Regulation of immune receptor kinase plasma membrane nanoscale organization by a plant peptide hormone and its receptors
Figures
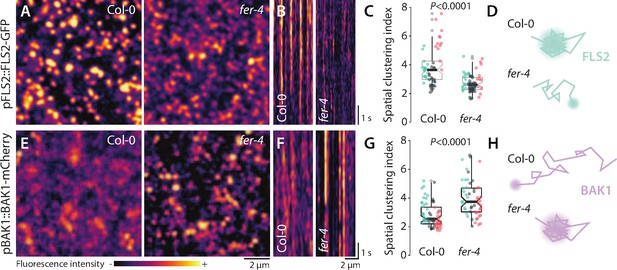
FER regulates the nanoscale organization of FLS2-GFP and BAK1-mCherry.
(A, E) FLS2-GFP and BAK1-mCherry nanodomain organization. Pictures are maximum projection of 20 variable angle total internal reflection fluorescence microscopy (VA-TIRFM) images obtained at 5 frames per second for FLS2-GFP (A) and 10 VA-TIRFM images obtained at 2.5 frames per second for BAK1-mCherry (E) in Col-0 and fer-4 cotyledon epidermal cells. (B, F) Representative kymograph showing lateral organization of FLS2-GFP (B) and BAK1-mCherry (F) overtime in Col-0 and fer-4. (C, G) Quantification of FLS2-GFP (C) and BAK1-mCherry (G) spatial clustering index. Graphs are notched box plots, scattered data points show measurements, colors indicate independent experiments, n = 16 cells for Col-0/pFLS2::FLS2-GFP; n = 31 cells for fer-4/pFLS2::FLS2-GFP, n = 23 cells for Col-0/pBAK1::BAK1-mCherry, n = 18 cells for fer-4/pBAK1::BAK1-mCherry. p-Values report two-tailed nonparametric Mann–Whitney test. (D, H) Graphical illustrations summarizing our observations for FLS2-GFP (D) and BAK1-mCherry (H) nanoscale dynamics.
-
Figure 1—source data 1
Source data points for the graphs in Figure 1C and G.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig1-data1-v2.xlsx
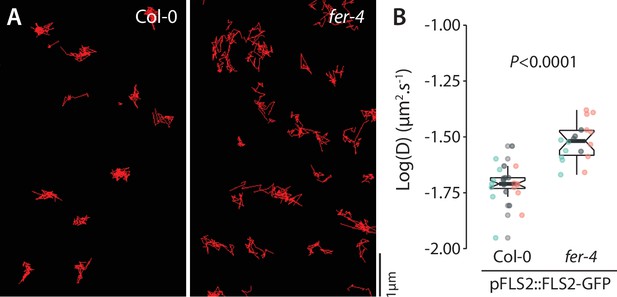
Analysis of FLS2-GFP single-particle dynamics in fer-4.
(A) Representative images of FLS2-GFP single particles tracked in Col-0 and fer-4 cotyledon epidermal cells of 5-day-old seedlings. (B) Quantification of FLS2-GFP diffusion coefficient (D). Graph is a notched box plot, scattered data points correspond to the average diffusion coefficient of single particles for each cell, colors indicate independent biological repetitions, n = 23 cells for Col-0/pFLS2::FLS2-GFP, n = 19 cells for fer-4/pFLS2::FLS2-GFP. p-Value reports nonparametric Mann–Whitney test.
-
Figure 1—figure supplement 1—source data 1
Source data points for the graph in Figure 1—figure supplement 1.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig1-figsupp1-data1-v2.xlsx
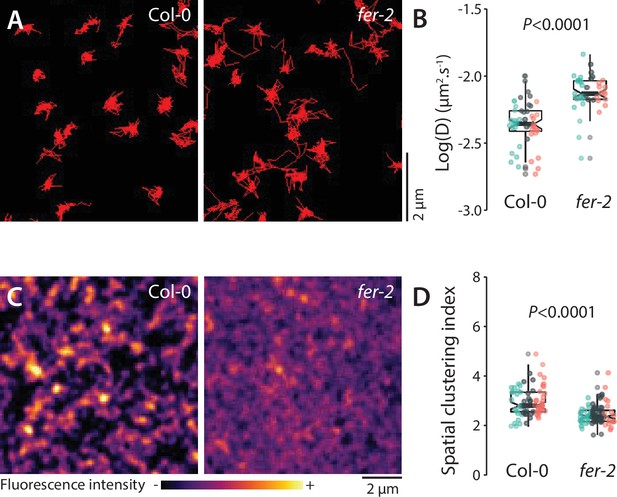
Analysis of FLS2-GFP organization and dynamics in fer-2.
(A) Representative images of FLS2-GFP single particles tracked in Col-0 and fer-2 cotyledon epidermal cells of 5-day-old seedlings. (B) Quantification of FLS2-GFP diffusion coefficient (D). Graph is a notched box plot, scattered data points correspond to the average diffusion coefficient of single particles for each cell, colors indicate independent experiments, n = 48 cells for Col-0/pFLS2::FLS2-GFP, n = 40 cells for fer-2/pFLS2::FLS2-GFP. p-Value reports nonparametric Mann–Whitney test. (C) Pictures are maximum projection images (20 variable angle total internal reflection fluorescence microscopy [VA-TIRFM] images obtained at 20 frames per second) of FLS2-GFP in Col-0 and fer-2 cotyledon epidermal cells. (D) Quantification of FLS2-GFP spatial clustering index in Col-0 and fer-2. Graph is a notched box plot, scattered data points show individual measurements, colors indicate independent experiments, n = 24 cells for Col-0/pFLS2::FLS2-GFP; n = 27 cells for fer-2/pFLS2::FLS2-GFP. p-Values report two-tailed nonparametric Mann–Whitney test.
-
Figure 1—figure supplement 2—source data 1
Source data points for the graphs in Figure 1—figure supplement 2B and D.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig1-figsupp2-data1-v2.xlsx
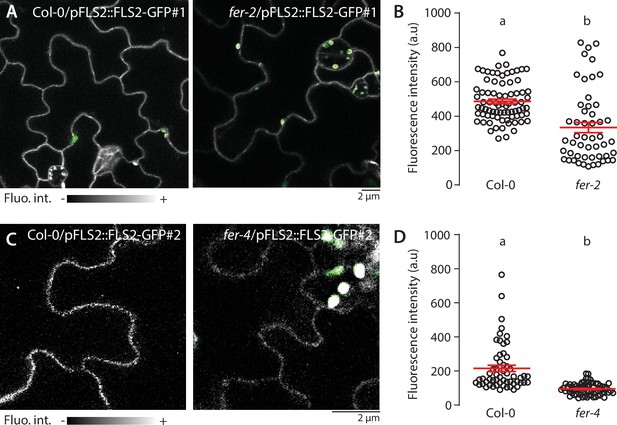
FLS2-GFP accumulation at the PM is altered in fer mutants.
(A–D) Representative confocal microscopy pictures of FLS2-GFP driven by its native promoter in Col-0 and fer-2 (A) and fer-4 (C) cotyledon epidermal cells of 5-day-old seedlings. Scale bar indicates 2 µm. Quantification of FLS2-GFP fluorescence intensity at the plasma membrane in Col-0 and fer-2 (B) and fer-4 (D) cotyledon epidermal cells of 5-day-old seedlings. Col-0/pFLS2::FLS2-GFP#1 and Col-0/pFLS2::FLS2-GFP#2 correspond to distinct lines. Data points correspond to individual and unique cell-cell border, n = 78 cell-cell border for Col-0/pFLS2::FLS2-GFP#1, n = 51 for fer-2/pFLS2::FLS2-GFP#1; n = 57 Col-0/pFLS2::FLS2-GFP#2 and n = 71 for fer-4/pFLS2::FLS2-GFP#2. Two-way Student’s t-test. Similar results were obtained in at least three independent experiments.
-
Figure 1—figure supplement 3—source data 1
Source data points for the graphs in Figure 1—figure supplement 3B and D.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig1-figsupp3-data1-v2.xlsx
Variable angle total internal reflection fluorescence microscopy (VA-TIRFM) imaging of FLS2-GFP in Col-0 and fer-4.
Representative stream images acquisition of FLS2-GFP particles observed at the surface of 5-day-old cotyledon epidermal cells by VA-TIRF microscopy at 5 frames per second.
Variable angle total internal reflection fluorescence microscopy (VA-TIRFM) imaging of BAK1-mCherry in Col-0 and fer-4 with or without RALF23 treatment.
Representative stream images acquisition of BAK1-mCherry observed at the surface of 5-day-old cotyledon epidermal cells by VA-TIRF microscopy at 2.5 frames per second, in Col-0 (A, B) and fer-4 (C, D) with (B, D) or without (A, C) 1 μM RALF23 treatment. Scale bar indicates 2 µm.
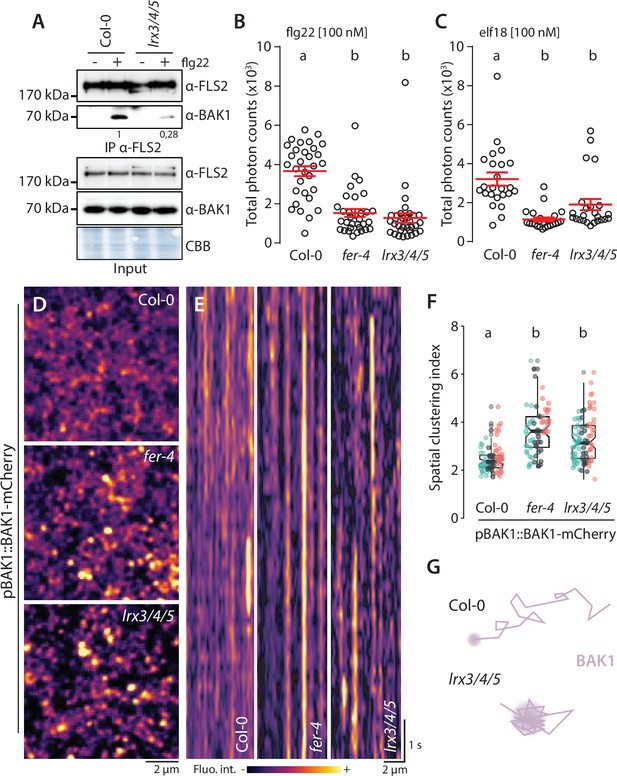
LRX3, LRX4, and LRX5 regulate pattern-triggered immunity (PTI) and BAK1-mCherry organization.
(A) flg22-induced FLS2-BAK1 complex formation. Immunoprecipitation of FLS2 in Arabidopsis Col-0 and lr3/4/5 seedlings either untreated or treated with 100 nM flg22 for 10 min. Blot stained with Coomassie brilliant blue (CBB) is presented to show equal loading. Western blots were probed with α-FLS2, α-BAK1, or α-FER antibodies. Numbers indicate quantification of BAK1 bands normalized based on the corresponding intensities of FLS2 bands and relative to the control Col-0. Similar results were obtained in at least three independent experiments. (B, C) Reactive oxygen species (ROS) production after elicitation with 100 nM elf18 (B) or 100 nM flg22 (C). Values are means of total photon counts over 40 min. Red crosses and red horizontal lines denote mean and SEM, n = 32. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.0001). (D) BAK1-mCherry nanodomain organization. Pictures are maximum projection images (10 variable angle total internal reflection fluorescence microscopy [VA-TIRFM] images obtained at 2.5 frames per second) of BAK1-mCherry in Col-0, fer-4, and lrx3/4/5 cotyledon epidermal cells. (E) Representative kymograph showing lateral organization of BAK1-mCherry overtime in Col-0, fer-4, and lrx3/4/5. (F) Quantification of BAK1-mCherry spatial clustering index. Graphs are notched box plots, scattered data points show measurements, colors indicate independent experiments, n = 26 cells for Col-0/pBAK1::BAK1-mCherry, n = 31 cells for fer-4/pBAK1::BAK1-mCherry, n = 28 cells for lrx3/4/5/pBAK1::BAK1-mCherry. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.0001). (G) Graphical illustration summarizing our observations for BAK1-mCherry nanoscale dynamics in lrx3/4/5.
-
Figure 2—source data 1
Source data points for the graphs in Figure 2B, C and F.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig2-data1-v2.xlsx
-
Figure 2—source data 2
Source blots images for the co-immunoprecipitation (co-IP) in Figure 2A.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig2-data2-v2.pdf
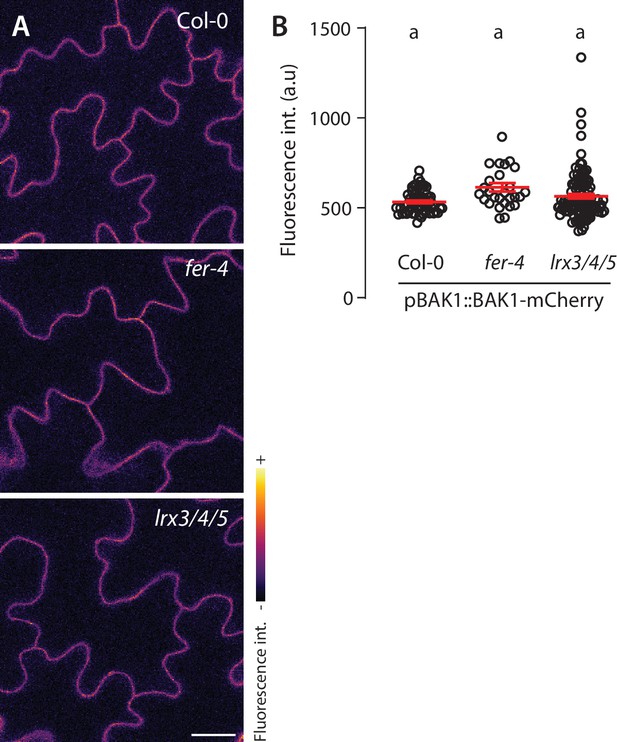
Subcellular localization of BAK1-mCherry in fer-4 and lrx3/4/5.
(A) Representative confocal microscopy pictures of BAK1-mCherry driven by its native promoter in Col-0, fer-4, and lrx3/4/5 cotyledon epidermal cells of 5-day-old seedlings. Scale bar indicates 20 µm. (B) Quantification of BAK1-mCherry fluorescence intensity at the plasma membrane. Data points correspond to individual and unique cell-cell border, n = 115 cell-cell border for Col-0/pBAK1::BAK1-mCherry, n = 27 for fer-4/pBAK1::BAK1-mCherry and n = 125 for lrx3/4/5/ pBAK1::BAK1-mCherry. Letters indicate Dunn’s multiple comparison statistical test.
-
Figure 2—figure supplement 1—source data 1
Source data points for the graph in Figure 2—figure supplement 1B.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig2-figsupp1-data1-v2.xlsx
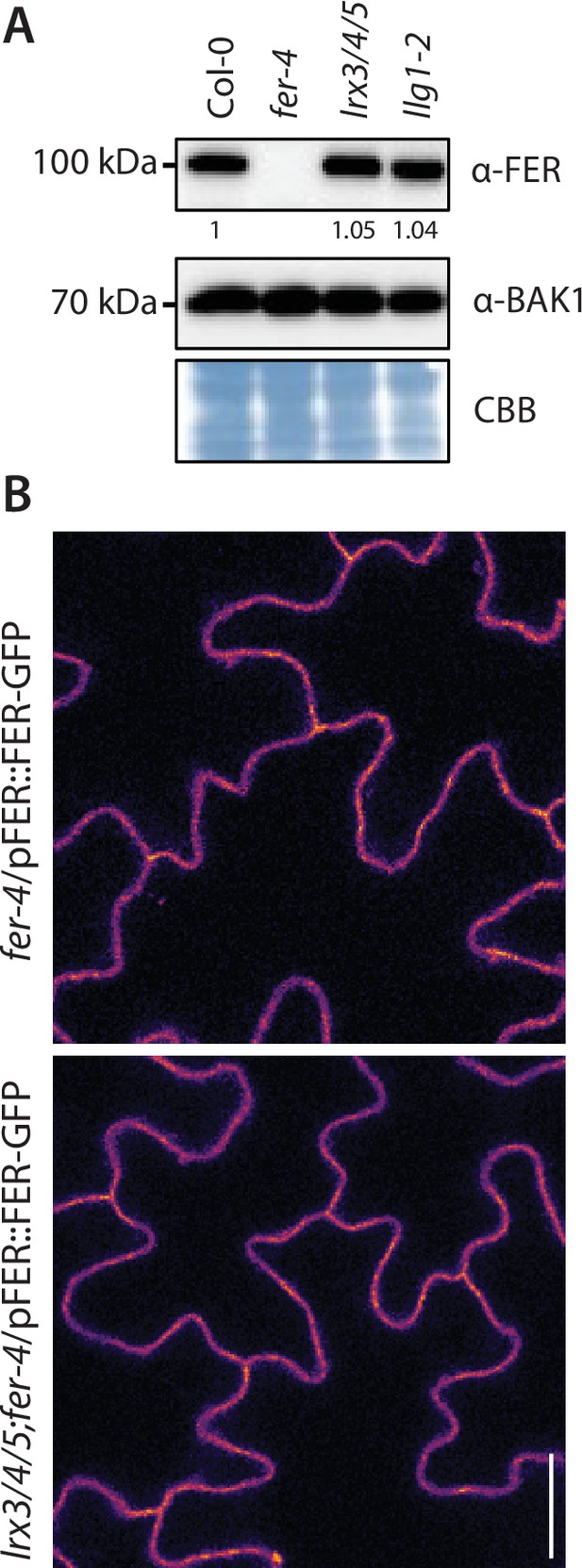
LRX3, LRX4, and LRX5 are dispensable for FER plasma membrane localization and accumulation.
(A) Accumulation of endogenous FER detected by western blot. Total proteins were extracted from 2-week-old seedlings. Blot stained with Coomassie brilliant blue (CBB) is presented to show equal loading. Numbers indicate quantification of FER bands normalized based on the BAK1 bands’ intensities for each condition and relative to the control Col-0. (B, C) Representative confocal microscopy pictures of FER-GFP driven by its native promoter in fer-4 and lrx3/4/5 cotyledon epidermal cells (B) of 5-day-old seedlings. Scale bar indicates 20 µm.
-
Figure 2—figure supplement 2—source data 1
Source blots images for Figure 2—figure supplement 2.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig2-figsupp2-data1-v2.pdf
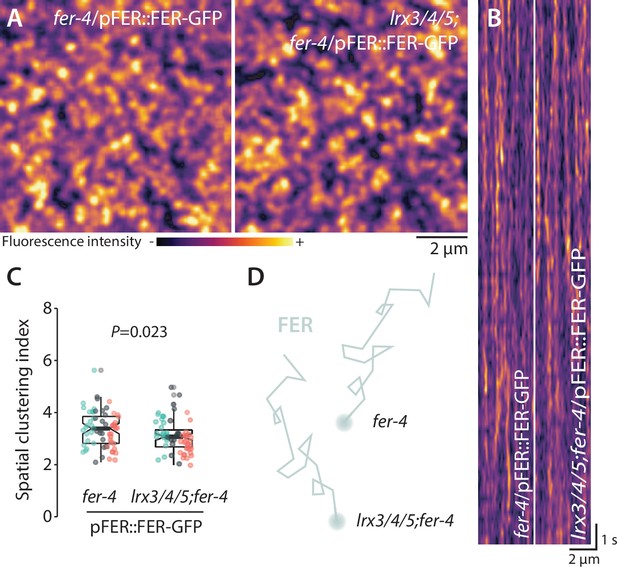
LRX3, LRX4, and LRX5 are dispensable for FER-GFP nanoscale organization.
(A) FER-GFP nanodomain organization. Pictures are maximum projection images (20 variable angle total internal reflection fluorescence microscopy [VA-TIRFM] images obtained at 5 frames per second) of FER-GFP in fer-4 and lrx3/4/5 cotyledon epidermal cells. (B) Representative kymograph showing lateral organization of FER-GFP overtime in fer-4 and lrx3/4/5. (C) Quantification of FER-GFP spatial clustering index. Graph is a notched box plot, scattered data points show individual measurements, colors indicate independent experiments, n = 21 cells for fer-4/pFER::FER-GFP; n = 21 cells for lrx3/4/5/fer-4/ pFER::FER-GFP. p-Value reports two-tailed nonparametric Mann–Whitney test. (D) Graphical illustration summarizing our observations for FER-GFP nanoscale dynamics.
-
Figure 2—figure supplement 3—source data 1
Source data points for the graph in Figure 2—figure supplement 3B.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig2-figsupp3-data1-v2.xlsx
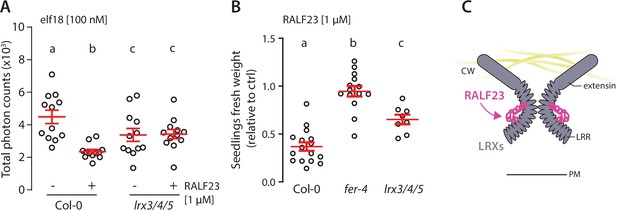
LRX3, LRX4, and LRX5 contribute to RALF23 responsiveness.
(A) Reactive oxygen species (ROS) production in Col-0 and lrx3/4/5 leaf discs treated with 100 nM elf18 and with or without 1 µM RALF23 co-treatment in 2 mM MES–KOH pH 5.8. Values are means of total photon counts over 40 min. Red crosses and red horizontal lines show mean and SEM, n = 12. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.0001). (B) Fresh weight of 12-day-old seedlings grown in the absence (mock) or presence of 1 µM of RALF23 peptide. Fresh weight is expressed as relative to the control mock condition. Similar results were obtained in at least three independent experiments. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.001). (C) Graphical illustration representing LRX3/4/5 as potential receptors for RALF23.
-
Figure 2—figure supplement 4—source data 1
Source data points for the graph in Figure 2—figure supplement 4A and B.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig2-figsupp4-data1-v2.xlsx
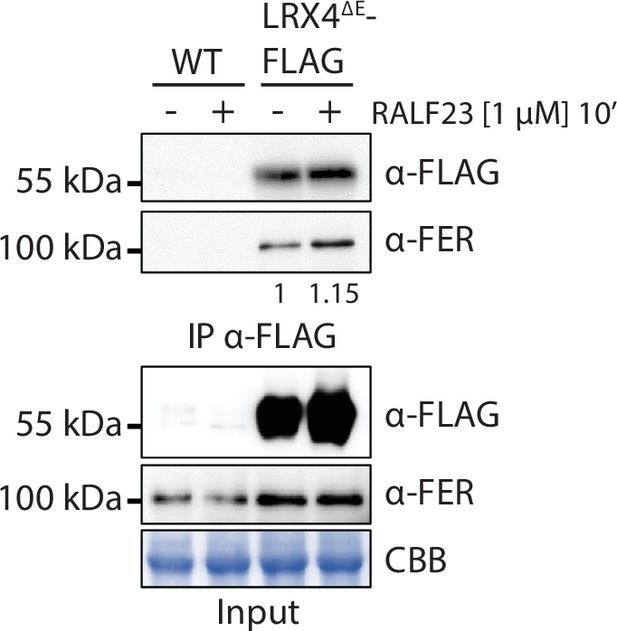
RALF23 does not modulate constitutive association between FER and LRX4.
Immunoprecipitation of LRX4ΔE-FLAG in Arabidopsis seedlings untreated or treated with 1 µM RALF23 for 10 min. Western blots were probed with α-FLAG or α-FER antibodies. Blot stained with Coomassie brilliant blue (CBB) is presented to show equal loading. Numbers indicate quantification of FER bands relative to the control without RALF23 treatment, normalized based on the FLAG bands’ intensities. Similar results were obtained in at least three independent experiments.
-
Figure 2—figure supplement 5—source data 1
Source blots images for the co-immunoprecipitation (co-IP) in Figure 2—figure supplement 5.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig2-figsupp5-data1-v2.pdf
Variable angle total internal reflection fluorescence microscopy (VA-TIRFM) imaging of BAK1-mCherry in Col-0 and lrx3/4/5.
Representative stream images acquisition of BAK1-mCherry observed at the surface of 5-day-old cotyledon epidermal cells by VA-TIRF microscopy at 2.5 frames per second, in Col-0 (A) and lrx3/4/5 (B). Scale bar indicates 2 µm.
Variable angle total internal reflection fluorescence microscopy (VA-TIRFM) imaging of FER-GFP in fer-4 and fer-4;lrx3/4/5.
Representative stream images acquisition of FER-GFP observed at the surface of 5-day-old cotyledon epidermal cells by VA-TIRF microscopy at 5 frames per second in fer-4 (A) and fer-4;lrx3/4/5 (B). Scale bar indicates 2 µm.
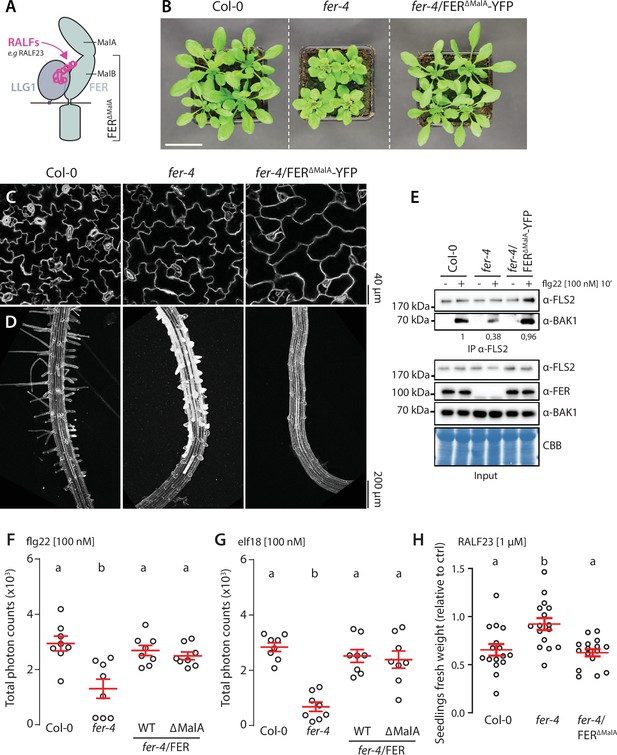
FER malectin A domain regulates cell morphogenesis not pattern-triggered immunity (PTI).
(A) Graphical representation of RALF23 perception by FER-LLG1 complex. (B) Morphology of 4-week-old Arabidopsis plants; scale bar indicates 5 cm. (C, D) Confocal microscopy pictures of 5-day-old seedlings cotyledon (C) and root (D) epidermal cells stained with propidium iodide. 3–4 seedlings per genotypes were observed per experiment. For each seedling, we observed the center of both cotyledons, and at the initiation site of root hairs. Similar results were obtained in at least three independent experiments. (E) Flg22-induced FLS2-BAK1 complex formation. Immunoprecipitation of FLS2 in Arabidopsis Col-0, fer-4, and fer-4/p35S::FER∆MalA-YFP seedlings that were either untreated or treated with 100 nM flg22 for 10 min. Blot stained with Coomassie brilliant blue (CBB) is presented to show equal loading. Western blots were probed with α-FLS2, α-BAK1, or α-FER antibodies. Numbers indicate quantification of BAK1 bands normalized based on the corresponding intensities of FLS2 bands and relative to the control Col-0 + flg22. Similar results were obtained in at least three independent experiments. (F, G) Reactive oxygen species (ROS) production after elicitation with 100 nM flg22 (F) or 100 nM elf18 (G). Values are means of total photon counts over 40 min, n = 8. Red crosses and red horizontal lines denote mean and SEM, respectively. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.0001). (H) Fresh weight of 12-day-old seedlings grown in the absence (mock) or presence of 1 µM of RALF23 peptide. Fresh weight is expressed as relative to the control mock treatment for each genotype. Similar results were obtained in at least three independent experiments. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.001).
-
Figure 3—source data 1
Source data points for the graphs in Figure 3F–H.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig3-data1-v2.xlsx
-
Figure 3—source data 2
Source blots images for the co-immunoprecipitation (co-IP) in Figure 3E.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig3-data2-v2.pdf
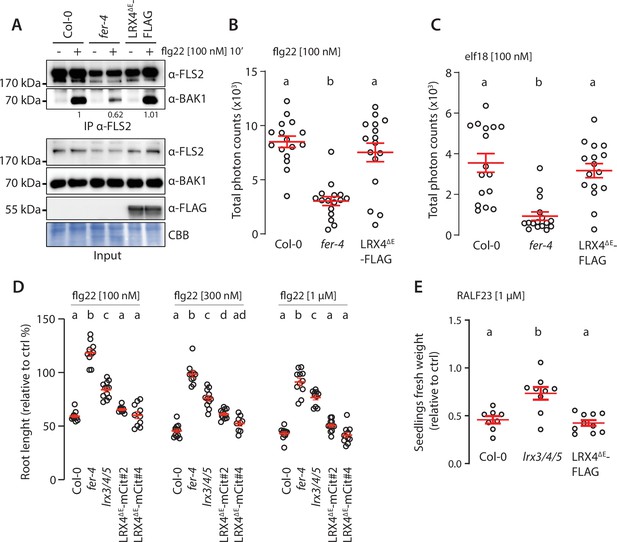
Overexpression of LRX4ΔE does not affect pattern-triggered immunity (PTI).
(A) Flg22-induced FLS2-BAK1 complex formation. Immunoprecipitation of FLS2 in Arabidopsis seedlings either untreated or treated with 100 nM flg22 for 10 min. Blot stained with Coomassie brilliant blue (CBB) is presented to show equal loading. Western blots were probed with α-FLS2 and α-BAK1 antibodies. Numbers indicate quantification of BAK1 bands normalized based on the corresponding intensities of FLS2 bands and relative to the control Col-0. (B, C) Reactive oxygen species (ROS) production after elicitation with 100 nM flg22 (B) or 100 nM elf18 (C). Values are means of total photon counts over 40 min. Red crosses and red horizontal lines denote mean and SEM. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.0001). Similar results were obtained in at least three independent experiments. (D) Root length of 6-day-old seedlings incubated for 3 days in liquid MS medium with or without indicated concentration of flg22. Red crosses and red horizontal lines denote mean and SEM. Conditions that do not share a letter are significantly different in Brown–Forsythe and Welch ANOVA multiple comparison test (p<0.001), n = 9–12 seedlings per condition. Similar results were obtained in at least three independent experiments. (E) Fresh weight of 12-day-old seedlings grown in the absence (mock) or presence of 1 µM of RALF23 peptide. Fresh weight is expressed as relative to the control mock condition. Similar results were obtained in at least three independent experiments. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.001).
-
Figure 3—figure supplement 1—source data 1
Source data points for the graphs in Figure 3—figure supplement 1B–E.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig3-figsupp1-data1-v2.xlsx
-
Figure 3—figure supplement 1—source data 2
Source blots images for the co-immunoprecipitation (co-IP) in Figure 3—figure supplement 1A.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig3-figsupp1-data2-v2.pdf
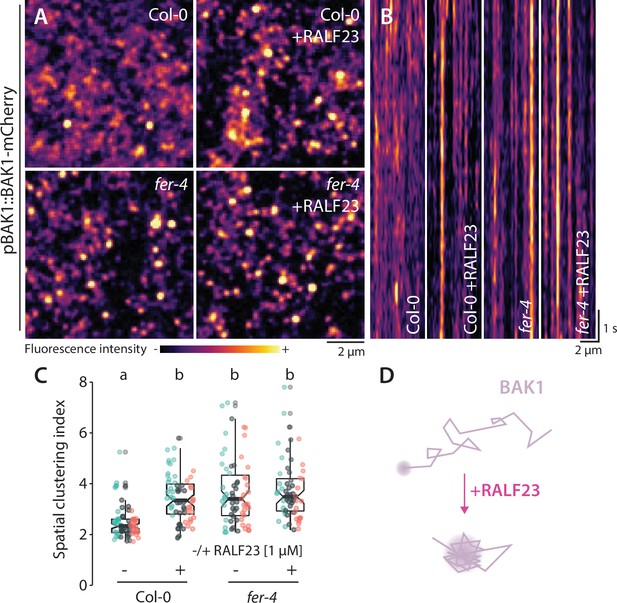
RALF23 perception regulates BAK1-mCherry organization.
(A) BAK1-mCherry nanodomain organization (pBAK1::BAK1-mCherry). Pictures are maximum projection images (10 variable angle total internal reflection fluorescence microscopy [VA-TIRFM] images obtained at 2.5 frames per second) of BAK1-mCherry in Col-0 and fer-4 cotyledon epidermal cells with or without 1 µM RALF23 treatment (2–30 min). (B) Representative kymograph showing lateral organization of BAK1-mCherry overtime in Col-0 and fer-4 with or without 1 µM RALF23 treatment. (C) Quantification of BAK1-mCherry spatial clustering index. Graphs are notched box plots, scattered data points show measurements, colors indicate independent experiments, n = 21 and n = 23 cells for Col-0/pBAK1::BAK1-mCherry with and without RALF23, respectively, n = 20 and n = 21 cells for fer-4/pBAK1::BAK1-mCherry with and without RALF23, respectively. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.0001). (D) Graphical illustration summarizing our observations for BAK1-mCherry nanoscale dynamics upon RALF23 treatment.
-
Figure 4—source data 1
Source data points for the graph in Figure 4C.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig4-data1-v2.xlsx
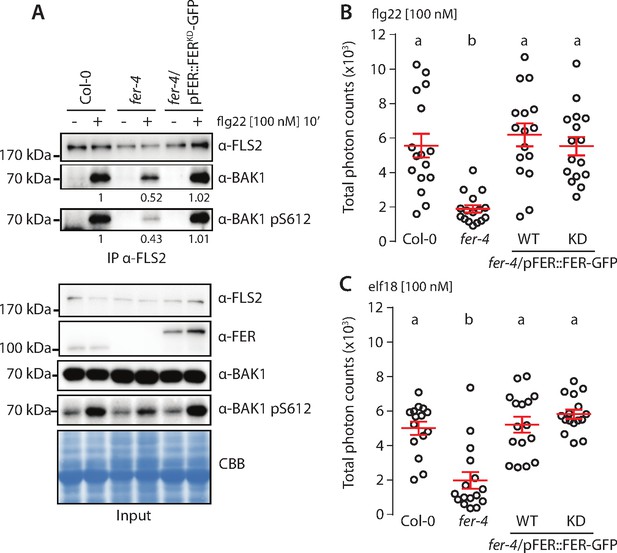
FER kinase activity is dispensable to support pattern-triggered immunity (PTI) signaling.
(A) Flg22-induced FLS2-BAK1 complex formation. Immunoprecipitation of FLS2 in Arabidopsis Col-0, fer-4, and fer-4/pFER::FERKD-GFP seedlings that were either untreated or treated with 100 nM flg22 for 10 min. Blot stained with Coomassie brilliant blue (CBB) is presented to show equal loading. Western blots were probed with α-FLS2, α-BAK1, α-BAK1-pS612, or α-FER antibodies. Numbers indicate bands’ quantification of BAK1 and BAK1-pS612 normalized based on the FLS2 bands’ intensities for each condition and relative to the control Col-0. (B, C) Reactive oxygen species (ROS) production after elicitation with 100 nM flg22 (A) or 100 nM elf18 (B). Values are means of total photon counts over 40 min, n = 16. Red crosses and red horizontal lines denote mean and SEM, respectively. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.0001).
-
Figure 4—figure supplement 1—source data 1
Source data points for the graphs in Figure 4—figure supplement 1B and C.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig4-figsupp1-data1-v2.xlsx
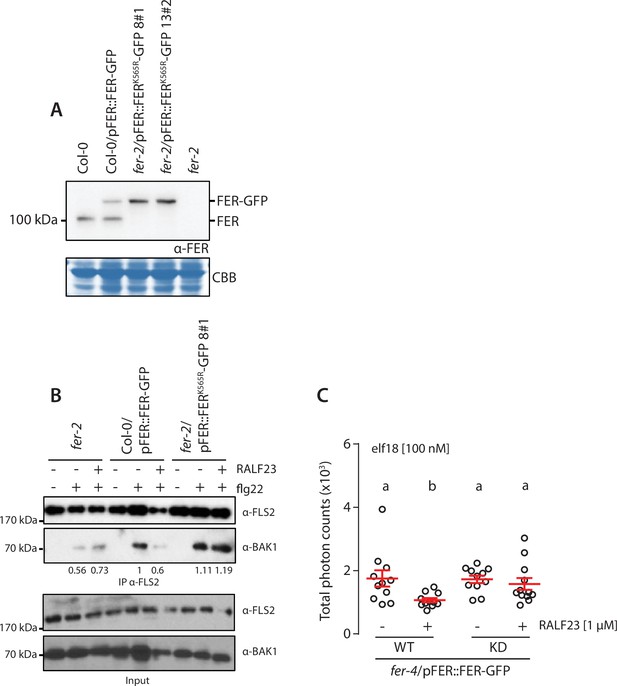
Inhibition of pattern-triggered immunity (PTI) signaling by RALF23 requires FER kinase activity.
(A) Accumulation of endogenous FER and FERK565R-GFP detected by western blot. Total proteins were extracted from 2-week-old seedlings. Blot stained with Coomassie brilliant blue (CBB) is presented to show loading. (B) Flg22-induced FLS2-BAK1 complex formation. Immunoprecipitation of FLS2 in Arabidopsis fer-2, Col-0/pFER::FERK565R-GFP, and fer-2/pFER::FERK565R-GFP seedlings that were either untreated or treated with 100 nM of flg22 and 1 µM RALF23 for 10 min. Western blots were probed with α-FLS2, α-BAK1 antibodies. Numbers indicate bands’ quantification of BAK1 normalized based on the FLS2 bands’ intensities for each condition and relative to the control Col-0/pFER::FERK565R-GFP. (C) Reactive oxygen species (ROS) production after elicitation with 100 nM elf18, with and without 1 µM RALF23 co-treatment. Values are means of total photon counts over 40 min, n = 10. Red crosses and red horizontal lines denote mean and SEM, respectively. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.05). Similar results were obtained in at least three independent experiments.
-
Figure 4—figure supplement 2—source data 1
Source data points for the graph in Figure 4—figure supplement 2C.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig4-figsupp2-data1-v2.xlsx
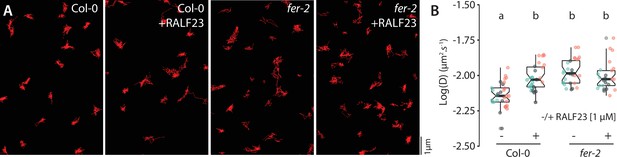
Analysis of FLS2-GFP single-particle dynamics upon RALF23 treatment.
(A) Representative images of FLS2-GFP single particles tracked in Col-0 and fer-2 cotyledon epidermal cells of 5-day-old seedlings with or without 1 µM RALF23 treatment. (B) Quantification of FLS2-GFP diffusion coefficient (D) in Col-0 and fer-2 with or without 1 µM RALF23 treatment. Graph is a notched box plot, scattered data points correspond to the average diffusion coefficient of single particles for each cell, colors indicate independent experiments, n = 27 and n = 29 cells for Col-0/pFLS2::FLS2-GFP with and without RALF23, respectively, n = 25 and n = 28 cells for fer-2/pFLS2::FLS2-GFP with and without RALF23, respectively. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.0001).
-
Figure 4—figure supplement 3—source data 1
Source data points for the graph in Figure 4—figure supplement 3B.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig4-figsupp3-data1-v2.xlsx
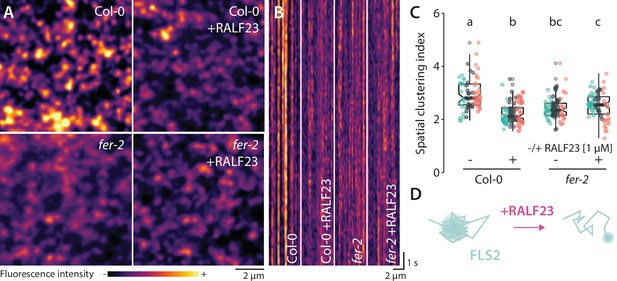
Analysis of FLS2-GFP organization upon RALF23 treatment.
(A) Pictures are maximum projection images (20 variable angle total internal reflection fluorescence microscopy [VA-TIRFM] images obtained at 20 frames per second) of FLS2-GFP in Col-0 and fer-2 cotyledon epidermal cells with or without 1 µM RALF23 treatment. (B) Representative kymograph showing lateral organization of FLS2-GFP overtime in Col-0 and fer-2 cotyledon epidermal cells with or without 1 µM RALF23 treatment. (C) Quantification of FLS2-GFP spatial clustering index. Graph is a notched box plot, scattered data points show individual measurements, colors indicate independent experiments, n = 29 and n = 24 cells for Col-0/pFLS2::FLS2-GFP with and without RALF23, respectively. n = 27 and n = 22 cells for fer-2/pFLS2::FLS2-GFP with and without RALF23, respectively. Conditions that do not share a letter are significantly different in Dunn’s multiple comparison test (p<0.0001). Similar results were obtained in at least three independent experiments. (D) Graphical illustration summarizing our observations for FLS2-GFP nanoscale dynamics upon RALF23 treatment.
-
Figure 4—figure supplement 4—source data 1
Source data points for the graph in Figure 4—figure supplement 4C.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig4-figsupp4-data1-v2.xlsx
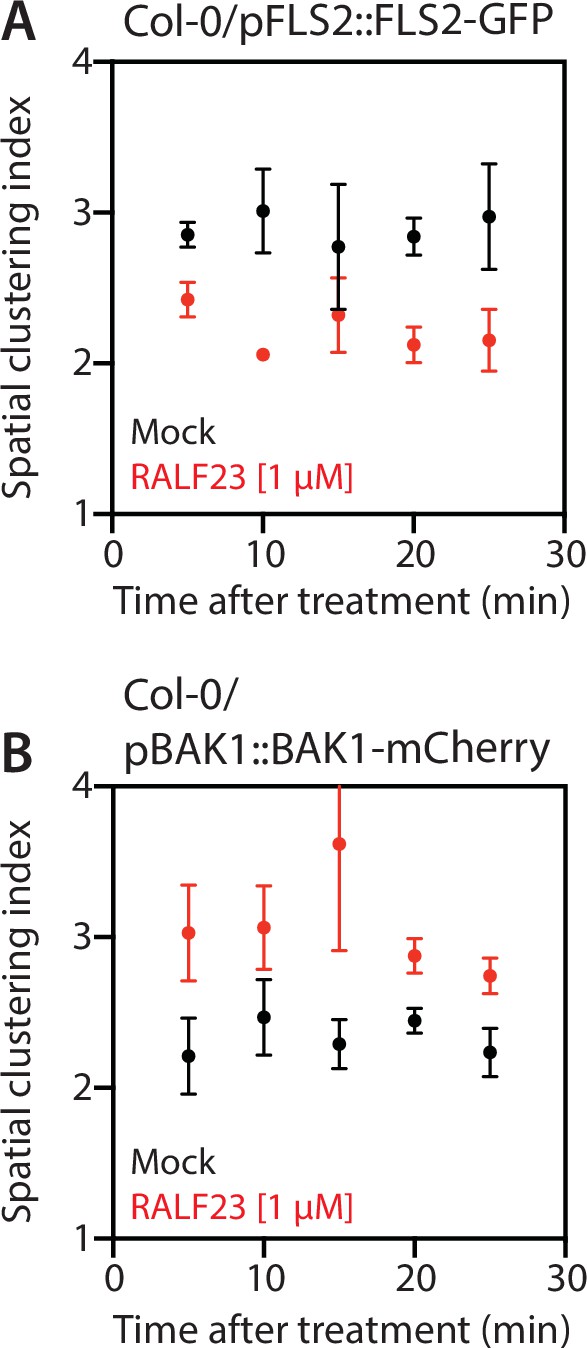
Time-resolved analysis of the spatial clustering index.
Data points correspond to the average of the spatial clustering index measured over 5 min windows for FLS2-GFP (A) and BAK1-mCherry (B) with or without 1 µM RALF23 treatment. The first data point (5 min) corresponds to the value obtained between 2 and 3 min post treatment. Acquisition approximately started 2 min after treatments, the time required for sample mounting.
-
Figure 4—figure supplement 5—source data 1
Source data points for the graphs in Figure 4—figure supplement 5A and B.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig4-figsupp5-data1-v2.xlsx
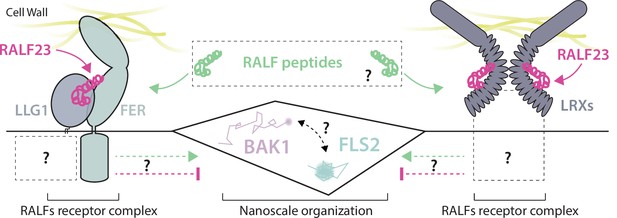
Working model for the regulation of FLS2 and BAK1 nanoscale organization by RALFs receptor complexes.
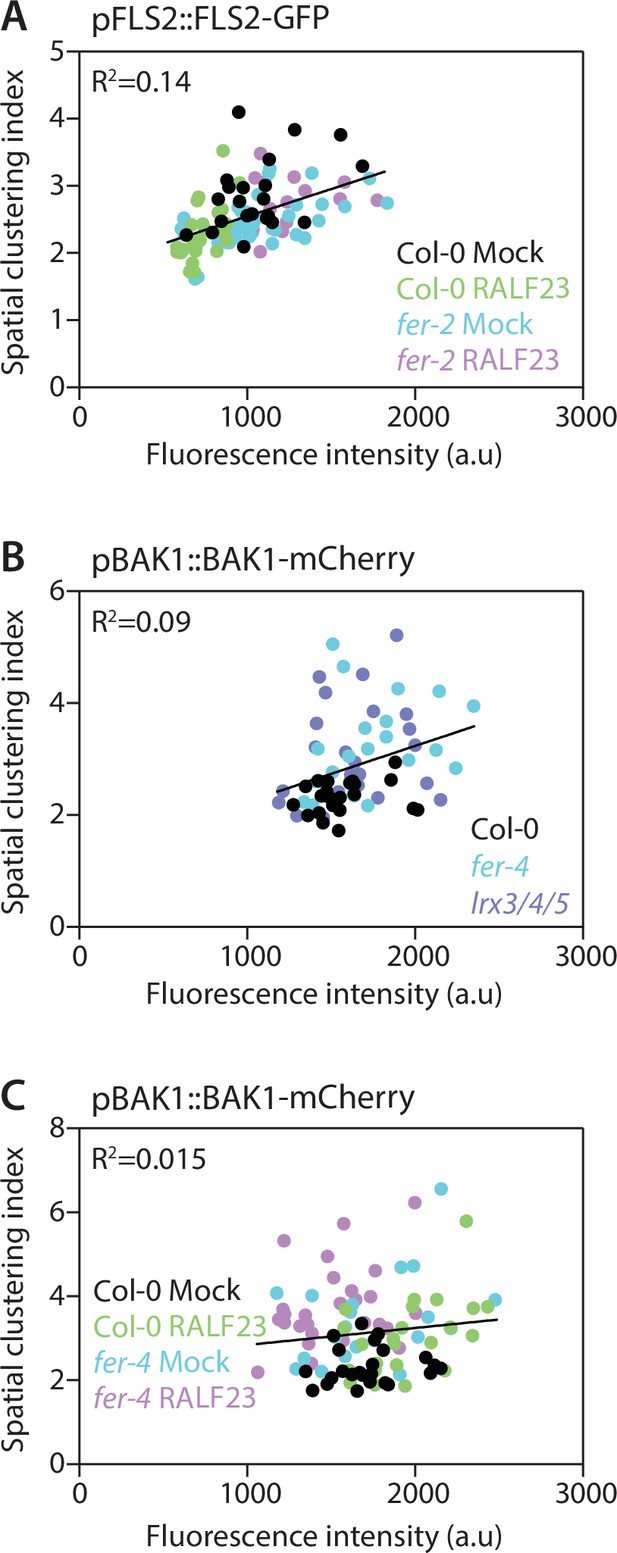
Linear regression analysis of the relationship between the spatial clustering index and fluorescence intensity.
Spatial clustering index is calculated as the ratio of the mean of the 5% highest values to the mean of 5% lowest values of fluorescence intensity obtained on a plot line. Here, spatial clustering index values were plotted against the mean fluorescence intensity of the corresponding plot line for FLS2-GFP (A) and BAK1-mCherry (B, C). Linear regression analyses show poor to no correlation between the spatial clustering index and fluorescence intensity (R2 ranking from 0.14 and 0.015).
-
Figure 4—figure supplement 7—source data 1
Source data points for the graphs in Figure 4—figure supplement 6A–C.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig4-figsupp7-data1-v2.xlsx
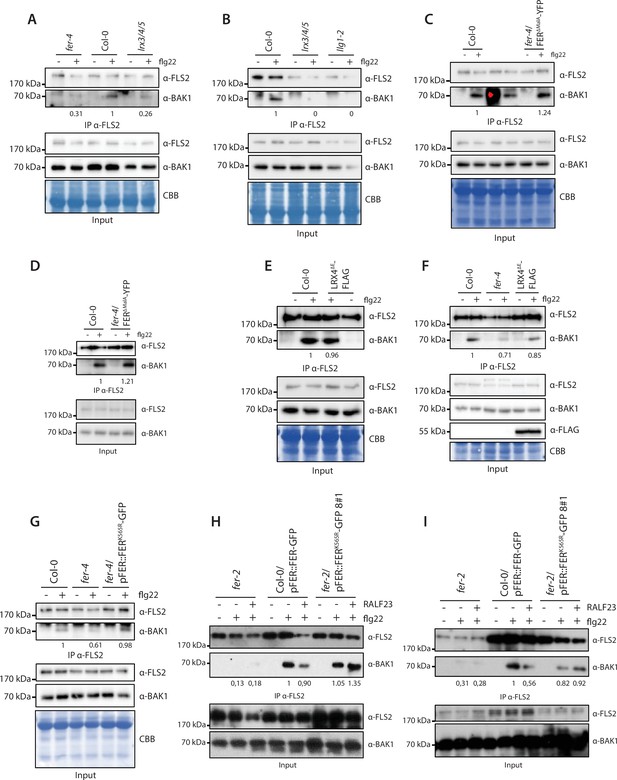
Replicates of co-immunoprecipitation experiments.
Replicate of immunoprecipitation experiments corresponding to Figure 2 (A, B), Figure 3 (C, D), Figure 3—figure supplement 1 (E, F), Figure 4—figure supplement 1 (G), and Figure 4—figure supplement 2 (H, I).
-
Figure 4—figure supplement 8—source data 1
Source blots images for the replicate of co-immunoprecipitation (co-IP) experiments.
- https://cdn.elifesciences.org/articles/74162/elife-74162-fig4-figsupp8-data1-v2.pdf
Variable angle total internal reflection fluorescence microscopy (VA-TIRFM) imaging of FLS2-GFP in Col-0 with or without RALF23 treatment.
Representative stream images acquisition of FLS2-GFP particles observed at the surface of 5-day-old cotyledon epidermal cells by VA-TIRF microscopy at 5 frames per second. Scale bar indicates 2 µm.
Variable angle total internal reflection fluorescence microscopy (VA-TIRFM) imaging of FLS2-GFP in fer-4 with or without RALF23 treatment.
Representative stream images acquisition of FLS2-GFP particles observed at the surface of 5-day-old cotyledon epidermal cells by VA-TIRF microscopy at 5 frames per second. Scale bar indicates 2 µm.
Variable angle total internal reflection fluorescence microscopy (VA-TIRFM) imaging of FLS2-GFP in Col-0 and fer-2 with or without RALF23 treatment.
Representative stream images acquisition of FLS2-GFP particles observed at the surface of 5-day-old cotyledon epidermal cells by VA-TIRF microscopy at 20 frames per second. Scale bar indicates 2 µm.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (Arabidopsis thaliana) | Col-0/pFLS2::FLS2-GFP#1 | Göhre et al., 2008 | See Materials and methods | |
| Genetic reagent (A. thaliana) | Col-0/pFLS2::FLS2-GFP#2 | This paper | See Materials and methods | |
| Genetic reagent (A. thaliana) | fer-2/pFLS2::FLS2-GFP#1 | Stegmann et al., 2017 | See Materials and methods | |
| Genetic reagent (A. thaliana) | fer-4/pFLS2::FLS2-GFP#2 | This paper | See Materials and methods | |
| Genetic reagent (A. thaliana) | fer-4 | Duan, 2010 | See Materials and methods | |
| Genetic reagent (A. thaliana) | fer-4/pFER::FER-GFP | Duan, 2010 | See Materials and methods | |
| Genetic reagent (A. thaliana) | fer-4/pFER::FERKD-GFP | Chakravorty et al., 2018 | See Materials and methods | |
| Genetic reagent (A. thaliana) | lrx3/4/5 | Dünser, 2019 | See Materials and methods | |
| Genetic reagent (A. thaliana) | p35S::LRX4ΔE-Citrine | Dünser, 2019 | See Materials and methods | |
| Genetic reagent (A. thaliana) | p35S::LRX4ΔE-FLAG | Dünser, 2019 | See Materials and methods | |
| Genetic reagent (A. thaliana) | Col-0/pBAK1::BAK1-mCherry | Bücherl et al., 2013 | See Materials and methods | |
| Genetic reagent (A. thaliana) | fer-4/ pBAK1::BAK1-mCherry | This paper | See Materials and methods | |
| Genetic reagent (A. thaliana) | lrx3/4/5/ pBAK1::BAK1-mCherry | This paper | See Materials and methods | |
| Genetic reagent (A. thaliana) | lrx3/4/5/ pFER::FER-GFP | This paper | See Materials and methods | |
| Antibody | anti-FLAG-HRP | Sigma-Aldrich | A8592 | WB (1:4000 dilution) |
| Antibody | Monoclonal rabbit anti-FLS2 | Chinchilla et al., 2007 | WB (1:1000 dilution) | |
| Antibody | Polyclonal rabbit anti-BAK1 | Roux, 2011 | WB (1:5000 dilution) | |
| Antibody | Polyclonal rabbit anti-BAK1 pS612 | Perraki, 2018 | WB (1:3000 dilution) | |
| Antibody | Polyclonal rabbit anti-FER | Xiao et al., 2019 | WB (1:2000 dilution) | |
| Antibody | Anti-rabbit IgG-HRP Trueblot | Rockland | 18-8816-31 | WB (1:10,000 dilution) |
| Peptide, recombinant protein | Flg22 | Synthesized by EZBiolab(purity >95%) | See Materials and methods | |
| Peptide, recombinant protein | Elf18 | Synthesized by EZBiolab(purity >95%) | See Materials and methods | |
| Peptide, recombinant protein | RALF23 | Synthesized by EZBiolab(purity >95%) | See Materials and methods | |
| Chemical compound, drug | GFP-Trap agarose beads | ChromoTek | See Materials and methods | |
| Chemical compound, drug | M2 anti-Flag affinity gel | Sigma-Aldrich | A2220-5ML | See Materials and methods |
| Chemical compound, drug | Anti-rabbit Trueblot agarose beads | eBioscience | SML1656 | See Materials and methods |
| Software, algorithm | Fiji | https://imagej.net/Fiji | See Materials and methods |
Additional files
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/74162/elife-74162-transrepform1-v2.pdf
-
Source data 1
Raw images of the western blots.
- https://cdn.elifesciences.org/articles/74162/elife-74162-supp1-v2.zip





