Variation in human herpesvirus 6B telomeric integration, excision, and transmission between tissues and individuals
Figures
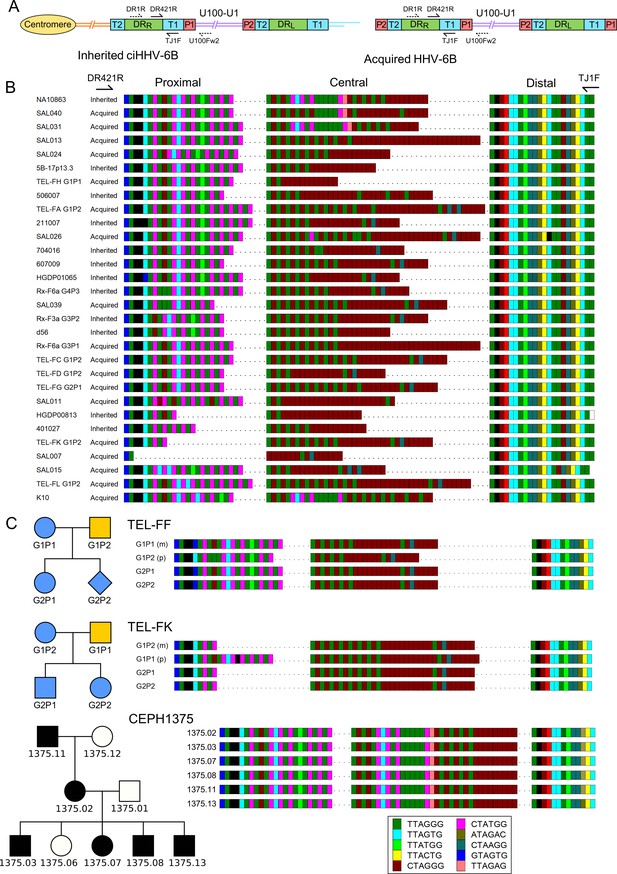
Characterization of the highly variable DRR-pvT1 in iciHHV-6B and acqHHV-6B genomes.
(A) Diagram showing the locations of PCR primers used to amplify the DRR-T1 region specifically (U100Fw2 and DR1R) and the nested primers used to reamplify and sequence pvT1 (DR421R and TJ1F) in the iciHHV-6B genome (left) and in the acqHHV-6B genome (right). (B) DRR-pvT1 repeat patterns are shown to demonstrate diversity among a subset of iciHHV-6B (inherited) and acqHHV-6B (acquired) genomes from various individuals. Telomere-like and degenerate repeats present in the HHV-6B pvT1 region are color-coded as shown in the key and as follows: dark green, TTAGGG; cyan, TTAGTG; yellow, TTACTG; lime green, TTATGG; brown, CTAGGG; pink, CTATGG; dark yellow, ATAGAC; teal, CTAAGG; blue, GTAGTG; peach, TTAGAG; red, GTCTGG. Black squares represent other less common degenerate repeats, white squares show where the sequence could not be determined accurately. Dashes between repeats were added to maximize alignment and to allow comparison between the sections of DRR-pvT1, labeled proximal, central (highly variable), and distal (highly conserved) with respect to the location of the DR421R primer. (C) DRR-pvT1 repeat patterns from acqHHV-6B in two families. None of the family members were iciHHV-6 carriers. In both families, the HHV-6B DRR-pvT1 region in the two children was the same as one parent, indicating that the children carried the same strain of the virus, whereas the second parent carried a different strain. Blue shapes indicate that the child had acqHHV-6B with the same pvT1 repeat map as the mother. Analysis of DRR-pvT1 in the CEPH1375 family showed stable inheritance of the same pvT1 repeat pattern across three generations. iciHHV-6B carriers in the CEPH1375 family are shown as filled black symbols.
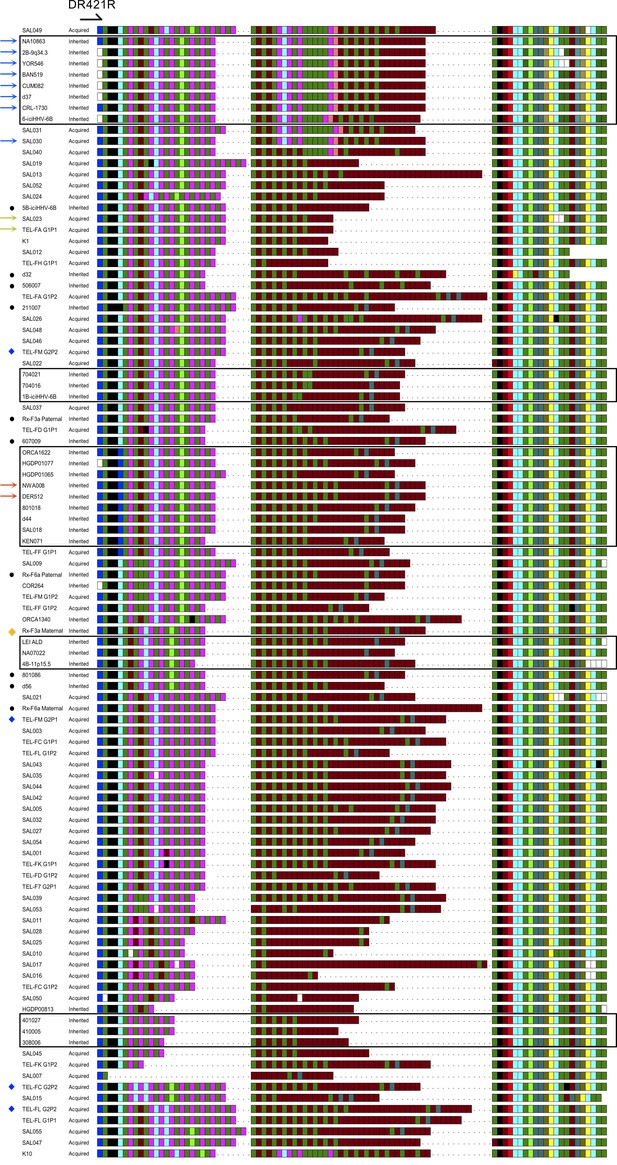
Complete set of pvT1 interspersion maps from DRR in iciHHV-6B (inherited) and acquired HHV-6B (non-inherited) strains.
The 102 DRR-pvT1 repeat patterns shown demonstrate the diversity among iciHHV-6B (inherited) and HHV-6B (acquired) genomes. 90 of the repeat patterns were unique. The TTAGGG and degenerate, telomere-like repeats present in the HHV-6B pvT1 region are color-coded as follows and as shown in Figure 1: dark green, TTAGGG; brown, CTAGGG; cyan, TTAGTG; yellow, TTACTG; dark yellow, ATAGAC; teal, CTAAGG; pink, CTATGG; lime green, TTATGG; blue, GTAGTG; peach, TTAGAG; red, GTCTGG. Black squares represent other, less common degenerate repeats, and white squares show where the sequence could not be determined accurately. Dashes between repeats were added to maximize alignment between samples allowing comparison between the proximal, central (highly variable), and distal (highly conserved) sections of DRR-pvT1. Repeat maps were grouped primarily based on the proximal section, then on less common features within the central or distal region, and finally by the length of the central region. 98 of the 102 DRR-pvT1 repeat patterns are from unrelated individuals and 4 are from children who have an acqHHV-6B that has a different DRR-pvT1 repeat pattern from their parents (blue diamonds). Colored arrows identify 12 repeats patterns found more than once in donors not known to be related. Among eight of the identical DRR-pvT1 repeat patterns (blue arrows), seven are in iciHHV-6B genomes predicted to be integrated in the 9q telomere and one in an acquired HHV-6B in SAL030. Two others, in NWA008 and DER512 from the UK, have a 17p iciHHV-6B integration and share identical DRR-pvT1 repeat patterns. In addition, the acquired HHV-6B strains in SAL023 and TEL-FA G1P1 share the same DRR-pvT1 repeat pattern. Black boxes surround repeat maps from iciHHV-6Bs predicted to have the same integration site, by whole viral genome sequence homology, fluorescent in situ hybridization (FISH), or subtelomere-iciHHV-6B junction sequence. Notably within integration groups there are often shared characteristic features in the DRR-pvT1 repeat patterns. Black dots indicate iciHHV-6B samples where DRR-pvT1 could not be used to predict the integration site confidently. For example, the DRR-pvT1 in iciHHV-6B in d32 has a unique distal section and is unlikely to share common ancestry with any of the other iciHHV-6B carriers analyzed. Yellow diamond identifies the DRR-pvT1 repeat pattern in family Rx-F3a, transmitted from iciHHV-6B mother to non-carrier son. This pvT1 repeat map is different from the other 101 maps shown here.
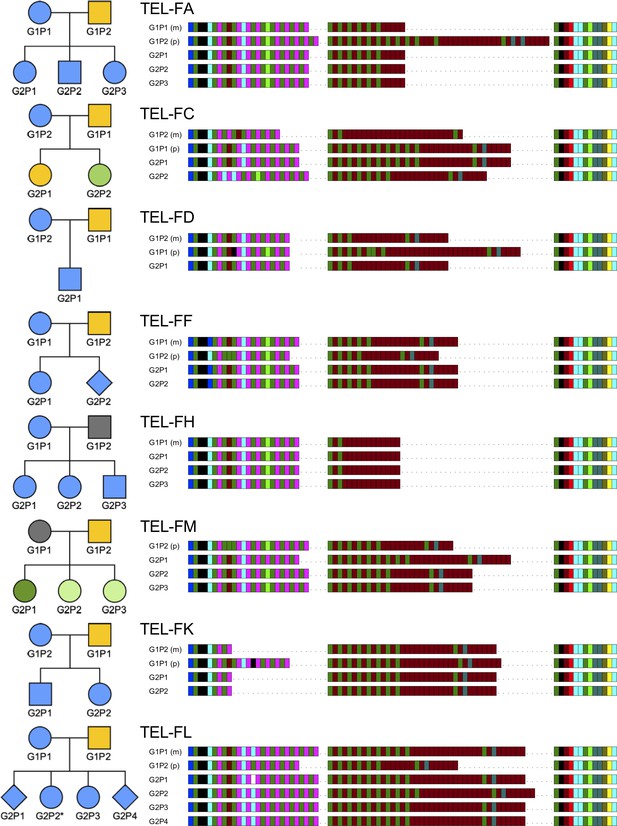
HHV-6B strain identification using DRR-pvT1 repeat patterns in eight families suggests that transmission of acqHHV-6B is predominantly from parents.
The key to the repeat pattern is the same as in Figure 1. Pedigree symbols shaded blue indicate the child(ren) had the same acqHHV-6B DRR-pvT1 repeat pattern as their mother and yellow indicates it was the same as their father. The pedigree symbols shaded gray identify individuals from whom the pvT1 region could not be amplified. The green symbols identify children that do not share a DRR-pvT1 map with either parent, and the dark green symbol for TEL-FM G2P1 indicates that this individual had a DRR-pvT1 that was different to their siblings and parents. TEL-FL G2P2 (asterisk) has a DRR-pvT1 repeat pattern that differs from that in their mother by a gain of two repeats (CTAGGG-TTAGGG) in the central section.
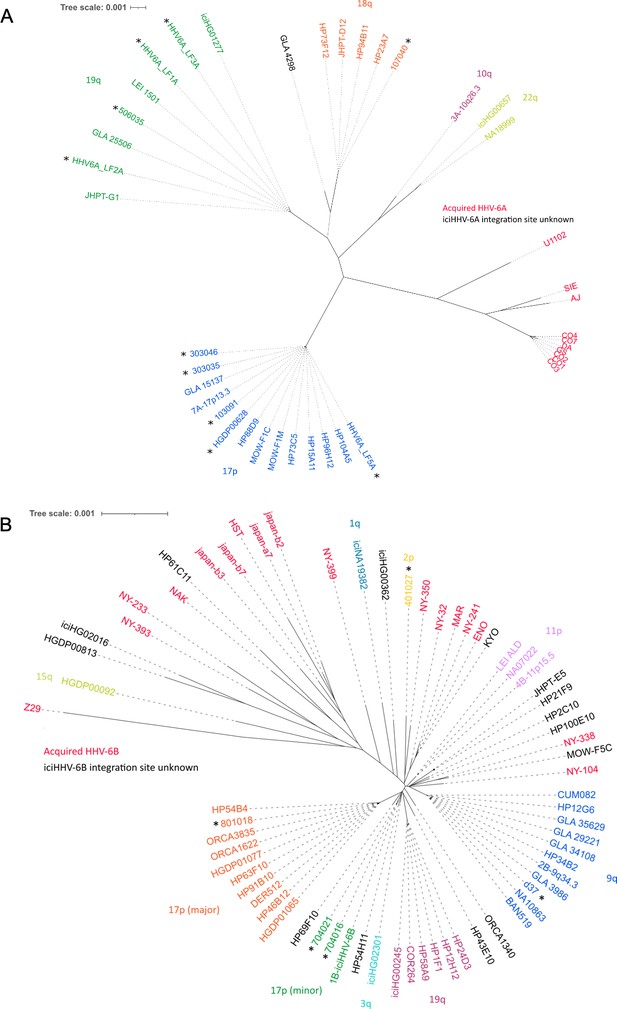
Distance-based phylogenetic networks for HHV-6A and HHV-6B.
(A) A phylogenetic network of 41 HHV-6A viral genome sequences. High-sequence homology to iciHHV-6A genomes for which the integration site had already been established allowed all but one (GLA_4298) iciHHV-6A genomes to be assigned to a telomeric integration site. The colors indicate a chromosomal integration as follows: green, 19q; orange, 18q; purple, 10q; yellow-green, 22q; blue, 17p. The name in black identifies the iciHHV-6A strain without a predicted chromosomal location. Non-inherited, acqHHV-6A strains are shown in red. iciHHV-6A genomes sequenced as part of this study are identified by a black asterisk. (B) A phylogenetic tree of 68 HHV-6B genome sequences (51 iciHHV-6B and 17 acqHHV-6B). iciHHV-6B genomes sequenced as part of this study are identified by a black asterisk. High-sequence homology to iciHHV-6B genomes with a known integration site (already established by fluorescent in situ hybridization [FISH]) allowed the majority of iciHHV-6B genomes to be assigned a telomeric chromosomal location. From the newly sequenced iciHHV-6B genomes, two new clades were identified and provisionally labeled as: 17p (minor) and 2p. The 17p (minor) clade includes the 1B-iciHHV-6B genome, previously a singleton in HHV-6B networks, and the iciHHV-6B genomes in 704021 and 704016. The subtelomere-iciHHV-6B junction sequences, amplified with the 17p311 and DR8F(A/B) primers, are similar for all three samples and distinct form the 17p (major) clade. The other new clade is characterized by the iciHHV-6B genome (401027). This group is provisionally labeled 2p because the subtelomere-iciHHV-6B junction was amplified by the 2p2 and DR8F(A/B) primers in 401027, and two other samples (410005 and 308006, see Figure 3C).
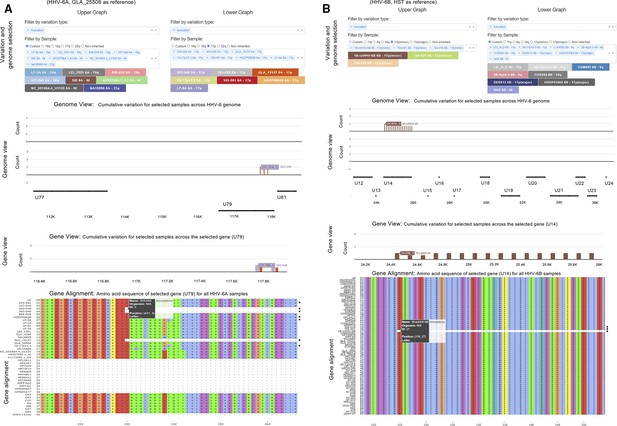
Examples of views taken from the HHV-6 Explorer.
The HHV-6 explorer allows different types of variation across iciHHV-6A/6B and acquired HHV-6A/6B genomes from different individuals to be compared against a reference genome. The user can select the size of overlapping or non-overlapping sliding windows in an upper and lower graph to allow easier comparison when a larger number of genomes are selected from the drop-down menus. After selecting the type(s) of variation and genomes to be explored, an upper and lower Genome View graph will be generated based on the user-selected sliding window size. The Genome View can show the specified type(s) of variation across the whole or part of the genome as a count of mutations per window size (100 bp, non-overlapping window in the examples shown). Cumulative mutations per window size are shown if multiple types of variation are selected. Each genome is assigned a unique color, and where part of sequence is missing, a value of negative one will be assigned for that particular window and the bar on the graph will be red. (A) In the examples shown, iciHHV-6A and acquired HHV-6A genomes have been compared to GLA_25506 (19q iciHHV-6A) as the reference. The left and right top panels allow different samples to be selected for comparison to the reference, and in both examples, truncations (i.e., a nonsense mutation encoding a premature stop codon) are displayed. In the examples shown, seven 17p iciHHV-6A genomes have been selected to be displayed on the lower graph and a variety of acquired HHV-6A and 18q, 19q, and 22q iciHHV-6A genomes have been selected to be displayed on the upper graph. The Genome View is zoomed in to show a region displaying truncations in U79 in three 17p iciHHV-6A genomes (303-046, 303-035, and GLA_15137). In the HHV-6 Explorer, the next graph generated is the Gene View where a single gene of interest can be selected to view variation at the amino acid level, U79 in this case. The final plot on the HHV-6 Explorer is the Gene Alignment, based on an alignment of all iciHHV-6A and acquired HHV-6A used to generate the phylogenetic networks. In this region of U79, the sequence assemblies are missing data in 13 of the viral genomes (dashes). The exact amino acid position of the premature stop codon for 303-035 is highlighted, underscored at amino acid 431. All other iciHHV-6A/HHV-6A genomes (for which sequence is available) lack this premature stop codon, including other 17p iciHHV-6A genomes (e.g., 7A-17p13.3), indicating that this mutation arose after integration. Black dots identify the 17p-iciHHV-6A samples with sequence across U79. (B) iciHHV-6B and acquired HHV-6B genomes have been compared to HST (acquired HHV-6B), a reference strain. The images taken from the HHV-6 Explorer are displayed as for panel (A). Members of the iciHHV-6B 17p (minor) integration group are displayed in the upper graphs, and a variety of other iciHHV-6B integrations and acquired HHV-6B genomes are displayed on the lower graphs. A premature stop codon is present at position 116 in U14 in 1B-iciHHV-6B but not in 704-021 or 704-016, which share the same common ancestor. Again, this indicates that the nonsense mutation arose after integration. Black dots identify the 17 p(minor)-iciHHV-6B genomes.
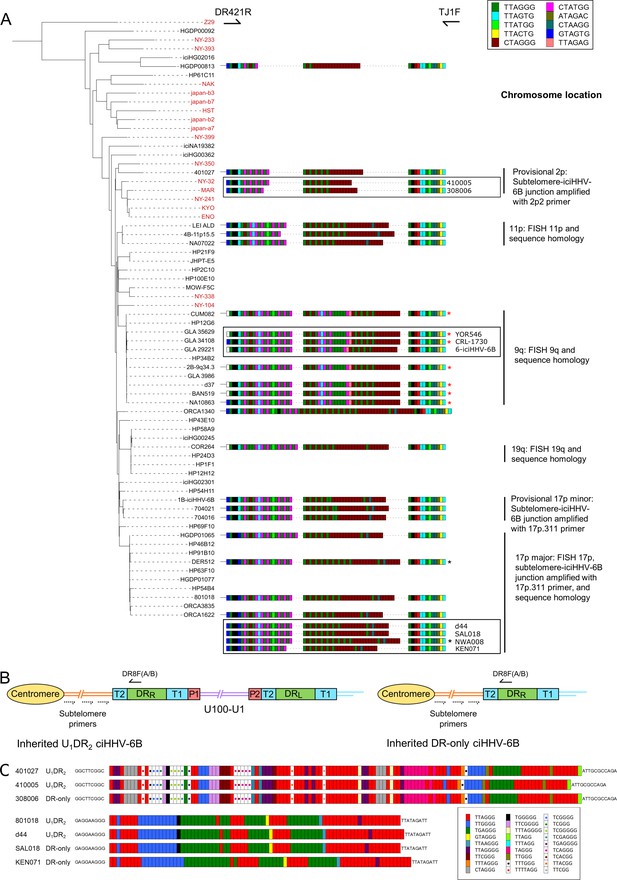
pvT1 repeat patterns are similar between strains of iciHHV-6B known to have the same ancestral integration site and those predicted to share that integration site.
(A) Distance-based phylogenetic tree of selected iciHHV-6B (black names) and acquired HHV-6B (red names) genomes. Where DNA was available, DRR-pvT1 repeat patterns from 19 iciHHV-6B samples in the phylogenetic tree are shown. The repeats in the pvT1 region are color-coded as for Figure 1 and shown in the key, top right. Black vertical bars on the right indicate groups of iciHHV-6Bs for which the integration site has been determined by fluorescent in situ hybridization (FISH), phylogeny, subtelomere-iciHHV-6B junction analysis, or a combination of these. In addition, red asterisks identify samples from the 9q-iciHHV-6B clade that share identical pvT1 repeat patterns. pvT1 repeat patterns are more variable between samples that belong to the 17p (major) iciHHV-6B clade with black asterisks identifying samples that share identical pvT1 patterns. DRR-pvT1 repeat patterns within black boxes identify iciHHV-6B samples for which the integration site has been predicted based on DRR-pvT1 similarity. For some of these samples, the integration site has been validated by subtelomere-iciHHV-6B junction analysis (410005, 308006, d44, SAL018, NWA008, and KEN071). (B) Line diagram showing the position of DR8F(A/B) and potential subtelomere primers used to amplify and sequence subtelomere-iciHHV-6B junctions. The subtelomere-iciHHV-6B integration site can be amplified whether the integrated viral genome is full length or iciHHV-6B-DR only. (C) Diagram showing the pattern of canonical (TTAGGG) and degenerate telomere-like repeat maps across the subtelomere-HHV-6B junction at two different chromosome ends. The subtelomere-HHV-6B repeat maps from 401027, 410005, and 308006 were generated following amplification with primers 2p2 and DR8F(A/B). They are very similar demonstrating common ancestry (top three repeat maps). The subtelomere-HHV-6B repeat maps from 801018, d44, SAL018, and KEN071 generated by amplification with 17p311 and DR8F(A/B) (bottom four maps) are very similar to the group of iciHHV-6B genomes in the 17p (major) clade some of which have been shown to be integrated in 17p. The 308006, SAL018, and KEN071 samples contain iciHHV-6B DR-only. The key (bottom right) shows the color-coding for the diverse degenerate repeats found across the subtelomere-HHV-6B repeat maps.
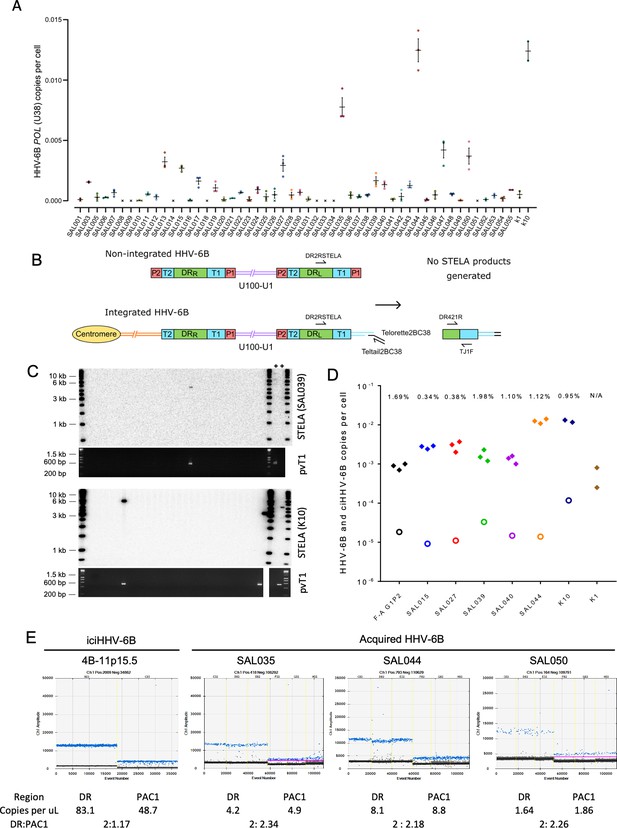
Evidence of telomere formation at the distal end of HHV-6B genomes in individuals with community-acquired HHV-6B.
(A) Graph showing HHV-6B copy number per cell estimated from HHV-6B POL (U38) vs. RPP30 duplex digital droplet PCR (ddPCR). Triplicate results are shown for saliva DNA from 52 individuals and kidney samples from 2 individuals. Crosses show where zero copies per cell were estimated for a single replicate, and black crosses indicate that zero copies per cell were estimated from all three replicates. The mean and standard error are shown. (B) Single TElomere Length Analysis (STELA) primers, DR2RSTELA and Telorette2BC38/TeltailBC38, that were used to amplify HHV-6B-associated telomeres are shown on an integrated HHV-6B genome. DR421R and TJ1F primers were used to amplify DRL-pvT1 in secondary nested PCRs. In non-integrated HHV-6B, STELA products cannot be generated as there is no HHV-6B-associated telomere. The copies of PAC1 (P1) and PAC2 (P2) are shown. (C) Examples of HHV-6 STELA from two DNA samples (SAL039 and K10) with acqHHV-6B. Upper panels show the outcome of 30 STELA reactions followed by agarose gel electrophoresis and Southern blot hybridization to radiolabeled (TTAGGG)n telomere-repeat probe. In a small number of reactions, an amplified HHV-6B-associated telomere band was detected. An aliquot of the products from each primary STELA PCR was diluted and used as input for a secondary, nested PCR to amplify DRL-pvT1. As shown in the agarose gel photograph below, DRL-pvT1-positive reactions corresponded to the STELA reactions that generated HHV-6B-associated telomere band. The 6-iciHHV-6B DNA was used as a positive control (+) for STELA and DRL-pvT1 secondary PCR. (D) Graph showing the total copy number of HHV-6B per cell measured by ddPCR (filled diamonds); the estimated number of copies of integrated HHV-6B per cell from STELA and pvT1 PCR (open circles). (E) Measuring the DR:PAC1 ratio by ddPCR. 1D EvaGreen ddPCR plots of one iciHHV-6B (4B-11p15.5) cell line and acqHHV-6B in three saliva samples (SAL035, SAL044, and SAL050). Triplicate reactions for a DR amplicon (DR6B-F and DR6B-R primers) and a PAC1 amplicon (PAC1F and PAC1R-33 primers) are shown for each sample. The absolute copy number of each amplicon per μL ddPCR reaction and the ratio between DR and PAC1 amplicons are shown below each plot.
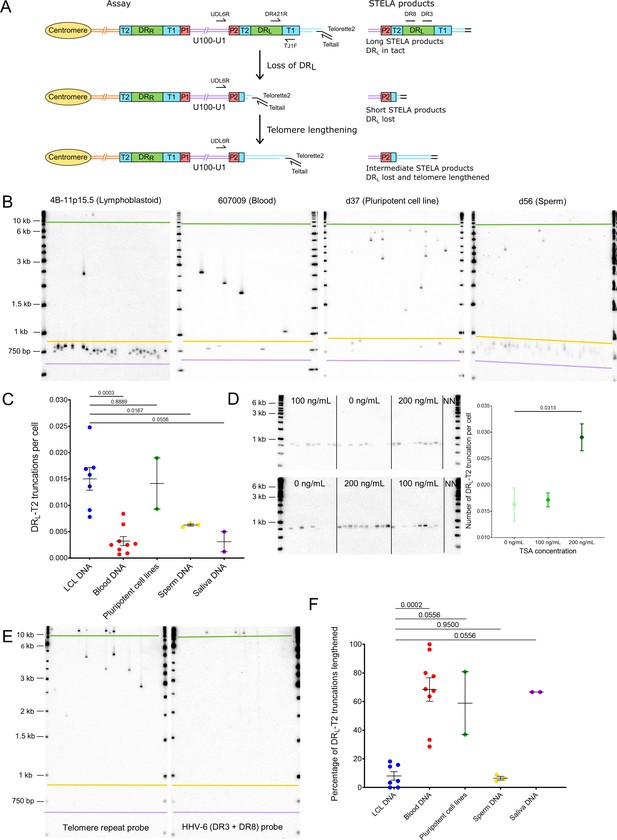
Quantifying iciHHV-6B truncations at DRL-T2 and assessment of the newly formed telomere length.
(A) Diagram showing the loss of DRL from iciHHV-6B, formation of a novel telomere at DRL-T2, and the potential lengthening of the newly formed telomere by a telomere maintenance mechanism. Single TElomere Length Analysis (STELA) primers (Teltail and Telorette 2) and the primer UDL6R, in the unique region of the iciHHV-6B genome, were used to amplify HHV-6B-associated telomeres. Long STELA products (>8.6 kb) are generated from intact copies of the iciHHV-6B genome and include amplification through DRL to the end of the telomere. Short STELA products (0.7–0.9 kb) are generated when DRL has been lost and a novel telomere has formed at DRL-T2. Intermediate sized STELA products are generated when the novel telomere formed at DRL-T2 has been lengthened. All three types of STELA amplicons contain TTAGGG telomere repeats and hybridize to a telomere repeat probe. The relative positions of the DR3 and DR8 probes used to detect DR sequences are shown. (B) Representative Southern blots exhibiting amplicons generated by HHV-6B STELA with the UDL6R flanking and Telorette 2 primers. Each blot displays products generated from 24 to 32 parallel STELA PCRs in different cell types from iciHHV-6B carriers. These include a lymphoblastoid cell line (LCL) from individual 4B-11p15.5; white blood cells from individual 607009; a pluripotent cell line d37; sperm DNA d56. Amplicons detected between the mauve and yellow lines are short STELA products (expected size range 0.7–0.9 kb). The bands detected between the yellow and green lines are intermediate length STELA products, representative of newly formed telomeres at DRL-T2 that have been lengthened. Long STELA amplicons, above the green line, are products generated from full-length copies of iciHHV-6B and include DRL. (C) Graph showing the estimated number of DRL-T2 truncations per cell, including those that have been lengthened, in DNA from different iciHHV-6B individuals and cell types. Means with standard error and the p-values from Mann–Whitney ranked sum test to compare truncation frequencies (LCL vs. other cell types) are shown. (D) Representative UDL6R-STELA blots from the 4B-11p15.5 cell line treated with 0, 100, and 200 ng/mL of trichostatin-A (10 parallel STELA PCRs per condition) and a graph showing the frequency of truncations per cell (mean and standard error). Data were derived from UDL6R-STELA conducted twice (technical replicates) using DNA extracted from treated and untreated cells grown in triplicate (biological replicates). p-values were from a Wilcoxon test. (E) DRL-T2 truncation assay on blood DNA from the 401027 iciHHV-6B carrier demonstrating that the long STELA products (above the green line) contain DR sequences that hybridize to the telomere repeat probe (left panel) and the combined DR3/DR8 probe (right panel). Whereas the intermediate length amplicons (between yellow and green lines) only hybridize to the telomere repeat probe (left panel) demonstrating that each amplicon represents newly formed telomere at DRL-T2 that has been lengthened. The two Southern blots display products detected across 24 parallel STELA PCRS. No short amplicons (0.7–0.9 kb) were detected in this replicate of the assay for this sample. (F) The graph shows the percentage of newly formed telomeres at DRL-T2 that were lengthened in samples from various unrelated iciHHV-6B carriers and cell types. Means with standard error and the p-values from Mann–Whitney ranked sum test to compare truncation frequencies (LCL vs. other cell types) are shown.
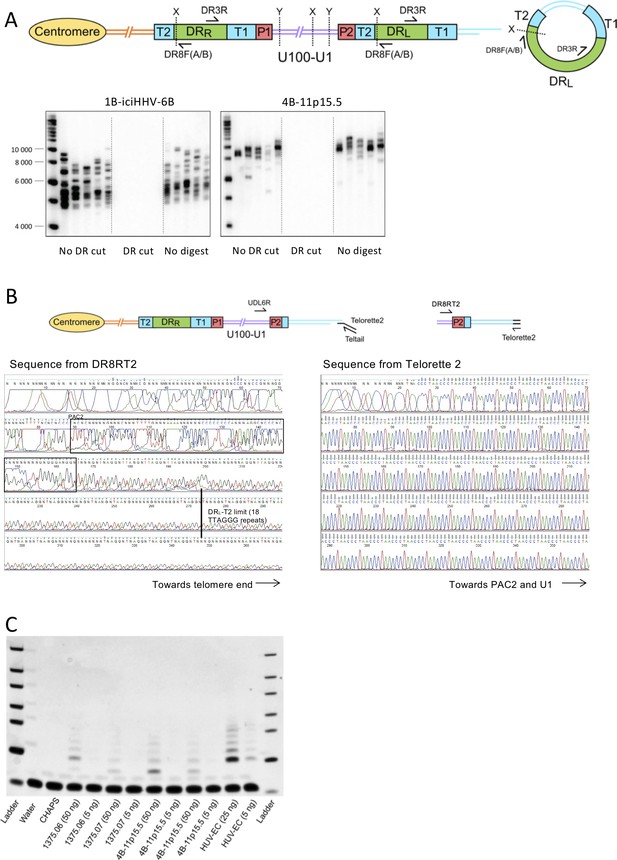
Detection of excised circular DR-only molecules, sequencing of a lengthened new telomere at DRL-T2, and detection of telomerase activity.
(A) Schematic of DR circle assay. DR circles were amplified using primers (DR8F(A/B) and DR3R) that should not generate a PCR product from full-length iciHHV-6B but should amplify across telomere repeats in a DR circle, the predicted reciprocal product of a t-loop mediated excision event at DRL-T2. Genomic DNA from two iciHHV-6B individuals (1B-iciHHV-6B and 4B-11p15.5) was digested with combinations of restriction enzymes that cut in the unique region and DR (X: SacI and PstI), only the unique region (Y: XbaI and ScaHF), or were not treated with restriction enzymes. Treated DNA was amplified using DR8F(A/B) and DR3R, and PCR products were size-separated and detected by Southern blot hybridized to a radiolabeled (TTAGGG)n telomere probe. Amplicons of variable length were detected reflecting different lengths of telomere repeat arrays expected in individual DR circles. Restriction digestion with SacI/PstI, including between DR8F(A/B) and T2, prevents amplification from DR circles. Importantly, the minimal size of amplicons detected was greater than the combined length of the flanking regions. Full-length 4B-11p15.5 had longer average telomere length at DRL-T1 than 1B-iciHHV-6B, which is consistent with the longer products generated in the DR circle assay. (B) The schematic shows how newly formed telomeres at DRL-T2 were detected and sequenced. UDL6R-STELA was used to amplify telomeres at DRL-T2, and it occasionally amplified telomeres that were longer than the length of DRL-T2. Six of these intermediate length STELA products from three different DNA samples were reamplified in a semi-nested, secondary PCR using primer DR8RT2 and Telorette 2. Following gel extraction, these products were Sanger sequenced using DR8RT2 or Telorette 2. The TTAGGG repeats were visualized with FinchTV and counted manually. Electropherograms from one reamplified molecule from NWA008 are shown. As expected, the PAC2 motif was present (boxed sequence). A black line at base 273 shows where DRL-T2 in this sample was expected to end (after 18 TTAGGG repeats). Over 100 telomere repeats were counted from Telorette 2, considerably more than the length of T2, showing that this telomere has been lengthened. (C) Detecting telomerase activity in iciHHV-6B lymphoblastoid cell lines. Telomere repeat amplification protocol (TRAP) was used to detect low levels of telomerase activity in various iciHHV-6B lymphoblastoid cell lines. Water and CHAPS were used as negative controls, and cell lysate from a telomerase positive HUV-EC cell line was used as a positive control.
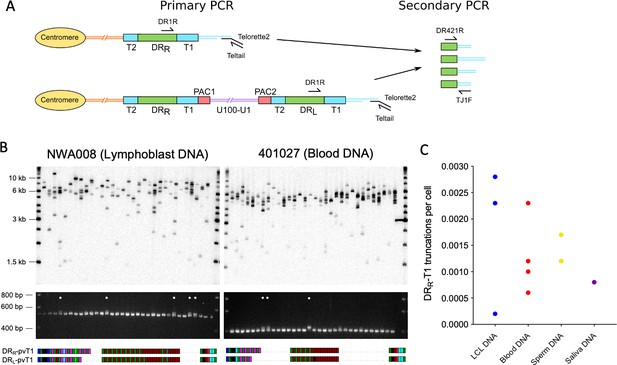
Measuring the frequency of iciHHV-6B truncations at DRR-T1 associated with novel telomere formation.
(A) Schematic of the two-step assay used to differentiate between iciHHV-6B-associated telomeres at DRR-T1 and DRL-T1. Telomeres at DRR-T1 and DRL-T1 were PCR amplified using the Single TElomere Length Analysis (STELA) primers, DR1R with Telorette 2/Teltail. Subsequently, secondary nested PCRs were used to amplify pvT1 (primers DR421 and TJ1F) followed by size separation to distinguish pvT1 from DRR-T1 or DRL-T1. (B) Detection of telomeres at DRR-T1 or DRL -T1 in two iciHHV-6B samples (NWA008 and 401027) with known length differences between pvT1 in the DRs. Top images show telomere amplicons derived from DRL and DRR in the iciHHV-6B samples. The telomeres were amplified in 32 parallel STELA PCRs using the DR1R flanking primer and detected by Southern blot hybridization to a radiolabeled (TTAGGG)n probe. Lower images show ethidium bromide-stained agarose gels of size-separated pvT1 sequences amplified from the corresponding STELA reaction in track shown in the panel above. White dots identify the PCR reactions that contain pvT1 from DRR-T1. Below each set of panels are the pvT1 repeat patterns from DRR-T1 or DRL-T1 for the corresponding sample, highlighting the differences. (C) Graph showing the frequency of truncations and new telomere formation at DRR-T1 in iciHHV-6B samples from unrelated individuals and cell types. The number of truncations per cell was calculated using the number of reactions from which DRR-pvT1 was amplified divided by the cell equivalents estimated from the total quantity of DNA screened (6.6 pg or 3.3 pg DNA per cell was assumed for diploid and haploid cells, respectively).

The majority of iciHHV-6B genomes show sequence differences between DRR-pvT1 and DRL-pvT1.
(A) DRR-T1 and DRL-T1 were specifically amplified by PCR using DR1R and U100Fw2 for DRR-T1, or by Single TElomere Length Analysis (STELA) using DR1R and STELA oligonucleotides (Teltail and Telorette 2) for DRL-T1. The products from DRR-T1 and DRL-T1-specific PCRs were used as input for secondary, nested PCR using DR421R and TJ1F to specifically amplify DRR-pvT1 and DRL-pvT1. Sanger sequencing was carried out using the TJ1F primer. (B) Sequences were manually examined to generate pvT1 repeat patterns. Repeats patterns were color coded and aligned as in Figure 1 and Figure 1—figure supplement 1. The majority of iciHHV-6B genomes (24/35, 68.6%) showed differences between DRR-pvT1 and DRL-pvT1, usually as loss or gain of a small number of hexameric telomere (TTAGGG) or degenerate telomere-like repeats. In a smaller number of cases, a single base change converted one repeat type to another. Regions containing CTAGGG repeats were most prone to loss or gain of repeats. Where differences between DRR-pvT1 and DRL-pvT1 were detected in multiple iciHHV-6B genomes from the same integration group (boxed), the differences were rarely the same between individuals. This indicates that the differences arose after integration. Interestingly, 9q iciHHV-6B genomes did not display any variation between DRR-pvT1 and DRL-pvT1.
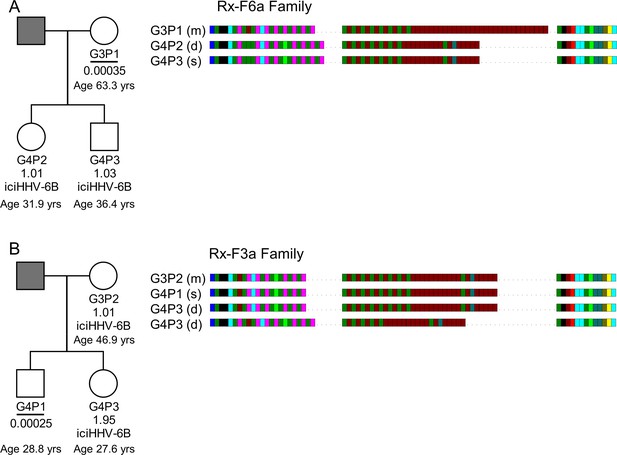
HHV-6B copy number and DRR-pvT1-repeat patterns can be used to identify potential iciHHV-6B reactivation and transmission within families.
(A) In family Rx-F6a, both children are iciHHV-6B carriers with approximately one copy per cell. They share the same DRR-pvT1 repeat map, presumably inherited from the father (gray filled square, not available for testing). The mother has a low level of acqHHV-6B in her saliva (0.00035 copies per cell) and the DRR-pvT1 repeat map is different from her children. (B) Evidence of iciHHV-6B reactivation in family Rx-F3a and transmission to non-iciHHV-6B son. Left shows the Rx-F3a family tree with HHV-6 copy number per cell in saliva DNA and iciHHV-6B carrier status. Father (gray filled square) was not available for testing. Right: the DRR-pvT1 repeat patterns from family members. Daughter (G4P3) has two copies of iciHHV-6B, one copy shares the same DRR-pvT1 as seen in her mother (G3P2) and a second copy has a different DRR-pvT1 repeat pattern assumed to have been inherited from her father. The son (G4P1) has a very low level of HHV-6B in his saliva with the same pvT1 repeat pattern as the maternal iciHHV-6B genome. The DRR-pvT1 repeat patterns are also labeled with m (present in mother), d (present in daughter), and s (present in son). The age of tested family members is stated.
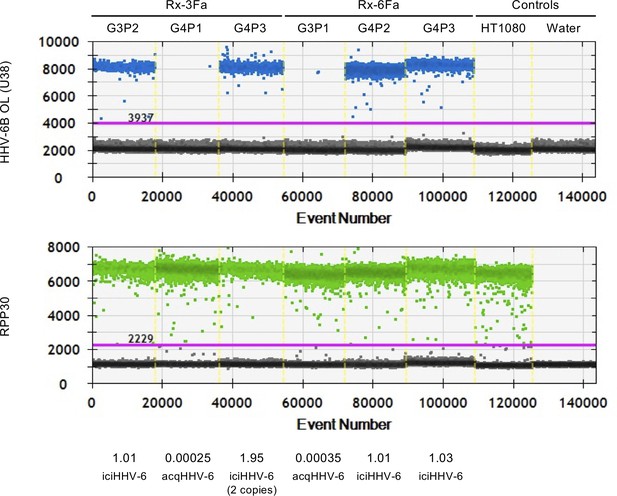
Quantification of HHV-6B using POL (U38) digital droplet PCR (ddPCR) in two families (Rx-3FA and Rx-6Fa).
HHV-6B copy number was estimated in Rx-3Fa and Rx-6Fa family members using a duplex hydrolysis probe ddPCR assay. 1D ddPCR plots for HHV-6B U38 and human reference RPP30 for three individuals in each family are shown. Each point represents a single droplet in which amplification of the target has occurred (blue or green) or has not occurred (gray). HHV-6B free cell line HT1080 and water were used as negative controls. Estimated copy numbers are shown below plots along with HHV-6B status.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (Homo sapiens) | Lymphoblastoid (LCL) | Various, Supplementary file 1 | Various, see Supplementary file 1 | |
| Cell line (Homo sapiens) | Pluripotent cell line | ATCC | CRL-1730(RRID:CVCL_2959) | See Supplementary file 1 |
| Cell line (Homo sapiens) | Pluripotent cell line | Dr Rai, University of Leicester | d37 | See Supplementary file 1 |
| Biological sample (Homo sapiens) | DNA from saliva | Various, see Supplementary file 1 | SAL001 to SAL055; TEL-FA(code) to TEL-FL(code); Rx-F6a; RX-F3a | |
| Biological sample (Homo sapiens) | DNA from blood | Various, see Supplementary file 1 | Various, see Supplementary file 1 | |
| Biological sample (Homo sapiens) | DNA from sperm | d32, d44, d56 | Various, see Supplementary file 1 | |
| Sequence-based reagent | Oligonucleotides and primers | See Supplementary file 7 | See Supplementary file 7 | |
| Sequence-based reagent | Hydrolysis probes | See Supplementary file 7 | See Supplementary file 7 | |
| Chemical compound, drug | Trichostatin A | Sigma-Aldrich | T8552 | |
| Commercial assay or kit | ddPCR Supermix for probes without dUTP | Bio-Rad Laboratories | 1863023 | |
| Commercial assay or kit | RPP30 primer/HEX labeled probe mix | Bio-Rad Laboratories | 10031243 | |
| Commercial assay or kit | ddPCR EvaGreen Supermix | Bio-Rad Laboratories | 1864033 | |
| Commercial assay or kit | Zymoclean gel DNA recovery kit | Zymo Research, Irvine, CA | D4002 | |
| Software, algorithm | HHV-6 Explorer | This paper; https://www.hhv6explorer.org/ | https://github.com/colinveal/HHV6-Explorer. | |
| Software, algorithm | Prism software (version 9.0) | GraphPad Software |
Additional files
-
Supplementary file 1
AcqHHV-6A/6B and iciHHV-6A/6B genomes included in study.
- https://cdn.elifesciences.org/articles/70452/elife-70452-supp1-v2.xlsx
-
Supplementary file 2
Estimated Time to Most Recent Common Ancestor (TMRCA) for carriers of iciHHV-6A and iciHHV-6B with different chromosomal locations.
- https://cdn.elifesciences.org/articles/70452/elife-70452-supp2-v2.docx
-
Supplementary file 3
Distinctive features of DRR-pvT1 repeat patterns associated with various iciHHV-6B phylogenetic clades.
- https://cdn.elifesciences.org/articles/70452/elife-70452-supp3-v2.docx
-
Supplementary file 4
Measuring the percentage of acquired HHV-6B with a telomere, as an indicator of integration.
- https://cdn.elifesciences.org/articles/70452/elife-70452-supp4-v2.docx
-
Supplementary file 5
Variation in the frequency of truncations at DRL-T2 and percentage lengthened between samples.
- https://cdn.elifesciences.org/articles/70452/elife-70452-supp5-v2.docx
-
Supplementary file 6
Measuring the frequency of truncations at DRR-T1 in various samples.
- https://cdn.elifesciences.org/articles/70452/elife-70452-supp6-v2.docx
-
Supplementary file 7
Primers used in this study, including primers used to generate overlapping amplicons for iciHHV-6A/6B genome sequencing.
- https://cdn.elifesciences.org/articles/70452/elife-70452-supp7-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/70452/elife-70452-transrepform1-v2.pdf





