The Shu complex prevents mutagenesis and cytotoxicity of single-strand specific alkylation lesions
Figures
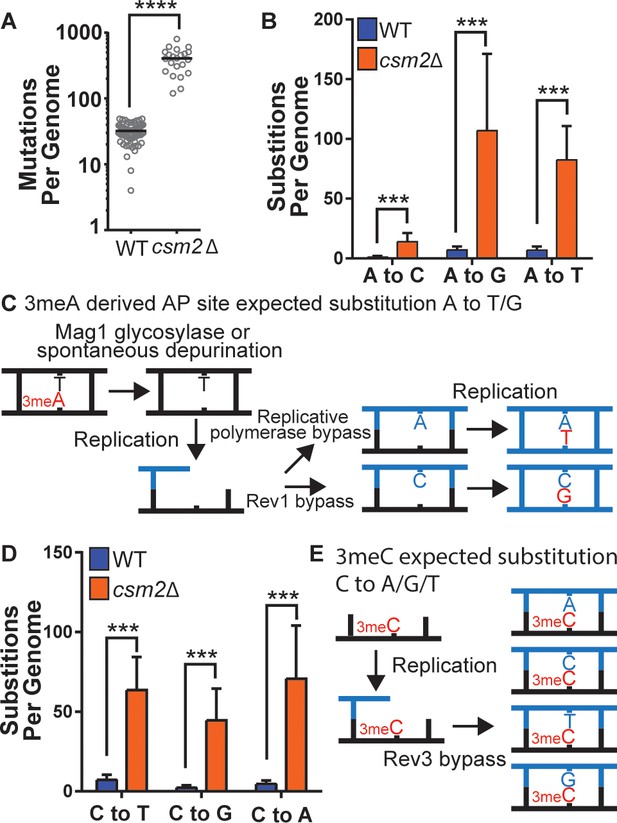
csm2Δ cells chronically exposed to methyl methanesulfonate (MMS) exhibit substitution patterns consistent with translesion synthesis (TLS) activity bypassing abasic (AP) sites and three-methyl cytosine (3meC).
WT and csm2Δ cells were chronically exposed to MMS. WT and csm2Δ cells were chronically exposed to 0.008% MMS by plating individual colonies onto rich medium containing MMS, after 2 days of growth, the colonies were plated onto fresh medium containing MMS for 10 passages. DNA was extracted from 75 WT and 22 csm2Δ clonal isolates and deep sequenced. (A) The number of mutations per genome for WT or csm2Δ cells chronically MMS-exposed. The horizontal bar indicates the median value for each genotype. **** indicates p < 0.0001 by Mann-Whitney test. (B) The average number of each type of A:T substitution per genome in MMS-treated WT and csm2Δ cells. Error bars indicate standard deviation among the samples in each group. *** indicates p < 0.0001 comparing the number of A:T substitutions per genome in WT and csm2∆ yeast by t-test. (C) Schematic of how 3meA-derived AP sites result in A to T/G mutations. 3meA is removed by the Mag1 glycosylase or it undergoes spontaneous depurination resulting in an AP site. During DNA replication, the replicative polymerase bypasses the AP site resulting in a T mutation, alternatively, Rev1 bypasses the AP site resulting in a G mutation. (D) The average number of each type of G:C substitution per genome in MMS-treated WT and csm2Δ cells. Error bars indicate standard deviation among the samples in each group. *** indicates p < 0.0001 comparing the number of G:C substitutions per genome in WT and csm2∆ yeast by t-test. (E) Schematic of how 3meC results in C to A/G/T base substitutions. 3meC occurs primarily in ssDNA and during replication, Rev3 mediated bypass results in incorporation of A, T, G, or C nucleotides.
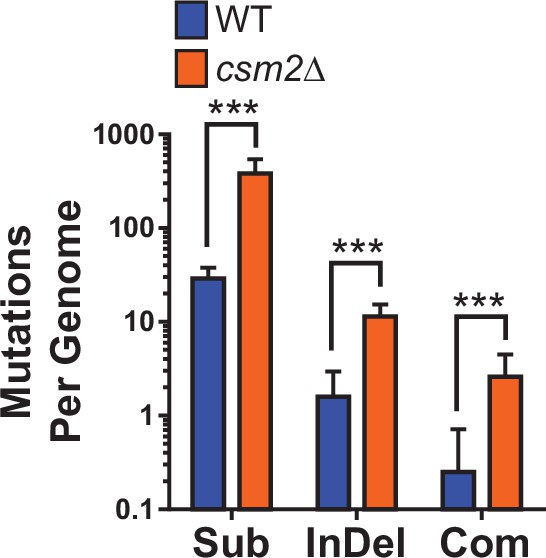
Density of methyl methanesulfonate (MMS)-induced mutations in diploid wild-type (WT) and csm2Δ/csm2Δ yeast.
The number of substitution (Sub), insertion/deletion mutations (InDel), and complex mutations (i.e. neighboring mutations separated by 10 bp or less; Com) in each sequenced genome of WT (blue) and csm2-deficient (orange) yeast (as described in Figure 1 and Supplementary file 1c). Bar heights indicate mean values and error bars are standard deviations. *** indicates p < 0.0001 by t-test.
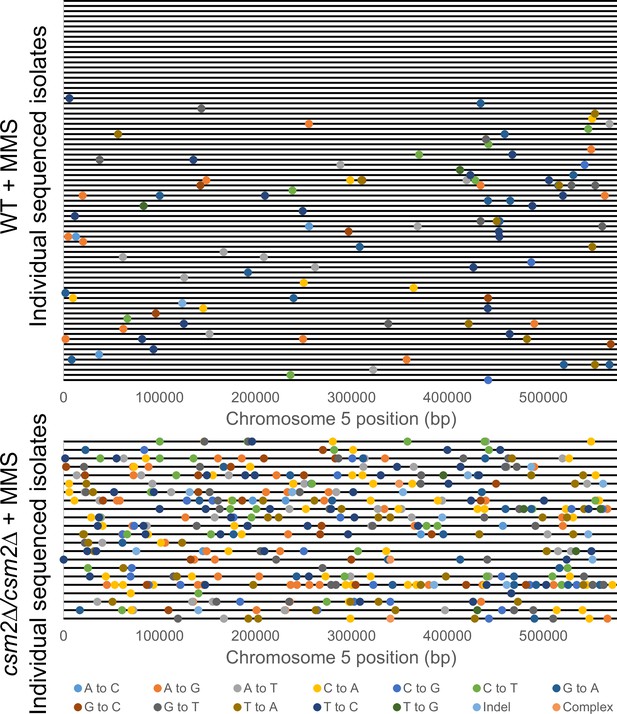
Location of methyl methanesulfonate (MMS)-induced mutations on Chr.5 of independently sequenced isolates of diploid wild-type (WT) and csm2Δ/csm2Δ yeast.
The position of mutations found on chromosome 5 of independent yeast isolates subjected to whole genome sequencing (from Figure 1 and Supplementary file 1c). Each horizontal black line indicates a different yeast isolate (upper panel = 75 WT yeast isolates; lower panel = 22 csm2-deficient yeast isolates). Filled circles indicate the position of mutations and are color-coded by mutation type.
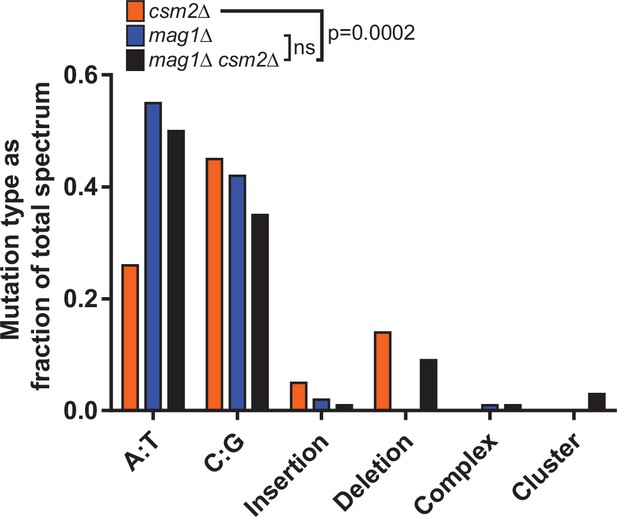
Spectra of methyl methanesulfonate (MMS)-induced mutations in the CAN1 gene in csm2Δ, mag1Δ, and mag1Δ csm2Δ yeast.
Independent canavanine-resistant isolates csm2Δ, mag1Δ, and mag1Δ csm2Δ yeast were obtained after exposure to MMS. Following genomic DNA isolation, the CAN1 gene of each isolate was PCR-amplified and sequenced. Bars indicate the fraction of the total mutation spectrum constituted by each mutation type for csm2Δ (orange), mag1Δ (blue), and mag1Δ csm2Δ (black) yeast. p = 0.0002 by chi-square analysis comparing the number of A:T substitutions, C:G substitutions, insertions, deletions, and complex, and clustered mutations between MMS-treated csm2Δ cells and mag1Δ csm2Δ cells. ns indicates ‘not significant’.
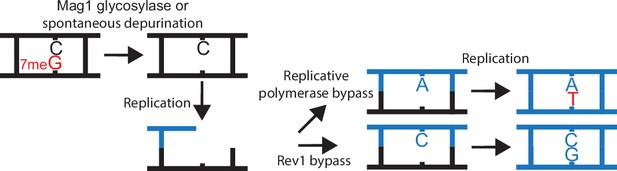
Schematic of how 7meG-derived abasic (AP) sites lead to G to T as the main substitution pattern.
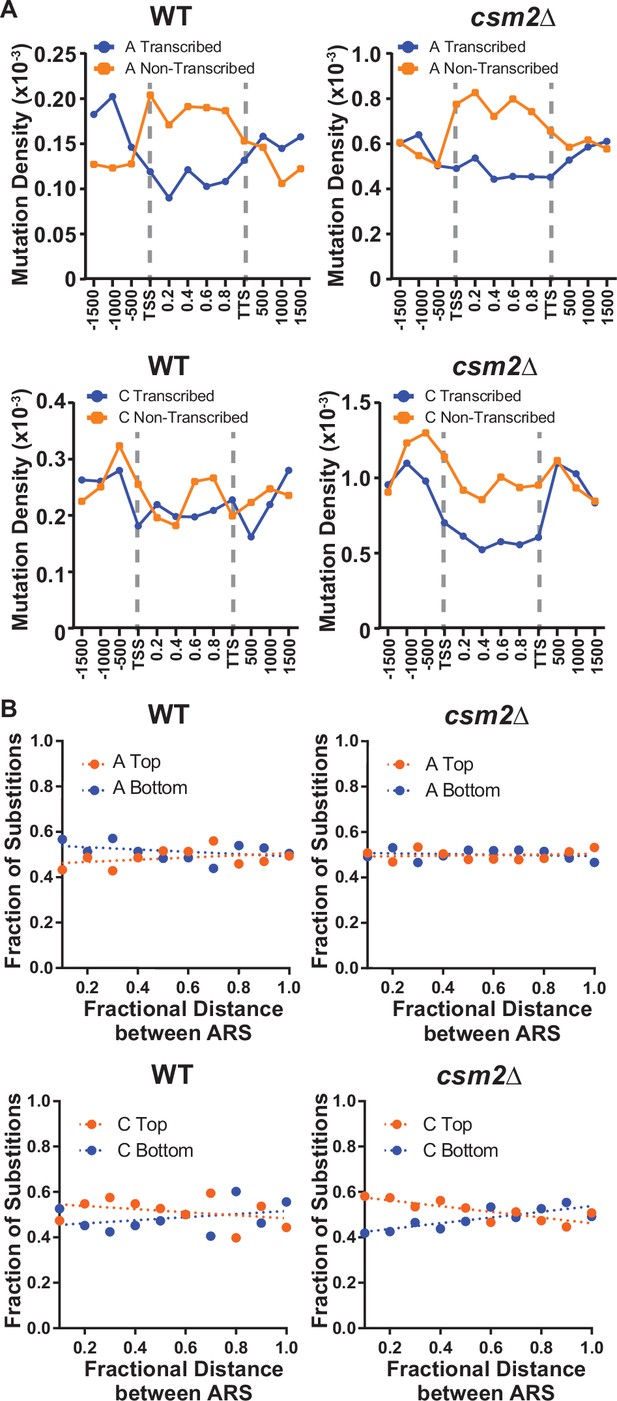
Transcriptional and replicative strand biases of methyl methanesulfonate (MMS)-induced substitutions.
(A) The density of A mutations (i.e. the fraction of A bases mutated) or C mutations on the transcribed (blue) and non-transcribed (orange) strand across yeast transcripts in wild-type (WT) and csm2∆ cells. Transcript regions were broken into fractional bins of 0.2 of the transcript length and the density of A or C mutations determined per bin. Three additional bins of 500 bp each were also included upstream of the transcription start site (TSS) and downstream of the transcription termination site (TTS). (B) The fraction of A and C mutations occurring on the top (orange) and bottom (blue) strands across replication units in the genomes of WT and csm2∆ cells. Replication units were broken into 0.1 fractional bins between neighboring origins of replication and the fraction of A mutations or C mutations associated with each strand were calculated per bin. The fraction of mutations for each strand across the replication unit was fitted with linear regression lines (dashed lines).
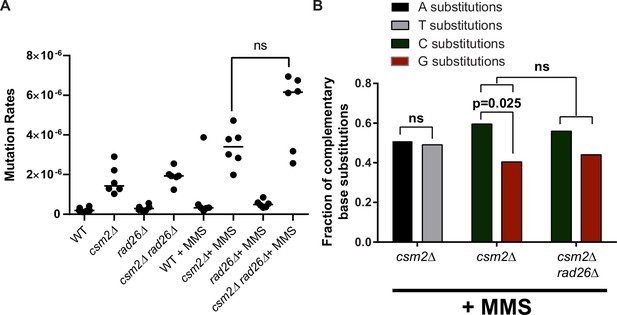
Effects of transcription-coupled nucleotide excision repair (TC-NER) deficiency on methyl methanesulfonate (MMS)-induced mutations in csm2Δ yeast.
(A) CAN1 mutation rates were measured in untreated and MMS-treated wild-type (WT), csm2Δ, rad26Δ, and csm2Δ rad26Δ yeast. Black circles indicate independent measurements and horizontal lines indicate median rates from six experiments. ns indicates no statistically significant difference in mutation rate was observed between the MMS-treated csm2Δ and csm2Δ rad26Δ cells by Mann-Whitney U-test. (B) Transcriptional strand biases for substitutions at A:T and C:G base pairs was assessed in MMS-treated csm2Δ and csm2Δ rad26Δ yeast by sequencing independent canavanine-resistant isolates for each genotype and calculating either the fraction of A:T substitutions that involve a mutated A or T base or the fraction of C:G substitutions that involve a mutated C or G base in the non-transcribed strand of CAN1. Nearly equal numbers of A and T bases in the non-transcribed strand of CAN1 were mutated in MMS-treated csm2Δ yeast (ns by one-sided G-test for goodness of fit predicting A base mutations would dominate on the non-transcribed strand). (C) Bases in the non-transcribed strand of CAN1 were elevated over G bases in csm2Δ yeast (p = 0.025 by one-sided G-test for goodness-of-fit predicting C base mutations would dominate on the non-transcribed strand). No statistical difference was observed in the transcriptional asymmetry favoring C base mutations in the non-transcribed strand between MMS-treated csm2Δ and csm2Δ rad26Δ cells (ns by Fisher’s exact test comparing the number of C- and G-based mutations in csm2Δ and csm2Δ rad26Δ genotypes).
-
Figure 2—figure supplement 1—source data 1
Numerical data corresponding to the graph in panel A.
- https://cdn.elifesciences.org/articles/68080/elife-68080-fig2-figsupp1-data1-v2.xlsx
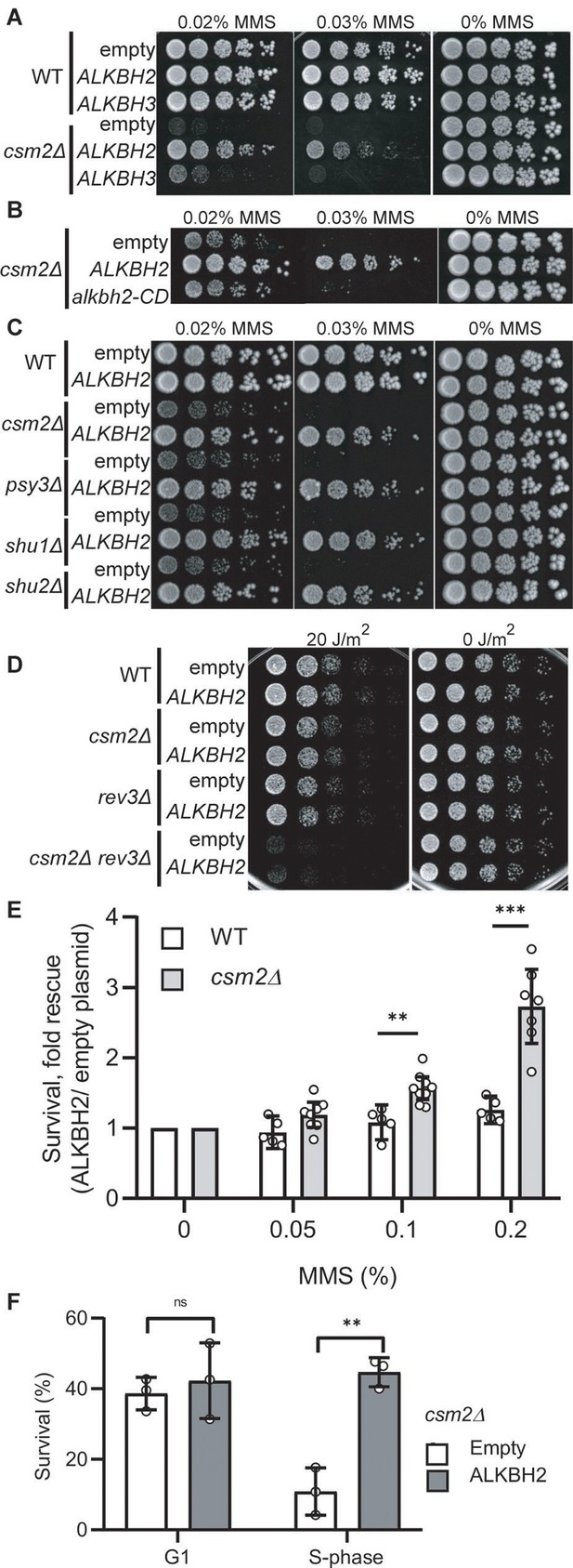
Expression of human ALKBH2 rescues the methyl methanesulfonate (MMS) sensitivity of csm2Δ cells.
(A) csm2Δ cells expressing ALKBH2 exhibit decreased MMS sensitivity. Fivefold serial dilution of wild-type (WT) or csm2∆ cells transformed with an empty plasmid, a plasmid expressing ALKBH2 or a plasmid expressing ALKBH3 onto rich YPD medium or YPD medium containing the indicated MMS concentration were incubated for 2 days at 30°C prior to being photographed. (B) The enzymatic activity of ALKBH2 is required for the rescue of the MMS sensitivity of csm2∆ cells. csm2∆ cells transformed with an empty plasmid, a plasmid expressing ALKBH2 or a plasmid expressing a catalytic dead ALKBH2 mutant were diluted and plated as described in (A) and incubated for 3 days at 30°C prior to being photographed. (C) ALKBH2 expression rescues the MMS sensitivity of cells with deletions of each Shu complex gene. WT, csm2∆, psy3∆, shu1∆, or shu2∆ cells transformed with an empty plasmid or a plasmid expressing ALKBH2 were fivefold serially diluted, plated, and analyzed as described in (B). (D) Expression of ALKBH2 does not rescue the increased ultraviolet (UV) sensitivity observed in csm2Δ rev3Δ double mutants. Fivefold serial dilution of WT, csm2Δ, rev3Δ, or rev3Δ csm2Δ cells were transformed with an empty plasmid or a plasmid expressing ALKBH2 and fivefold serially diluted onto rich YPD or rich YPD medium exposed to 20 J/m2 UV, and incubated for 2 days at 30°C prior to being photographed. An untreated plate (0 J/m2) serves as a loading control. (E) csm2Δ cells expressing ALKBH2 exhibit increased survival after acute MMS treatment. YPD liquid cultures of WT or csm2∆ cells transformed with an empty plasmid or a plasmid expressing ALKBH2 were treated with the indicated concentration of MMS following plating onto rich YPD medium. Colony number was assessed after incubation for 2 days at 30°C. Fold rescue of cellular survival represents the ratio of the survival of cells expressing ALKBH2 relative to the survival of cells expressing the empty plasmid. Survival represents the number of colonies as a percentage of the colonies obtained without MMS treatment. The individual and mean values from five to nine experiments were plotted. Error bars indicate 95% confidence intervals. The p-values between WT and csm2∆ cells treated with 0.1% MMS and 0.2% MMS were calculated using an unpaired two-tailed Student’s t-test and were p ≤ 0.01 and p ≤ 0.001, respectively. (F) S-phase csm2∆ cells expressing ALKBH2 exhibit increased survival after acute MMS treatment. WT or csm2∆ cells were synchronized on G1 with alpha factor and either released from G1 arrest or maintained in G1 in the presence or absence of 0.1% MMS. Cells were plated after 30 min of treatment and the colony number was assessed after incubation for 2 days at 30°C. Survival is calculated as described in (E). The mean values from three experiments were plotted with standard deviations. The p-values between control (empty plasmid) and ALKBH2 expressing cells were calculated using an unpaired two-tailed Student’s t-test and were p > 0.05 (n.s.) and p ≤ 0.001 for the G1- and S-phase cells, respectively.
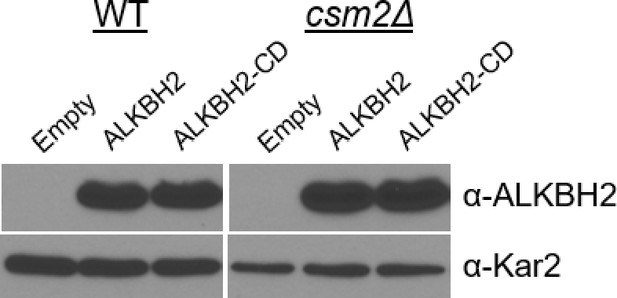
Protein blot analysis of ALKBH2 and ALKBH2-CD expression in wild-type (WT) and csm2∆ cells.
WT and csm2∆ strains expressing an empty plasmid (pAG416GPD-ccdB) or a plasmid (pAG416GPD-ccdB) expressing either ALKBH2 or ALKBH2-CD were analyzed for ALKBH2 protein levels. Protein extracts from equal cell numbers were analyzed by Western blot for ALKBH2 (α-ALKBH2) or Kar2 (α-Kar2) expression as a loading control.
-
Figure 3—figure supplement 1—source data 1
Unprocessed images of the blots.
- https://cdn.elifesciences.org/articles/68080/elife-68080-fig3-figsupp1-data1-v2.zip
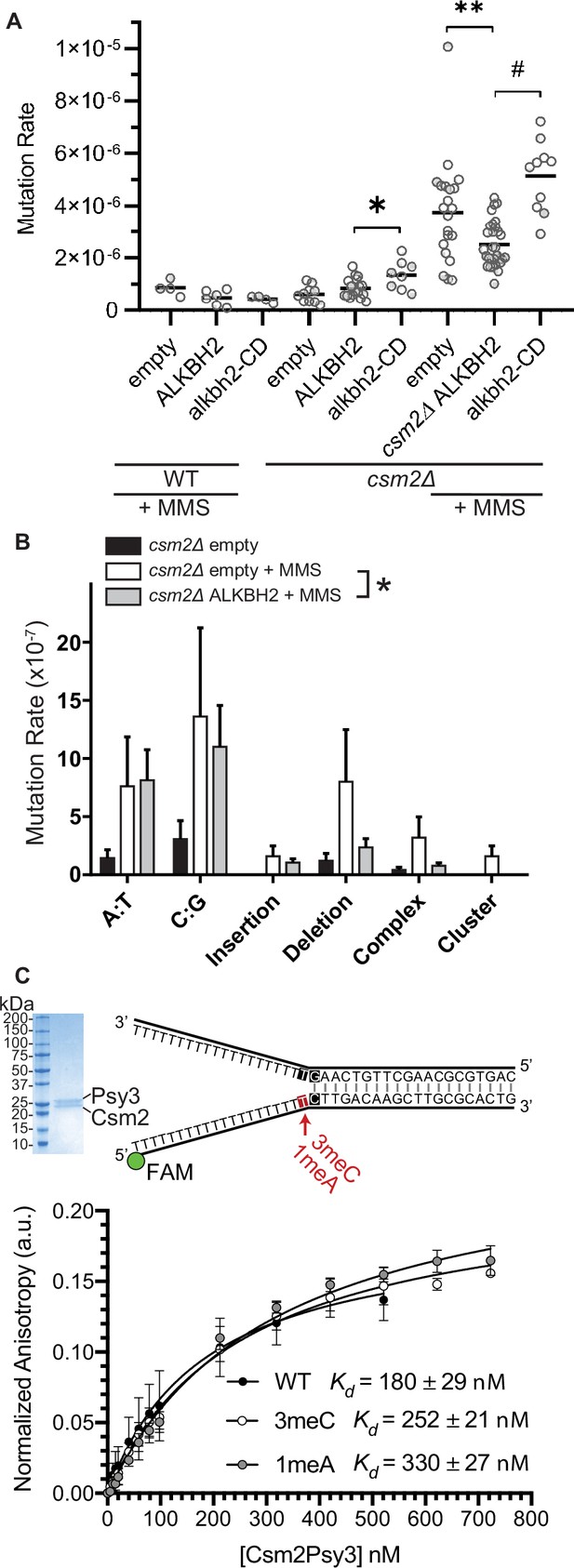
ALKBH2 expression rescues the 3meC-induced mutagenesis observed in methyl methanesulfonate (MMS) exposed csm2Δ cells.
(A) csm2Δ cells expressing ALKBH2 exhibit reduced MMS-induced mutation rate. Spontaneous and MMS-induced mutation rates at the CAN1 locus were measured in wild-type (WT) (MMS treated only) and csm2∆ cells transformed with an empty plasmid, a plasmid-expressing ALKBH2 or alkbh2-CD. Each measurement represents a separate experiment (circle) and the median value (horizontal bar) of 4–22 experiments were plotted. The p-values were calculated using a Mann-Whitney ranked sum test and were as follows: p = 0.0667 for treated WT empty and WT ALKBH2; p = 0.1327 for untreated csm2∆ empty and csm2∆ ALKBH2; p = 0.0221 for untreated csm2∆ ALKBH2 and csm2∆ alkbh2-CD (*); p = 0.0092 for treated csm2∆ empty and csm2∆ ALKBH2 (**); and p < 0.0001 for treated csm2∆ ALKBH2 and csm2∆ alkbh2-CD (#). (B) Sequencing of the CAN1 gene in canavanine-resistant colonies was used to calculate the frequency of MMS-induced substitutions, insertions, deletions, and complex mutations in csm2∆ cells transformed with either an empty vector or ALKBH2 expression vector. ALKBH2 expression significantly alters the MMS-induced mutation spectra (* indicates p = 0.015 by chi-square analysis comparing the number of A:T substitutions, C:G substitutions, insertions, deletions, complex, and clustered mutations between MMS-treated csm2∆ cells containing an empty vector or ALKBH2 expression vector). The spectrum contains a reduction in C:G substitutions, deletions, and complex mutations, while A:T substitutions and insertions are unchanged. (C) The Csm2-Psy3 protein binds to a double-flap DNA substrate containing unmodified, 3meC, and 1meA lesions. A Coomassie-stained SDS-PAGE gel of recombinant Csm2-Psy3, which run at 27.7 and 28 kDa, respectively. Equilibrium binding titrations were performed by titrating Csm2-Psy3 into the FAM-labeled double-flap substrate in unmodified (Kd = 180 ± 29), 3meC containing (Kd = 252 ± 21) and 1meA (Kd = 313 ± 27) containing lesions at the indicated position, and anisotropy was measured. Experiments were performed with three protein preparations and standard deviations plotted. The data were fit to a quadratic equation (one-site binding model assumed) and dissociation constants (Kd) were calculated.
-
Figure 4—source data 1
Numerical data corresponding to the graph in panel A.
- https://cdn.elifesciences.org/articles/68080/elife-68080-fig4-data1-v2.xlsx
-
Figure 4—source data 2
Unprocessed image of the gel in panel C.
- https://cdn.elifesciences.org/articles/68080/elife-68080-fig4-data2-v2.zip
-
Figure 4—source data 3
Unprocessed anisotropy values.
- https://cdn.elifesciences.org/articles/68080/elife-68080-fig4-data3-v2.zip
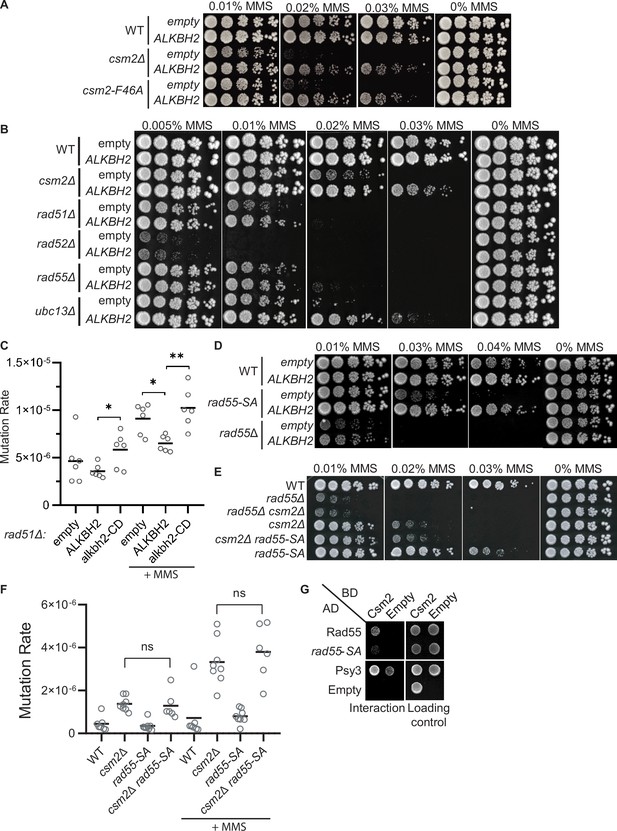
3meC are bypassed by the error-free post-replication repair (PRR) pathway.
(A) ALKBH2 rescues the methyl methanesulfonate (MMS) sensitivity of a csm2-F46A mutant, which is deficient for its Rad51 mediator activity. Fivefold serial dilutions of wild-type (WT), csm2Δ, or csm2-F46A cells transformed with an empty plasmid or a plasmid-expressing ALKBH2, were platted onto rich YPD medium or YPD medium containing the indicated MMS concentration and incubated for 3 days at 30°C prior to being photographed. (B) Unlike PRR mutant UBC13, expression of ALKBH2 mildly rescues the MMS sensitivity of homologous recombination (HR) factors, RAD51, RAD52, and RAD55. Fivefold serial dilution of WT, csm2Δ, rad51Δ, rad52Δ, rad55Δ, or ubc13Δ transformed with an empty plasmid or a plasmid-expressing ALKBH2 were fivefold serially diluted, plated, and analyzed as described in (A). (C) rad51Δ cells expressing ALKBH2 exhibit reduced MMS-induced mutation rate. Spontaneous and MMS-induced mutation rates at the CAN1 locus were measured in rad51∆ cells transformed with an empty plasmid, a plasmid-expressing ALKBH2 or alkbh2-CD. Each measurement represents a separate experiment (circle) and the median value (horizontal bar) of six experiments were plotted. The p-values were calculated using a Mann-Whitney ranked sum test and were as follows: p = 0.5887 for untreated empty and ALKBH2; p = 0.026 untreated ALKBH2 and alkbh2-CD (*), p = 0.026 for MMS-treated rad51∆ empty and ALKBH2 (*); p = 0.0043 for MMS-treated rad51∆ ALKBH2 and alkbh2-CD (**). (D) rad55-S2,8,14A (rad55-SA) cells expressing ALKBH2 exhibit decreased MMS sensitivity. WT, rad55-S2,8,14A, or rad55Δ cells transformed with an empty plasmid or a plasmid-expressing ALKBH2 were fivefold serially diluted, plated, and analyzed as described in (A). (E) csm2Δ is epistatic to rad55-S2,8,14A (rad55-SA) for MMS damage. Cells with the indicated genotypes were fivefold serially diluted and plated as described in (A), and incubated for 2 days at 30°C prior to being photographed. (F) The mutation rate of a csm2∆ rad55-S2,8,14A (csm2∆ rad55-SA) double mutant is the same as a csm2∆ cell. Spontaneous and MMS-induced mutation rates at the CAN1 locus were measured in WT, csm2∆, rad55-SA, and csm2∆ rad55-SA cells. Each measurement represents a separate experiment (circle) and the median value (horizontal bar) of six to eight experiments were plotted. The p-values were calculated using a Mann-Whitney ranked sum test and were p = 0.3238 and p = 0.3965 for csm2∆ and csm2∆ rad55-SA untreated or MMS-treated, respectively (not significant, ns). (G) rad55-S2,8,14A (rad55-SA) exhibit an impaired yeast-2-hybrid (Y2H) interaction with Csm2. Y2H analysis of pGAD-RAD55, rad55-S2,8,14A, PSY3, or pGAD-C1 (Empty) with pGBD-RAD57, CSM2, pGBD-C1 (Empty). A Y2H interaction is indicated by plating equal cell numbers on SC medium lacking histidine, tryptophan, and leucine. Equal cell loading is determined by plating on synthetic complete (SC) medium lacking tryptophan and leucine used to select for the pGAD (AD) and pGBD (BD) plasmids.
-
Figure 5—source data 1
Numerical data corresponding to the graph in panel C.
- https://cdn.elifesciences.org/articles/68080/elife-68080-fig5-data1-v2.xlsx
-
Figure 5—source data 2
Numerical data corresponding to the graph in panel F.
- https://cdn.elifesciences.org/articles/68080/elife-68080-fig5-data2-v2.xlsx
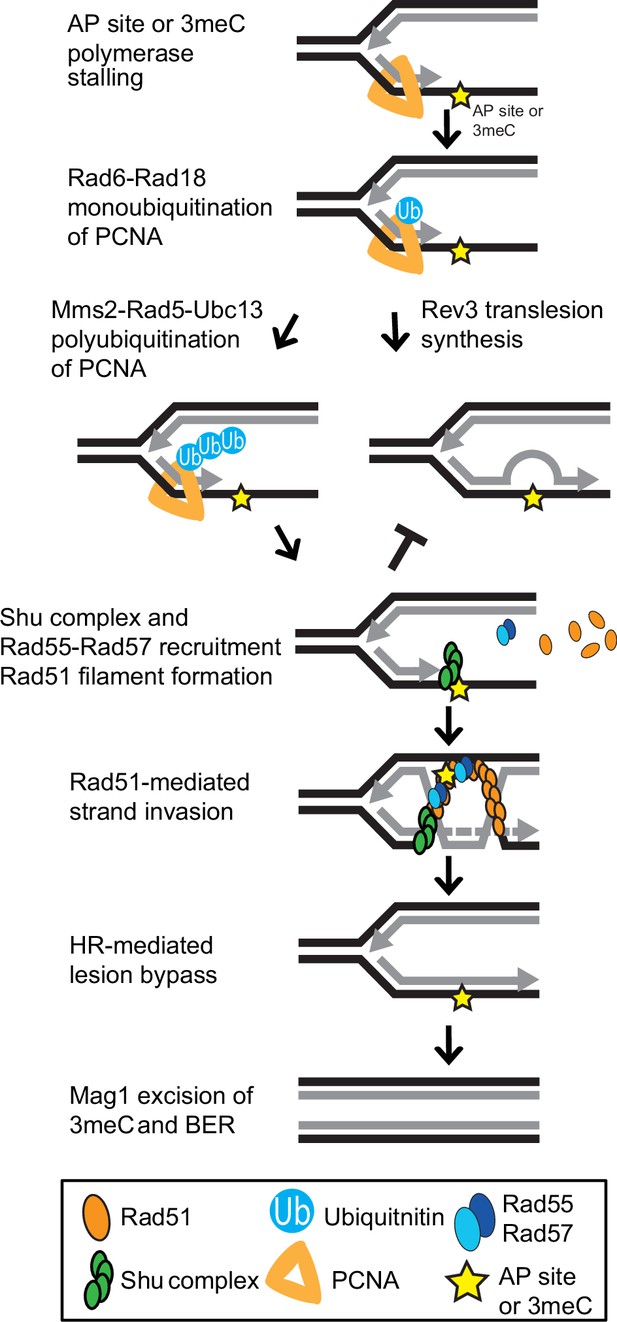
Model of Shu complex-mediated error-free bypass of abasic (AP) sites and 3meC.
MMS-induced AP sites and 3meC (yellow star) arising at DNA replication intermediates at ssDNA can stall the replicative polymerase. Replication fork stalling leads to PCNA (orange triangle) K63-linked polyubiquitination of lysine 164 (K164) by the sequential activities of the Rad6-Rad18 and Mms2-Rad5-Ubc18 complexes. When an AP site or 3meC forms, the Shu complex (green ovals) promotes Rad55-Rad57 (blue ovals) recruitment and Rad51 filament formation (orange ovals). Thus, enabling Rad51-mediated HR with the newly synthesized sister chromatid. Importantly, the Shu complex activity prevents mutagenesis from TLS-mediated error-prone bypass of 3meC. After DNA synthesis using the undamaged sister chromatid as a template, the HR intermediates are resolved. The error-free bypass of 3meC enables S-phase completion in a timely manner. Finally, after replication is completed, 3meC are likely recognized and excised by the Mag1 glycosylase, which initiates the BER-mediated repair.
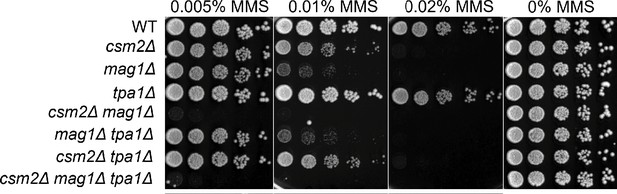
TPA1 does not genetically interact with CSM2 or MAG1 for methyl methanesulfonate (MMS) damage.
Fivefold serial dilution of cells, with the indicated genotypes, was transformed onto rich YPD medium or rich YPD medium containing the indicated MMS concentration was incubated for 2 days at 30°C prior to being photographed.
Additional files
-
Supplementary file 1
(a): List of strains used in this study (b): PCR oligonucleotides used in this study (c): Unique mutations in diploid wild-type (WT) and csm2∆/csm2∆ methyl methanesulfonate (MMS) exposed yeast (d): Primers used for CAN1 sequencing (e): CAN1 sequencing from untreated and methyl methanesulfonate (MMS)-treated yeast.
- https://cdn.elifesciences.org/articles/68080/elife-68080-supp1-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/68080/elife-68080-transrepform1-v2.docx





