Mutational sources of trans-regulatory variation affecting gene expression in Saccharomyces cerevisiae
Figures
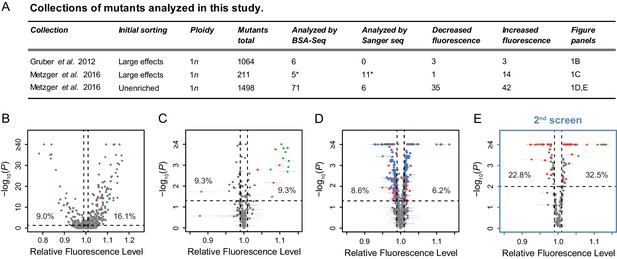
Mutant strains analyzed with altered expression of a PTDH3-YFP reporter gene.
(A) Summary of the three previously published collections of S. cerevisiae mutants obtained by ethyl methanesulfonate (EMS) mutagenesis of a haploid strain expressing a yellow fluorescent protein (YFP) under control of the TDH3 promoter. *One mutant is included in both columns because it was analyzed both by BSA-Seq and Sanger sequencing. (B–D) Previously published fluorescence levels (x-axis) and statistical significance of the difference in median fluorescence between each mutant and the un-mutagenized progenitor strain (y-axis) are shown for mutants analyzed in (B) Gruber et al., 2012 and (C,D) Metzger et al., 2016. (B) Collection of 1064 mutants from Gruber et al., 2012 enriched for mutations causing large fluorescence changes. p-values were computed using Z-tests in this study, based on one measure of fluorescence for each mutant and 30 measures of fluorescence for the progenitor strain. (C) Collection of 211 mutants from Metzger et al., 2016 enriched for mutations causing large fluorescence changes. (D) Collection of 1498 mutants from Metzger et al., 2016 obtained irrespective of their fluorescence levels (unenriched mutants). (E) A new fluorescence dataset for 197 unenriched mutants from Metzger et al., 2016 (blue in panel D) that were reanalyzed in a 2nd screen as part of this study. (C–E) Four replicate populations were analyzed for each mutant. Error bars show 95% confidence intervals of fluorescence levels measured among these replicates. p-values were obtained using the permutation tests described in Methods. (B–E) Mutants analyzed by BSA-Seq are highlighted in red. All of these mutants showed fluorescence changes greater than 0.01 (vertical dotted lines) and p-value below 0.05 (horizontal dotted lines); percentages of all mutants that met these selection criteria in each collection are also shown. Mutants selected for Sanger sequencing of the ADE4, ADE5, and/or ADE6 candidate genes are highlighted in green. The mutant analyzed with both BSA-seq and Sanger sequencing is both red and green in panel (C). Two mutants selected for Sanger sequencing of the ADE2 gene are highlighted in purple, one in (D) and one in (E).

Diagram showing the number of mutant strains and mutations considered at each step of the study.
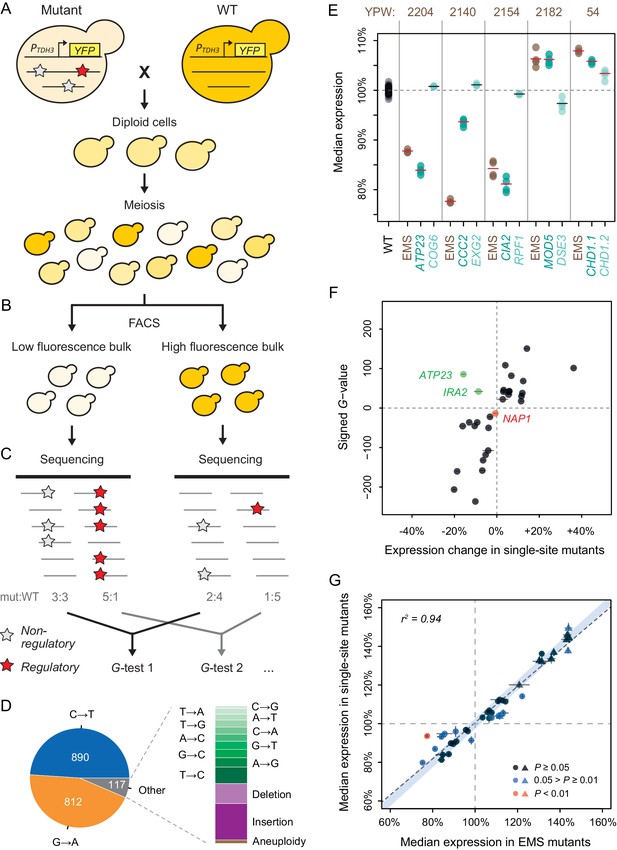
Genetic mapping and functional testing of trans-regulatory mutations affecting PTDH3-YFP expression.
(A–C) Overview of the BSA-Seq approach. (A) Crossing scheme used to map mutations in each EMS mutant strain by crossing to an un-mutagenized strain expressing PTDH3-YFP. Stars indicate hypothetical mutations. (B) Isolation of two bulks of haploid segregants with high and low fluorescence levels (see Methods). (C) Estimation of allele frequencies in each bulk using high-throughput sequencing. A mutation without effect on fluorescence is found at similar frequencies in the two bulks (white stars). A mutation affecting fluorescence or genetically linked to a mutation affecting fluorescence is found at different frequencies between the two bulks (red stars). (D) Type of mutations identified in BSA-Seq data for the 76 mutants from Metzger et al., 2016. (E) Median expression of PTDH3-YFP is shown for the wild-type (WT) progenitor strain (black), for five EMS mutants (brown) with two linked mutations associated with fluorescence in BSA-Seq data and for 10 single-site mutants (turquoise) carrying one of the two linked mutations in the five EMS mutants. Single-site mutants are grouped in pairs next to the EMS mutant carrying the same mutations and are named after the gene that they affect. Expression levels are expressed relative to the wild-type progenitor strain. For each strain, dots represent the median expression measured for each replicate population and tick marks represent the mean of median expression from replicate populations. (F) Effects of mutations associated with fluorescence in BSA-Seq experiments tested in single-site mutants. X-axis: Effect of each mutation on expression measured in a single site mutant and relative to the wild-type progenitor strain. Error bars are 95% confidence intervals obtained from at least four replicate populations. Y-axis: G statistics of the tests used to compare the frequencies of each mutation between the two bulks in BSA-Seq experiments, with a negative sign if the mutation was more frequent in the low fluorescence bulk and a positive sign if the mutation was more frequent in the high fluorescence bulk. One single-site mutant (NAP1, red) showed no significant change in expression relative to the wild-type progenitor strain (t-test, p-value > 0.05); the mutation it carries is therefore considered to be a false positive in the BSA-seq data. For two other single-site mutants (ATP23 and IRA2, green), the expression changes were not in the same direction as predicted by the signed G-values. (G) PTDH3-YFP expression levels in single-site mutants and in EMS mutants sharing the same mutation. Data points represent median expression levels of 40 EMS mutants (x-axis) and 40 single-site mutants (y-axis) measured by flow cytometry in four replicate populations. Circles: mutations identified by BSA-Seq. Triangles: mutations identified by sequencing candidate genes. Error bars: 95% confidence intervals of expression levels obtained from replicate populations. Data points are colored based on the p-values of permutation tests used to assess the statistical significance of expression differences between each single site mutant and the EMS mutant carrying the same mutation (see Figure 2—figure supplement 5 for details). The light blue area represents the 95% confidence interval of expression differences between genetically identical samples across the whole range of median expression values. This confidence interval was calculated from a null distribution described in Figure 2—figure supplement 5A. (E–G) Expression levels are expressed on a scale linearly related to YFP mRNA levels and relative to the median expression of the wild-type progenitor strain (see Materials and methods).

Number of mutations per strain identified from BSA-Seq data.
Data from 76 EMS mutants from Metzger et al., 2016 are shown. Vertical dotted line: mean number of mutations per strain (23.9). Blue dots and line: Poisson distribution with λ = 23.9 and k = 76 representing the expected numbers of mutations per line if mutations had the same probability of occurring in all mutant lines.

Magnitude of expression changes in EMS mutants depending on the number of mutations associated with fluorescence in BSA-Seq experiments.
Individual data points represent absolute differences between the median expression levels of EMS mutants and of the un-mutagenized progenitor strain averaged among four replicate populations. Mutations that were associated with fluorescence only because of genetic linkage (i.e. without additional evidence of affecting expression) were not counted (see Supplementary file 4). Blue dots: mutants with decreased expression relative to the progenitor strain. Red dots: mutants with increased expression relative to the progenitor strain. Using Mann-Whitney-Wilcoxon tests, the magnitude of expression changes was found to be significantly lower for mutants without any mutation associated with fluorescence than for mutants with 1 (p = 5.3 x 10−5) or 2 (p = 0.018) mutations associated with fluorescence.

Relationship between the number of mutations per EMS mutant strain and the absolute expression change relative to the progenitor strain.
This relationship is shown for EMS mutants without any mutation associated with fluorescence in BSA-Seq data (green dots and green regression line) as well as for EMS mutants with at least one mutation associated with fluorescence in BSA-Seq data (gray dots and gray regression line). Mutations that were associated with fluorescence only because of genetic linkage and without other evidence of affecting expression were excluded (see Supplementary file 4). F-tests were used to assess the statistical significance of linear regressions. A significant relationship was observed between the number of mutations per mutant strain and the absolute expression change only when no mutation was associated with fluorescence (r2 = 0.127, p-value = 0.03). This observation supports the hypothesis that several mutations with small effects could collectively contribute to the expression change observed in mutants for which no mutation was associated with fluorescence. The small effects of these mutations would explain why they were not associated with fluorescence in the BSA-Seq analyses.

Effects of individual mutations in purine biosynthesis genes on YFP expression levels differ among promoters.
Each dot indicates the median fluorescence level of at least 5 x 104 cells for each genotype averaged across three experimental replicates. Error bars represent median absolute deviation across replicates. Dots are grouped along the x-axis based on the yeast promoter used to drive YFP expression (PGPD1, PRNR1, PSTM1, and PTDH3), with ‘None’ corresponding to the autofluorescence measured in a strain without a fluorescent reporter gene. The color of each dot indicates which mutation was introduced in one of the genes involved in de novo purine synthesis (ADE2, ADE5 or ADE6), with the specific mutation introduced indicated in the key. The goal of this experiment was to determine whether the regulatory mutations identified in purine synthesis genes altered PTDH3-YFP expression at the transcriptional or post-transcriptional level. If the mutations acted post-transcriptionally, their effect on fluorescence level should be the same among strains with different promoters driving YFP expression because they all produce the same YFP transcript. However, we observed that the mutations in purine synthesis genes increased fluorescence level when YFP expression was driven by the TDH3 or the GPD1 promoter but not when YFP expression was driven by the RNR1 or the STM1 promoter, indicating that the effects of these mutations on YFP expression were promoter specific.

Factors contributing to expression differences observed between EMS and single-site mutants.
(A) Distribution of absolute expression differences observed between EMS and single-site mutants (bars). To assess the statistical significance of these expression differences, we estimated the magnitude of expression differences expected to arise by chance between genetically identical strains grown at different positions of a 96-well plate (red line). This null distribution was obtained from the differences in expression measured for 10,440 pairs of the un-mutagenized progenitor strain grown at different well positions in four replicate populations. We next randomly permuted 105 times the expression values between (i) each pair of EMS and single-site mutants and (ii) random pairs of the progenitor strain to calculate the one-sided p-value for each pair of mutants (i.e. the proportion of randomized expression differences greater than the observed expression difference). After Benjamini-Hochberg correction for multiple testing, we found that the expression difference between the single-site mutant and the EMS mutant carrying the same mutation was statistically significant (adjusted p-value < 0.05) for 14 out of the 40 pairs of mutants (35%, red and blue bars), but highly significant (adjusted p-value < 0.01) for only one pair (2.5%, red bar). Because mutant strains were exposed to the same micro-environmental and technical variation as the control samples used to establish the null distribution, these sources of variation are unlikely to explain the significant differences of expression observed between EMS and single-site mutants. Panels (B–F) test three other hypotheses to explain expression differences observed between single-site and EMS mutants. (B) Hypothesis 1: expression differences between EMS and single-site mutants are explained by differences in expression noise (i.e. the variability of expression observed among genetically identical cells grown in the same environment) among mutants. To test this hypothesis, we compared the expression noise measured by flow cytometry for each EMS mutant (x-axis) to the absolute difference of median expression levels between this EMS mutant and the corresponding single-site mutant (y-axis). We observed no significant correlation between the two parameters (r = 0.06, p-value = 0.71), indicating that expression noise is unlikely to explain expression differences between EMS and single-site mutants. Expression noise was calculated for each sample as the standard deviation of expression among cells divided by the median expression and it is reported as the average value among four replicate populations relative to the expression noise of the wild-type progenitor strain. Dot colors: p-values as shown in panel A. Dot shapes: circles represent mutations identified by BSA-seq; triangles represent mutations identified by sequencing candidate genes. Error bars: 95% confidence intervals calculated from four replicate populations. (C–D) Hypothesis 2: expression differences between EMS and single-site mutants are explained by additional mutations present in the EMS mutants. (C) Testing effects of additional mutations associated with fluorescence: boxplot comparing the magnitude of expression differences when only one mutation was associated with fluorescence and when more than one mutation was associated with fluorescence in BSA-Seq experiments. The fact that no statistical difference was observed between the two classes (Mann-Whitney-Wilcoxon test, p = 0.192) suggests that expression differences between EMS and single-site mutants were not likely to be caused by additional mutations associated with fluorescence in the BSA-Seq data. (D) Testing effects of additional mutations with statistical support for an association with fluorescence below the significance threshold. Expression difference between EMS and single-site mutants (x-axis) was compared to the highest G-value that was below our significance threshold for considering a mutation to be associated with fluorescence in the BSA-Seq data from each mutant (y-axis). A significant correlation was observed between the two parameters (Pearson’s r = 0.48; p = 0.02), suggesting that some mutations with associations below our detection threshold in the BSA-Seq experiments might contribute to expression differences observed between EMS and single-site mutants. Dots represent individual pairs of EMS and single-site mutants sharing the same mutation (with random jitter). The red line represents the linear regression of the y-axis parameter on the x-axis parameter. (E–F) Hypothesis 3: expression differences between EMS and single-site mutants are explained by secondary mutation(s) or epigenetic changes that occurred during construction of single-site mutants. To test this hypothesis, we isolated two independent clones for 26 single-site mutants after transformation of the progenitor strain and measured the expression difference between the two clones. (E) A positive correlation was observed between the expression difference between EMS and single-site mutants (x-axis) and the expression difference between the two independent clones for each single-site mutant (y-axis). This positive correlation indicates that mutations with larger expression differences between the single-site and EMS mutants tended to also show larger expression differences between independent transformants. Dot colors: p-values as shown in panel A. (F) Boxplot also showed that the average magnitude of expression differences between independent clones was higher for single site mutants with a statistically significant expression difference between the single-site and EMS mutant sharing the same mutation (Mann-Whitney-Wilcoxon test, p = 0.008). Results from E and F suggest that secondary mutation(s) and/or epigenetic changes that unintentionally occurred in some of the single-site mutant clones likely contributed to expression differences between some EMS and single-site mutants. It is important to emphasize, however, that these expression differences were small in magnitude and that overall the expression level of single-site mutants was strongly correlated with the expression level of EMS mutants (Figure 3).
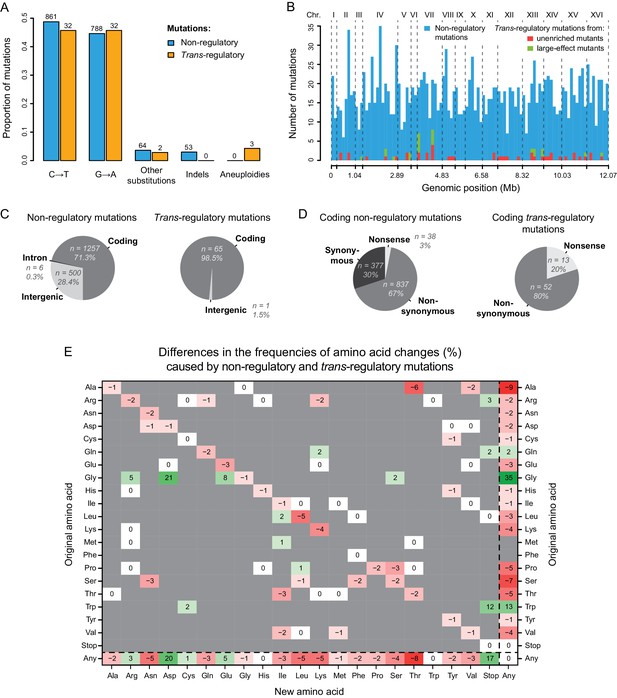
Contrasting properties of trans-regulatory and non-regulatory mutations.
(A) Proportions of different types of mutations in a set of 1766 non-regulatory mutations (blue) and in a set of 69 trans-regulatory mutations (orange). Numbers of mutations are indicated above bars. (B) Distributions of non-regulatory and trans-regulatory point mutations along the yeast genome. A total of 1766 non-regulatory mutations are shown in blue, 44 trans-regulatory mutations that were identified from the collections of unenriched mutants in Metzger et al., 2016 are shown in red and 22 trans-regulatory mutations that were identified from the collections of mutants enriched for large expression changes in Gruber et al., 2012 and in Metzger et al., 2016 are shown in green. (C) Proportions of non-regulatory (left) and trans-regulatory (right) mutations affecting either coding sequences, introns or intergenic regions. (D) Proportions of coding non-regulatory (left) and coding trans-regulatory (right) mutations that either introduce an early stop codon (nonsense), that substitute one amino acid for another (nonsynonymous) or that do not change the amino acid sequence (synonymous). (E) Frequency of all amino acid changes induced by trans-regulatory mutations as compared to non-regulatory mutations. Each entry of the table represents the difference of frequency (percentage) between non-regulatory and trans-regulatory mutations that are changing the amino acid shown on the y-axis into the amino acid shown on the x-axis. For instance, the −6 on the first row indicates that the proportion of mutations changing an Alanine into a Threonine is 6% lower among trans-regulatory mutations than among non-regulatory mutations. Shades of red: amino acid changes underrepresented in the set of trans-regulatory mutations. Shades of green: amino acid changes overrepresented in the set of trans-regulatory mutations. White: amino acid changes equally represented in the trans-regulatory and non-regulatory sets of mutations. Gray: amino acid changes not observed in the sets of trans-regulatory and non-regulatory mutations. (B–E) The three aneuploidies were excluded for these plots. (D,E) Non-coding mutations were excluded for these plots.

Contrasting properties of non-regulatory and trans-regulatory mutations identified by BSA-Seq and of trans-regulatory mutations identified by Sanger sequencing of candidate genes.
(A) Proportions of different types of mutations observed among 1766 non-regulatory mutations (blue), among 52 trans-regulatory mutations identified by BSA-Seq (red) and among 17 trans-regulatory mutations identified by Sanger sequencing of candidate genes. Numbers of mutations are indicated above bars. (B) Distributions of non-regulatory and trans-regulatory point mutations along the yeast genome. A total of 1766 non-regulatory mutations are shown in blue, 49 trans-regulatory mutations that were identified by BSA-Seq are shown in red and 17 trans-regulatory mutations that were identified by Sanger sequencing are shown in green. (C) Proportions of non-regulatory mutations (left), trans-regulatory mutations identified by BSA-Seq (upper right) and trans-regulatory mutations identified by Sanger sequencing (bottom right) that affect either coding sequences, introns or intergenic regions. (D) Proportions of coding non-regulatory mutations (left), coding trans-regulatory mutations identified by BSA-Seq (upper right) and coding trans-regulatory mutations identified by Sanger sequencing (bottom right) that either introduce an early stop codon (nonsense), that substitute one amino acid for another (nonsynonymous) or that do not change the amino acid sequence (synonymous). (E) Frequency of all amino acid changes induced by trans-regulatory mutations identified by BSA-Seq as compared to non-regulatory mutations. Each entry of the table represents the difference of frequency (percentage) between non-regulatory and trans-regulatory mutations that are changing the amino acid shown on the y-axis into the amino acid shown on the x-axis. Shades of red: amino acid changes underrepresented in the set of trans-regulatory mutations identified by BSA-Seq. Shades of green: amino acid changes overrepresented in the set of trans-regulatory mutations identified by BSA-Seq. White: amino acid changes equally represented in the trans-regulatory and non-regulatory sets of mutations. Gray: amino acid changes not observed in the sets of trans-regulatory and non-regulatory mutations. (B–E) The three aneuploidies were excluded for these plots. (D,E) Non-coding mutations were excluded for these plots.

Distributions of trans-regulatory and non-regulatory mutations among chromosomes.
1766 non-regulatory mutations are shown in blue and 69 trans-regulatory mutations are shown in orange, among which 52 mutations were identified by BSA-Seq (shown in red) and 17 mutations were identified by Sanger sequencing of candidate genes (shown in green). Trans-regulatory mutations were significantly enriched on chromosome VII that contained the purine biosynthesis genes ADE5 and ADE6 in which several mutations were identified (24.3% of trans-regulatory mutations located on chromosome VII vs 9.3% of non-regulatory mutations; G-test, p = 3.4 x 10−4). Trans-regulatory mutations were also enriched on chromosome XIII that contained the purine synthesis gene ADE4, although this enrichment was not statistically significant (13.0% of trans-regulatory mutations located on chromosome XIII vs 7.8% of non-regulatory mutations; G-test, p = 0.15).

Statistical significance of the enrichment and depletion of amino acid changes induced by trans-regulatory mutations.
Permutations tests were used to assess the statistical significance of the frequency differences between non-regulatory and trans-regulatory mutations shown on Figure 3E. Each number represents the negative logarithm (base-10) of the p-value obtained using a permutation test to compare the frequency of changing the amino acid on the y-axis to the amino acid shown on the x-axis between non-regulatory and trans-regulatory mutations. Green color intensity scales with the negative logarithm of p-values. White: amino acid changes equally represented in the trans-regulatory and non-regulatory sets of mutations. Gray: amino acid changes not observed in the sets of trans-regulatory and non-regulatory mutations.

Statistical significance of the enrichment and depletion of amino acid changes induced by trans-regulatory mutations identified by BSA-Seq.
Permutations tests were used to assess the statistical significance of the frequency differences between non-regulatory and trans-regulatory mutations shown on Figure 3—figure supplement 1E. Each number represents the negative logarithm (base-10) of the p-value obtained using a permutation test to compare the frequency of changing the amino acid on the y-axis to the amino acid shown on the x-axis between non-regulatory and trans-regulatory mutations. Green color intensity scales with the negative logarithm of p-values. White: amino acid changes equally represented in the trans-regulatory and non-regulatory sets of mutations. Gray: amino acid changes not observed in the sets of trans-regulatory and non-regulatory mutations.
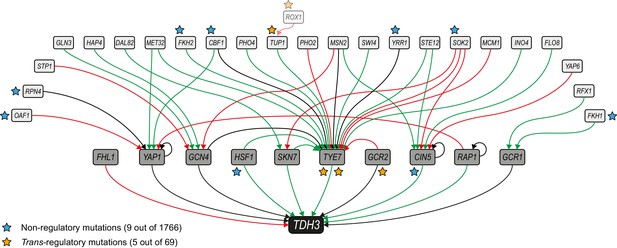
Mutations mapping to a predicted TDH3 regulatory network.
The network of inferred interactions between TDH3 and transcription factors regulating its expression was established using the YEASTRACT repository (Teixeira et al., 2018). First level regulators (dark gray boxes) are transcription factors with evidence of binding to the TDH3 promoter and regulating its expression. Second level regulators (light gray boxes) are transcription factors with evidence of binding to the promoter of at least one first level regulator and regulating its expression. Green arrows: evidence for activation of expression. Red arrows: evidence for inhibition of expression. Black arrows: unknown direction of regulation. Non-regulatory and trans-regulatory mutations identified in the network are represented by blue and orange stars, respectively, near the affected genes. ROX1, inferred to be a third level regulator, is also shown because a trans-regulatory mutation was identified in its coding sequence.
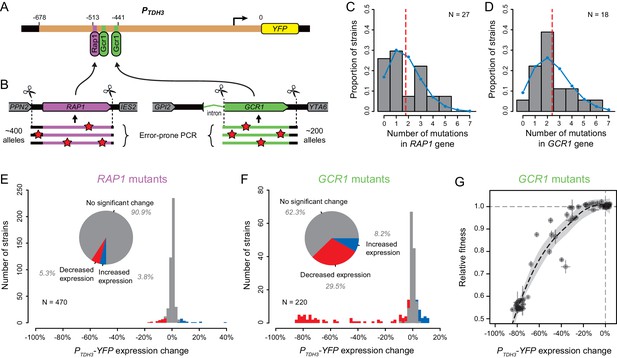
Impact of mutations in two direct regulators of the TDH3 promoter.
(A) Schematics of the PTDH3-YFP reporter gene with locations of three known binding sites for transcription factors Rap1p (purple) and Gcr1p (green) shown in the TDH3 promoter. (B) Regions of RAP1 (purple) and GCR1 (green) genes that were subjected to random mutagenesis using error-prone PCR. 470 RAP1 mutants and 220 GCR1 mutants were obtained by integration of random PCR fragments at the native RAP1 or GCR1 loci using CRISPR/Cas9 allelic replacement. (C–D) Distributions of the number of mutations per strain identified by Sanger sequencing the mutated regions of (C) RAP1 in 27 strains or (D) GCR1 in 18 strains. These data are shown in histograms. Blue curves: Poisson distribution with the same mean as observed in data. Red dotted line: Mean number of mutations among sequenced strains. (E–F) Distributions of PTDH3-YFP expression changes relative to the un-mutagenized reporter strain measured in four replicate samples for (E) the 470 RAP1 mutants or (F) the 220 GCR1 mutants. Fluorescence measures were transformed to be linearly related with YFP mRNA levels (see Methods). Red bars: Mutants with significant decrease in median expression greater than 3% relative to the un-mutagenized strain (permutation test, p < 0.05). Blue bars: Mutants with significant increase in median expression greater than 3% relative to the un-mutagenized strain (permutation test, p < 0.05). Pie charts: Proportions of mutants with significant increase in expression (blue), significant decrease in expression (red) and no significant change in expression (gray) relative to the un-mutagenized strain. (G) Relationship between changes in PTDH3-YFP expression levels (x-axis) and fitness (y-axis) measured in 62 GCR1 mutants. Expression changes and fitness are both expressed relative to the un-mutagenized strain. Gray dotted lines: Expression change and fitness of the un-mutagenized strain. Error bars: 95% confidence intervals of expression changes and fitness measures obtained from four replicate populations of each mutant. The black dotted line represents a LOESS regression of fitness on median expression with a smoothing parameter of 1% and 95% confidence intervals of the estimates shown as a gray shaded area.
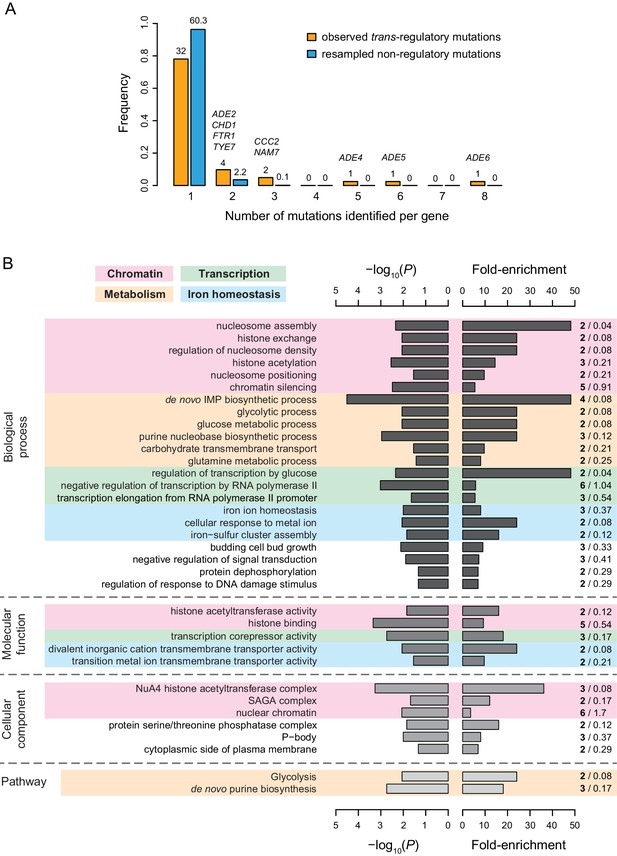
Properties of genes with coding mutations altering PTDH3-YFP expression level.
(A) Proportion of genes with one or more mutations identified among EMS mutants. Mutations in intergenic regions were excluded from this analysis. Orange bars include genes harboring one or more of the 65 trans-regulatory mutations identified in coding sequences. Blue bars include genes harboring one or more of 65 non-regulatory mutations randomly chosen among the set of 1095 non-regulatory mutations observed in coding sequences. The number of genes hit by 1–8 mutations is indicated above the corresponding bar. For blue bars, this number represents the mean number of genes obtained from 1000 random sets of 65 non-regulatory mutations. The names of genes with at least two trans-regulatory mutations identified among mutants are indicated above the bars. FTR1 and CCC2 are involved in iron homeostasis, ADE2,4,5,6 are involved in de novo purine biosynthesis, NAM7 is involved in nonsense-mediated mRNA decay, CHD1 is involved in chromatin regulation and TYE7 encodes a transcription factor regulating TDH3 expression. (B) Summary of gene ontology (GO) enrichment analysis performed with PANTHER tool (http://www.pantherdb.org/). Fisher’s exact tests were used to evaluate the overrepresentation of GO terms among the 42 genes affected by one or more of the 66 trans-regulatory mutations in coding sequences relative to the 1043 genes affected by one or more of the 1251 non-regulatory mutations in coding sequences. The descriptions shown on the left correspond to GO terms with a p-value < 0.05 (left bars), a fold-enrichment > 3 (right bars) and that are not parents to other GO terms in the ontology hierarchy (i.e. GO terms that are the most specific). A more complete list of enriched GO terms can be found in Supplementary file 8. Shades of gray represent different categories of GO terms (from darkest to lightest: biological processes, molecular functions and cellular components) or PANTHER pathways (lightest gray). Fold-enrichment was calculated as the observed number of genes with a particular GO term in the set of genes affected by trans-regulatory mutations (bold numbers on the right) divided by an expected number of genes obtained from the number of genes with the same GO term in the set of genes affected by non-regulatory mutations (regular numbers on the right). Four groups of GO terms and pathways involved in similar processes are represented by colored areas: chromatin (pink), metabolism (orange), transcription (green), and iron homeostasis (blue).
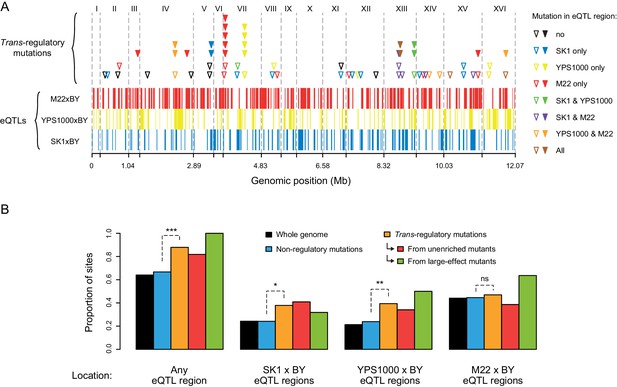
Overrepresentation of trans-regulatory mutations in eQTLs regions.
(A) Overlap of 66 trans-regulatory point mutations and 317 eQTL regions along the yeast genome. eQTL regions were identified by BSA-Seq in Metzger and Wittkopp, 2019 from three crosses of a laboratory strain (BY) to each of three strains expressing PTDH3-YFP in the genetic background of different S. cerevisiae isolates: SK1 (eQTL regions represented by blue bars), YPS1000 (eQTL regions represented by yellow bars) and M22 (eQTL regions represented by red bars). Triangles indicate the genomic locations of trans-regulatory mutations, with open triangles representing mutations identified in mutants from the unenriched collection and filled triangles representing mutations identified in mutants enriched for large effects. Triangles are colored depending on the overlap between mutations and eQTL regions: black if the mutation is outside of any eQTL region, blue if the mutation lies in an eQTL region only identified from SK1xBY, yellow if the mutation lies in an eQTL region only identified from YPS1000xBY, red if the mutation lies in an eQTL region only identified from M22xBY, green if the mutation lies in two overlapping eQTL regions identified from SK1xBY and YPS1000xBY, purple if the mutation lies in two overlapping eQTL regions identified from SK1xBY and M22xBY, orange if the mutation lies in two overlapping eQTL regions identified from M22xBY and YPS1000xBY and brown if the mutation lies in three overlapping eQTL regions identified from the three crosses. (B) Proportions of non-regulatory and trans-regulatory mutations located in eQTL regions. Black bars: proportions of sites among the 12.07 Mb yeast genome. Blue bars: proportions of the 1759 non-regulatory point mutations. Orange bars: proportions of the 66 trans-regulatory mutations (excluding aneuploidies). Red bars: proportions of the 44 trans-regulatory mutations identified in mutants from the unenriched collection. Green bars: proportions of the 22 trans-regulatory mutations identified in mutants enriched for large effects. The proportions of non-regulatory and trans-regulatory mutations in eQTL regions were compared using G-tests (***: p < 0.001, **: 0.001 < p < 0.01, *: 0.01 < p < 0.05, ns: p > 0.05).

Proportions of different categories of non-regulatory mutations and trans-regulatory mutations located in eQTLs regions.
Black bars: proportions of all sites among the 12.07 Mb yeast genome. Medium blue bars: proportions of the 1759 non-regulatory point mutations. Light blue bars: proportions of non-regulatory mutations at sites for which the total sequencing depth was below the median sequencing depth of the corresponding library in BSA-Seq data. Dark blue bars: proportions of non-regulatory mutations at sites for which the total sequencing depth was equal or above the median sequencing depth of the corresponding library in BSA-Seq data. Orange bars: proportions of the 66 trans-regulatory mutations (excluding aneuploidies). Red bars: proportions of the 49 trans-regulatory mutations identified by BSA-Seq. Green bars: proportions of the 17 trans-regulatory mutations identified by Sanger sequencing of candidate genes. The proportions of non-regulatory and trans-regulatory mutations in eQTL regions were compared using G-tests (***: p < 0.001, **: 0.001 < p < 0.01, *: 0.01 < p < 0.05, ns: p > 0.05).
Additional files
-
Source code 1
R scripts used for the analysis of flow cytometry data.
- https://cdn.elifesciences.org/articles/67806/elife-67806-code1-v1.txt.zip
-
Source code 2
R scripts used for the analysis of BSA-Seq data and for comparing the properties of trans-regulatory and non-regulatory mutations.
- https://cdn.elifesciences.org/articles/67806/elife-67806-code2-v1.txt.zip
-
Source code 3
R script used to annotate variants identified in BSA-Seq data.
- https://cdn.elifesciences.org/articles/67806/elife-67806-code3-v1.txt.zip
-
Source code 4
PBS script used to process FASTQ files.
- https://cdn.elifesciences.org/articles/67806/elife-67806-code4-v1.txt.zip
-
Source data 1
Compressed folder including 34.
Source Data files in txt format that contain quantitative data displayed on Figure 1B–E, Figure 2E–G, Figure 3A,B,E, Figure 5C–G, Figure 6, Figure 7, Figure 2—figure supplements 1–5, Figure 3—figure supplements 1–4 and Figure 7—figure supplement 1.
- https://cdn.elifesciences.org/articles/67806/elife-67806-data1-v1.zip
-
Supplementary file 1
Sequencing depth in BSA-seq data.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp1-v1.xls
-
Supplementary file 2
List of all mutations identified by BSA-Seq or Sanger sequencing in this study.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp2-v1.xls
-
Supplementary file 3
Statistical associations between aneuploidies and fluorescence level.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp3-v1.docx
-
Supplementary file 4
Linked mutations associated with fluorescence level in BSA-Seq experiments.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp4-v1.xls
-
Supplementary file 5
Mutations identified by Sanger sequencing of candidate genes.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp5-v1.xls
-
Supplementary file 6
Mutations tested in single-site mutants.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp6-v1.xls
-
Supplementary file 7
Mutations associated with fluorescence level in BSA-Seq experiments.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp7-v1.xls
-
Supplementary file 8
Targeted mutagenesis of RAP1 residues making direct contact with DNA.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp8-v1.xls
-
Supplementary file 9
List of GO terms overrepresented in genes hit by causative mutations relative to genes hit by neutral mutations.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp9-v1.xls
-
Supplementary file 10
Mutations located in the coding sequence of glucose signaling genes.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp10-v1.xls
-
Supplementary file 11
Trans-regulatory effects of mutations in purine biosynthesis genes or iron homeostasis genes.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp11-v1.xls
-
Supplementary file 12
Files used as inputs for analyses performed with the PBS script (Source code 4) and R scripts (Source code 1–3).
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp12-v1.bz2
-
Supplementary file 13
List of DNA libraries grouped by sequencing runs.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp13-v1.xls
-
Supplementary file 14
List of oligonucleotides used in this study.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp14-v1.xls
-
Supplementary file 15
Construction of single-site mutant strains.
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp15-v1.xls
-
Supplementary file 16
Phenotypes of RAP1 mutants (expression) and GCR1 mutants (expression and fitness).
- https://cdn.elifesciences.org/articles/67806/elife-67806-supp16-v1.xls
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/67806/elife-67806-transrepform-v1.docx





