The Arabidopsis SAC9 enzyme is enriched in a cortical population of early endosomes and restricts PI(4,5)P2 at the plasma membrane
Figures
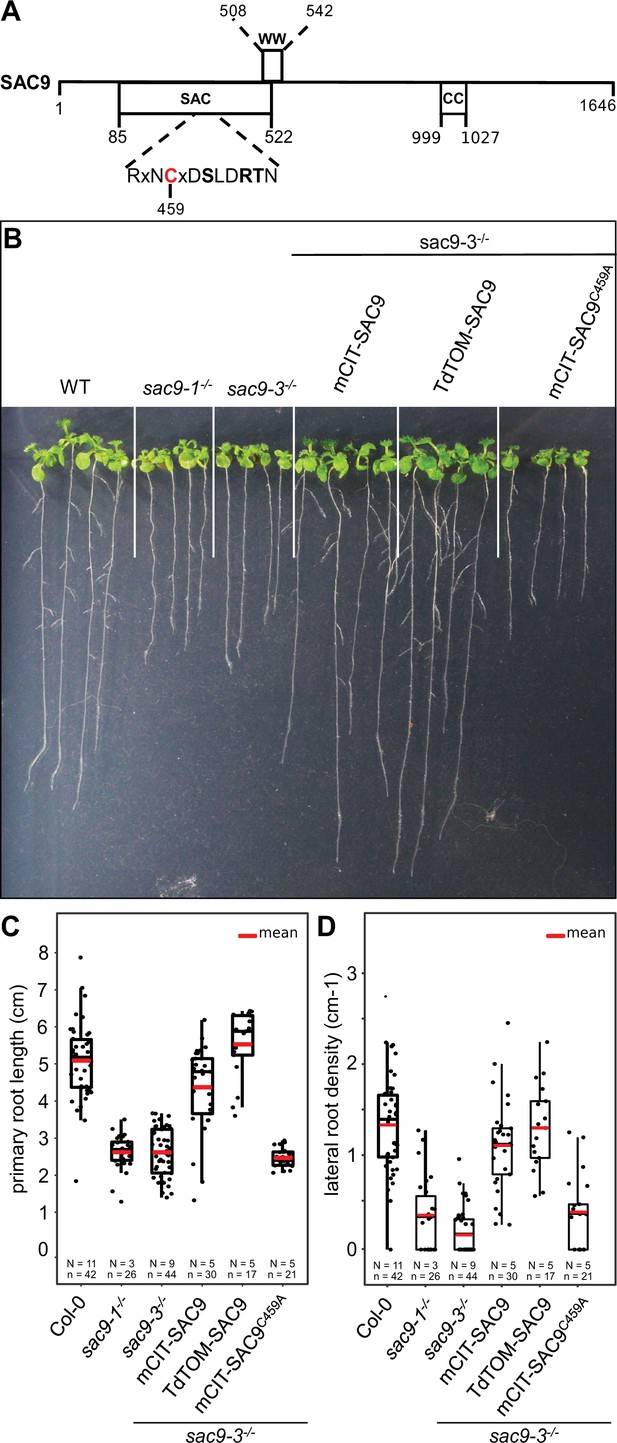
Structure-function analysis of SUPPRESSOR OF ACTIN9 (SAC9).
(A) Schematic representation of SAC9 protein. The SAC catalytic domain, as well as the WW domain and the coil-coiled domain, are represented. (B) Representative images of the macroscopic phenotype observed in (i) wild-type (Col-0), (ii) sac9-1-/- and sac9-3-/- loss of function mutants, (iii) sac9-3-/- complemented lines expressing full-length genomic DNA encoding SAC9 fused to yellow (mCIT-SAC9, line #1000-9-1) or red (TdTOM-SAC9, line #987-5-4) fluorescent proteins and a mutated version of the putative catalytic cysteine residue within the C-x(5)-R-[TS] catalytic motif in the SAC domain (mCIT-SAC9C459A, line #1354-12-14; right panel). Pictures are taken 12 days post germination (dpg). Note that a second independent transgenic line is presented for each construct in Figure 1—figure supplement 1. (C) Quantification of primary root length in sac9-3-/- homozygous mutants expressing mCIT-SAC9 and mCIT-SAC9C459A under the control of the native promoter (SAC9prom) and TdTOM-SAC9 under the expression of the Ubq10 promoter. Wild-type (Col-0) seedlings and two independent mutant alleles of SAC9, sac9-1-/-, and sac9-3-/-, are used as controls. (D) Same as (C) but for the quantification of the lateral root density (ratio of the number of lateral roots to primary root length). In the plots, middle horizontal bars represent the median, while the bottom and top of each box represent the 25th and 75th percentiles, respectively. At most, the whiskers extend to 1.5 times the interquartile range, excluding data beyond. For range of value under 1,5 IQR, whiskers represent the range of maximum and minimum values. Details for statistical analysis can be found in the Methods section and Supplementary file 1C. N=number of replicates; n=number of roots.

Complementation analysis of the sac9-3 mutant phenotype.
(A) Representative images of the complementation analysis of the sac9-3 mutant. For each construct, 24 primary T1 transformants were selected. Independent T2 and T3, T4 plants were obtained for subsequent analysis. Line 1000-9-1; 987-5-4; 1354-12-14 used for the quantitative analysis of sac9 complementation are shown in Figure 1. Example of the phenotype observed in other independent transgenic lines 1000–10 h2; 987-14-5; 1354-15-11, as well as two independent transgenic lines 1365-1-5-3 and 1356-4-1 are presented. (B) Immunoblot showing the expression of the various SUPPRESSOR OF ACTIN9 (SAC9) variants in independent transgenic lines (the number of the lines shown corresponds to the line presented in panel A and in Figure 1). mCIT-SAC9 and its respective mutants were detected by immunoprecipitation using an anti-GFP antibody. Ponceau S staining serves as a loading control. The experiment was repeated three times with similar results. N=number of roots, n=number of cells.
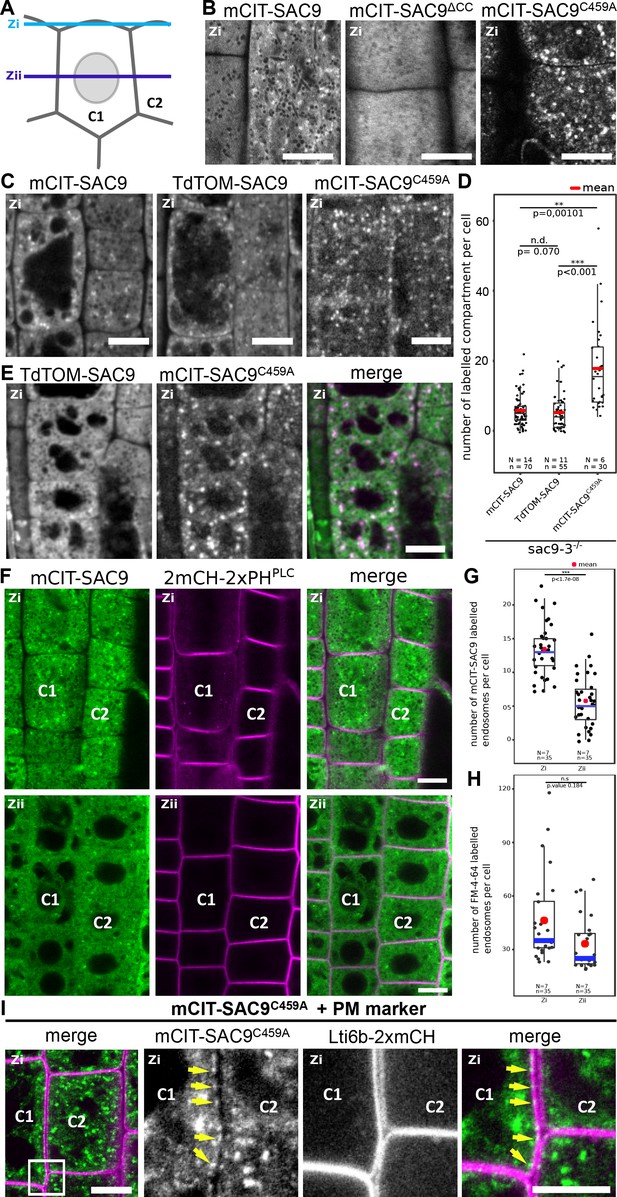
SUPPRESSOR OF ACTIN9 (SAC9) localizes in the cytosol and in intracellular compartment enriched at the cell cortex.
(A) Schematic representation of two root epidermal cells (cell #1 indicated as C1 and cell #2 indicated as C2) imaged at two different focal planes (cortical plane, close to the plasma membrane and designated as Zi and a median plane, in the middle of the cell designated as Zii). Note that the C1/C2 and Zi/Zii notations are used consistently according throughout the figures. (B) Confocal images of the subcortical part (i.e. Zi) of the Arabidopsis root epidermis expressing mCIT-SAC9, mCIT-SAC9∆CC mutated in the predicted coil-coiled domain, and mCIT-SAC9C459A mutated in its putative catalytic cysteine, under the control of SAC9 native promoter (SAC9prom). (C) Representative images of the fluorescent signal observe in sac9-3-/- mutant expressing mCIT-SAC9, TdTOM-SAC9, and mCIT-SAC9C459A. (D) Comparison of the number of labeled intracellular compartments per cell in sac9-3-/- root epidermis expressing mCIT-SAC9 and TdTOM-SAC9, mCIT-SAC9C459A. (E) Colocalization analysis on Ub10pro:TdTOM-SAC9 (magenta) and SAC9pro:CIT-SAC9C459A (green). (F) Confocal images of the subcortical part of the Arabidopsis root epidermis (upper panels, Zi) or the center of the cell (lower panels, Zii) expressing mCIT-SAC9 together with the PI(4,5)P2 biosensor 2mCH-2xPHPLC (note that the cells shown in Zi and Zii are the same, just on different focal plane). (G), Quantification of the number of endosomes labeled by mCIT-SAC9 observed per cell at two different focal planes (Zi and Zii). (H), Quantification of the number of endosomes labeled by FM4-64 observed per cell at two different focal planes (Zi and Zii). (I), Co-visualization of SAC9pro:mCIT-SAC9C459A (green) together with the plasma membrane marker Lti6b-2xmCH (magenta) in epidermal root cells taken in the Zi focal plane. The inset shows a magnification, with the yellow arrows indicating some of the intracellular compartments decorated by mCIT-SAC9C459A observed at the close vicinity to the labeled plasma membrane in cell C1. Scale bar: 10 µm in A-F; 5 µm in G (left panel) and 2 µm in the close-up of G (right panel). In the plot, middle horizontal bars represent the median, while the bottom and top of each box represent the 25th and 75th percentiles, respectively. At most, the whiskers extend to 1.5 times the interquartile range, excluding data beyond. For range of value under 1,5 IQR, whiskers represent the range of maximum and minimum values. Details of the statistical analysis could be found in Supplementary file 1D. The plane (Zi or Zii) in each image is mentioned, and the image display is representative for the plane used for the analysis. N=number of replicates; n=number of roots.
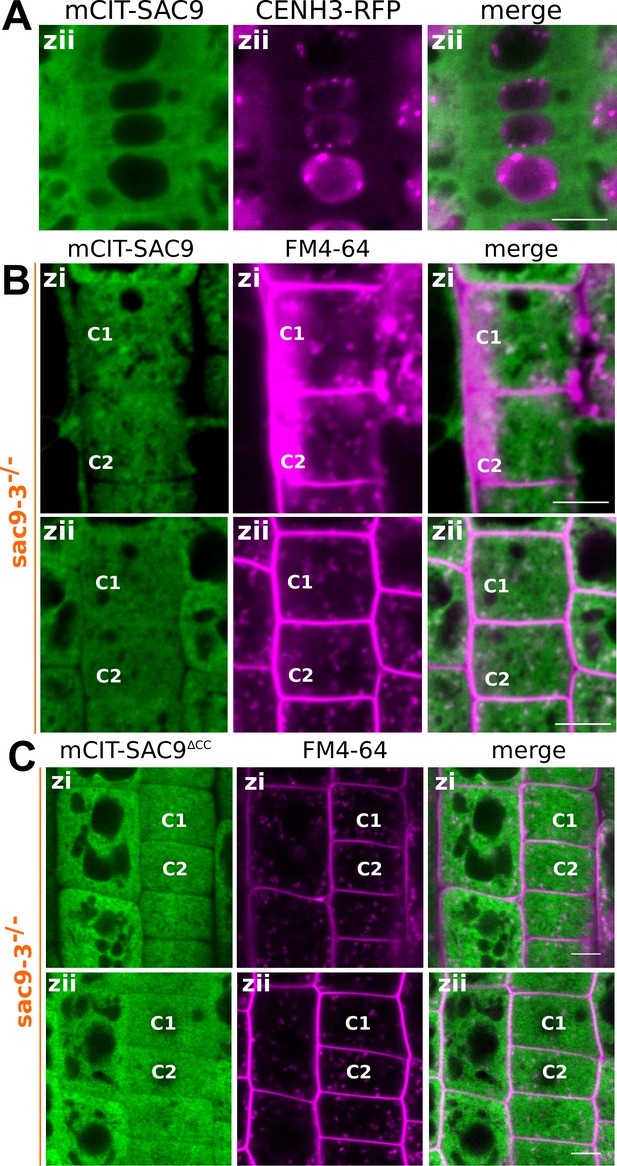
Subcellular localization of SAC9 alleles at different depths in the cell.
(A) Confocal imaging of mCIT-SAC9 (green) together with the nuclear marker CENH3-RFP (magenta). Scale bars: 10 µm. (B) Representative images of the signal corresponding to mCIT-SAC9 and FM4-64 at two focal planes (Zi and Zii) for a same cell (see C1 and C2 as examples). (C) Representative images of the signal corresponding to mCIT-SAC9∆CC and FM4-64 at two focal planes (Zi and Zii) for a same cell (see C1 and C2 as examples). (C) Quantification of the number of endosomes labeled by FM4-64 at two focal planes (Zi and Zii) for a same cell.
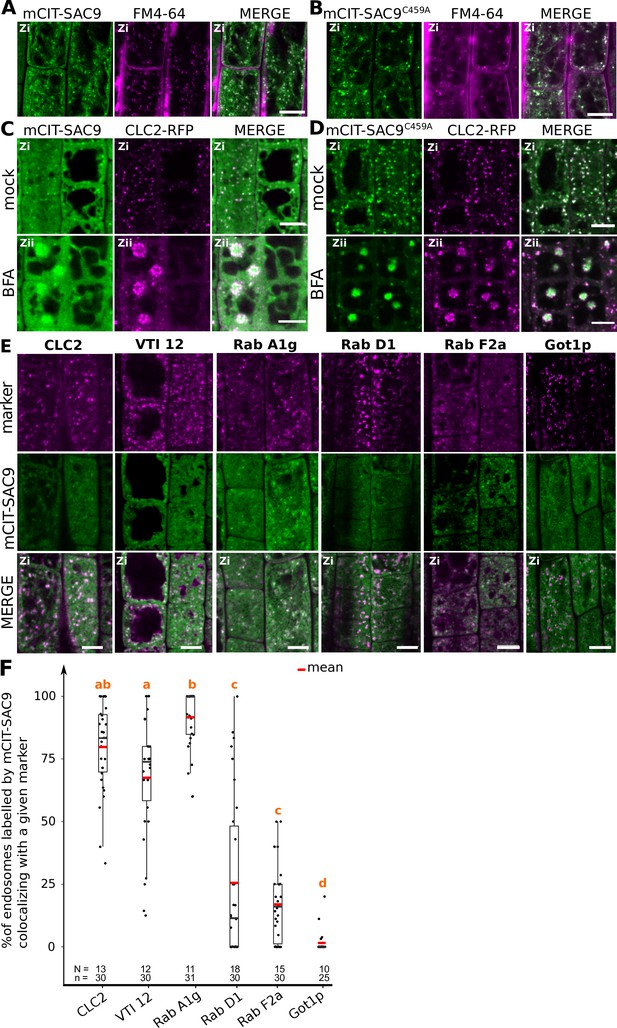
SUPPRESSOR OF ACTIN9 (SAC9) localizes to a subpopulation of TGN/EE.
(A, B) Confocal images of SAC9pro:mCIT-SAC9 (A, green) or SAC9pro:mCIT- SAC9C459A (B, green) in vivo in Arabidopsis root epidermis together with the endocytic tracer FM4-64 (magenta). (C, D) Confocal images of Arabidopsis root epidermis co-expressing SAC9pro:mCIT-SAC9 (C, green) or SAC9pro:mCIT- SAC9C459A (D, green) together Clathrin Light Chain 2 (CLC2) fused to RFP (CLC2-RFP; magenta). Upper panel: fluorescent signals observed in the mock treatment; Lower panel: fluorescent signals observed after 50 µM 60 min BFA treatment. (E) Confocal images of Arabidopsis root epidermis co-expressing SAC9pro:mCIT-SAC9 (green) and TGN markers CLC2-RFP, mCH-VTI12, mCH-RabA1g, mCH-RabD1, the late endosome/pre-vacuolar compartment (LE/MVB) marker mCH-RabF2a/Rha1 and the Golgi marker mCH-Got1p (magenta). (F) Percentage of colocalization between mCIT-SAC9 and a given endosomal compartment marker per cell, at zi. N=number of roots, n=number of cells. In the plots, middle horizontal bars represent the median, while the bottom and top of each box represent the 25th and 75th percentiles, respectively. At most, the whiskers extend to 1.5 times the interquartile range, excluding data beyond. For range of value under 1,5 IQR, whiskers represent the range of maximum and minimum values. The plane (Zi or Zii) in each image is mentioned, and the image display is representative for the plane used for the analysis. Scale bars: 10 µm.
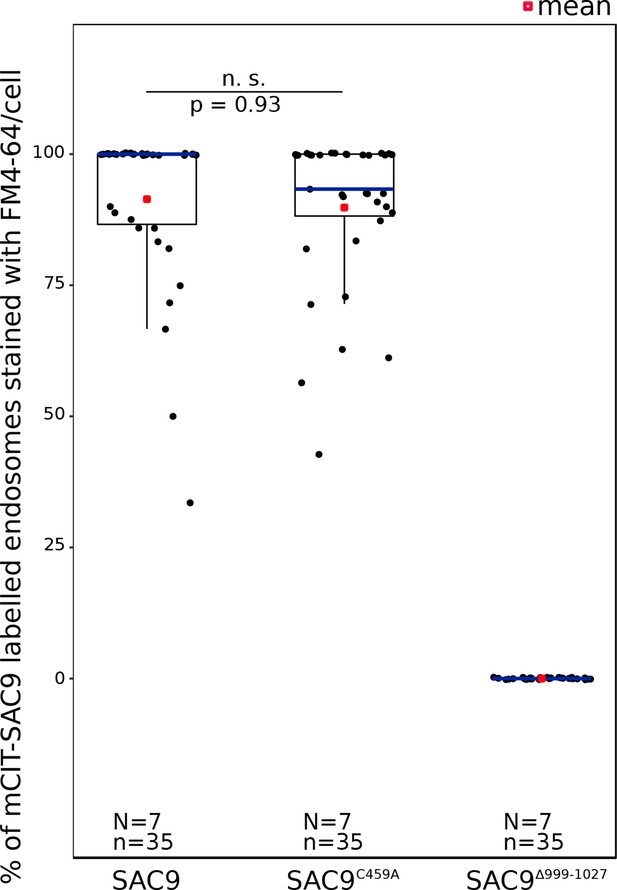
Quantification of the number of endosomes per cell labeled by mCIT-SAC9 variants which colocalized with FM4-64 at Zi.
In the plots, middle horizontal bars represent the median, while the bottom and top of each box represent the 25th and 75th percentiles, respectively. At most, the whiskers extend to 1.5 times the interquartile range, excluding data beyond. For range of value under 1,5 IQR, whiskers represent the range of maximum and minimum values. Details of the statistics are presented in Supplementary file 1K. N=number of roots, n=number of cells.
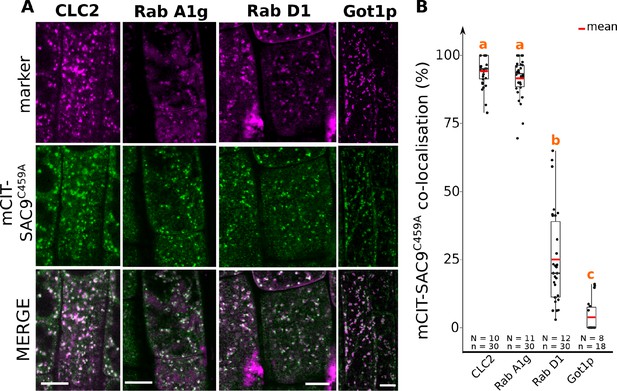
Confocal images of SAC9C459A with endosomal markers.
(A) Confocal images of Arabidopsis root epidermis co-expressing SAC9pro:mCIT-SAC9C459A (green) and markers such as mCH-RabA1g; mCH-RabD1, mCH-Got1p (magenta). The plane (Zi or Zii) in each image is mentioned, and the image display is representative for the plane used for the analysis. (B) Percentage of colocalization between mCIT-SAC9C459A and a given endosomal compartment marker at Zi. Similar letters correspond to markers with non-significantly different percentages of colocalization with mCIT-SAC9C459A (one-way ANOVA and post hoc Tuckey HSD pairwise test with a 95% confidence level). In the plots, middle horizontal bars represent the median, while the bottom and top of each box represent the 25th and 75th percentiles, respectively. At most, the whiskers extend to 1.5 times the interquartile range, excluding data beyond. For range of value under 1,5 IQR, whiskers represent the range of maximum and minimum values. Scale bars: 10 µm. N=number of roots, n=number of cells.
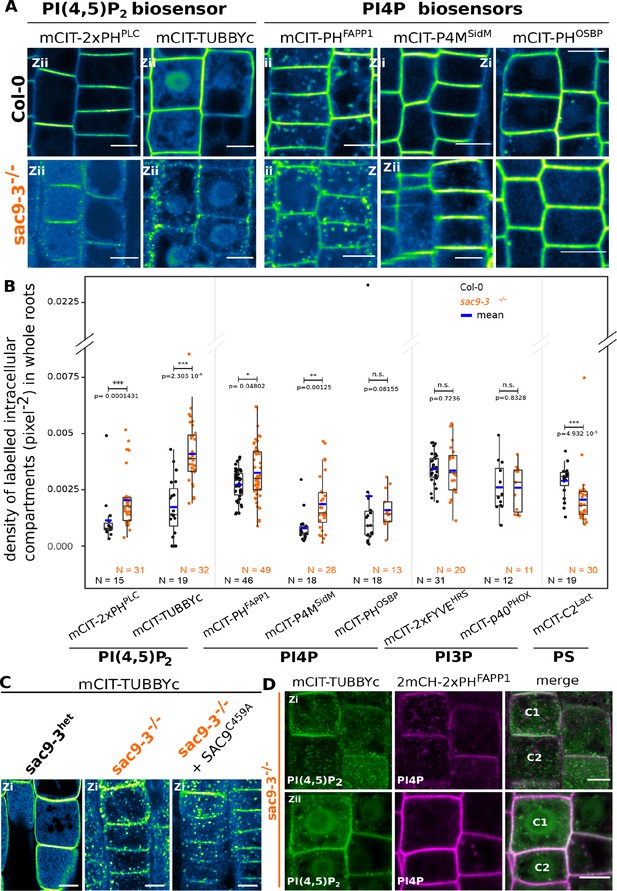
SUPPRESSOR OF ACTIN9 (SAC9) restricts PI(4,5)P2 at the plasma membrane.
(A) Confocal images of Arabidopsis root epidermis expressing mCIT-tagged sensors in WT (Col-0) and sac9-3-/- genetic backgrounds. Fluorescence intensity is color-coded (green fire blue scale). (B) Quantification of density of labeled intracellular compartments (pixel–2) in whole roots epidermis expressing mCIT-tagged lipid sensors in wild-type (Col-0) and sac9-3-/-. Non-parametric Wilcoxon rank sum tests with Bonferroni correction. In the plots, middle horizontal bars represent the median, while the bottom and top of each box represent the 25th and 75th percentiles, respectively. At most, the whiskers extend to 1.5 times the interquartile range, excluding data beyond. For range of value under 1,5 IQR, whiskers represent the range of maximum and minimum values. Details for statistical analysis can be found in the Methods section and Supplementary file 1. N=number of replicates; n=number of roots. (C) Confocal images of mCIT-TUBBYc in root epidermal cell of sac9-3 heterozygous, sac9-3-/- (left panel) or sac9-3-/- expressing TdTOM-SAC9C459A (right panel); N=3 replicates, n=47 roots. (D) Confocal images of sac9-3-/- Arabidopsis root epidermal cell (C1 and C2) expressing mCIT-TUBBYc (green) and 2xmCH-PHFAPP1 (magenta) at their cortex (upper panel, Zi) and at their center (bottom panel, Zii); N=3 biological replicates, n=49 roots. Scale bar: 10 µm. The plane (Zi or Zii) in each image is mentioned, and the image display is representative for the plane used for the analysis. Note that in panel D, the same cells are shown in the upper and lower panel, but at different focal planes.
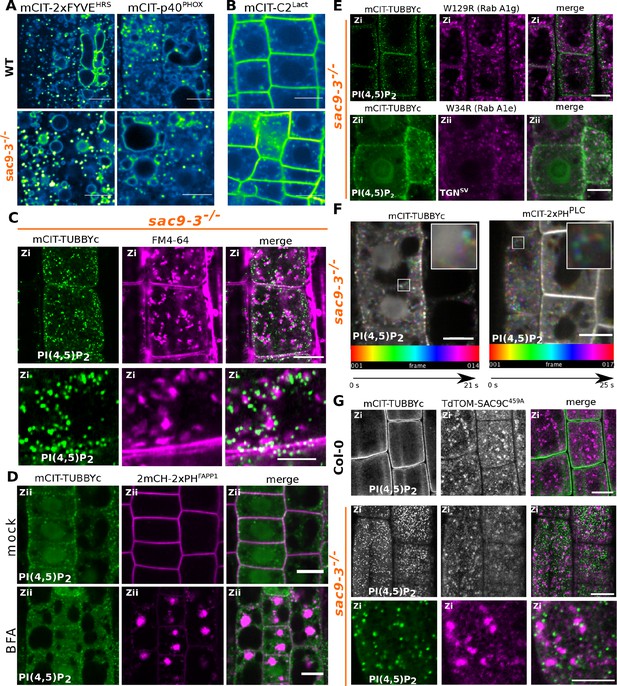
Confocal images of Arabidopsis root epidermis expressing various phosphoinositide biosensors in WT and sac9-3-/-.
(A) Representative confocal images of Arabidopsis root epidermis expressing mCIT-tagged lipid sensors for PI3P in wild-type (Col-0) and sac9-3-/- genetic backgrounds. (B) Confocal images of Arabidopsis root epidermis expressing mCIT-tagged lipid sensors for phosphatidylserine (PS) in wild-type (Col-0) and sac9-3-/- genetic backgrounds. Fluorescence intensity is color-coded (green fire blue scale). (C) Confocal images of sac9-3-/- Arabidopsis root epidermis expressing mCIT-TUBBYc (green) and in vivo stained with FM4-64 endocytic tracer (magenta); N=3 replicates, n=10 roots. (D) Confocal images of sac9-3-/- root epidermal cells co-expressing mCIT-TUBBYc (green) and 2xmCH-PHFAPP1 (magenta). Upper panel: fluorescent signals observed in the mock treatment; lower panel: fluorescent signal observed after 50 µM 60 min BFA treatment at the Zii plane. N=3 replicates, n=12 roots. (E) Confocal images of sac9-3-/- root cells where mCIT-TUBBYc (green) was covisualized together with various TGN markers (magenta) in vivo in transgenic lines expressing W129R (mCH-RabA1g) or W34R(mCH-RabA1e). N=3 replicates, n=15 roots. (F) Temporal-color coded XY 2D image of mCIT-2xPHPLC and mCIT-TUBBYc in sac9-3-/- root epidermis (Fiji Spectrum Temporal-color code). (G) Confocal images of mCIT-TUBBYc (green) were co-visualized together with tdTOM-SAC9C459A (magenta) in vivo in Col-0 WT and sac9-3-/-; N=3 replicates, n=13 roots. Scale bar: 10 µm.
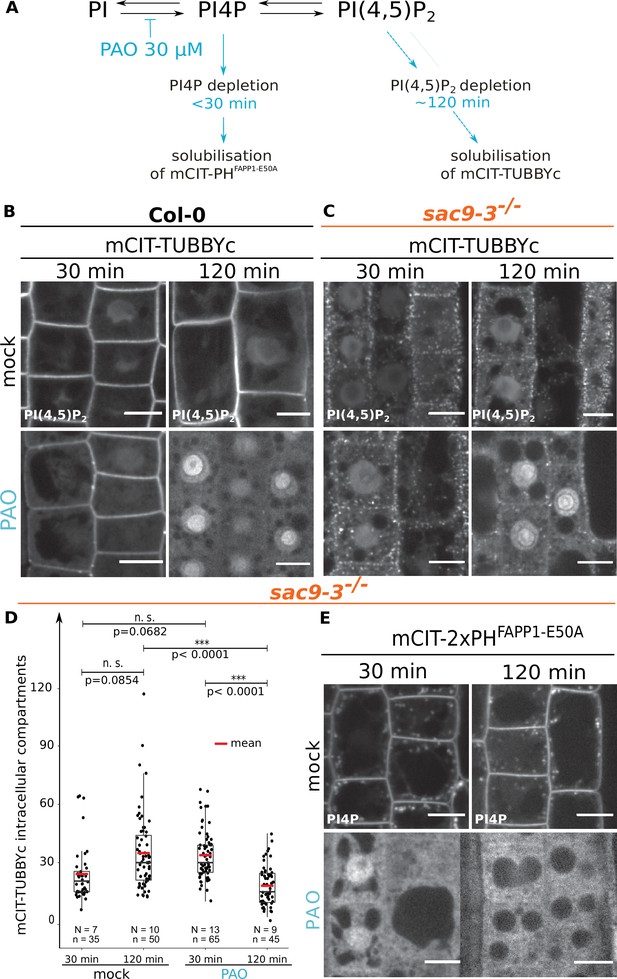
The intracellular accumulation of mCITRINE-TUBBYc observed in sac9 is sensitive to PI-4 kinase inhibition.
(A) Schematic representation of the mode of action of the PAO on the PI4P and PI(4,5)P2 metabolism based on previously published work (Doumane et al., 2020; Platre et al., 2018). (B) Confocal images of WT root epidermal cells expressing mCIT-TUBBYc PI(4,5)P2 biosensor that underwent mock, or 30 µM PAO treatments for 30- or 120 min. (C) Confocal images of sac9-3-/- root epidermal cells expressing mCIT-TUBBYc PI(4,5)P2 biosensor that underwent mock, or 30 µM PAO treatments for 30- or 120 min. (D) Quantification of the number of mCIT-TUBBYc labeled intracellular compartments per cell in sac9-3-/- following mock, or 30 µM PAO treatments for 30- or 120 min. Generalized linear mixed effect model (Poisson family) to account for root as a random factor, and two sided post hoc lsmeans pair-wise test with a 95% confidence level. In the plot, middle horizontal bars represent the median, while the bottom and top of each box represent the 25th and 75th percentiles, respectively. At most, the whiskers extend to 1.5 times the interquartile range, excluding data beyond. For range of value under 1,5 IQR, whiskers represent the range of maximum and minimum values. All statistical tests were two-sided. Details of the statistical analysis could be found in Supplementary file 1L. (E) Confocal images of sac9-3-/- root epidermal cells expressing mCIT-PHFAPP1-E50A PI4P biosensor that underwent mock or 30 µM PAO treatments for 30- or 120 min. Scale bar: 10 µm. N=number of roots, n=numbers of cells.
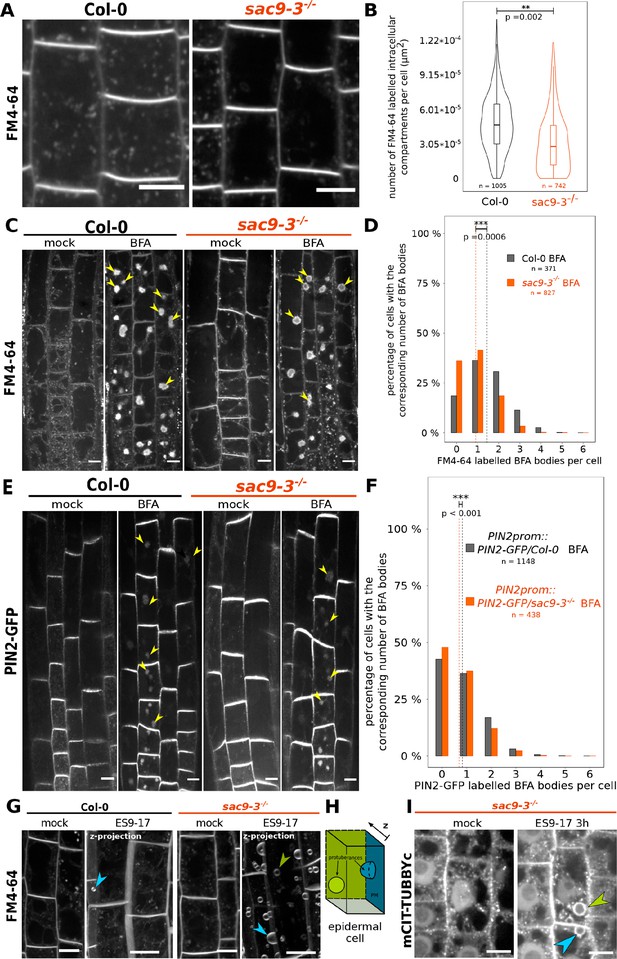
The endocytic flux is perturbed in the sac9 mutant.
(A) Representative images of Col-0 and sac9-3-/- seedlings treated for 30 min with FM4-64, which is endocytosed and labels endocytic intracellular compartments. Scale bars: 10 µm. (B) Quantification from the experiment shown in (A). Violin and box plots quantifying the number of FM4-64 labeled intracellular compartments in Col-0 and sac9-3-/-. (C) Representative images of root epidermis following BFA and FM4-64 treatment of Col-0 and sac9-3-/- seedlings. Examples of FM4-64 labeled BFA bodies are pointed out (green arrowheads). (D) Quantification from the experiment shown in (C). For Col-0 and sac9-3-/-, the proportion (%) of cells containing from none to six BFA bodies is displayed. Dotted line: means. (E) Representative images following BFA treatment of PIN2-GFP/WT and PIN2-GFP/sac9-3-/- seedlings. Examples of PIN2-GFP labeled BFA bodies are pointed out (green arrowheads). Scale bars: 10 µm. (F) Quantification from the experiment shown in (E). For PIN2-GFP/WT and PIN2-GFP/sac9-3-/-, the proportion (%) of cells containing from none to six BFA bodies is displayed. Dotted line: means. (G) Over-sensitivity of sac9-3-/- to prolonged inhibition of endocytosis. Seedlings were treated 180 min with 30 µM ES9-17 or DMSO (mock), FM4-64 being added after 30 min (150 min of exposure). The picture shown after the ES9-17 treatment are the results of the projection of a z-stack. (H) ES9-17 treatments led to dome-shaped plasma membrane invagination. blue arrowheads: invaginations with an obvious connection to the plasma membrane; green arrowheads: invaginations without a clear connection to the plasma membrane (often connected to medullar plasma membrane). N=2 biological replicates, n=8 roots. (I) In sac9-3 mutant, ES9-17 treatments led to dome-shaped plasma membrane invagination labeled by the PI(4,5)P2 biosensor mCIT-TUBBYc; N=2 biological replicates, n=10 roots. Blue arrowheads: invaginations with an obvious connection to the plasma membrane; green arrowheads: invaginations without a clear connection to the plasma membrane (often connected to medullar plasma membrane). Scale bars: 10 µm. Details of the statistical analysis could be found in Supplementary file 1C. N=3 biological replicates, n=number of cells.
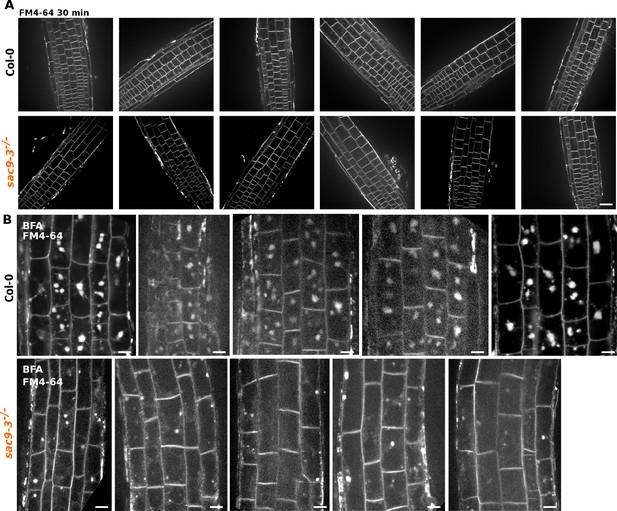
Example of FM4-64 labeling in Col-0 and sac9-3 mutant.
(A) Example of confocal images used for the number of FM4-64 labeled intracellular compartments per cell in Col-0 and sac9-3-/-. Scale bar: 75 µm. (B) Example of confocal images used for the number of cell with the corresponding number of BFA bodies stained by FM4-64 in Col-0 and sac9-3-/-. Scale bar: 10 µm.
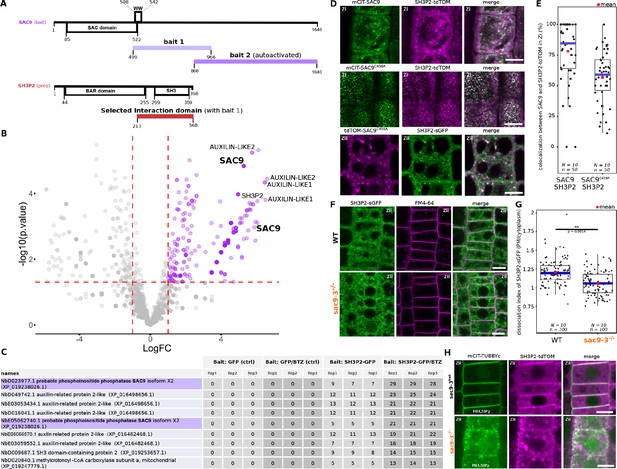
SUPPRESSOR OF ACTIN9 (SAC9) interact and colocalize with SH3P2 and regulates it subcellular distribution.
(A) Schematic representation of the yeast two-hybrid screen using SAC9 as a bait, where SH3P2 was found as a protein partner. The selected interaction domain (interacting with bait 1) corresponds to the amino acid sequence shared by all the eleven prey fragments matching the same reference protein. Note that one other fragment of SAC9 was intended to be screened for interacting proteins (bait 2), but the construct was autoactivate in yeast. For design purpose, the scale between SAC9 and SH3P2 was not respected. (B) Volcano plot of SH3P2-GFP IP-MS after transient expression in N. benthamiana. X-axis displays the log fold-change (log(FC)) of proteins in SH3P2-GFP compared to the control (GFP). The Y-axis shows the statistical significance, log10(p-value). Dashed lines represent threshold for log(FC)>1 and p-value <0.05. (C) Table showing the peptide counts of the top hits obtained in the IP-MS (n=3 independent pooled biological replicates), including the controls GFP and GFP/BTZ. Note that multiple protein isoforms of SAC9 are present in the N. benthamiana annotated proteome (Kourelis et al., 2019b), and two were found in the IP-MS as SAC9 interactors. (D), Representatives images of the colocalization between mCIT-SAC9 and SH3P2-tdTOM in a proximal (Zi) or distal (Zii) region from the plasma membrane of Arabidopsis epidermal root cells. Scale = 5 µm. (E) Quantification of the number of SAC9-labeled endosomes colocalizing with SH3P2 signal per cell. N=number of replicates; n=number of roots. (F), Representatives images of the SH3P2-sGFP localization (green) in WT and sac9-3 mutant in which the plasma membrane was labeled using FM4-64 (magenta). (G) Quantification of the dissociation index of SH3P2-sGFP in WT and sac9-3 mutant in which the plasma membrane was labeled using FM4-64. N=number of replicates; n=number of roots. (H), Representative images of the localization of SH3P2pro:SH3P2-tdTOM transformed in sac9-3 heterozygous plant expressing the PI(4,5)P2 biosensor mCIT-TUBBYc. N=2 biological replicates, n=50 roots. In the plots, middle horizontal bars represent the median, while the bottom and top of each box represent the 25th and 75th percentiles, respectively. At most, the whiskers extend to 1.5 times the interquartile range, excluding data beyond. For range of value under 1,5 IQR, whiskers represent the range of maximum and minimum values. Details of the statistical analysis could be found in Supplementary file 1H.
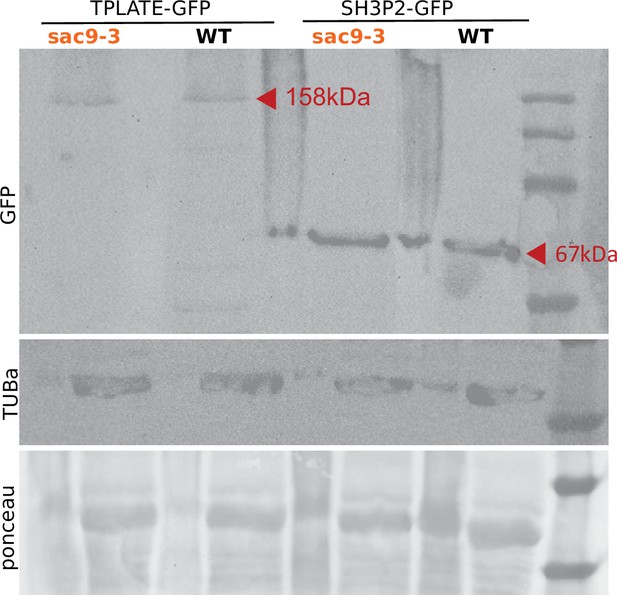
Western blot analysis of TPLATE-GFP and SH3P2-GFP in WT and sac9-3-/-.
The expression of the transgene was detected using an anti-GFP antibody. Ponceau S staining as well as western blot using the anti-tubulinα antibody served as a loading control. The experiment was repeated three times with similar results. Note that the narrow lanes correspond to the signal obtained with the crude extracts, for which the detection of TPLATE-GFP was not optimal and therefore was not used to analyze the results.
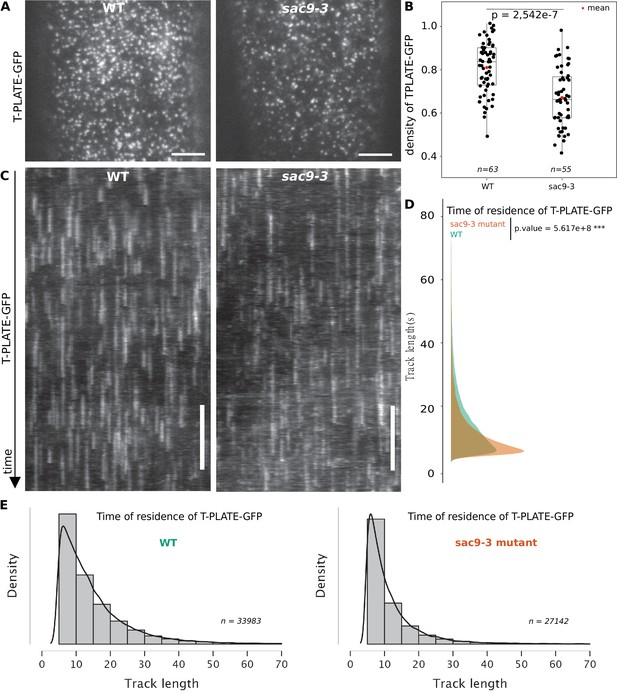
TPLATE dynamics and density at the plasma membrane is disturbed in the sac9 mutant.
(A) Representatives images of the TPLATE-GFP localization in WT and sac9-3 mutant at the plasma membrane observed by TIRF microscopy in etiolated hypocotyl. Scale = 5 µm. (B) Quantification of the density at the plasma membrane of TPLATE-GFP in WT and sac9-3 mutant observed by TIRF microscopy in etiolated hypocotyl. Only one of the three replicates are represented. In the plot, middle horizontal bars represent the median, while the bottom and top of each box represent the 25th and 75th percentiles, respectively. At most, the whiskers extend to 1.5 times the interquartile range, excluding data beyond. For range of value under 1,5 IQR, whiskers represent the range of maximum and minimum values. N=13 plants n=63 cells for the WT; N=11, n=55 for sac9-3. (C) Representatives kymograph of the TPLATE-GFP dynamics at the plasma membrane in WT and sac9-3 mutant observed by TIRF microscopy in etiolated hypocotyl over 5 min; scale = 60 s. (D), Frequency distribution of the lifetimes of TPLATE-GFP tracks in WT (green) and sac9-3 mutant (oranges). Results from an independent samples T-Test followed by Mann-Whitney U test is presented (see Supplementary file 1L for details). (E), Histograms of median normalized fluorescence of TPLATE-GFP in WT (left) and sac9-3 mutant (right) representing the density of tracks per track length. Details of the statistical analysis could be found in Supplementary file 1J. For WT, 21 cells from N=10 plants were used, n=33,983 tracks; For sac9, N=13 n=27,142 tracks.
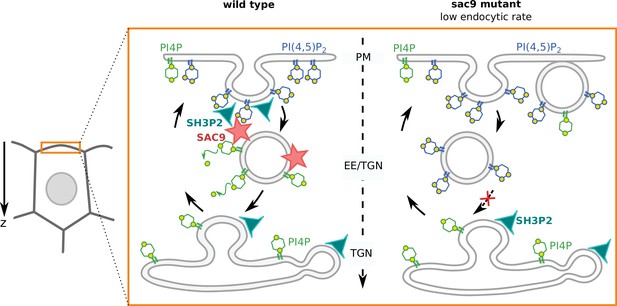
Model for the mode of action of SUPPRESSOR OF ACTIN9 (SAC9) in regulating PI(4,5)P2 subcellular patterning.
In wild-type plants, SAC9 restricts the localization of PI(4,5)P2 at the plasma membrane allowing endocytic processes to occur. In the absence of SAC9, endocytosis PI(4,5)P2 accumulates in atypical endomembrane compartments that are no longer able to fuse with the TGN possibly because of their abnormal anionic lipid signature. SAC9 interacts with SH3P2 close to the plasma membrane. The defects in PI(4,5)P2 patterning in absence of SAC9 leads to decreasing in the endocytic rate and the formation of membrane protuberances in contact with the plasma membrane. Note that in this model, the observed increase in intracellular PI4P levels as measured by probes is not included.
Videos
Time-lapse imaging of mCIT-SAC9 using spinning disk confocal microscope (1 s per frame).
Time-lapse imaging of mCIT-2xPHPLC in sac9 mutant (1 s per frame) using spinning disk confocal microscope.
Time-lapse imaging of TPLATE-GFP at the plasma membrane of WT and in sac9-3 mutant using TIRF microscopy.
Time lapses were acquired during 300 time-points for 300 s (acquisition time 500 ms).
Additional files
-
MDAR checklist
- https://cdn.elifesciences.org/articles/73837/elife-73837-mdarchecklist1-v2.docx
-
Supplementary file 1
Ressources and statistics.
(A) Reagent and resources. (B) Raw data from ULTImate Y2H SCREEN Arabidopsis thaliana - SAC9 (aa 499–966) vs Universal Arabidopsis Normalized_dT. (C) Details of the statistics corresponding to Figure 1C, D. (D) Details of the statistics corresponding to Figure 2D, G. (E) Details of the statistics corresponding to Figure 3G-F. (F) Details of the statistics corresponding to Figure 4B. (G) Details of the statistics corresponding to Figure 5. (H) Details of the statistics corresponding to Figure 6E, G. (I) Details of the statistics corresponding to Figure 7B. (J) Details of the statistics corresponding to Figure 1—figure supplement 1. (K) Details of the statistics corresponding to Figure 3—figure supplement 2.
- https://cdn.elifesciences.org/articles/73837/elife-73837-supp1-v2.xlsx





