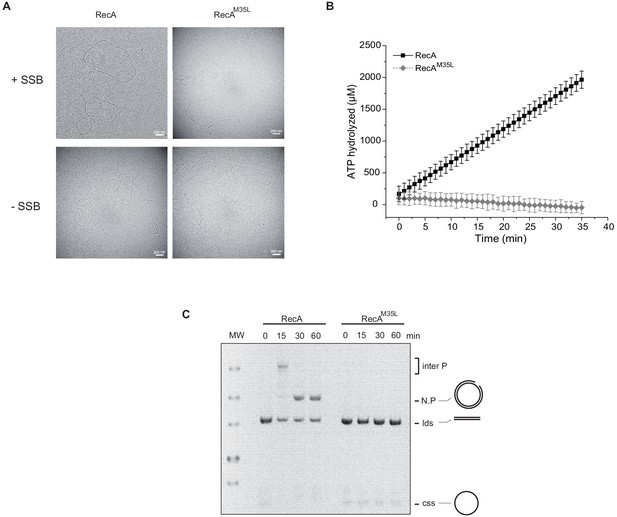
Importance of the residue Met35 of RecA.
(A) Electron microscopy of RecA filament. Filament formation was studied using RecA and RecAM35L as indicated at the top of the pictures. (B) RecAM35L was unable to hydrolyze ATP. (C) RecAM35L was unable to invade and/or catalyze DNA strand exchange.
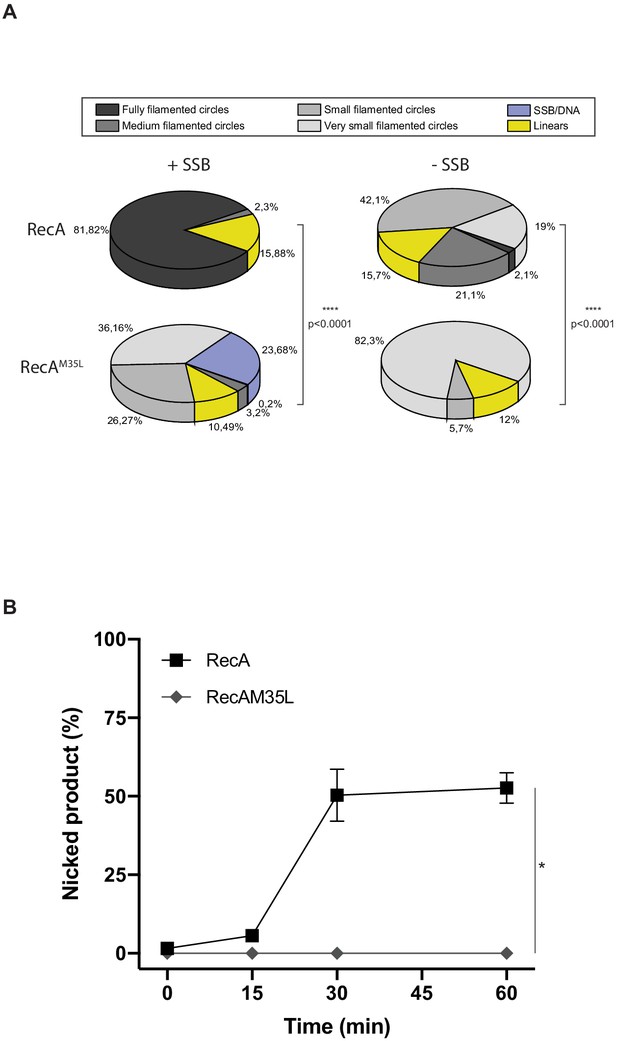
Quantification analysis of data from Figure 5.
(A) EM quantification analysis, and DNA molecules observed by EM from panel (A) were manually counted. For each condition, 2–4 squares of three or four grids of the replicates were fully counted. RecA nucleofilament/DNA molecules were categorized into six groups: fully filamented circles, medium filamented circles, small filamented circles, very small filamented circles, linear (broken circles), and single-strand binding protein (SSB)/DNA (circular single-strand DNA SSB coated). The proportion of each categories expressed in percentage is represented. Asterisks indicate a statistically significant difference, **p≤0.01, ***p≤0.001, and ****p≤0.0001 (Mann–Whitney U test). The numbers of molecules counted were respectively n:1588 (RecA + SSB), n:1956 (RecAM35L + SSB), n:1693 (RecA – SSB), and n:1438 (RecAM35L – SSB). (B) RecAM35L was unable to invade and/or catalyze DNA strand exchange. *p≤0.05 (Mann–Whitney U test).
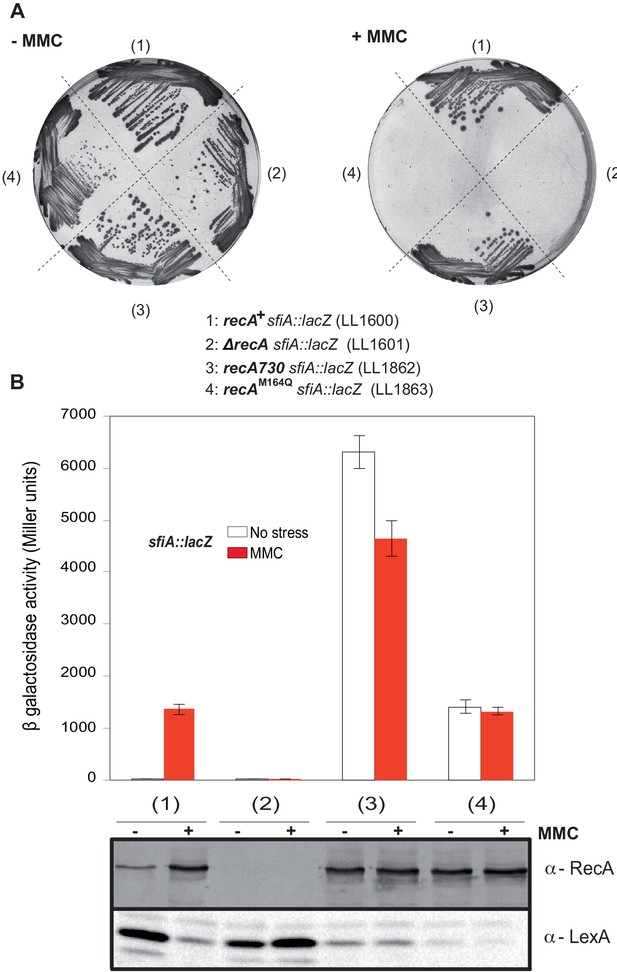
Characterization of the recA-M164Q chromosomal allele and comparison with recA730.
The sfiA::lacZ fusion was inserted in wild type (WT) (LL1600), ∆recA (LL1601), recA730 (LL1862), and recA-M164Q (LL1863) strains. (A) Strains tested, as indicated under the figure, were incubated on LB plates in the presence or absence of mitomycin C (MMC). (B) SOS induction after 2 hr exposure to MMC was monitored by assaying sfiA::lacZ expression (top panel) and western blot anti-RecA and anti-LexA analyses (bottom panel).
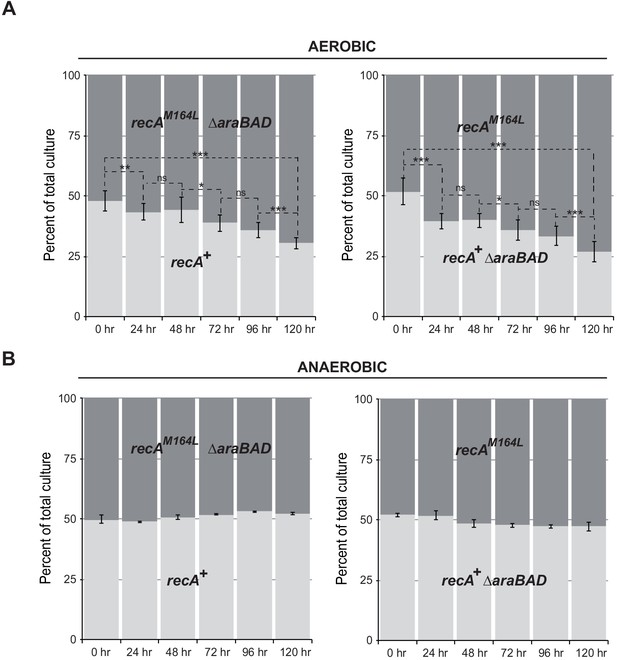
recA-M164L chromosomal allele enhances growth competition of E. coli in mixed aerobic culture.
(A) In aerobic condition, recA-M164L outcompetes wild type (WT). E. coli recAWT (MG1655) and recA-M164L (LL1781) mutant strains were grown overnight in (LB)-rich medium. Cultures were then diluted into fresh LB medium and used in co-cultures with recAWT and recA-M164L ∆araBAD (LL1976) or recAWT ∆araBAD (LL1974) and recA-M164L (LL1781). ∆araBAD is a neutral mutation used to differentiate strains in the competition assay; loss of the araBAD operon results in red colonies on tetrazolium arabinose medium. The cell density was adjusted to obtain a 1:1 ratio of each of the two strains. The two strains were then grown together in fresh LB medium over 24 hr periods. Total cell titers were determined by serial dilutions on LB-tetrazolium arabinose agar plates after 0, 24, 48, 72, 96, and 120 hr of culture. (B) In anaerobic condition, a 1:1 ratio of recA-M164L and recAWT remains over time. The same protocol was used under anaerobic condition with LB containing 0.2% glucose. Asterisks indicate a statistically significant difference, *p≤0.05; **p≤0.01, and ***p≤0.001, and ns indicates a not statistically significant difference (Mann–Whitney U test).
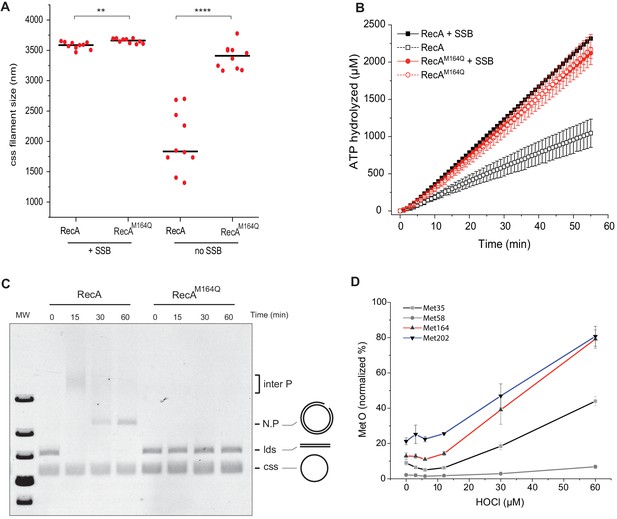
Biochemistry characterization of the RecAM164Q, a position highly prone to oxidation.
(A) Size measurement of cssDNA-RecA and cssDNA-RecAM164Q filament observed by electron microscopy in the absence or presence of single-strand binding protein (SSB). Red dots indicate the size of each individual filament, and black lines indicate the median values. Asterisks indicate a statistically significant difference, **p≤0.01 and ****p≤0.0001 (Mann–Whitney) ; n = 10. (B) ATP hydrolytic activity of RecA and RecAM164Q in the presence or absence of SSB. (C) RecAM164Q is unable to invade and/or catalyze DNA strand exchange. (D) Purified RecA protein was treated with increasing concentration of HOCl followed by mass spectrometry analysis. The relative percentage of Met-O was determined. Met residues showed intrinsic sensitivity to HOCl depending upon their position in the polypeptide. Error bar ± means s.d., n = 3. css: circular single strand.
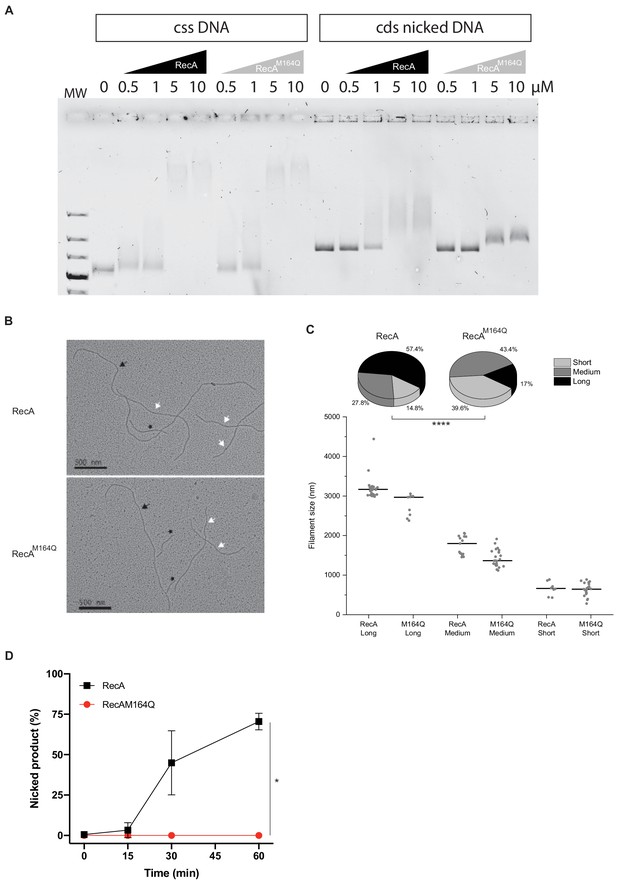
RecAM164Q double-strand affinity is reduced.
(A) Electrophoresis mobility shift assay in the presence of circular single-strand DNA (cssDNA) or circular double strand (cds) nicked DNA. Various concentrations of RecA or RecAM164Q are used (top). With the cssDNA substrate, the DNA band starts to shift at 0.5 μM of either RecA or RecAM164Q and a strong shift is observed for 5 and 10 μM of RecA proteins. With cds nicked DNA, the shift starts in the presence of 1 μM RecA and increases for 5 and 10 μM. In contrast, for RecAM164Q the shift is reduced and starts at 5 μM. (B, C) Electron microscopy of RecA filament formation on linear double- strand DNA, and measurement and distribution of the filaments sizes. Images were selected to have the three types of filaments, long (black arrow), medium (white arrow), and short (black star). RecA forms mostly long and medium filaments, while RecAM164Q forms mostly short and medium filaments. Asterisks indicate a statistically significant difference, ****p≤0.0001 (Mann–Whitney U test). The numbers of molecules counted were respectively n:54 (RecA) and n:53 (RecAM164Q). (D) Quantification of Figure 8C. This analysis was carried out on three independents experiments, *p≤0.05 (Mann–Whitney U test).
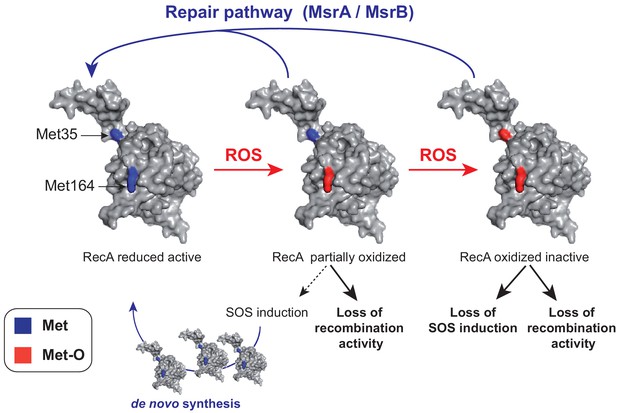
A model of homeostasis control of RecA under oxidative stress.
RecA structural model is shown with the two Met residues (Met35 and Met164) analyzed in the present work. The RecA partially oxidized form contains a Met164-oxidized residue while the RecA-oxidized inactive form contains both Met35- and Met164-oxidized residues. The MsrA/B repair pathway allows to maintain the pool of functional RecA by repair, and the SOS induction pathway allows to replenish the pool of functional RecA by de novo synthesis.

Amino acid sequence alignment of RecA/Rad51.
The protein sequence comparison between the bacterial RecA consensus sequence (RecA-bact) obtained from Lusetti and Cox, 2002, the E. coli RecA sequence (RecA-E. coli), and the human and S. cerevisiae Rad51 sequences (Rad51-H. sapiens and Rad51-S. cerevisiae) was performed using Clustal Omega (https://www.ebi.ac.uk/Tools/msa/clustalo/). Methionine residues (Met35, Met58, Met164, Met202) are colored in red above the sequence. * indicates positions that have a single, fully conserved residue, : indicates conservation between groups of strongly similar properties, and . indicates conservation between groups of weakly similar properties.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information | |
|---|---|---|---|---|---|
| Strain, strain background (Escherichia coli) | MG1655 | LCB Collection | N/A | ||
| Strain, strain background (Escherichia coli) | BE152 | This study | N/A | ΔmsrA ΔmsrB | |
| Strain, strain background (Escherichia coli) | CH011 | This study | N/A | ΔmsrA::FRT ΔmsrB::FRT | |
| Strain, strain background (Escherichia coli) | CH074 | This study | N/A | ΔrecA::kan | |
| Strain, strain background (Escherichia coli) | LL401 | Loiseau et al., 2007 | N/A | arap::erpA | |
| Strain, strain background (Escherichia coli) | LL1594 | This study | N/A | ΔrecA::spc | |
| Strain, strain background (Escherichia coli) | BE007 | Ezraty et al., 2014 | N/A | Hpx- = ΔkatE ΔkatG ahpC::kan | |
| Strain, strain background (Escherichia coli) | BE080 | This study | N/A | BE007 ΔmsrA ΔmsrB | |
| Strain, strain background (Escherichia coli) | LL1609 | This study | N/A | BE007 ΔrecA::spec | |
| Strain, strain background (Escherichia coli) | BE032 | This study | N/A | BE007 ΔuvrA | |
| Strain, strain background (Escherichia coli) | BE033 | This study | N/A | BE007 ΔuvrA ΔmsrA ΔmsrB | |
| Strain, strain background (Escherichia coli) | sfiA::lacZ | Grompone et al., 2004 | N/A | ΔlacZ::cat sfiA::Mud lacZ cat-bla | |
| Strain, strain background (Escherichia coli) | CH039 | This study | N/A | CH011 ΔlacZ::cat sfiA::Mud lacZ cat-bla | |
| Strain, strain background (Escherichia coli) | LL1713 | This study | N/A | recA::gfp-cat | |
| Strain, strain background (Escherichia coli) | EAW831 | This study | N/A | sfiA ::FRT recAM164Q | |
| Strain, strain background (Escherichia coli) | EAW287 | Robinson et al., 2015 | N/A | sfiA ::FRT recAE38K::kan (recA730) | |
| Strain, strain background (Escherichia coli) | LL1601 | This study | N/A | MG1655 ΔlacZ::cat sfiA::Mud lacZ cat-bla ΔrecA::spc | |
| Strain, strain background (Escherichia coli) | LL1862 | This study | N/A | MG1655 ΔlacZ::cat sfiA::Mud lacZ cat-bla recAE38K::kan (recA730) | |
| Strain, strain background (Escherichia coli) | LL1863 | This study | N/A | MG1655 ΔlacZ::cat sfiA::Mud lacZ cat-bla + recAM164Q | |
| Strain, strain background (Escherichia coli) | LL1976 | This study | N/A | MG1655 recAM164L ∆araBAD::kan | |
| Strain, strain background (Escherichia coli) | LL1781 | This study | N/A | MG1655 recAM164L | |
| Strain, strain background (Escherichia coli) | LL1974 | This study | N/A | MG1655 ∆araBAD::kan | |
| Strain, strain background (Escherichia coli) | BW25113 | LCB Collection | N/A | F-, ∆(araD-araB)567, lacZ4787(del)::rrnB-3, LAM-, rph-1, ∆(rhaD-rhaB)568, hsdR514 | |
| Strain, strain background (Escherichia coli) | STL2669 | A gift from S Lovett | N/A | (ΔrecA-srlR)306::Tn10 TetRxonA2 (exoI-) | |
| Strain, strain background (Escherichia coli) | DH5α | LCB Collection | N/A | F– endA1 glnV44 thi-1 recA1 relA1 gyrA96 deoR nupG purB20 φ80dlacZΔM15 Δ(lacZYA-argF)U169, hsdR17(rK–mK+), λ– | |
| Plasmids | pGEMT-easy | Promega | A1360 | ‘High’ copy vector, cloning site into lacZ ApR | |
| Plasmids | pUC18 | LCB Collection | N/A | High copy vector with the small size MCS inverted as pUC19 ApR | |
| Plasmids | pMsrA | LCB Collection | N/A | pUC18 - msrA+ | |
| Plasmids | pMsrB | LCB Collection | N/A | pUC18 - msrB+ | |
| Plasmids | pRecA | A gift from M. Modesti | N/A | pUC18 - recA+ | |
| Plasmids | pBBR | LCB Collection | N/A | ‘Medium-low’ copy vector pBBR-MCS2 + cat HindIII CmR KanR | |
| Plasmids | pBBR-RecA | This study | N/A | pBBR - recA SacII/SpeI | |
| Plasmids | pBBR-RecAM35L | This study | N/A | pBBR - recAM35L SacII/SpeI | |
| Plasmids | pBBR-RecAM164L | This study | N/A | pBBR - recAM164L SacII/SpeI | |
| Plasmids | pBBR-RecAM35Q | This study | N/A | pBBR - recAM35Q SacII/SpeI | |
| Plasmids | pBBR-RecAM164Q | This study | N/A | pBBR - recAM164Q EcoRI | |
| Plasmids | pEAW1078 | This study | N/A | Derived from pEAW260 - recAM35L ApR | |
| Plasmids | pCJH0002 | This study | N/A | Derived from pEAW260 - recAM164Q ApR | |
| Plasmids | pT7pol26 | Cox Lab Collection | N/A | Carries the T7 RNA polymerase under the control of a lac promoter | |
| Sequenced-based reagent | recA for | This study | PCR primers | CGGTGCGTCGTCAGGCTACTGCGT | |
| Sequenced-based reagent | recA rev | This study | PCR primers | GCCAGAATGCGTACCGCACGAT | |
| Sequenced-based reagent | M35L for | This study | PCR primers | GACCGTTCCCTGGATGTGGAAACCATCTCTACCGGTT CGCTTTCACTGGATATCGCGCTT | |
| Sequenced-based reagent | M35L rev | This study | PCR primers | GGTTTCCACATCCAGGGAACGGTCTTCA CCCAGGCGCATG | |
| Sequenced-based reagent | M35Q for | This study | PCR primers | GACCGTTCCCAGGATGTGGAAACCATCTCTACCGG TTCGCTTTCACTGGATATCGCG | |
| Sequenced-based reagent | M35Q rev | This study | PCR primers | TTCCACATCCTGGGAACGGTCTTCACCCAGGCGCAT GATGGAGCC | |
| Sequenced-based reagent | M164L for | This study | PCR primers | GGCGACTCTCACCTGGGCCTTGCGGCACGTATGATGAGCCAGGCG | |
| Sequenced-based reagent | M164L rev | This study | PCR primers | TGCCGCAAGGCCCAGGTGAGAGTCGCCGATTTCGCCTTCGATTTC | |
| Sequenced-based reagent | M164Q for | This study | PCR primers | GGCGACTCTCACCAGGGCCTTGCGGCACGTATGATGAGCCAGGCG | |
| Sequenced-based reagent | M164Q rev | This study | PCR primers | TGCCGCAAGGCCCTGGTGAGAGTCGCCGATTTCGCCTTCGATTTC | |
| Sequenced-based reagent | T7 pro | This study | PCR primers | TAATACGACTCACTATAGGG | |
| Sequenced-based reagent | recA*-M35L-rev | This study | PCR primers | GTAGAGATGGTTT CCACATCCAGGGAA CGGTCTTC | |
| Sequenced-based reagent | recA*-M164Q-rev | This study | PCR primers | TGCCGCAAGGCCCT GGTGAGAGTCGCCG | |
| Sequenced-based reagent | 5for recA M165L SceI K7kan | This study | PCR primers | GGCGACTCTCACCTGGG CCTTGCG GCACGTATGATGAGCCAGGC GTGTAGGCTGGAGCTGCTTC | |
| Sequenced-based reagent | 3rev recA K7Kan | This study | PCR primers | AATTTTCATACGGATCTGGTTGAT GAAGATCAGCAGCGTCCT ATTACCCTGTTATCCCTACC TTAGTTCCTATTCCGAAGTTC | |
| Sequenced-based reagent | 5-recA-GFP | This study | PCR primers | GATGATAGCGAAGGCGTAGCAGAAACTAACGAAGA TTTTTAATAAGAAGGAGATATACATATGAG | |
| Sequenced-based reagent | 3-recA-K7Cat | This study | PCR primers | AGGGCCGCAGATGCGACCCTTGTGTATCAAACAAGA CGATTATCACTTATTCAGGCGTA | |
| Sequenced-based reagent | 5-K7wanner-spc | This study | PCR primers | TGTGTAGGCTGGAGCTGCTTCGAAGTTCCTATACTT TCTAGCAGGAGGAATTCACCATGAGTGAAAAAGTGCCCGCC | |
| Sequenced-based reagent | 3-K7wanner-spc | This study | PCR primers | CATATGAATATCCTCCTTAGTTCCTATTCCGAAGT TCCTATTCTCTAGAAAG TATCATTGGCTGGCA CCAAGCAGTTTA | |
| Sequenced-based reagent | 5-wanner-recA | This study | PCR primers | ATTGACTATCCGGTATTACCCGGCATGACAGGAGTAAAAGTGTAGGCTGGAGCTGCTTC | |
| Sequenced-based reagent | 3-wanner-recA | This study | PCR primers | AGGGCCGCAGATGCGACCCTTGTGTATCAAACAAGACGACATATGAATATCCTCCTTA | |
| Antibody | Anti-RecA (Rabbit polyclonal) | Abcam | 63797 | WB (1∶20,000) | |
| Antibody | Anti-LexA (Rabbit polyclonal) | P. Moreau (LCB) | WB (1∶5,000) | ||
| Antibody | Goat anti-rabbit IgG-Alk Phos (polyclonal) | Chemicon International | AP132A | WB (1 :10,000) | |
| Antibody | Goat anti-rabbit HRP (polyclonal) | Promega | W4018 | WB (1 :10,000) | |
| Commercial assay or kit | ECL western blotting kit | Thermo Scientific | 32106 | ||
| Commercial assay or kit | Gel purified using kit | Promega | A9282 | ||
| Commercial assay or kit | Protein assay kit | Bio-Rad | 23236 | ||
| Software, algorithm | QI-Macros software | KnowWare International, Inc, CO | |||
| Chemical compound, drug | ProteaseMAX | Promega | V2071 | ||
| Chemical compound, drug | Trypsin Gold | Promega | V5280 | ||
| Chemical compound, drug | Sodium hypochlorite | Acros Organics | AC219255000 | ||
| Chemical compound, drug | Phusion enzyme | Thermo Scientific | F531S | ||
| Chemical compound, drug | LB (Lysogeny Broth) | Difco | 244620 | ||
| Other | Vinyl Anaerobic Chambers | Coy | Coy Laboratory Products Inc, USA | coylab.com | |
| Other | Typhoon FLA-9000 | GE Healthcare | FLA9000 | ||
| Other | Thermo Scientific | F531S | |||
Sequences of RecA peptides containing Met residues and m/z expected for the different oxidation states.
| Peptide sequence | Reduced | 1× Ox | 2× Ox | 3× Ox |
|---|---|---|---|---|
| MMSQAMR | 427.687 +2 | 435.684 +2 | 443.681 +2 | 451.678 +2 |
| AEIEGEIGDSHMGLAAR | 585.949 +3 | 591.280 +3 | – | – |
| IGVMFGNPETTTGGNALK | 903.956 +2 | 911.953 +2 | – | – |
| SMDVETISTGSLSLDIALGAGGLPMGR | 1324.667 +2 | 1332.664 +2 | 1340.661 +2 | – |





