A universal pocket in fatty acyl-AMP ligases ensures redirection of fatty acid pool away from coenzyme A-based activation
Figures
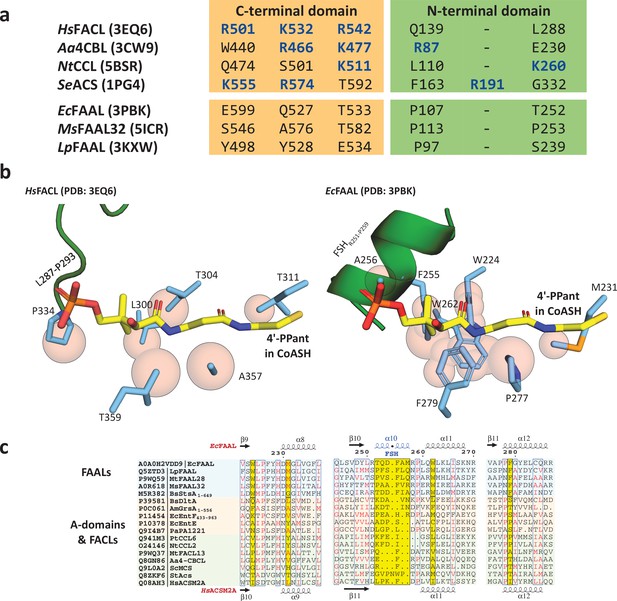
The presence of conserved negative selection elements and absence of positive selection elements in canonical coenzyme A (CoA)-binding pocket of fatty acyl-AMP ligases (FAALs) can prevent CoA binding.
(a) The CoA-interacting residues in fatty acyl/aryl-CoA ligases (FACLs) (HsFACL, Aa4CBL, NtCCL, and SeACS) and the structurally analogous residues in FAALs (EcFAAL, MsFAAL32, and LpFAAL) are tabulated. The positively charged residues (blue) mainly are from the C-terminal domain (orange) and occasionally from the N-terminal domain (green). (b) The residues (cyan) in the vicinity (≤2.5 Å) of the bound CoA (yellow; HsFACL; PDB: 3EQ6) are shown as van der Waals spheres (light red) to compare the canonical pocket in FAALs and FACLs. The adenosine 3',5'-bisphosphate moiety of CoA is omitted in the representation for clarity. A unique FAAL-specific helix (FSHR251-P259) at the entry of the canonical pocket shown in cartoon representation (green) is replaced by a loop (L287-P293) in FACLs. (c) The canonical pocket-obstructing features seen in FAALs (light cyan) are highlighted in yellow in the structure-based sequence alignment and compared to other representative members of the ANL superfamily (FACLs in light green and A-domains in light orange). The secondary structures of EcFAAL (PDB: 3PBK) and HsFACL (PDB: 3EQ6) are marked at the top and bottom of the alignment, respectively.
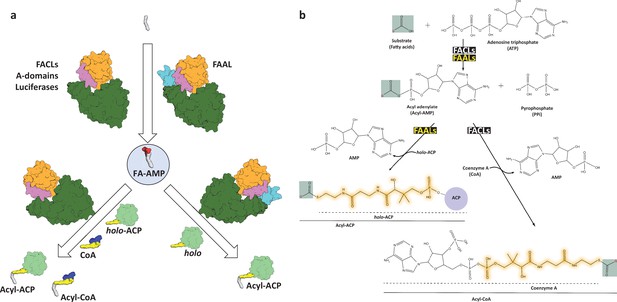
A schematic showing the structural and biochemical differences between fatty acyl-AMP ligases (FAALs) and fatty acyl/aryl-CoA ligases (FACLs).
(a) An overview of the characteristic enzymology presented by fatty acyl-AMP ligases (FAALs) and fatty acyl/aryl-CoA ligases (FACLs) leading to the dichotomy of the fate of free fatty acids. FAALs and FACLs both produce the fatty acyl-AMP from fatty acid (gray) but transfer the intermediate to different acceptors. Such a differential activity is despite two critical similarities; first, the overall structural similarity (N-terminal domain in green; C-terminal domain in orange connected by the A8-motif in violet), except an insertion in FAALs (cyan). Second, the differential activity is despite the chemical identity of the substrates that react during the second step, where coenzyme A (CoA) used by FACLs is used to post-translationally modify apo-ACP to holo-ACP (light green) that is utilized by FAALs. FAALs are highly specific in diverting the intermediate fatty acyl-adenylate to the holo-ACP acceptor and then divert the fate of the pool of free fatty acids towards various pathways that work independently of CoA. (b) The chemical structures of the molecules involved in the reaction are depicted using a general substrate (shown as G). The first step of the reaction with a molecule of adenosine triphosphate (ATP) is identical for both FAALs and FACLs that result in the formation of acyl-adenylate or acyl-AMP and a concomitant release of pyrophosphate (PPi). In the second step, FAALs facilitate the transfer of acyl group to 4'-phosphopantetheine (highlighted in orange) tethered holo-ACP to form acyl-ACP while FACLs transfer the acyl group to 4'-phosphopantetheine linked to CoA to form acyl-CoA. In both instances, a single molecule of AMP is released.
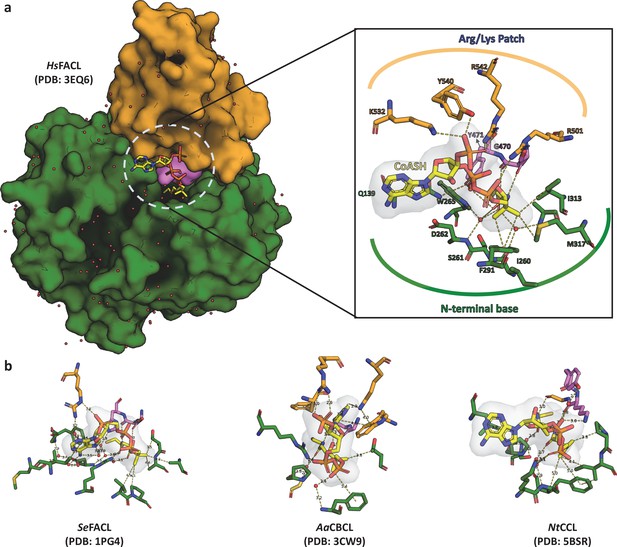
Structural anlaysis of the interactions between FACLs and the bound coenzyme A shows high degree of plasticity.
(a) A surface representation of coenzyme A (CoA)-bound HsFACL (PDB: 3EQ6) is shown with CoA (yellow) as a stick model sandwiched between the N-terminal (green) and C-terminal domain (orange) along with the connecting A8-motif (violet). The inset highlights the network of interactions between the CoA and the protein molecule, which can be grouped into three categories. (i) Arg/Lys clamp: structurally conserved Arg/Lys residues from the C-terminal domain (occasionally from the N-terminal domain) make ionic interactions with phosphates of CoA. The sequence positions of these positively charged residues need not be conserved. (ii) PPant clamp: residues from the highly conserved A8-motif form hydrogen bonds with the 4'-phosphopantetheine arm. (iii) N-terminal clamp: a series of water-mediated interactions with residues from the N-terminal and CoA. (b) A stick model of the CoA and residues of the protein forming a network of interactions is shown from three diverse CoA-bound fatty acyl/aryl-CoA ligases (FACL) structures, namely acetyl-CoA synthetase (SeFACL; PDB: 1PG4), 4-chloro benzoyl-CoA ligase (AaFACL; PDB: 3CW9), and coumarate-CoA ligase (NtCCL; PDB: 5BSR). The residues from the C-terminal domain are shown in orange, those from the N-terminal domain are shown in green, and residues from A8-motif are shown in violet. These interactions highlight the plasticity of the CoA-binding pocket with limited or no specific residues that impart specificity for CoA.
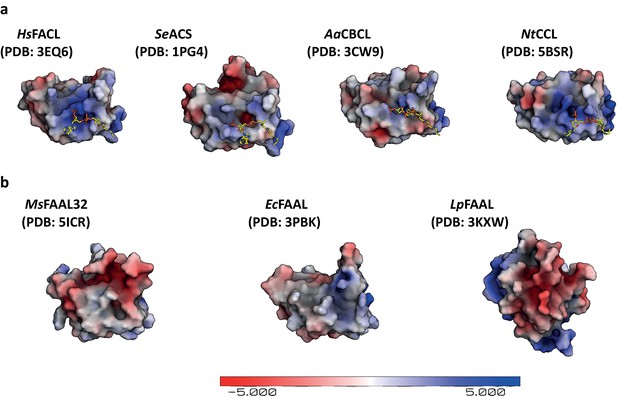
A surface represenation of the C-terminal domains coloured based on electrostaic potential, from FACLs and FAALs.
(a) An electrostatic map of the C-terminal domain from coenzyme A (CoA)-bound fatty acyl/aryl-CoA ligases (FACLs) structures is shown along with a stick model for CoA, which shows that there are positive patches that assist in holding the phosphates of CoA. (b) The electrostatic potential map of the crystal structures of the C-terminal domain of fatty acyl-AMP ligases (FAALs) is shown for comparison, where the structurally analogous region binding to CoA is devoid of positively charged patch as seen in the case of FACLs. The electrostatic map was generated using the APBS module in PyMOL (Schrödinger, LLC, 2015).
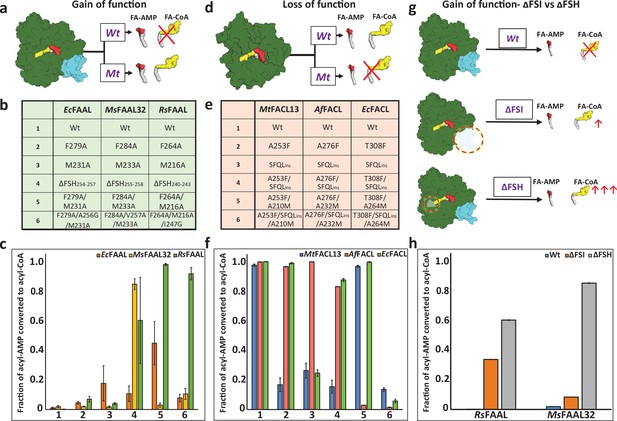
Biochemical analysis of ‘gain of function’ mutants of fatty acyl-AMP ligases (FAALs) and ‘loss of function’ mutants of fatty acyl/aryl-CoA ligases (FACLs).
(a) The biochemical activity of a wild-type (Wt) FAAL as compared to its mutants (Mt) is schematically represented. (b) Various ‘gain of function’ mutations of FAALs (EcFAAL, MsFAAL32, and RsFAAL) are tabulated as the following: row 1: wild-type protein; rows 2 and 3: single-point mutations; row 4: deletion of FAAL-specific helix (FSH); rows 5 and 6: combinations of these mutations. (c) The fraction of acyl-AMP converted to acyl-CoA by wild-type and various mutants (numbered on the x-axis according to b) of FAALs is represented as a bar graph along with standard error of the mean. (d) The biochemical activity of a wild-type (Wt) FACL as compared to its mutants (Mt) is schematically represented. (e) The various ‘loss of function’ mutations generated in FACLs (MtFACL, AfFACL, and EcFACL) are tabulated as the following: row 1: wild-type protein; row 2: single-point mutations; row 3: insertion of FSH; rows 4–6: combinations of these mutations. (f) The fraction of acyl-AMP converted to acyl-CoA by wild-type and various mutants (numbered on the x-axis according to e) of FACLs is represented as a bar graph along with standard error of the mean. (g) A comparison of the gain of function from ΔFSH mutation with ΔFSI mutation in FAALs is schematically represented. (h) The fraction of acyl-AMP converted to acyl-CoA by wild-type, ΔFSH, and ΔFSI mutants of RsFAAL and MsFAAL32 (represented on the x-axis) is represented as a bar graph along with standard error of the mean. The original uncropped images of these experiments are presented as Figure 2—source data 1. The intensity values used to compute the bar graphs are provided as an Excel file as the following: c = Figure 2—source data 2 f = Figure 2—source data 3; h = Figure 2—source data 4.
-
Figure 2—source data 1
The original uncropped radio-TLC images presented here were used to assess the ‘gain of function’ and ‘loss of function’ of fatty acyl-AMP ligases (FAALs) and fatty acyl/aryl-CoA ligases (FACLs) respectively following mutations in their canonical coenzyme A (CoA)-binding pocket.
It was also used to compare the gain of function obtained from deletion of the previously annotated FAAL-specific insertion (ΔFSI) and FAAL-specific helix (ΔFSH). Wild-type FAAL, wild-type FACL, and reaction lacking any protein were used as controls. All the TLCs were marked at the origin, where the reaction mix containing 1-14C fatty acids was spotted. The products acyl-CoA band and the acyl-AMP band along with the free fatty acid band are visualized owing to the radio-labeled fatty acid. (A) A representative image of TLC showing the canonical pocket mutations leading to the gain of function in EcFAAL. (B) A representative image of TLC showing the canonical pocket mutations leading to the gain of function in MsFAAL32. (C) A representative image of TLC showing the canonical pocket mutations leading to the gain of function in RsFAAL. (D) A representative image of TLC showing the canonical pocket mutations leading to the loss of function in MtFACL13. (E) A representative image of TLC showing the canonical pocket mutations leading to the loss of function in AfFACL. (F) A representative image of TLC showing the canonical pocket mutations leading to the loss of function in EcFACL. (G) A representative image of TLC showing the gain of function by ΔFSI mutation in RsFAAL. (H) A representative image of TLC showing the gain of function by ΔFSI mutation in MsFAAL32. Several mutations were generated in this study, which had multiple issues including protein stability, poor or complete loss of biochemical activity, etc., hence were not analyzed further and such mutations are marked by a red asterisk as ‘not part of the study.’ These original uncropped images of radio-TLC are source data for Figure 2.
- https://cdn.elifesciences.org/articles/70067/elife-70067-fig2-data1-v2.pdf
-
Figure 2—source data 2
The excel file is a tabulation of the raw values of the acyl-AMP and acyl-CoA formed in the gain-of-function experiments in EcFAAL, MsFAAL32, and RsFAAL, which are then converted to fraction of the total acyl-AMP (TA) formed to the remaining acyl-AMP (A) and the acyl-AMP converted to acyl-CoA (T).
These values were obtained from densitometric quantification of the TLC images, which were used to plot Figure 2c.
- https://cdn.elifesciences.org/articles/70067/elife-70067-fig2-data2-v2.xlsx
-
Figure 2—source data 3
The excel file is a tabulation of the raw values of the acyl-AMP and acyl-CoA formed in the loss-of-function experiments in MtFACL13, AfFACL, and EcFACL, which are then converted to fraction of the total acyl-AMP (TA) formed to the remaining acyl-AMP (A) and the acyl-AMP converted to acyl-CoA (T).
These values were obtained from densitometric quantification of the TLC images, which were used to plot Figure 2f.
- https://cdn.elifesciences.org/articles/70067/elife-70067-fig2-data3-v2.xlsx
-
Figure 2—source data 4
The excel file is a tabulation of the raw values of the acyl-AMP and acyl-CoA formed in the experiment comparing the gain of function in fatty acyl-AMP ligases (FAALs) with deletion of FAAL-specific insertion (ΔFSI) to deletion of FAAL-specific helix (ΔFSH).
The values are then converted to fraction of the total acyl-AMP (TA) formed to the remaining acyl-AMP (A) and the acyl-AMP converted to acyl-CoA (T).
These values were obtained from densitometric quantification of the TLC images, which were used to plot Figure 2h.
- https://cdn.elifesciences.org/articles/70067/elife-70067-fig2-data4-v2.xlsx
-
Figure 2—source data 5
The original uncropped radio-TLC showing ‘gain of function’ and ‘loss of function’ of FAALs and FACLs.
- https://cdn.elifesciences.org/articles/70067/elife-70067-fig2-data5-v2.zip
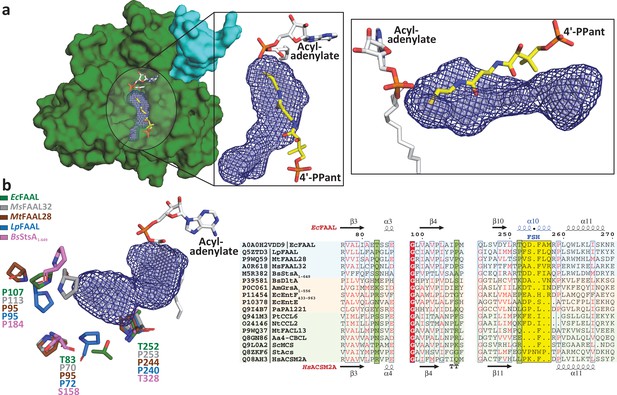
Fatty acyl-AMP ligases (FAALs) have an alternative pocket in the N-terminal domain distinct from the canonical coenzyme A (CoA)-binding pocket and lined by conserved prolines.
(a) The analysis of N-terminal domain of EcFAAL (PDB: 3PBK) (green: surface representation) with pocket finding algorithms identified a new pocket (dark blue: mesh representation; light blue: spheres representation). Also shown are the FAAL-specific insertion (cyan), an EcFAAL-bound lauryl-adenylate (gray: sticks representation) and the 4'-PPant moiety (yellow: sticks representation) of a CoA bound to HsFACL (PDB: 3EQ6) in the canonical pocket. The inset shows two different orientations (top view and lateral view) of the distinct alternative pocket poised against the acyl-adenylate and compared to the 4'-PPant of a CoA in canonical pocket. (b) Various residues (stick representation) of FAALs residing at the entrance of the unique pocket (dark blue: mesh representation; light blue: spheres representation) are identified through structural comparison. A structure-based sequence alignment of these residues (highlighted in pale green) with other representative members of the ANL superfamily reveals that FAALs have a higher frequency of prolines (occasionally Thr/Ser) than other members of the superfamily.
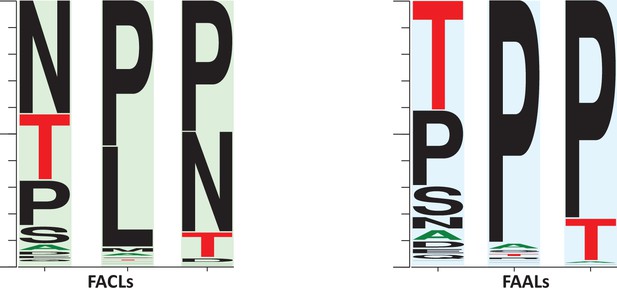

A comparison of the frequency of the prolines surrounding the newly identified alternative pocket in fatty acyl-AMP ligases (FAALs) (highlighted in blue) against the frequency of prolines in fatty acyl/aryl-CoA ligases (FACLs) (highlighted in green).
The representation indicates that the regions are highly variable. The conservation of prolines is higher in FAALs than FACLs. It may be occasionally replaced with threonine or serine in case of FAALs, and in case of FACLs it is replaced by leucine or asparagine. The frequency plot was generated from the sequence alignment using WebLogo (Crooks et al., 2004).
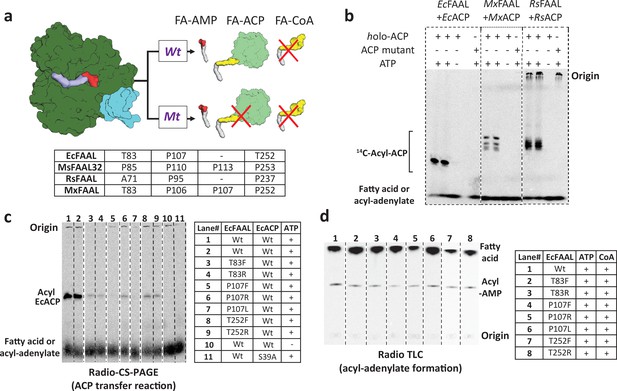
The alternative pocket identified in fatty acyl-AMP ligases (FAALs) is a functional pocket that assists in catalysis by accommodating 4'-PPant tethered to ACP.
(a) A schematic representation of the acyl-transfer function in a wild-type (Wt) FAAL is compared to a mutant protein (Mt) that blocks the alternative pocket. (b) A representative radio conformationally sensitive urea-PAGE (radio-CS-PAGE) gel shows the acyl-transfer reaction by wild-type FAALs on their respective holo-ACP. The radio-CS-PAGE gel shows the successful transfer of the activated fatty acyl-AMP to holo-ACP using three pairs of FAAL-ACP systems (EcFAAL-EcACP, MxFAAL-MxACP, and RsFAAL-RsACP), which is observed as a band due to the incorporated radiolabeled fatty acid. The absence of a band is indicative of failure to transfer as seen in the control reactions lacking ATP or the presence of a mutant ACP (conserved serine mutated to alanine) lacking the 4'-PPant arm. Samples are loaded at the point marked origin and the bands below contain either unreacted radiolabeled fatty acid or fatty acyl-AMP. (c) The radio-CS-PAGE shows that mutations of the residues lining the alternative pocket to Arg/Phe show a reduced or negligible acyl-ACP formation as the bands are diminished or absent. (d) A radio-TLC shows the acyl-AMP formation ability of various FAAL mutants indicated by the bright band in the center after the sample was loaded at the origin and developed until the edge of TLC where unused radiolabeled fatty acid is observed. The radio-TLC shows that loss of acyl-ACP formation is not due to affected acyl-adenylate formation. The original uncropped images of these experiments are presented as Figure 4—source data 1.
-
Figure 4—source data 1
The radio conformationally sensitive urea-PAGE (radio-CS-PAGE) images presented here were used to show the optimization of fatty acyl-AMP ligase (FAAL)-dependent acyl transfer on holo-ACP using three different FAAL-ACP systems (EcFAAL-EcACP, RsFAAL-RsACP, and MxFAAL-MxACP).
These were also used to assess the impact of mutations in the alternative pocket of EcFAAL to form acyl-EcACP. Wild-type FAAL, reaction lacking ATP, and reaction containing a mutant ACP, lacking 4'-phosphopantetheine arm, were used as controls. All the radio-CS-PAGE were marked at the origin, where the reaction mix containing 1-14C fatty acids was loaded without boiling. The acyl-ACP(s), along with unreacted fatty acids or acyl-AMP(s), could only be visualized owing to the radiolabeled fatty acid. The unreacted fatty acids or acyl-AMP(s) appeared as a diffused band at the bottom of the radio-CS-PAGE. The radio-TLC images presented here were used to assess the ability of EcFAAL and its mutations in the alternate pocket to form acyl-AMP. Wild-type EcFAAL, wild-type MtFACL13, and reaction lacking any protein were used as controls. All the TLCs were marked at the origin, where the reaction mix containing 1-14C fatty acids was spotted. The products, acyl-CoA band and the acyl-AMP band, along with the free fatty acid band, were visualized owing to the radiolabeled fatty acid. (A) A representative image showing the optimization and validation of the modified radio-CS-PAGE in three pairs of FAAL-ACP systems, namely EcFAAL-EcACP, RsFAAL-RsACP, and MxFAAL-MxACP. (B) A representative TLC image showing that alternate pocket mutations of EcFAAL have minimal or no effect on the acyl-AMP formation. (C) A representative image of radio-CS-PAGE showing that acyl-EcACP formation is severely attenuated by mutations in the alternate pocket of EcFAAL. Several mutations of the alternate pocket of EcFAAL were generated in this study, which had multiple issues including protein stability, poor or complete loss of biochemical activity, etc., hence were not analyzed further, and these mutations are marked by a red asterisk as ‘not part of the study.’ These original uncropped images of radio-CS-PAGE and radio-TLC are source data for Figure 4.
- https://cdn.elifesciences.org/articles/70067/elife-70067-fig4-data1-v2.pdf
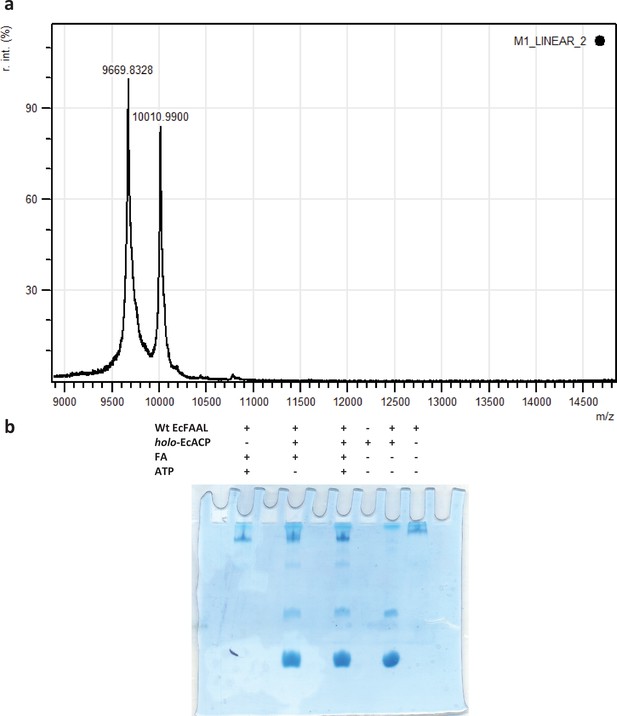
Typically, acyl-transfer reactions with ACP are assessed using an SDS-PAGE (coupled to radio-labeled or fluorescently labeled substrates) or conformationally sensitive urea-PAGE or HPLC-coupled to mass spectrometry.
SDS-PAGE-based assays are suitable for ACPs that are fused to polyketide synthase/nonribosomal peptide synthetase (PKS/NRPS) modules, which was used for MsFAAL32-MsPKS131-1042 in this study. This strategy is not suitable for smaller-sized molecules such as ACP (<10 kDa), which often migrate at the edges along with residual labeled fatty acid and acyl-adenylates. The assay also requires converting apo-ACP to holo-ACP using an enzyme, Sfp, from Bacillus subtilis. (a) A representative image of the mass-spectrometry analysis (AB-SCIEX 4800 MALDI-TOF and using 5 mg/ml sinapinic acid in 50% acetonitrile and 0.1% trifluoroacetic acid as matrix) of Sfp catalyzed post-translational modification of EcACP (9.6 kDa) to form holo-ACP (10 kDa). The analysis revealed that less than 50% of the apo-ACP is only converted to holo-ACP. (b) A representative image of Coomassie-stained conformationally sensitive urea-PAGE (15% acrylamide and 2.5 M urea) of the EcFAAL catalyzed acyl-transfer on holo-EcACP. The Coomassie-stained gel does not reveal the classic separation of holo-ACP and acyl-ACP. The uncropped images of b are provided as Figure 4—figure supplement 1—source data 1.
-
Figure 4—figure supplement 1—source data 1
A representative image of Coomassie-stained conformationally sensitive urea-PAGE (15% acrylamide and 2.5 M urea) of the EcFAAL catalyzed acyl-transfer on holo-EcACP.
The Coomassie-stained gel does not reveal the classic separation of holo-ACP, acyl-ACP, and apo-ACP.
- https://cdn.elifesciences.org/articles/70067/elife-70067-fig4-figsupp1-data1-v2.pdf
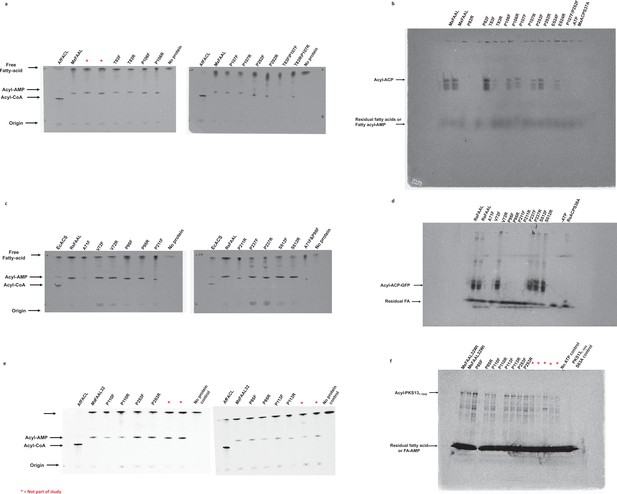
The radio-TLC images presented here were used to assess the ability of fatty acyl-AMP ligases (FAALs), MxFAAL (A), RsFAAL (B), and MsFAAL32 (C), with mutations in the alternate pocket to form acyl-AMP.
Wild-type FAAL, wild-type fatty acyl/aryl-CoA ligases (FACL), and reaction lacking any protein were used as controls. All the TLCs were marked at the origin, where the reaction mix containing 1-14C fatty acids was spotted. The products acyl-CoA band and the acyl-AMP band along with the free fatty acid band were visualized owing to the radiolabeled fatty acid. The radio conformationally sensitive urea-PAGE (radio-CS-PAGE) images presented here were used to assess the ability of FAALs, MxFAAL (D) and RsFAAL (E), with mutations in the alternate pocket to form acyl-ACP with their respective ACPs (RsACP and MxACP). Wild-type FAAL, reaction lacking ATP, and reaction containing a mutant ACP lacking 4'-phosphopantetheine arm were used as controls. All the radio-CS-PAGE were marked at the origin, where the reaction mix containing 1-14C fatty acids was loaded without boiling. The acyl-ACP along with unreacted fatty acids or acyl-AMP can only be visualized owing to the radiolabeled fatty acid. The unreacted fatty acids or acyl-AMP appeared as a diffused band at the bottom of the radio-CS-PAGE. Multiple bands in the modified radio-CS-PAGE may represent either the degradation products of MxACP-GFP or the multiple unfolded forms of MxACP-GFP in the presence of urea in the gel. Radio-SDS-PAGE images presented here were used to assess the ability of MsFAAL32 and its mutations in the alternate pocket to form acyl-MsPKS131-1042 (F). Wild-type MsFAAL32, reaction lacking ATP, and reaction containing a mutant ACP lacking 4'-phosphopantetheine arm were used as controls. The radio-SDS-PAGE was marked at the origin, where the reaction mix containing 1-14C fatty acids was loaded without boiling. The acyl-MsPKS131-1042 along with unreacted fatty acids or acyl-AMP can only be visualized owing to the radiolabeled fatty acid. The unreacted fatty acids or acyl-AMP appeared as a diffused band at the bottom of the radio-SDS-PAGE. Multiple bands in the radio-SDS-PAGE may represent either the degradation products of MsPKS131-1042 or the multiple unfolded forms of MsPKS131-1042 in the presence of SDS in the gel. Several mutations of the alternate pocket of different FAALs were generated in this study, which had multiple issues including protein stability, poor or complete loss of biochemical activity, etc., hence were not discussed further, and these mutations are marked by a red asterisk as ‘not part of the study.’ The original, uncropped images of the radio-TLC, radio-CS-PAGE, and radio-SDS-PAGE, which were part of the study, are presented here as Figure 4—figure supplement 2—source data 1.
-
Figure 4—figure supplement 2—source data 1
The FAAL-ACP pairs, RsFAAL-RsACP and MxFAAL-MxACP, along with mutations in the alternate pocket of the respective fatty acyl-AMP ligases (FAALs), were assessed using modified radio conformationally sensitive urea-PAGE (CS-PAGE) while MsFAAL32-MsPKS1-1042 was assessed using a radio-SDS-PAGE.
All the TLCs are marked with the origin (where 20 μL reaction mix is spotted), the acyl-CoA band (closest to origin), the acyl-AMP band (closest to the solvent front), and the free fatty acid band (near the solvent front). All the modified radio-CS-PAGE are marked with the acyl-ACP band and a diffused band, which may be free fatty acids or the acyl-AMP formed in the reaction. (A) Representative TLC images showing that the acyl-AMP formation is minimally or not affected by mutations in the alternate pocket of MxFAAL. (B) A representative modified radio-CS-PAGE image showing the acyl-MxACP formation by alternate pocket mutants of MxFAAL. The multiple bands in the modified radio-CS-PAGE may represent either the degradation products of MxACP-GFP or the multiple unfolded forms of MxACP-GFP in the presence of urea in the gel. (C) Representative TLC images showing the acyl-AMP formation by alternate pocket mutants of RsFAAL. (D) A representative modified radio-CS-PAGE image showing the acyl-RsACP formation by alternate pocket mutants of RsFAAL. (E) Representative TLC images showing the acyl-AMP formation by alternate pocket mutants of MsFAAL32. (F) A representative radio-SDS-PAGE showing the acyl-MsPKS131-1042 formation by the alternate pocket mutants of MsFAAL32. Several mutations of the alternate pocket of different FAALs were generated in this study, which had multiple issues including protein stability, poor or complete loss of biochemical activity, etc., hence were not discussed further, and these mutations are marked by a red asterisk as ‘not part of the study.’.
- https://cdn.elifesciences.org/articles/70067/elife-70067-fig4-figsupp2-data1-v2.zip
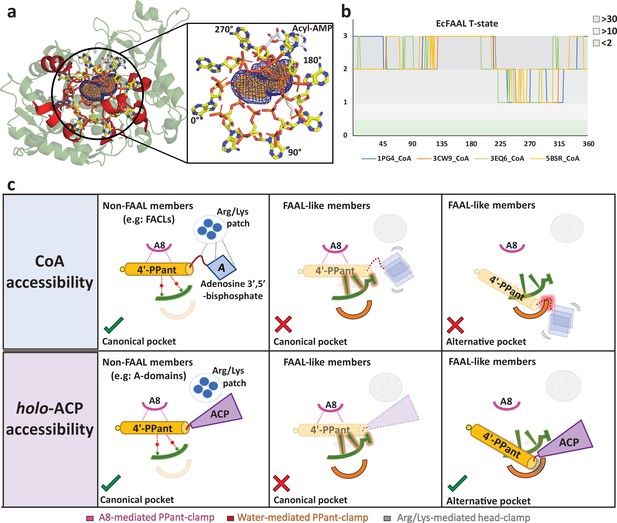
The alternative pocket in fatty acyl-AMP ligases (FAALs) is highly selective, and its unique architecture negatively selects for coenzyme A (CoA).
(a) A ‘mast’ (the 4’-PPant arm) is aligned along the longest length of the predicted pocket (blue: mesh representation) and the ‘flag’ (adenosine 3',5'-bisphosphate) is rotated to generate theoretical orientations of the ‘flag.’ These conformations of HsFACL (PDB: 3EQ6) are shown (yellow: stick format) at 45° intervals. The regions of the EcFAAL (cartoon representation) showing clashes are highlighted (red). (b) The count of atoms (C, N, O, Cα, and Cβ) of EcFAAL showing van der Waals short contacts (≤0.25 Å) with the conformations of the ‘flag’ is scored (short contacts ≥30 = 3; ≥10 = 2; ≥2 = 1 and <2 = 0). These scores (y-axis) are plotted against the specific rotation angles of the conformation, where the short contact is observed (x-axis). (c) The universal CoA-rejection mechanism is schematically summarized. The 4’-PPant (light orange) of both CoA and holo-ACP (purple) bind to the canonical pocket of non-FAAL members through multiple interactions (dotted lines). Interactions between Arg/Lys patch (circle: blue) and the adenosine 3',5'-bisphosphate moiety (rhombus: blue) acts as positive selection. The negative selection elements of the canonical pocket clash (highlighted in red) with the adenosine 3',5'-bisphosphate in the alternative pocket. The absence of Arg/Lys patch (dotted circle) fails to provide stability to the adenosine 3',5'-bisphosphate moiety (dotted rhombus; blue). The 4'-PPant (yellow) tethered to ACP (purple) is only accepted in the alternative pocket, which is absent in non-FAAL members, as it shows no clashes. The ranked count of atoms used to generate the plot 5b is provided as an Excel file, Figure 5—source data 1.
-
Figure 5—source data 1
The Excel sheet tabulates the number of clashes observed between various conformations of coenzyme A (CoA) and fatty acyl-AMP ligases (FAALs) (main-chain atoms: N, C, O, Cα, and Cβ).
The bound CoA from the crystal structures of SeACS (PDB: 1PG4), AaCBCL (PDB: 3CW9), HsFACL (PDB: 3EQ6), and NtCCL (PDB: 5BSR) were used for the starting conformation of the head group of CoA. These crystal structures were then superimposed over the crystal structures of representative FAALs in adenylation competent state (A-state), EcFAAL (PDB: 3PBK), and MsFAAL32 (PDB: 5ICR), which were also used to generate the thioesterification competent state (T-state) by superimposing the C-terminal domain separately. The number of clashes was computed for every 1° rotation of the head group and then ranked as 3 if number ≥30, as 2 if number ≥10, and as 0 if number ≥1. These ranked values were used to plot Figure 5b and Figure 5—figure supplement 2a.
- https://cdn.elifesciences.org/articles/70067/elife-70067-fig5-data1-v2.xlsx
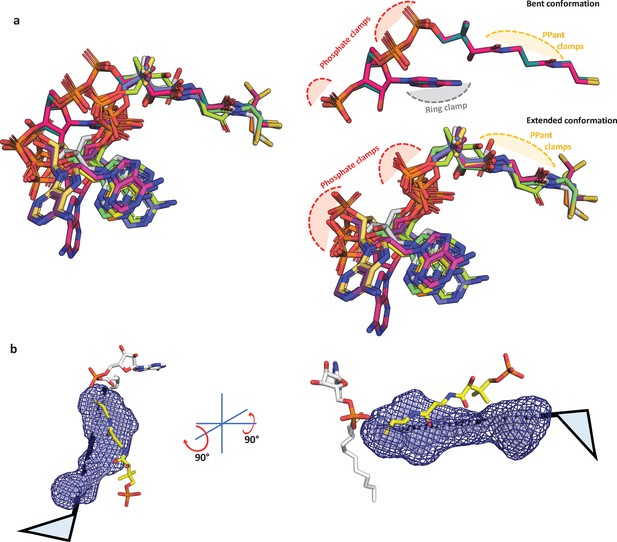
Structural analysis of known coenzyme A conformations and a schematic showing accomodation of 4'-phosphopantetheine arm in the alterative pocket.
(a) The available coenzyme A (CoA)-bound crystal structures of the ANL superfamily members were taken, which include SeACS (1PG4, two protomers; 2P2F, two protomers; 2P2J, two protomers), HsACSM2A (3EQ6, two protomers), NtCCL (5BSR, one protomer), DsPrpE (5GXD, one protomer), and Aa4CBL (3CW9, two protomers). These were aligned based on the maximum-common-substructure method as implemented in mcsalign plugin of PyMOL (Schrödinger, LLC, 2015) along the length of the 4'-PPant. The superposition reveals that CoA structures are bound to ANL superfamily members mainly in an extended conformation (adenine ring away from the 4'-PPant arm) and rarely in bent conformation (adenine ring in proximity to 4'-PPant arm). The difference is due to the conformational freedom around the phosphodiester bonds; however, despite the variations, the phosphates tend to spatially organize in the similar positions available for the phosphate clamping arginine or lysine residues. The bent conformation has an additional contact point, ring clamp, where the adenine ring is held through aromatic amino acids, while the adenine ring is free in the extended conformation of CoA with orientations of adenine ring having a degree of variability but restricted to a small zone. The 4'-PPant arm is presented in a fully extended form in both the subsets of CoA and the PPant clamp atoms presented for hydrogen bonding with the A8 motif and water molecules. (b) In the absence of any structural information on how the 4'-PPant may be accommodated in the pocket, it is safe to assume the 4'-PPant is accommodated within the identified pocket analogous to the ‘mast’ of the flag. The flag here represents the head group, which can be rotated to identify any possible orientation that can be accommodated within the alternative pocket. All the known conformations of head groups from CoA-bound ANL superfamily members serve as a template in this case.
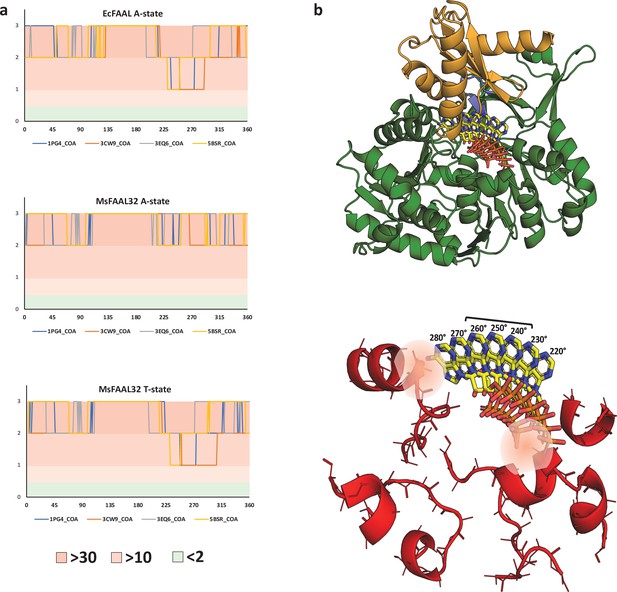
The clash score analysis of various conformations of adenosine 3',5'-bisphosphate shows the incompatibility of coenzyme A binding in alternative pocket.
(a) The variation of clash score with an angle for the possible adenylation and putative thioesterification conformations of the two known fatty acyl-AMP ligases (FAALs) structures (EcFAAL; PDB: 3PBK; and MsFAAL32; PDB: 5ICR) is presented. It is very clear that none of the conformations, A-state or T-state, of both the proteins, EcFAAL and MsFAAL32, are capable of accommodating any of the known conformations of adenosine 3',5'-bisphosphate moiety of coenzyme A (CoA). A large number of the sampled conformations present clashes with more than 30 main-chain atoms even with a conservative distance of ≤0.25 Å. The plot for EcFAAL in A-state and T-state is comparable in the zones of least resistance, which is also true for MsFAAL32 T-state, suggesting that these are conformation-independent. This observation indicates that the clashes can primarily be attributed to the N-terminal itself and the unique location of the pocket. It should be noted that the plot for MsFAAL32 A-state is unique because of the alternative pocket being occluded by a FAAL32-specific insertion described by Aldrich and co-workers. (b) The T-state EcFAAL was modeled by superposing the C-terminal domain with HsACSM2A and is shown in cartoon form (N-terminal in green and C-terminal in orange) along with theoretical orientations of the head group CoA from 220° to 280°, which is shown in stick representation (yellow). The major elements of the N-terminal domain in EcFAAL that contribute to clashes are shown in a cartoon representation (red) along with the stick representation of the main-chain atoms and Cβ. The conformations of the head group from 220° to 280°, where only a few (2–10) clashes are observed, particularly the region spanned between 240° and 260°. The zone of least resistance is largely owing to a vacant space between N- and C-terminal domains without any residues to hold the phosphates or adenine ring. The space occupied by these conformations is close to the typical structural site for CoA accommodation in fatty acyl/aryl-CoA ligases (FACLs) except that the adenine ring is oriented away from the protein. The typical Arg/Lys residues that bind phosphates are absent in FAALs, and hence these conformations cannot be expected to be stabilized by phosphate clamps. The orientations with inward-facing adenine ring resemble the bent conformation of CoA, a rare conformation in ANL superfamily members, where the adenine ring is held by a conserved tryptophan as seen in AaCBL (PDB: 3CW9). The conserved tryptophan residue is absent in most ANL superfamily members including FAALs and FACLs, and hence these conformations cannot be expected to be stabilized by stacking interactions with adenine. Therefore, in the 220°–280° region even though the rejection by negative selection is weak, there is no scope of positive selection that renders the region incompatible with CoA binding. The ranked clash score used to generate the plot in (a) is tabulated in an Excel file, Figure 5—source data 1.
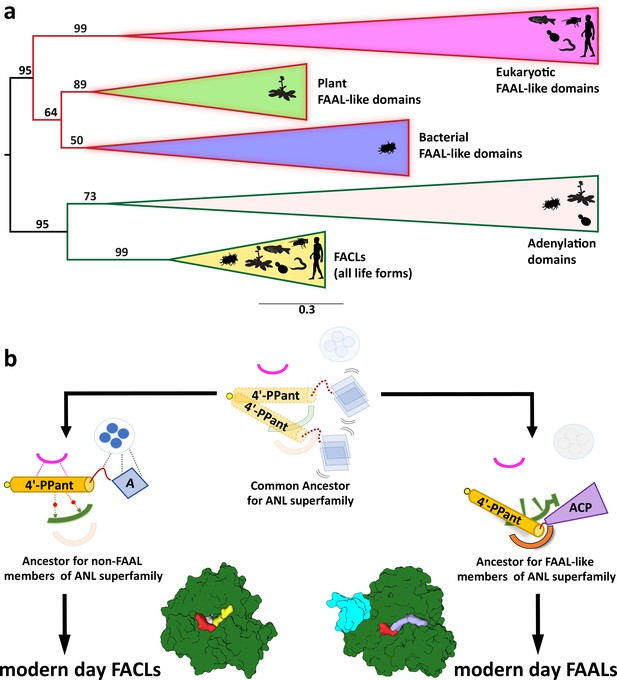
The coenzyme A (CoA)-rejection elements in fatty acyl-AMP ligases (FAALs) are conserved in all forms of life, and therefore, FAALs and fatty acyl/aryl-CoA ligases (FACLs) have parallelly evolved from a common ancestor of the ANL superfamily.
(a) A clustering diagram with the bootstrapping values for all the FAAL-like sequences is presented. The ANL superfamily members have two major divergent classes, viz., CoA-rejecting FAAL-like sequences and CoA-accepting non-FAAL members (FACLs and A-domains). A graphical representation of the distribution of these members is shown using pictorial representation of the organisms, which include bacteria, plants, yeast, worm, fly, and human. (b) The ancestral fold of the ANL superfamily may have been a loose organization of peptide scaffolds working with thiol-containing molecules such as pantetheine, etc. The ancestral fold accumulated various mutations, resulting into the ancestor of modern-day acceptor promiscuity lacking FAAL-like forms and acceptor promiscuity containing non-FAAL members. The evolution of acceptor promiscuity spectrum may have been driven based on their participation in bulk metabolic reactions. FAALs did not participate in bulk metabolic reactions and hence dedicated themselves to their cognate ACP partners to redirect small molecules to specific pathways. The non-FAAL members of the ANL superfamily such as FACLs participate in bulk reactions, where minor cross-reaction products are observed.
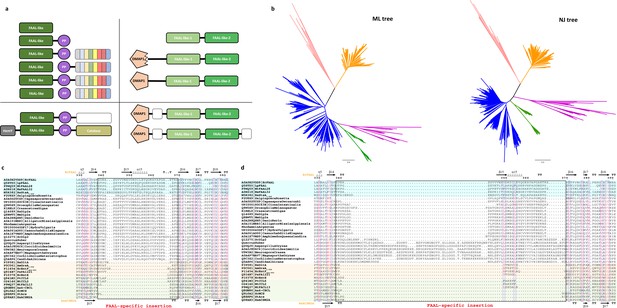
The fatty acyl-AMP ligases (FAAL)-like sequences were identified and aligned to generate a structure-based alignment as described in the Materials and methods section.
The sequences whose structures are available are colored (FAALs in blue, A-domains in light red, and fatty acyl/aryl-CoA ligases [FACLs] in green) and the secondary structures of EcFAAL (PDB: 3PBK; top) along with HsACSM2A (PDB: 3EQ6) are also included. The FAAL-specific insertion region is marked as a black rectangle and labeled accordingly (red). (a) A survey of the various domain organizations of various FAAL-like sequences is depicted as polyketide synthase/nonribosomal peptide synthetase (PKS/NRPS)-related domain organizations (top-left panel) and PKS/NRPS-independent domain organizations (top-right panel), while the bottom panels represent rare/miscellaneous domain organizations. The PKS/NRPS-related domain organizations are seen in bacteria, plants, and few lower eukaryotes including amoebozoa (e.g., Acanthamoeba), Haptista (e.g., Emiliania), and few fungi where bacterial/plant FAAL-like domains (dark green) are associated with a carrier protein domain (violet) and different domains of NRPS/PKS systems (variegated color). The bacterial/plant FAAL-like domains are occasionally seen in different domain contexts such as alpha aminoadipate reductase domains or α/β-hydrolases (white dotted box) in few bacterial, plant, and lower eukaryotic species or sandwiched between HemY (gray) and catalase (olive) domains exclusively found in plants. The PKS/NRPS-independent domain organizations are almost exclusively seen in opisthokonts and rarely in few plants. The FAAL-like sequences of opisthokonts are related to typical bacterial/plant FAAL-like sequence but show two different levels of sequence divergence, namely FAAL-like-1 (pale green) and FAAL-like-2 (green), where the lighter shade indicates the extent of divergence. All the identified opisthokonta FAAL-like sequences are always found as a tandem fusion of FAAL-like-1 and FAAL-like-2. This didomain organization is seen in choanoflagellates (e.g., Monosiga, Salpingoeca), while in higher forms it is almost exclusively seen as a fusion of didomain architecture with a DMAP1-binding domain (pentagon in light red) forming a highly conserved three-domain architecture. In fungi (and few plant species such as Quercus and Rhodamnia), the sequence similarity to DMAP1-binding domain is poor, indicated by a serrated pentagon, unlike higher forms. Additional domains such as ARG80, RAMP, PHA03247 family of domains (dotted white box) are occasionally fused to the conserved three-domain architecture. (b) A better sequence relationship with FAALs along with subtle differences in various motifs in opisthokont FAAL-like domains is commensurate with the clustering behavior in the phylogenetic tree generated by different methods. The maximum-likelihood tree implicates a closer relationship of opisthokont FAAL-like domains (pink) to bacterial FAALs (blue) and plant FAALs (green) unlike the NJ-tree; nevertheless, the partitioning of all FAALs away from A-domains (peach) or FACLs (orange) is noted throughout. (c) The FAAL-specific insertion region of FAAL-like-2 shows a comparable length with that of prokaryotic FAALs. (d) The FAAL-specific insertion region of FAAL-like-1 in fungi shows an unusually long insertion as compared to the insertion in prokaryotic FAALs, while the higher forms show comparable length.
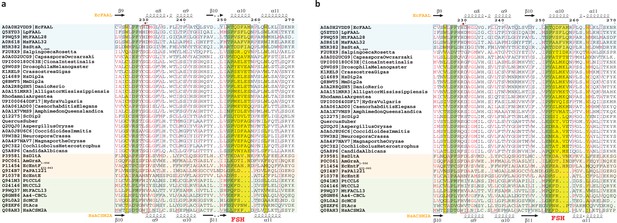

The fatty acyl-AMP ligases (FAAL)-like sequences were identified and aligned to generate a structure-based alignment as described in the Materials and methods section.
The sequences whose structures are available are colored (FAALs in blue, A-domains in light red, and fatty acyl/aryl-CoA ligases [FACLs] in green) and the secondary structures of EcFAAL (PDB: 3PBK; top) along with HsACSM2A (PDB: 3EQ6) are also included. The FAAL-specific helix (FSH) region is marked as a yellow rectangle and labeled accordingly (red). The bulky residues contributing to the CoA-rejection mechanism are highlighted in yellow. The structural sites that correspond to the prolines of the unique alternative pocket in FAALs are highlighted in green. The A4 motif, which is proposed to harbor the residues important for the catalysis of thioesterification, is marked in a red rectangle along with a star. (a) The FSH region of FAAL-like-2 is comparable to that of prokaryotic FAALs, while the additional bulky residues such as Trp/Phe (succeeding FSH) are conserved in higher opisthokonts but not fungal FAAL-like-2 sequences, whereas the Trp (preceding FSH) is completely absent in both fungi and higher opisthokonts. The prolines of the alternative pocket away from FSH appear in a circular mutation in fungi but in no other opisthokonts, while the prolines of the alternative pocket proximal to FSH appear conserved in all opisthokonts. The A4 motif of the FAAL-like-2 is a conserved Cys in higher opisthokont; however, no such conservation in noted in any of the fungal FAAL-like-2 domains. (b) The FSH region of FAAL-like-2 is comparable to that of prokaryotic FAALs, while the additional bulky residues such as Trp/Phe (succeeding FSH) are conserved in higher opisthokonts but not fungal FAAL-like-2 sequences, whereas the Trp/Tyr (preceding FSH) is present in fungi but no other opisthokonts. Similar to FAAL-like-2 domains, the prolines of the alternative pocket away from FSH appear in a circular mutation in fungi but no other opisthokonts, while the prolines of the alternative pocket proximal to FSH appear in circular mutation in higher opisthokonts but not fungi. The A4 motif of the FAAL-like-2 is a conserved Arg in higher opisthokont; however, in the fungal FAAL-like-2 domains, they may be Gln/Glu/His.

The fatty acyl-AMP ligases (FAAL)-like sequences were identified and aligned to generate a structure-based alignment as described in the Materials and methods section.
The sequences whose structures are available are colored (FAALs in blue, A-domains in light red, and fatty acyl/aryl-CoA ligases [FACLs] in green) and the secondary structures of EcFAAL (PDB: 3PBK; top) along with HsACSM2A (PDB: 3EQ6) are also included. The A8-motif and A5-motif, which are proposed to assist domain alternation and binding ribose of ATP respectively, are marked in a red rectangle along with a star(s). The phylogenetic tree was generated as described in the Materials and methods section. The A8-motif of FAAL-like-2 (a) and FAAL-like-1 (b) of higher opisthokonts is comparable in length to a typical A8-motif of ANL superfamily unlike the A8-motif of fungal FAAL-like-1 domains. However, in terms of sequence conservation, both FAAL-like-1 or FAAL-like-2 neither resemble FAALs nor other members of the superfamily. The A5-motif of fungal FAAL-like-2 (c) has a conserved Asp just like all members of the superfamily, but it has been replaced by a conserved Tyr in higher opisthokonts. The A5-motif of all FAAL-like-1 (d) has the conserved Asp replaced by a conserved Leu in all opisthokonts.
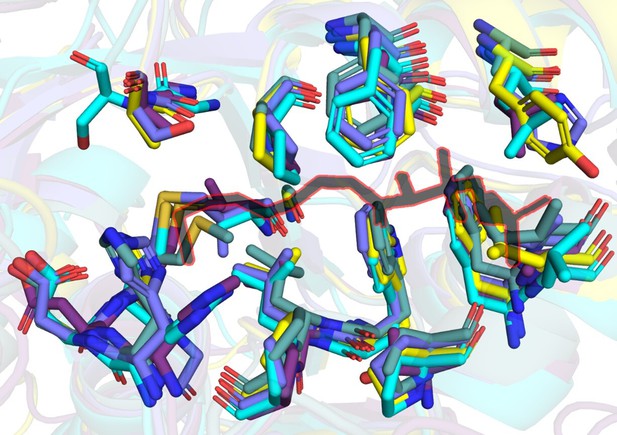
A snapshot of residues (stick representation) in FAALs at 4-5 Å distance from Coenzyme A (black colour outlined in red) bound to SeACS (PDB: 1PG4).
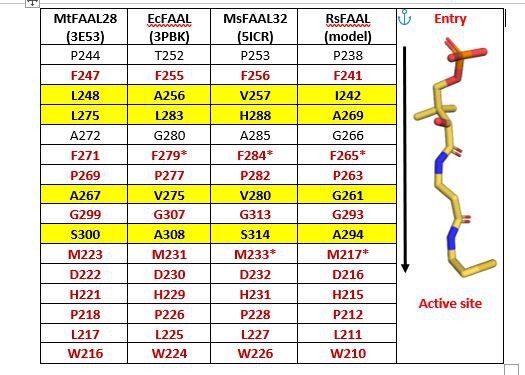
A tabulation of residues in representative FAALs at 4-5 Å distance from Coenzyme A bound to SeACS (PDB: 1PG4).
The stick representation of CoA (yellow) is shown along the table with the entry of the pocket and active site marked.
Additional files
-
Supplementary file 1
List of PDBs of the ANL superfamily family members curated from the RCSB-PDB database.
- https://cdn.elifesciences.org/articles/70067/elife-70067-supp1-v2.docx
-
Supplementary file 2
A tabulation of the number of atoms from the N-terminal domain of fatty acyl-AMP ligases (FAALs) and fatty acyl/aryl-CoA ligases (FACLs) (excluding hydrogens) at a defined distance from the atoms in multiple conformations of coenzyme A (CoA) seen in the CoA-bound structures of FACLs.
The number of atoms of the protein at a clashing distance is an indicator of the space available in the canonical pocket. A higher number of atoms in FAALs indicates the limited space available in the pocket, while the lower number indicates that it is more accommodative in the case of FACLs. Crystal structures of known FACLs have at least one atom.
- https://cdn.elifesciences.org/articles/70067/elife-70067-supp2-v2.docx
-
Supplementary file 3
A tabulation of well-characterized biosynthetic clusters containing fatty acyl-AMP ligases (FAALs) adjacent to polyketide synthase /nonribosomal peptide synthetase (PKS/NRPS) that synthesize important bioactive metabolites.
The phyla of the eubacteria where these clusters are found, the name of the metabolites, the GenBank ID of the biosynthesis cluster, the FAAL (stand-alone or fused) along with their GenBank ID (common name included) and the references where they have been described are detailed.
- https://cdn.elifesciences.org/articles/70067/elife-70067-supp3-v2.docx
-
Supplementary file 4
A tabulation of the various fatty acyl-AMP ligases (FAAL)-like domains and their domain organizations identified in the different lineages of eukaryotes as per the taxonomic distribution provided at NCBI Taxonomy ( http://www.ncbi.nlm.nih.gov/taxonomy).
- https://cdn.elifesciences.org/articles/70067/elife-70067-supp4-v2.docx
-
Supplementary file 5
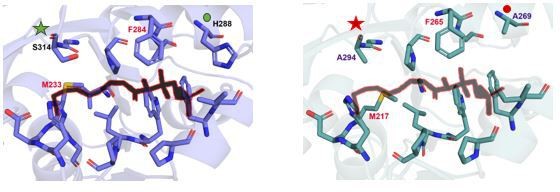
(A) Residues from representative fatty acyl/aryl-CoA ligases (FACLs) (HsFACL, 3EQ6; SeACS, 1PG4; MtFACL13, 3R44; AfFACL, 3G7S; EcFACL, homology model from AlphaFold Protein Structure Database) and fatty acyl-AMP ligases (FAALs) (MtFAAL28, 3E53; MsFAAL32, 5ICR; EcFAAL, 3PBK; RsFAAL, homology model generated using MODELLER) at 4.5 Å distance from coenzyme A (CoA) bound to SeACS (PDB: 1PG4) and HsFACL (PDB: 3EQ6) are identified by structural superposition and tabulated. The FAALs and FACLs included here are those that were used for biochemical analysis along with the availability of their crystal structures. The stick representation of CoA (yellow) is shown along the table with entry and active site marked. The variability of residues along the CoA-binding site results in differential orientation of 4'-phosphopantetheine arm during catalysis. For instance, compare the biochemical profiles of F284A/M233A mutation in MsFAAL32 and F265A/M217A in RsFAAL with respect to residues in the vicinity of the residues chosen for mutation. The conserved phenylalanine in FAALs (F284 of MsFAAL32 and F265 in RsFAAL) is flanked by different residues (H288 in MsFAAL32 and A269 in RsFAAL). Similarly, the conserved methionine of FAALs (M233 of MsFAAL32 and M217 of RsFAAL) is abutted by different residues (S314 in MsFAAL32 and A294 in RsFAAL). (B) A tabulation summarizing the biochemical profiles of various mutations generated in the study. Residue numbers for EcFAAL re used as reference in tables for the ‘gain of function in FAALs through mutations in canonical pocket’ and ‘loss of function in FAALs through mutations in alternate pocket,’ while MtFACL13 is used as reference in tables for the ‘loss of function in FAALs through mutations in canonical pocket.’ A tick (✓) mark indicates that the biochemical activity has been performed in accordance with the hypothesis, which is mentioned in the title of each of the table. A hyphen (-) sign indicates that the mutation did not work as per the hypothesis while 'NA' indicates experiment not performed due to protein expression-related problems.
- https://cdn.elifesciences.org/articles/70067/elife-70067-supp5-v2.docx
-
Supplementary file 6
Primers used for cloning and generating mutants of proteins in this study.
- https://cdn.elifesciences.org/articles/70067/elife-70067-supp6-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/70067/elife-70067-transrepform1-v2.docx