Mechanically transduced immunosorbent assay to measure protein-protein interactions
Peer review process
This article was accepted for publication as part of eLife's original publishing model.
History
- Version of Record published
- Accepted
- Received
- Preprint posted
Decision letter
-
Raymond E GoldsteinReviewing Editor; University of Cambridge, United Kingdom
-
Cynthia WolbergerSenior Editor; Johns Hopkins University School of Medicine, United States
-
Erika EiserReviewer; University of Cambridge, United Kingdom
In the interests of transparency, eLife publishes the most substantive revision requests and the accompanying author responses.
Acceptance summary:
This article introduces a new experimental method that enables the direct measurements of weak interactions between proteins. It is based on densely attaching one type of protein (antigen) to a ferromagnetic microsphere, which can then bind to another protein that has been attached to a flat surface, and applying an external, rotating magnetic field to force the particle to roll across the surface, where its motion is slowed by binding and unbinding of antigens. The sensitivity of the method suggests that it may prove useful in the study of weak protein-protein interactions.
Decision letter after peer review:
Thank you for sending your article entitled "Mechanically Transduced Immunosorbent Assay To Measure Protein-Protein Interactions" for peer review at eLife. Your article is being evaluated by 2 peer reviewers, and the evaluation is being overseen by a Reviewing Editor and Cynthia Wolberger as the Senior Editor. The following individual involved in reviewer of your submission has agreed to reveal their identity: Erika Eiser (Reviewer #3).
As you can see from the reviews below, the reviewers agree that the method you have described has interesting potential, particularly in the small amount of material needed for the assay. However, there are concerns about the fact that the method itself has been published already some time ago, lessening the novelty of the publication. Focusing on the applications of the method would in principle be acceptable, but the concerns about the dynamic range of the method, and the important need for validation need to be addressed (perhaps requiring new experiments to do so) before we can reach a decision on your paper.
Reviewer #1:
The authors are seeking to develop a method for measuring the binding affinities between biomolecules using low amounts of material. The method described in this manuscript uses a magnetic field to cause a ferromagnetic particle to roll along a surface. When the particle is modified with one biomolecule and the surface is modified with another, there is an increase in friction if the two molecules interact. The degree of interaction is proportional to the amount of friction.
Strengths
– The method requires a minimal amount of material.
– The rolling parameter can be measured for many ferromagnetic particles simultaneously, leading to robust statistical analyses.
Weaknesses
– The method itself has already been described in a paper published by one of the corresponding authors in 2014.
– Insufficient detail is provided on the method and output. Raw data showing that the rolling particle is displaced in the presence of a binding partner on the surface is missing.
– The Kd cannot be directly obtained from the rolling parameter.
– Unlike methods such as surface plasmon resonance and biolayer interferometry, which use a comparably low amount of material, this method requires both binding partners to be affinity-tagged. The tags and the immobilization could obscure native interactions.
– Biomolecular interactions have a wide range of affinities that span over 15 orders of magnitude (1 M to 10-15 M). Yet, the dynamic range of the rolling parameter is only one order of magnitude (0.081 for Kd of 1 M and 0.918 for Kd of 10-15M). As a result, the rolling parameter values cluster together when correlated with Kds on a logarithmic scale. Subtle differences in binding affinity can therefore be lost.
– For any given system, the Kd must be measured using an orthogonal technique in order to establish binding affinities that can be correlated with the measured rolling parameter. More specifically, the rolling parameter values are not generalizable from system to system.
– The schemes presented in Figure 1 are difficult to interpret. Perhaps it would be better show a video of the particles rolling along a surface with no binding partner and another video when there is a binding partner. In the absence of these raw data for each system, the binding measurements are not compelling.
– The authors spend a great deal of time comparing their method to NMR, why? Other measurements such as SPR and BLI would be more appropriate comparisons.
– In general, I found the way in which the manuscript was written to be very misleading. After reading the Intro, I was expecting the rest of the paper would focus on the development of the method. Then, in lines 184-185 on page 6, it becomes clear that the method was already published 7 years ago by one of the corresponding authors. Thus, the paper is less about the method and more about the application.
– Please remove sentences like "It is difficult to overstate how transformative this will be for the study of PPIs." The method is not a direct measurement of binding affinity. The dynamic range of the measurement does not correlate with the range of biomolecular binding affinities. Multiple affinity tags must be used and orthogonal binding measurements must be made in order to interpret the rolling parameter values.
Reviewer #3:
In this article the authors introduce a new experimental method, named mechanically transduced immunosorbent (METRIS) assay, enabling the direct measurements of weak interactions between proteins. Such protein-protein interactions (PPT) are typically very weak (on the order of a fraction of the thermal energy, kBT). This new technique is based on densely attaching one type of protein (antigen) to a ferromagnetic micron bead, which can then bind to another protein, that has been attached to a flat surface. By applying an external, rotating magnetic field, the probe particles will be driven to perform a rolling motion on the flat surface. In order to see any displacement of the probe particle the antigens need to detach on one side and bind on the opposing side in the direction of the motion induced by the magnetic field. This apparent friction informs us on the strength of the PPT. The authors have tested a number of important PPT's – the non-covalent Interactions between Ubiquitin-like Domains and Ube2D1 is one example.
An important strength of the new technique the authors introduce is the sensitivity of their new method – it exceeds that of common methods by far, as they were able to measure protein-protein interactions at much lower concentrations (2 orders of magnitude) than is typically done. This allows a systematic determination of many interaction potentials between the complex configurations of proteins as function of solvent parameters and I guess also as function of temperature, which could be interesting new the denaturation region. In addition, the affinity or attractive interactions between specific proteins or RNA and proteins may change as function of concentration: As the reaction constant or KD value depends directly on the total concentration in the system the up- and down regulation processes of proteins may be better understood. The authors demonstrate that they can achieve such a sensitivity, which is well explained in the results and discussion.
The only point that needs to be addressed is the interpretation of the data. While the systems and references are very well presented and researched it would be helpful to the reader to explain in more detail the actual measurements and in particular their interpretation and relation between the displacements measured and the KD and Gibbs free energy differences.
My main suggestions are of technical nature concerning the interpretation of the results and the underlying theory.
In the present description (line 1567) the authors relate the displacement of the probe bead to a parameter they introduce as rolling parameter RP: zeta = δ x/(pi times the diameter of the bead times the frequency of the oscillating magnetic field). First I would call RP = zeta, or only RP unless I have not understood the difference. However, my main questions is how did the authors derive this expression in equation 1. It is clear that zeta must be dimensionless, hence it is presented as displacement divided a velocity that is multiplied with the actuation time. This velocity is the rolling velocity = angular velocity times the radius of the probe colloid. It is confusing what π is doing there. The authors should double check the equation and also explain to the reader why you introduce this parameter. Moreover, the radius of the colloids sured here should be given.
If I understand right, this zeta or rolling parameter is at the same time normalised against the streptavidin-biotin binding, which is known to be the strongest non-covalent binding in the system. If the probe particle is brought in contact and assuming that there is full coverage of avidin and biotin on the opposing surfaces, the effective contact area will depend on the size of the particle, and the binding strength with be more than 100 kBT. This means there is full traction and now slip or friction between the probe particle and the support surface. But with such strong binding energies, this means the fields applied must be very large and as the particle roller over the surface the biotin or avidin on one of the surfaces must be ripped off. Such behaviour was observed previously in AFM contact experiments, e.g. by the group of Herman Gaup in the 90's. First, the authors need to give details about the reproducibility of these reference measurements, in particular, the applied Forces (e.g. Magnetic field) grafting density and effective number of bonds in the contact area (which also may be influenced by the roughness of the colloidal bead surface) need to be detailed. Secondly, the authors do not discuss or present any calibration of the friction or rolling parameter. It is not necessarily a linear function between zero (no interaction and thus no friction) and one (full sticking/traction like when riding a bicycle).
In this context, it is important to look also into the displacement velocity. In these measurements the friction relies on the fact whether the protein-protein interactions are easily pealed off on one side and possibly re-established on the front of the rolling motion. But this will depend on how fast the rolling motion is. Again, AFM work by Gaup and in particular by Evan Evans (Annu. Rev. Biophys. Biomol. Struct. 2001. 30:105-28 – which should be referenced) showed that the force or interactions measured depend on how fast one pulls on the protein-protein link. Hence, the authors should give much more experimental detail and relation to known measurements. The numbers presented here for the Binding energies and their relation to the rolling parameters must be clarified as well, so that it would be possible for others to reproduce these measurements.
To summarise, the technique and the measured parameters need better elucidation and validation to be useful to others. In Particular friction is not necessarily the right terminology used in these measurements.
https://doi.org/10.7554/eLife.67525.sa1Author response
Reviewer #1:
The authors are seeking to develop a method for measuring the binding affinities between biomolecules using low amounts of material. The method described in this manuscript uses a magnetic field to cause a ferromagnetic particle to roll along a surface. When the particle is modified with one biomolecule and the surface is modified with another, there is an increase in friction if the two molecules interact. The degree of interaction is proportional to the amount of friction.
Strengths
– The method requires a minimal amount of material.
– The rolling parameter can be measured for many ferromagnetic particles simultaneously, leading to robust statistical analyses.
We thank the reviewer for these comments and agree that these are some of the practical strengths of this new technique, in addition to the high resolution, simplicity, and ability to measure weak interactions using minimal amount of material. Furthermore, by immobilizing both binding partners, the assay is not dependent on mass action, and therefore, interactions of any strength can be measured with a small quantity of protein.
Weaknesses
– The method itself has already been described in a paper published by one of the corresponding authors in 2014.
We appreciate the reviewer for pointing out this potential concern with the manuscript. We understand that the previous wording of the manuscript, particularly the following phrasing, might have accidently been misleading:
"Previous studies have demonstrated the utilized of the METRIS assay to measure rolling parameters for a variety of interactions (e.g., Protein A-Fc, histidine-metal, and Protein-PIP lipid interactions) [42-44]."
While the statement could be misinterpreted to mean that the assay was previously developed, the PRL paper was actually focused on trying to achieve artificial chemotactic directed motion based on differences in the effective friction coefficient across a substrate. This was accomplished by creating a gradient in the density of binding sites by passivating a biotin substrate with a droplet of avidin and allowing it to evaporate and leave coffee-ring-like gradients in the density of binding sites. This paper was a physics paper, and we weren't interested or aware of the possibility of deducing binding affinity. It was only after this paper that we recognized the potential of using the concept of effective friction to detect differences in binding affinity by measuring translational displacement. There are also several critical differences between the 2014 PRL paper and this manuscript. The 2014 PRL paper utilized superparamagnetic walkers composed of two 3 μm diameter polystyrene and iron oxide beads which align their magnetic moments in the presence of a rotating magnetic field produced by two coils that were stochastically actuated. We were not focused on measuring the biotin-streptavidin interaction, and in fact, we could not measure the biotin-streptavidin interaction without heavy passivation due to the limited magnetic torque that could be applied due to the small magnetic moment of our superparamagnetic particles. The current technique utilizes a single roller 10μms in diameter, a glass substrate functionalized with avidin, and a 3D Helmholtz coil like apparatus to drive walker motion. Additionally, this current study is exclusively focused on correlating differences in the translational displacement (via introduction of a rolling parameter) to binding affinity and extrapolating an estimated Kd for PPIs. Given these significant differences, we have revised the manuscript to indicate that the foundations for this technique to be applied were previously published. The other citations in this section were to a thesis and a patent, which are not peer reviewed publications, which have some of the experimental optimization, but were not able to measure protein-protein affinities. However, this previously cited works were investigating a primarily physics phenomenon, whereas here we have further developed this technique to measure differences in binding affinity.
We would like to thank the reviewer for pointing out this potential area of confusion, and we have significantly changed the manuscript, spending more time discussing the experimental results of the streptavidin-avidin streptavidin-biotin experiment in Figure 1, since this is the first reporting of that experiment. Overall, this new format really improves the readability and clarity of the manuscript.
– Insufficient detail is provided on the method and output. Raw data showing that the rolling particle is displaced in the presence of a binding partner on the surface is missing.
We appreciate this comment and agree that being able to visualize and work with the raw data would be very helpful to understand the technique and potential applications. To that end, we have made significant changes to the manuscript to illustrate more raw data in the presence of non-binding, extremely strong binding (biotin-streptavidin), and other binding PPI partners with differing affinity.
1. We have made raw videos from some of the conditions we tested available as a supplemental video. These raw videos should be very helpful in visualizing the differences in the mechanical signal observed, translational displacement due to the presence of the binding interaction.
2. For figure 2,3,4, we have included a panel which shows the average distance that a roller translates per actuation.
3. We have also included some animations that illustrate the translational displacement of the beads, in micrometers, for the different interactions to illustrate even small differences in binding affinity can be deduced visually by the differences in the translational displacement of the rollers.
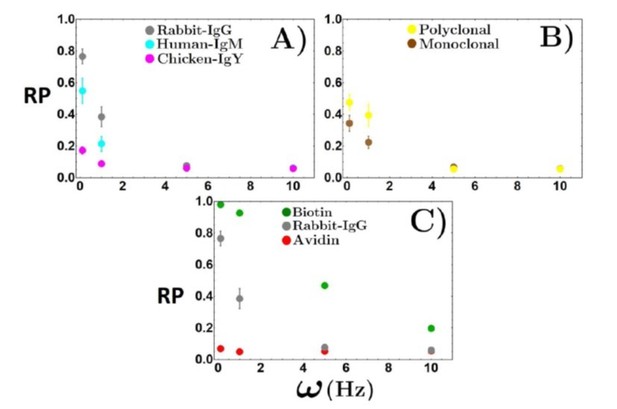
4. We have included histograms of the rolling parameters of the beads for all interactions, and these show clear differences. As seen for all the interactions (shown in Author response image 1), the distributions are Gaussian, but we can see some stark differences in the width of the interactions, particularly the PPI interactions.
5. We have substantially revised figure 1 to include clockwise and counter-clockwise actuation for Streptavidin-Biotin and Streptavidin-Avidin (Figure 1C), More detail about the roller trajectory (Figure 1D), and included the rolling parameter distribution
We believe this additional raw data will help clarify how this technique's mechanical signal output is utilized to differentiate between binding affinities, and we hope that these videos illustrate the novel nature and ease of use of this technique. If there are any other raw data the reviewers think should be included in the manuscript, we are happy to include it.
– The Kd cannot be directly obtained from the rolling parameter.
While it is true that the Kd cannot be directly obtained in isolation at this time, this fact does not take away from the importance of this technique and possible implications. With further work and more data, it will be possible in the future to derive a mathematical model to calculate Kd directly or develop a fundamental theoretical model that will enable one to directly measure binding affinity from the rolling parameter value. Nonetheless, this is the first report of a technique that can be utilized to deduce differences in binding affinity from a mechanical signal, the translational displacement. Also, even without a Kd based on a standard curve, related molecules' binding strengths can still be measured with reasonable accuracy using the ΔΔG method as described in the manuscript.
– Unlike methods such as surface plasmon resonance and biolayer interferometry, which use a comparably low amount of material, this method requires both binding partners to be affinity-tagged. The tags and the immobilization could obscure native interactions.
We agree that METRIS requires both proteins be immobilized with high-affinity tags, which could obstruct some biological interactions. However, this issue is not unique to METRIS. Both SPR and BLI have one protein immobilized, usually through an affinity tag (not necessarily a high-affinity tag) or surface adsorption, which can also block interactions. In this study, we present two methods of tagging the protein at different locations, through a BirA tag and with maleamide-biotin, which in this study showed comparable RPs, (Supplemental Figure 3) so this gives the user flexibility in how to achieve immobilization, and there are many other possible immobilization strategies that we are testing. Importantly, this also indicates that we can see similar results from both approaches. We also have added some additional text in the Discussion section of the manuscript about this. For the interactions we measured in our study, we have no indication that tagging blocked any of the interactions, given the strong correlation of our data to other studies that measured these protein-protein interactions.
Additionally, immobilizing both proteins affords advantages to SPR and BLI because we use significantly less material than either BLI and SPR. However, in BLI and SPR, the soluble analyte is still dependent on mass action and thus can require high concentrations of protein to measure weaker affinities, still consuming a lot of materials and also leading to potential thermodynamic non-ideality of the soluble analyte. We have a more detailed comparison between SPR/BLI and METRIS below and directly compare BLI and METRIS for low-affinity binders.
– Biomolecular interactions have a wide range of affinities that span over 15 orders of magnitude (1 M to 10-15 M). Yet, the dynamic range of the rolling parameter is only one order of magnitude (0.081 for Kd of 1 M and 0.918 for Kd of 10-15M). As a result, the rolling parameter values cluster together when correlated with Kds on a logarithmic scale. Subtle differences in binding affinity can therefore be lost.
We believe that this is one of the key properties that makes the METRIS technique valuable as a bioassay platform, and can clarify this point. While the rolling parameter only spans one order of magnitude, our manuscript and other preliminary data illustrate that METRIS can measure affinities spanning 15 orders of magnitude, but is also sensitive enough to measure differences as low as 2-fold like we observed when measuring the Orc1-BAH interaction with methylated peptides (See Figure 3). Thus, we do not believe we are losing subtle differences in binding. This accuracy is due to the very small uncertainty or error that we report and this may be because of some confusion with the definition of the rolling parameter, which we have clarified in the manuscript in response to one of the comments by Reviewer #3 (see below). The rolling parameter is the ratio of the observed, experimental translational displacement of the roller, Δx, to the maximum theoretical displacement of a rolling sphere. The maximum displacement of a rolling sphere that hinges perfectly on a single point will simply be the circumference of the sphere (πD, where D is the diameter of the sphere typically 10 μm for our rollers, but we explicitly measure the diameter for every roller) times the number of rotations which is governed by the rotational frequency of the magnetic field, ω, which is 1Hz for this manuscript, times the actuation period in seconds, τ, which is 5s for this experiment. Thus, the maximum theoretical displacement is simply πDωτ. A rolling parameter of 1 would equal 157 µm, where a rolling parameter of 0.082, the null binding measurement in this manuscript, would correspond to a distance of 13 µm. We can measure the translational displacement and the diameter of the particle with high accuracy using a custom-written Mathematica analysis script in conjunction with Able Particle tracker, and there is virtually no uncertainty for the actuation period and the rotational frequency of the magnetic field in this technique. The recorded images have a resolution of approximately 1μm per pixel, and our image analysis software has subpixel resolution so we can measure subtle differences in displacement on the order of 100s of nanometers. Couple this with the fact that we measure on average 20-30 rollers. Each roller rolls approximately 36 times, 5 seconds each roll, meaning we roll the entire sphere 5 complete rotations, we have incredible statistical averaging. This METRIS technique and probes are extremely sensitive and can distinguish forces on the order of 10s of femtonewtons, as previously reported (PRL 2014), and we have added this to the main text of the manuscript.
This is also somewhat akin to thermodynamics, while biological interactions span 15 orders of magnitude, the ΔG/mol between the bound and unbound states only span ~2 orders of magnitude (~20 kcal/mol). Moreover, we would like to highlight that for these interactions, which are very weak and can be impossible to measure with other traditional techniques like BLI, METRIS is indeed sensitive enough to distinguish and measure a wide range of binding affinities with high resolution.
– For any given system, the Kd must be measured using an orthogonal technique in order to establish binding affinities that can be correlated with the measured rolling parameter. More specifically, the rolling parameter values are not generalizable from system to system.
We agree with the reviewer's comment. As we acknowledge in the text, this is one of the limitations of METRIS at this point, as we do not have a robust theoretical model that allows for the direct calculation of binding affinity from the rolling parameter. We are continuing to work on this aspect of the project. Still, at this point, even an empirical and orthogonal calculation of binding affinities using METRIS is valuable at this point for the following reasons:
(1) This technique does not utilize a large amount of material, so even at this stage, METRIS can be very useful as a quick diagnostic technique for binding partners that are difficult to produce in large quantities.
(2) METRIS is relatively simple to perform, and one can quickly obtain at the very least a qualitative comparison between several different binders within a family of binders/interactions.
(3) METRIS is well suited to measure very weak interactions, and as we demonstrated in this paper – even when studying characterized systems, we can still produce biological insights into interactions. Furthermore, even without an empirically determined Kd using an orthogonal system, you can still derive relatively accurate ΔΔG using the standard curve from our aggregate data for a series of family members.
(4) If a user establishes a baseline, the results can be highly quantitative compared to other methods for the reasons described.
(5) METRIS can still be very useful even as a qualitative bioassay screener, especially as we improve throughput.
Additionally, similar calibration curves are required to get quantitative results from ELISA measurements; however, our sensitivity and reproducibility is well beyond the ability of any quantitative ELISA with significantly less material utilized.
– The schemes presented in Figure 1 are difficult to interpret. Perhaps it would be better show a video of the particles rolling along a surface with no binding partner and another video when there is a binding partner. In the absence of these raw data for each system, the binding measurements are not compelling.
We really appreciate this comment from the reviewer, and we have made the recommended changes in Figure 1 and included more supplemental videos and supplemental figures.
– The authors spend a great deal of time comparing their method to NMR, why? Other measurements such as SPR and BLI would be more appropriate comparisons.
In the manuscript, we focused on NMR because it is the standard for measuring very weak interactions. However, the reviewer is correct that SPR and BLI can use small quantities of materials and may be more similar to METRIS, given that one of the interacting partners is immobilized. We have added some text to the manuscript to address this point.
Additionally, METRIS has significant advantages to SPR and BLI, especially when examining very weak binders. Both SPR and BLI are dependent on mass action, so the soluble analyte still must be above the Kd, and for weak binding, this still requires a significant amount of protein. For example, in another unpublished study where we used METRIS in combination with BLI to measure weak binding to ubiquitin for three mutants, we needed 5.5 mLs of ubiquitin at 5mM (~200 mg) to measure affinities in the 10-4-10-3 range (see Author response image 2). For very weak binders, it is not possible to measure the Kd. When fitting BLI data, it is critical to establish the endpoint, i.e., saturate the binding curve, and one often needs very high concentrations of proteins. With BLI, we could not accurately measure the Kd values for the weakest binders (compare results from METRIS and BLI for mutants 2 and 3 in Author response image 2C) because we did not saturate the binding curves, and at higher ubiquitin concentrations, thermodynamic non-ideality occurs (Author response image 2C and D). With METRIS, we could accurately measure these affinities with 30 pmols of protein total, and we could even measure the affinity for mutant 3, which we couldn't fit with BLI (Author response image 2 A-C). For these BLI measurements, we needed 23 µmols of material – ~100,000 times more material than we needed for METRIS. For most proteins, it is impossible to obtain that much protein at that high of a concentration. Moreover, the BLI signal is noisy and is dependent on accurately measuring the ubiquitin concentration (Figure 2D).
Additionally, for BLI and SPR, it is difficult to distinguish between background binding to the chip when studying weak interactions, where for METRIS, we have good sensitivity between the null and weak binding range.
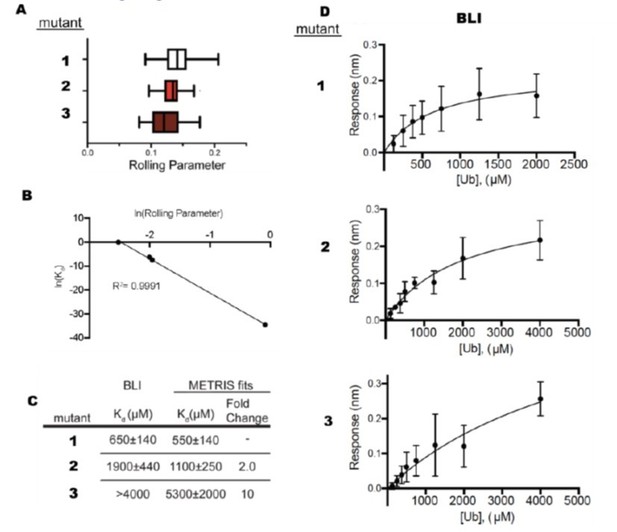
Comparison Between BLI and METRIS.
(A) Rolling parameter measurements for 3 mutants that bind ubiquitin. (B) Fit of METRIS to Kd values. (C) Kdvalues obtained from METRIS and BLI. (D) BLI data for each of the mutants.
For SPR, sometimes you can derive binding values below the Kd, but in the scenarios, only a fraction of the chip is bound, and depending on your signal, you may not be able to detect binding. Also, the signal for BLI and SPR is dependent on a significant size change (i.e., change the refractive index), so for some binding experiments, like a peptide/small molecule to a protein, the signal can be very small. Given these issues, we think that METRIS has significant advantages to BLI and SPR. We have incorporated this discussion into the manuscript and highlighted the utility of METRIS as a new technique.
– In general, I found the way in which the manuscript was written to be very misleading. After reading the Intro, I was expecting the rest of the paper would focus on the development of the method. Then, in lines 184-185 on page 6, it becomes clear that the method was already published 7 years ago by one of the corresponding authors. Thus, the paper is less about the method and more about the application.
We appreciate the reviewer's comment, and this is due to some confusion about how the manuscript was written. Above, we described the key differences between the 2014 PRL paper, and we have provided more background about how this method is different from previously described techniques. We have also made changes to the manuscript to avoid this potential source of confusion.
– Please remove sentences like "It is difficult to overstate how transformative this will be for the study of PPIs." The method is not a direct measurement of binding affinity. The dynamic range of the measurement does not correlate with the range of biomolecular binding affinities. Multiple affinity tags must be used and orthogonal binding measurements must be made in order to interpret the rolling parameter values.
We have removed the claims. We were attempting to highlight the importance of the METRIS technique but appreciate this comment and have made subsequent changes to the manuscript to ensure the claims made are reasonable, accurate, and not overstated
Reviewer #3:
In this article the authors introduce a new experimental method, named mechanically transduced immunosorbent (METRIS) assay, enabling the direct measurements of weak interactions between proteins. Such protein-protein interactions (PPT) are typically very weak (on the order of a fraction of the thermal energy, kBT). This new technique is based on densely attaching one type of protein (antigen) to a ferromagnetic micron bead, which can then bind to another protein, that has been attached to a flat surface. By applying an external, rotating magnetic field, the probe particles will be driven to perform a rolling motion on the flat surface. In order to see any displacement of the probe particle the antigens need to detach on one side and bind on the opposing side in the direction of the motion induced by the magnetic field. This apparent friction informs us on the strength of the PPT. The authors have tested a number of important PPT's – the non-covalent Interactions between Ubiquitin-like Domains and Ube2D1 is one example.
An important strength of the new technique the authors introduce is the sensitivity of their new method – it exceeds that of common methods by far, as they were able to measure protein-protein interactions at much lower concentrations (2 orders of magnitude) than is typically done. This allows a systematic determination of many interaction potentials between the complex configurations of proteins as function of solvent parameters and I guess also as function of temperature, which could be interesting new the denaturation region. In addition, the affinity or attractive interactions between specific proteins or RNA and proteins may change as function of concentration: As the reaction constant or KD value depends directly on the total concentration in the system the up- and down regulation processes of proteins may be better understood. The authors demonstrate that they can achieve such a sensitivity, which is well explained in the results and discussion.
We truly appreciate the comments made by the reviewer and appreciate the insight into the METRIS technique. We agree that METRIS will allow scientists to study interactions in different ways than other methods and we are excited to test some of the experiments that the reviewer mentions above.
The only point that needs to be addressed is the interpretation of the data. While the systems and references are very well presented and researched it would be helpful to the reader to explain in more detail the actual measurements and in particular their interpretation and relation between the displacements measured and the KD and Gibbs free energy differences.
My main suggestions are of technical nature concerning the interpretation of the results and the underlying theory.
In the present description (line 1567) the authors relate the displacement of the probe bead to a parameter they introduce as rolling parameter RP: zeta = δ x/(pi times the diameter of the bead times the frequency of the oscillating magnetic field). First I would call RP = zeta, or only RP unless I have not understood the difference. However, my main questions is how did the authors derive this expression in equation 1. It is clear that zeta must be dimensionless, hence it is presented as displacement divided a velocity that is multiplied with the actuation time. This velocity is the rolling velocity = angular velocity times the radius of the probe colloid. It is confusing what π is doing there. The authors should double check the equation and also explain to the reader why you introduce this parameter. Moreover, the radius of the colloids sured here should be given.
This is a great comment. We have made changes in the text that correct the description of the rolling parameter, which initially and mistakenly stated normalization by the theoretical velocity, which was not correct. We have also changed the labeling of the zeta to the RP. The RP is the ratio of the actual translational displacement of the roller, Δx, divided by the maximum theoretical displacement of a rolling sphere on a substrate. One can imagine that the maximum distance a sphere can roll on a substrate would be to hinge on a single point and then roll perfectly on this point, whereby the sphere would translate a distance equivalent to the sphere's circumference. In this scenario, our sphere or roller undergoes 5 complete rolls per actuation period as we kept the rotational frequency of the magnetic field constant at 1Hz and the time of rolling was 5 seconds, so the roller undergoes 5 complete rotations per roll. So, the maximum theoretical translational displacement of a sphere would be the circumference of our sphere π D, multiplied by the actuation time τ, and the rotational frequency of the magnetic field ω. We introduce this parameter because, as the reviewer points out, the translational displacement of the roller depends on several factors beyond that of binding partners, and those include D, τ, and ω. We introduced the rolling parameter given this dimensionless number allows us to provide the reader with an apples-to-apples comparison of how the translational displacement varies across each PPI. Additionally, and pragmatically, although our particle diameters are nominally approximately 10 μm, they tend to vary slightly, but these differences affect the rolling parameter (which is why we explicitly measure each roller diameter) and thus the extrapolated binding affinity. The introduction and creation of the rolling parameter is essential to provide an accurate comparison of the effective friction induced by binding and, therefore, the translational displacement of the rollers. Without such a parameter, the extrapolation to a binding affinity would be impossible or, at the very least, much less robust.
If I understand right, this zeta or rolling parameter is at the same time normalised against the streptavidin-biotin binding, which is known to be the strongest non-covalent binding in the system. If the probe particle is brought in contact and assuming that there is full coverage of avidin and biotin on the opposing surfaces, the effective contact area will depend on the size of the particle, and the binding strength with be more than 100 kBT. This means there is full traction and now slip or friction between the probe particle and the support surface. But with such strong binding energies, this means the fields applied must be very large and as the particle roller over the surface the biotin or avidin on one of the surfaces must be ripped off. Such behaviour was observed previously in AFM contact experiments, e.g. by the group of Herman Gaup in the 90's. First, the authors need to give details about the reproducibility of these reference measurements, in particular, the applied Forces (e.g. Magnetic field) grafting density and effective number of bonds in the contact area (which also may be influenced by the roughness of the colloidal bead surface) need to be detailed. Secondly, the authors do not discuss or present any calibration of the friction or rolling parameter. It is not necessarily a linear function between zero (no interaction and thus no friction) and one (full sticking/traction like when riding a bicycle).
We appreciate the reviewer’s keen insight. Indeed, we calibrate our METRIS platform by looking at the RP of a non-interacting binding pair and that of the strongest non-covalent interaction biotin and streptavidin. As we mention in the manuscript, we fully coat our probe and substrate with more than 50X the theoretical binding capacity to ensure a fully coated substrate and particle. We do this to ensure that the density of binders remains as constant as possible, but as the diameter of the particles is slightly different, the number of binders will change slightly, hence why we also introduce the rolling parameter to account for these differences. The density of biotin-binding sites per roller is on the order of 6 x 108 biotin molecules per roller, which corresponds to a density of biotin-binding sites on the order of 5 x 1010 binding sites per mm2, which is of similar density to the density of streptavidin binding sites is reported in the manuscript. The actual footprint of the sphere on the substrate is not a trivial calculation, but assuming that footprint is 1% of the projected area, that would result in approximately 105 potential binding sites, but due to the size of tagged PPI, this number of binding interactions is likely less. Regardless to break these interactions will require an applied magnetic torque, τ, which is defined as the m x B where m is the magnetic moment, and B is the applied magnetic field strength. As described in the manuscript, the applied magnetic field strength is approximately 10mT, and the magnetic moment is on the order of 10-11 A M-2 this would produce a torque large enough to break the biotin-streptavidin bond. In our previous study in 2014, we studied a system composed of walking streptavidin-coated superparamagnetic dimers, 3 μm in diameter each, the biotin-streptavidin bond was not able to be broken without heavily passivating the biotin substrate with avidin particles and coating the walkers with biotin-PEG. This combination reduced the number of biotin-binding sites, and due to the PEG brush, the biotin-streptavidin interaction strength was reduced because compressing the polymer PEG brush is entropically unfavorable. The superparamagnetic particles were unable to break the biotin-streptavidin interaction even when using approximately the same magnetic field strength because the magnetic moment of the walkers is more than an order of magnitude less than that of the ferromagnetic rollers. Thus, the torque is more than an order of magnitude less.
However, despite this increase in the magnetic torque, we are not stripping the biotin from the substrate in this system. This is simply not possible in the system that we are using because the biotin or the streptavidin is covalently linked to the glass substrate, and we are not breaking covalent bonds, just breaking the biotin-streptavidin interaction. If the biotin or avidin was being ripped off from the substrate and remained stuck on the roller, there is no way for these ligands to re-attach to the substrate. Thus, if this is indeed the mechanism when the rollers are actuated in the reverse direction, we should see a drastic decrease in the rolling parameter as one of the binding partners has been removed. However, this is not observed, as seen in Author response image 3 and in our supplemental movies. This is distinct from scenario where binding partners are being ripped from the surface. Take for example the data from a pre-print of a system consisting of a supported lipid bilayer that has biotinylated lipid bilayers. The supported lipid bilayer is not covalently attached to the substrate and in here the binding affinity of the biotin-streptavidin interaction is enough to rip the lipids from the substrate and you can see in Author response image 3B as the lipids are ripped from the bilayer the rolling parameter decreases as the roller continues to roll, thus illustrating that in this system in the manuscript we are not ripping off binding partners from the substrate.
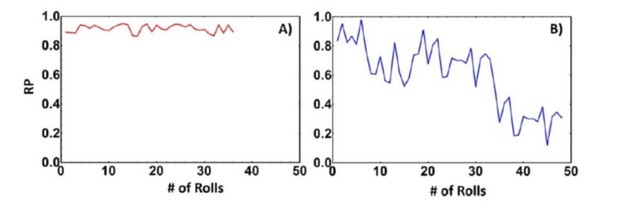
Rolling parameter of biotin-streptavidin interaction on avidin substrate (A) and a biotinylated supported lipid bilayer (B).
Finally, in terms of calibrating the RP to friction, we clarify that the RP gives us an indication of the effective friction induced by binding interactions, and we correlate RP to the binding affinity. We appreciate the reviewer for this comment and have updated the manuscript to make this distinction more clear. We do not assume that the RP and friction scale linearly and have changed the manuscript. We also want to thank the reviewer for the comments and hope that these experimental details the distributions of the rolling parameter values given in Figure 1 illustrate the robust and reproducible nature of the technique.
In this context, it is important to look also into the displacement velocity. In these measurements the friction relies on the fact whether the protein-protein interactions are easily pealed off on one side and possibly re-established on the front of the rolling motion. But this will depend on how fast the rolling motion is. Again, AFM work by Gaup and in particular by Evan Evans (Annu. Rev. Biophys. Biomol. Struct. 2001. 30:105-28 – which should be referenced) showed that the force or interactions measured depend on how fast one pulls on the protein-protein link. Hence, the authors should give much more experimental detail and relation to known measurements. The numbers presented here for the Binding energies and their relation to the rolling parameters must be clarified as well, so that it would be possible for others to reproduce these measurements.
We thank the reviewer for pointing us to the excellent manuscript by Evan Evans and also pointing us to relevant papers from the field of force spectroscopy. We have added these citations to the manuscript and have also added a section in the discussion about force spectroscopy and compared it to METRIS. We would also like to point the reviewer to the previous comments regarding the interactions being peeled off the substrate. In terms of the protein-protein interactions begin broken, as the roller rolls, this does indeed depend on the rotational frequency of the magnetic field. In this study, the rotational frequency is kept fixed at 1Hz as this has been determined as an appropriately slow rotational frequency to allow for binding events. If one increases the magnetic field's rotational frequency, there will be less time for binding, and the effective friction will decrease, as you can see in Author response image 4. Here we find that at 5Hz, most interactions are no longer measurable as the bead is rotating too fast, and binding cannot occur because there is not sufficient time for binding. In fact, only the biotin-streptavidin interaction is observed at 10Hz, but as you can see, the rolling parameter has decreased drastically. We hope this new information makes it easier for comparisons.
To summarise, the technique and the measured parameters need better elucidation and validation to be useful to others. In Particular friction is not necessarily the right terminology used in these measurements.
We have made the appropriate changes to the manuscript and included the information suggested above. We have also changed the term friction to effective friction induced by binding to be more accurate. However, the reviewer is correct that the particle is essentially slipping along and what we are measuring is how effectively the rotational torque is converted to rotational motion. Thus, we have simplified this concept and describe it as effective friction. Again, we thank the reviewer for these comments, which have improved the quality of the manuscript.
https://doi.org/10.7554/eLife.67525.sa2