Augmenter of liver regeneration regulates cellular iron homeostasis by modulating mitochondrial transport of ATP-binding cassette B8
Figures
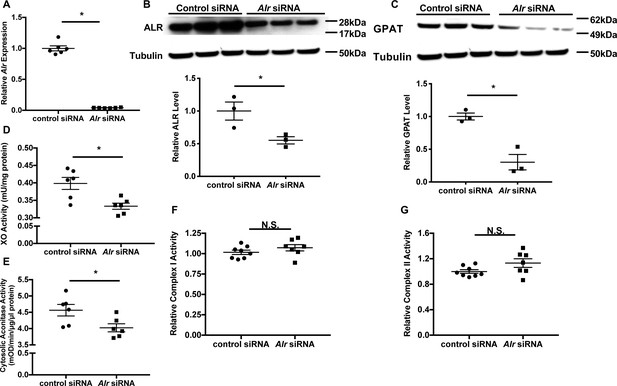
Downregulation of Alr results in cytoplasmic Fe/S cluster maturation defects.
(A) mRNA (n = 6) and (B) protein (n = 3) levels of Alr in wild type (WT) mouse embryonic fibroblasts (MEFs) with or without Alr downregulation. GPAT protein levels (n = 3, C), xanthine oxidase (XO, n = 6, D), and cytosolic aconitase (n = 6, E) activities in cells treated with Alr siRNA. Mitochondrial complex I (n = 7–8, F) and complex II activity (n = 7–8, G) in WT MEFs with Alr downregulation. Cells were harvested 48 hr after transfection for RNA analysis and 72 hr after transfection for protein analysis. Quantification of western blotting images is shown in the same panel. Data are presented as mean ± standard error mean (SEM). *p<0.05 by ANOVA. N.S. = not significant.
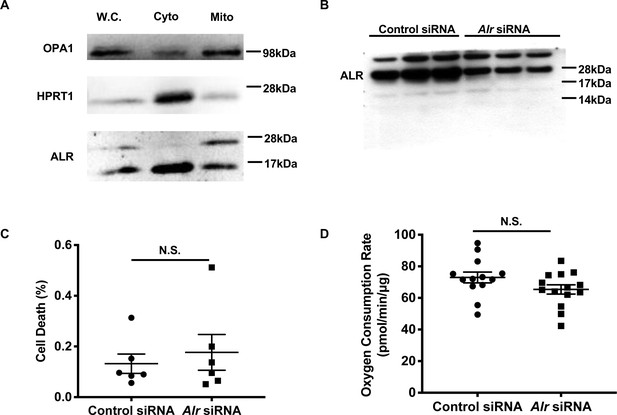
Acute downregulation of Alr does not affect cell viability or mitochondrial respiration.
(A) Representative western blotting of total cell lysate, cytosolic and mitochondrial fraction from MEFs demonstrate the localization of long and short isoforms of ALR. OPA1 serves as a mitochondrial fraction marker while HPRT1 serves as a cytosolic fraction marker. W.C. = whole cell lysate. Cyto = cytosolic fraction. Mito = mitochondrial fraction. (B) Uncropped western blot from Figure 1B demonstrating the effect of siRNA on full-length and short isoforms of ALR. (C) Cell death in cells treated with control or Alr siRNA and assayed 72 hr after transfection using propidium iodide and Hoechst 33342 staining. (D) Mitochondrial oxygen consumptions assayed 60 hr after cells are treated with control or Alr siRNA in the presence of glucose and L-glutamine. Data are presented as mean ± SEM. N.S. = not significant by ANOVA.
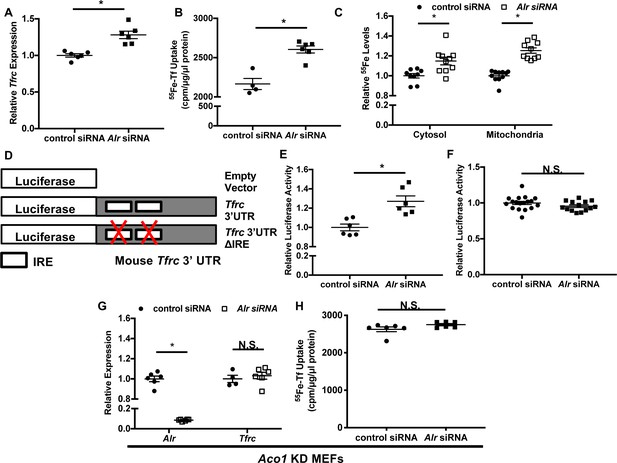
Downregulation of ALR increases Tfrc levels and cellular iron content through IRP1.
Tfrc mRNA levels (A) and transferrin-dependent iron uptake (B) in wild type (WT) mouse embryonic fibroblasts (MEFs) with Alr downregulation (n = 5–6). (C) Steady-state 55Fe levels in cytosolic and mitochondrial fraction from cells treated with Alr siRNA (n = 6). (D) Schematic representation of Tfrc 3’UTR reporter constructs. (E) Full-length Tfrc 3’UTR reporter activities were measured in WT MEFs with ALR downregulation (n = 6). (F) IRE-deleted Tfrc 3’UTR reporter activities in WT MEFs with ALR downregulation (n = 14–18). (G) Alr and Tfrc mRNA levels in Aco1 knockdown (KD) MEFs treated with Alr siRNA (n = 4–6). (H) TFRC-mediated 55Fe uptake in Aco1 KD MEFs treated with Alr siRNA (n = 6). Experiments were run 48 hr after siRNA transfection for mRNA levels and 60 hr after transfection for steady state 55Fe levels and transferrin-mediated iron uptake. For luciferase assay, cells were sequentially transfected with siRNA and reporter construct (24 hr apart), and assayed 24 hr after transfection of reporter constructs. Data are presented as mean ± SEM. *p<0.05 by ANOVA. N.S. = not significant.
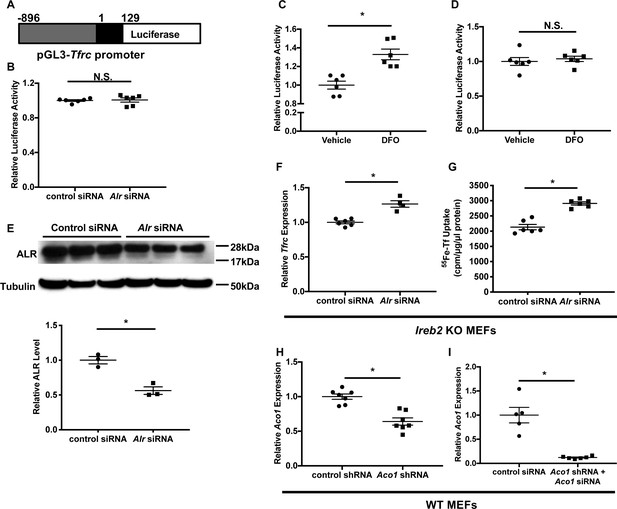
ALR regulates Tfrc mRNA levels through a posttranscriptional mechanism independent of IRP2.
(A) Schematic representation of the Tfrc promoter reporter construct. +1 represents transcription start site on Tfrc genomic sequence. (B) Tfrc promoter activities are measured in wild type (WT) mouse embryonic fibroblasts (MEFs) with Alr downregulation (n = 6). Cells were first transfected with Alr siRNA and luciferase construct was transfected 24 hr later. 24 hr after luciferase transfection, cells were harvested for luciferase assay. Luciferase activity of full-length Tfrc 3’UTR (n = 6, C) and Tfrc 3’UTR ΔIRE (n = 6, D) constructs in cells treated with vehicle or 150 μM DFO for 16 hr. (E) Alr protein levels in Ireb2 KO MEFs treated with Alr siRNA (n = 3). Densitometry analysis is shown below the western blots. (F) Tfrc mRNA level in Ireb2 knockout (KO) MEFs treated with Alr siRNA and harvested 48 hr after transfection (n = 4–6). (G) 55Fe-transferrin uptake in Irrb2 KO MEFs treated with Alr siRNA (n = 6). Experiment was performed 60 hr after transfection. (H) Aco1 mRNA levels in WT MEFs stably expressing Aco1 shRNA at baseline (n = 6). (I) Aco1 mRNA levels in WT MEFs stably expressing Aco1 shRNA and transfected with Aco1 siRNA (n = 5–6). mRNA levels were assayed 48 hr after siRNA transfection. Data are presented as mean ± SEM.* p<0.05 by ANOVA. N.S. = not significant.
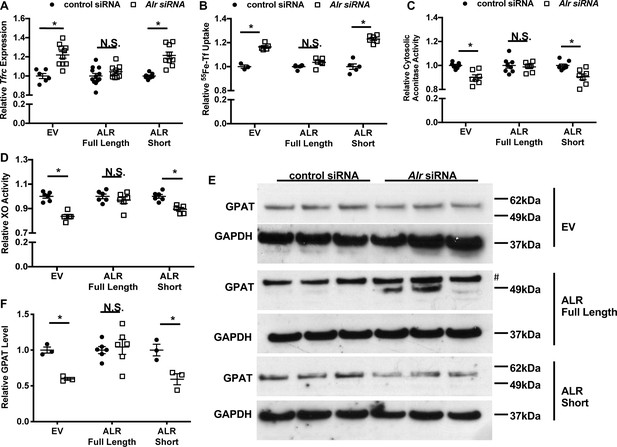
Overexpression of full-length but not cytosolic isoform of ALR rescues the iron-sulfur cluster maturation defects from ALR downregulation.
Tfrc mRNA levels (n = 6, A) and transferrin-dependent iron uptake (n = 4–6, B) were measured in wild type (WT) mouse embryonic fibroblasts (MEFs) with endogenous Alr downregulation and concurrent overexpression of different ALR isoforms. Cytosolic aconitase (n = 7–8, C) and XO activities (n = 5–6, D), and GPAT protein levels (n = 3–6, E) are measured in cells with downregulation of endogenous ALR and overexpression of various ALR isoforms. # indicates the band corresponds to GPAT protein. (F) Quantification of panel E. EV = empty vector. ALR short = cytosolic isoform of ALR lacking the first 80 amino acids. Data are presented as mean ± SEM. Cells were infected with lentivirus overexpressing the respective ALR isoforms, and transfected with 3’UTR-specific ALR siRNA 24 hr after lentivirus infection. mRNA levels were assayed 48 hr after siRNA transfection, and protein levels and cytosolic XO activity were assayed 72 hr after siRNA transfection. *p<0.05 by ANOVA with post hoc Tukey comparison. N.S. = not significant.
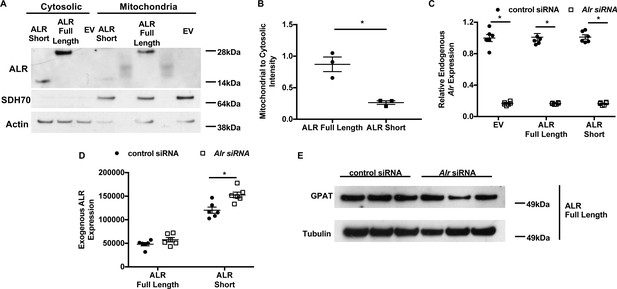
Validation of isoform-specific overexpression of ALR with concurrent downregulation of endogenous Alr.
(A) Representative western blotting results showing the subcellular localization of full-length and cytosolic isoform of ALR. (B) Densitometry analysis mitochondrial localization of each construct shown in panel A (n = 3). (C) Expression of endogenous Alr in wild type (WT) mouse embryonic fibroblasts (MEFs) treated with Alr siRNA targeting 3’UTR of the endogenous locus and overexpressing indicated constructs (n = 6). (D) mRNA levels of overexpressed ALR in WT MEFs treated with Alr siRNA targeting 3’UTR of the endogenous locus and overexpressing indicated constructs (n = 6). (E) Additional western blot showing GPAT protein level in MEF cells with overexpression of full-length isoform of ALR and treated with indicated siRNA. EV = empty vector. ALR short = truncated ALR lacking the first 80 amino acids. SDH70 = 70 kDa subunit of succinate dehydrogenase, a mitochondrial marker. Data are presented as mean ± SEM.* p<0.05 by ANOVA.
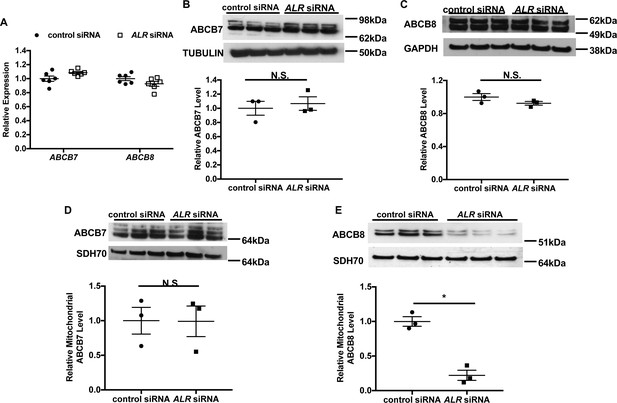
Downregulation of ALR in HEK293 cells reduces mitochondrial ABCB8 protein level.
(A) mRNA levels of ABCB7 and ABCB8 in HEK293 cells with ALR downregulation harvested 48 hr after siRNA transfection (n = 6). Total cellular levels of ABCB7 (B) and ABCB8 (C) in HEK293 cells with ALR downregulation. Mitochondrial levels of ABCB7 (D) and ABCB8 (E) in HEK293 cells with ALR downregulation. Samples for protein analysis were harvested 72 hr after transfection. Densitometry analysis is shown together with representative images. n = 3 for panels B–E. SDH70 = 70 kDa subunit of mitochondrial succinate dehydrogenase, a mitochondrial loading control. Data are presented as mean ± SEM. *p<0.05 by ANOVA. N.S. = not significant.

Downregulation of ALR in HEK293 cells results in cytosolic Fe/S cluster maturation defects.
(A) ALR mRNA levels in HEK293 cells treated with ALR siRNA and harvested 48 hr after transfection (n = 6). (B) Representative western blot showing ALR protein levels in HEK293 cells treated with ALR siRNA (n = 3). (C) Representative western blot showing GPAT protein levels in HEK293 cells treated with ALR siRNA (n = 3). (D) Densitometry analysis of western blots shown in in panel C. Samples for protein analysis were harvested 72 hr after transfection. (E) Tfrc mRNA levels in wild type (WT) mouse embryonic fibroblasts (MEFs) overexpressing ABCB7 and/or treated with Alr siRNA (n = 5–6). Cells were sequentially transfected with ABCB7 overexpression plasmid and Alr siRNA, and harvested 48 hr after siRNA transfection. Data are presented as mean ± SEM.* p<0.05 by ANOVA.
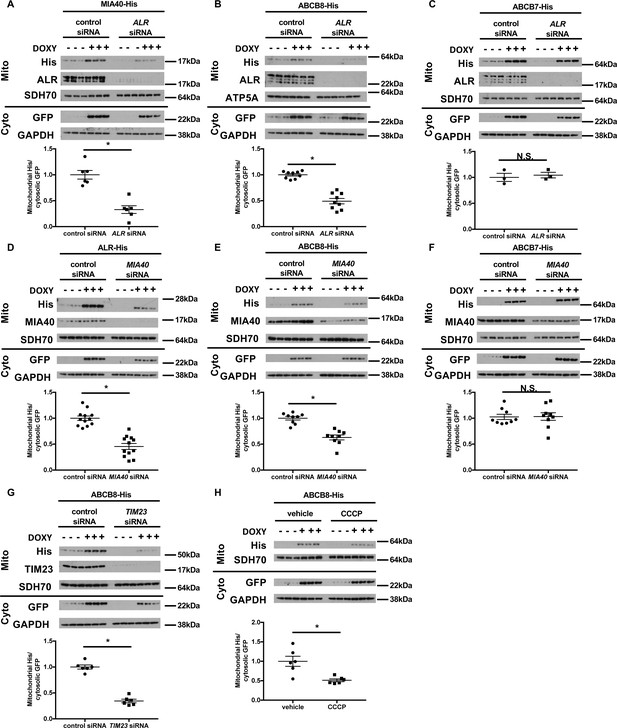
Mitochondrial import of ABCB8 requires ALR, MIA40, TIM23, and mitochondrial membrane potential.
Imported levels of MIA40-His (n = 6, A), whose mitochondrial translocation depends on Mia40/ALR protein transport pathway, ABCB8-His (n = 9, B), and ABCB7-His (n = 3, C) in cells treated with ALR siRNA with and without doxycycline (DOXY) treatment for 6 hr. Imported levels of ALR-His (n = 12, D), ABCB8-His (n = 9, E), and ABCB7-His (n = 9, F) in cells with or without MIA40 downregulation with and without DOXY treatment for 6 hr. (G) Imported levels of ABCB8-His in cells with or without MIA40 downregulation with and without DOXY treatment for 6 hr (n = 6). (H) Imported levels of ABCB8-His in cells treated with carbonyl cyanide 3-chlorophenylhydrazone (CCCP) and/or DOXY (n = 6). Representative western blotting results and densitometry analyses are shown in each panel. Mito = mitochondrial fraction. Cyto = cytosolic fraction. SDH70 = 70 kDa subunit of succinate dehydrogenase. ATP5A = ATP synthase subunit 5A. Cells were first transfected with siRNA and then treated with doxycycline when indicated 48 hr later. Data are presented as mean ± SEM. *p<0.05 by ANOVA. N.S. = not significant.
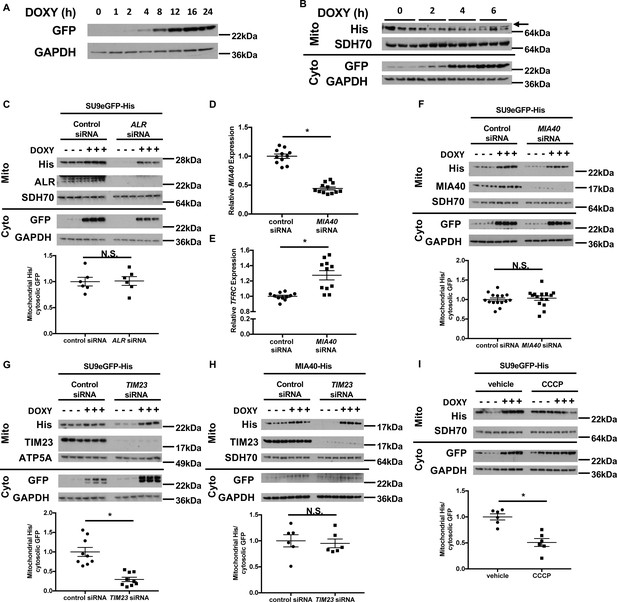
Downregulation of ALR or MIA40 does not affect mitochondrial import of SU9-eGFP.
(A) Representative western blot showing time-dependent accumulation of GFP protein in cells stably expressing rtTA3 and transfected with doxycycline (DOXY)-inducible GFP expression plasmid. (B) Representative western blot images showing time-dependent accumulation of GFP in the cytosol and ABCB7 in the mitochondria in cells stably expressing rtTA3 and transfected with the bicistronic doxycycline-inducible plasmid containing GFP and ABCB7. Arrow indicates the correct band for ABCB7. (C) Imported levels of SU9eGFP-His in cells with or without ALR downregulation with and without doxycycline treatment (n = 6). MIA40 (D) and TFRC (E) mRNA levels in HEK293 cells treated with MIA40 siRNA and harvested 48 hr later (n = 11–12). (F) Imported levels of SU9eGFP-His in cells with or without MIA40 downregulation with and without doxycycline treatment (n = 15). (G) Imported levels of SU9eGFP-His in cells with or without TIM23 downregulation with and without doxycycline treatment (n = 9). (H) Imported levels of MIA40-His in cells with or without TIM23 downregulation with and without doxycycline treatment (n = 6). (I) Imported levels of SU9eGFP-His in cells treated with carbonyl cyanide 3-chlorophenylhydrazone (CCCP) and/or DOXY treatment (n = 6). Mito = mitochondrial fraction. Cyto = cytosolic fraction. SDH70 = 70 kDa subunit of mitochondrial succinate dehydrogenase. ATP5A = ATP synthase subunit 5A. Both of them served as mitochondrial loading control. For panels C, F-I, cells were first transfected with siRNA and then treated with doxycycline when indicated 48 hr later. Data are presented as mean ± SEM.* p<0.05 by ANOVA. N.S. = not significant.
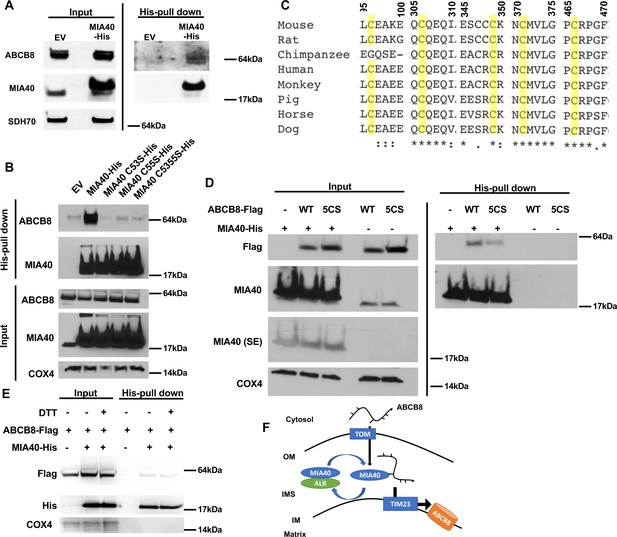
ABCB8 interacts with Mia40 during its transport into mitochondria.
(A) Representative western blot image of mitochondrial lysates and His-tag pulldown fraction from cells overexpressing His-tagged MIA40 or empty vector. (B) Co-immunoprecipitation of His-tagged MIA40 and endogenous ABCB8 in mitochondrial fraction from cells overexpressing indicated constructs. (C) Clustal Omega alignment of ABCB8 protein sequences from higher vertebrate showing the location of five conserved cysteines (highlighted in yellow). (D) Co-immunoprecipitation of His-tagged MIA40 and Flag-tagged ABCB8 in mitochondrial fraction from cells overexpressing indicated constructs. (E) Co-immunoprecipitation of His-tagged MIA40 and Flag-tagged ABCB8 in mitochondrial fraction in the presence and absence of 100 mM dithiothreitol (DTT). (F) Schematic representation of proposed model in which ABCB8 peptide interacts with the MIA40/ALR protein import machinery prior to insertion into inner mitochondrial membrane through TIM23 complex. EV = empty vector. 5CS = ABCB8 Flag construct with mutations of five conserved cysteines to serine. SE = short exposure. OM = outer mitochondrial membrane. IMS = intermembrane space. IM = inner mitochondrial membrane. WT = wild type.
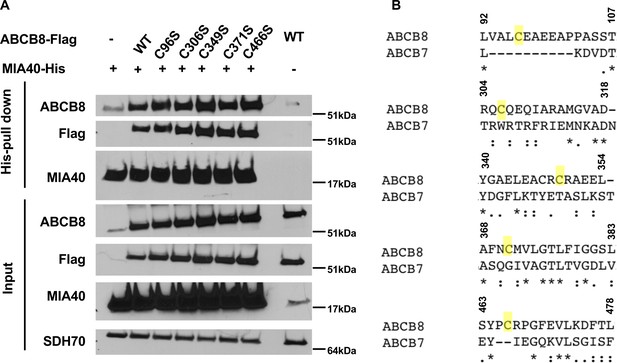
Individual mutation of the five conserved cysteines in ABCB8 minimally impacts its interaction with MIA40.
(A) Western blot results showing co-immunoprecipitation of His-tagged MIA40 and various mutants of Flag-tagged ABCB8 in cells overexpressing indicated constructs. (B) Clustal Omega alignment between human ABCB7 and ABCB8 protein sequences demonstrates lack of conservation of the cysteines. WT = wild type.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Antibody | Rabbit polyclonal anti-GPAT antibody | Dr. Roland Lill, University of Marburg | N/A | WB(1:1000) |
| Antibody | Rabbit polyclonal anti-ABCB8 antibody | Lab generated. For reference, see Ardehali et al 2004. | N/A | WB(1:1000) |
| Antibody | Rabbit polyclonal anti-GAPDH antibody | Abcam | ab9485 | WB(1:6000) |
| Antibody | Rabbit polyclonal anti-ALR antibody | ProteinTech | 11293–1-AP | WB(1:1000-1:4000) |
| Antibody | Rabbit polyclonal anti-ABCB7 antibody | ProteinTech | 11158–1-AP | WB(1:500) |
| Antibody | Rabbit polyclonal anti-alpha tubulin antibody | Abcam | ab4074 | WB(1:2000) |
| Antibody | Mouse monoclonal anti-SDH70 antibody (2E3GC12FB2AE2) | Invitrogen | 459200 | WB(1:7000) |
| Antibody | Mouse monoclonal anti-6x-His antibody | ProteinTech | 66005–1-Ig | WB(1:4000-1:6000) |
| Antibody | Mouse monoclonal anti-GFP antibody | ProteinTech | 66002–1-Ig | WB(1:4000-1:6000) |
| Antibody | Mouse monoclonal anti-ATP5A antibody [15H4C4] - Mitochondrial Marker | Abcam | ab14748 | WB(1:4000) |
| Antibody | Rabbit polyclonal anti-Mia40 antibody | ProteinTech | 21090–1-AP | WB(1:4000) |
| Antibody | Rabbit polyclonal anti-Tim23 antibody | ProteinTech | 11123–1-AP | WB(1:4000) |
| Antibody | Mouse monoclonal Flag M2 antibody | Sigma | F1804 | WB(1:1000-1:4000) |
| Antibody | Mouse monoclonal anti-COX4 antibody [1A12A12] | Abcam | ab110272 | WB(1:1000) |
| Antibody | Rabbit polyclonal anti-beta actin antibody | Cell Signaling | 4967 | WB(1:2000) |
| Antibody | HRP-conjugated donkey polyclonal anti-rabbit IgG antibody | Jackson ImmunoResearch | 711-035-152 | WB(1:2000-1:12000) |
| Antibody | HRP-conjugated donkey polyclonal anti-mouse IgG antibody | Jackson ImmunoResearch | 715-035-150 | WB(1:2000-1:12000) |
| Chemical compound, drug | 55Fe radionuclide. | Perkin Elmer | NEZ043002MC | |
| Chemical compound, drug | RNA-Stat60 | Teltest | Cs-502 | |
| Chemical compound, drug | Glycogen | Life Technologies | AM9510 | |
| Chemical compound, drug | Lipofectamine 2000 | Life Technologies | 11668019 | |
| Chemical compound, drug | Apo-transferrin, human | Sigma | T1147 | |
| Chemical compound, drug | Dharmafect Transfection Reagent I | Dharmacon | T-2001–03 | |
| Chemical compound, drug | Deferoxamine mesylate | Sigma | D9533-1G | (150 µM) |
| Chemical compound, drug | Doxycycline | Fisher Scientific | AC446060050 | (1 µg/mL) |
| Chemical compound, drug | Blasticidin | Life Technologies | R21001 | (10 µg/mL) |
| Chemical compound, drug | Puromycin | Sigma | P8833-25MG | (6 µg/mL) |
| Chemical compound, drug | DMEM, 4.5 g/L glucose with L-glutamine and sodium pyruvate | Corning | 10-013CV | |
| Chemical compound, drug | MEM | Corning | 15-010CV | |
| Chemical compound, drug | Sodium pyruvate solution | Corning | 25–000 CI | |
| Commercial assay, kit | qScript cDNA Synthesis Kit | Quanta | 95047–500 | |
| Commercial assay, kit | PerfeCTa SYBR Green FastMix | Quanta | 95074–05K | |
| Commercial assay, kit | Dual-Luciferase Reporter Assay System | Promega | E1980 | |
| Commercial assay, kit | BCA Protein Assay Kit | Pierce | 23225 | |
| Commercial assay, kit | Amplex Red Xanthine/Xanthine Oxidase Assay Kit | ThermoFisher | A22182 | |
| Commercial assay, kit | Complex I Enzyme Activity Microplate Assay Kit | Abcam | ab109721 | |
| Commercial assay, kit | Complex II Enzyme Activity Microplate Assay Kit | Abcam | ab109908 | |
| Commercial assay, kit | Mitochondria Isolation Kit for Cultured Cells | Thermo Scientific | 89874 | |
| Commercial assay, kit | Aconitase Activity Assay Kit | Abcam | ab109712 | |
| Commercial assay, kit | Seahorse Extracellular FluxPak | Agilent | 102416–100 | |
| Cell line (human) | HEK293 cells | ATCC | CRL-1573 | |
| Transfected construct (human) | HEK293 cells stably expressing rtTA3 | This manuscript. | N/A | Cells were stably infected with rtTA3 lentivirus made from pLenti CMV rtTA3 Blast (w756-1) plasmid |
| Cell line (mouse) | WT and IRP2 KO mouse embryonic fibroblasts | Dr. Tracey Rouault NICHD, NIH | N/A | |
| Transfected construct (mouse) | WT MEFs stably expressing Irebp1 shRNA | This manuscript | N/A | Cells were stably infected with lentivirus expressing mouse Irebp1 shRNA made from rtTA3 lentivirus made from pGIPZ-shIrebp1 |
| Sequence-based reagents | siGenome siRNA against mouse GFER | Dharmacon | MQ-065586-01-0002 | |
| Sequence-based reagents | Negative Control siRNA | Qiagen | 1027310 | |
| Sequence-based reagents | siGenome siRNA against human GFER | Dharmacon | MQ-065586-01-0002 | |
| Sequence-based reagents | siGenome siRNA against mouse IRP1 | Dharmacon | MQ-043541-01-0002 | |
| Sequence-based reagents | siRNA against mouse ALR 3’UTR | Qiagen | SI01011199, SI01011220 | |
| Sequence-based reagents | Primer for qRT-PCR (Supplementary file 1) | This manuscript | N/A | Please see Supplementary file 1 |
| Recombinant DNA reagent | pGIPZ-shIrebp1 | OpenBioSystems | V2LMM_82191 | |
| Recombinant DNA reagent | pRL-TK (Renilla Luciferase) | Promega | E2241 | Luciferase reporter construct expressing Renilla luciferase |
| Recombinant DNA reagent | pMIR-REPORT | Ambion | AM5795 | Empty 3’UTR luciferase reporter construct expressing Firefly luciferase |
| Recombinant DNA reagent | pGL3 Basic Firefly Reporter construct | Promega | E1751 | Promoter luciferase reporter construct expressing Firefly luciferase |
| Recombinant DNA reagent | Mouse Tfrc 3’-UTR luciferase and mutants | This manuscript and Bayeva et al., 2012 | N/A | |
| Recombinant DNA reagent | Mouse Tfrc promoter luciferase construct | This manuscript | N/A | |
| Recombinant DNA reagent | pBI-MCS-EGFP | Addgene | 16542 | |
| Recombinant DNA reagent | pLenti CMV rtTA3 Blast (w756-1) | Addgene | 26429 | |
| Software, algorithm | GraphPad Prism | GraphPad | Version 9 | |
| Software, algorithm | ImageJ | NIH | 1.53 c |
Additional files
-
Supplementary file 1
Primer sequences for quantitative RT PCR.
The protocol for qRT-PCR were described in 'Materials and methods' section.
- https://cdn.elifesciences.org/articles/65158/elife-65158-supp1-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/65158/elife-65158-transrepform-v2.docx





