Hierarchical architecture of dopaminergic circuits enables second-order conditioning in Drosophila
Figures
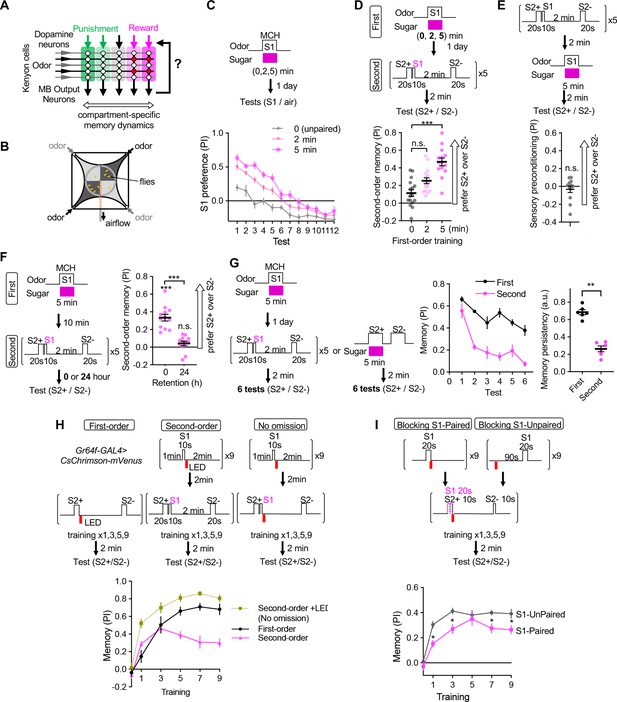
Appetitive olfactory second-order conditioning in Drosophila.
(A) A simplified diagram of the mushroom body circuit. Identity of odors are encoded by patterns of activity in ~2000 Kenyon cells. Contingent activity of Kenyon cells and dopamine release leads to plasticity of excitatory synapses from Kenyon cells to MB output neurons with compartment-specific dynamics. (B) A diagram of the four-armed olfactory arena. Flies were confined in the 9 cm diameter circular area above the LED board. For odor-sugar conditioning, flies were first trained in a tube by pairing an odor with dried sugar paper, and then introduced to the olfactory arena. Performance index was calculated by counting the number of flies in each quadrant (see Methods). (C) Dynamics of MCH preference after various 2 or 5 min of first-order conditioning with sugar. Flies were trained after 40–48 hr of starvation and memories were tested 20–24 hr later without feeding in between by examining preference to MCH over air for 12 times. Unpaired group received 5 min of sugar 2 min prior to 5 min exposure to MCH. Mean performance index of the first 5 tests after 5 min training was higher than that of 2 min. p<0.01; unpaired t-test; N=10–12. (D) Second-order memory performance by wild type flies. n.s., not significant (p=0.152); ***, p<0.0001; Dunn’s multiple comparison tests following Kruskal-Wallis test; N=14–16. Means and SEMs are displayed with individual data points. (E) The odor preference following the sensory preconditioning protocol, in which the order of the first and second-order conditioning was swapped. n.s., not significantly different from the chance level; Wilcocxon signed-rank test; N=12. (F) Retention of second-order memory. After 24 hr, the second-order memory decayed to the chance level. ***, p<0.001; Wilcocxon signed-rank test or Mann-Whiteney test; N=12. (G) Odor preference between two S2 odors after the second-order or first-order conditioning was measured for six times by alternative position of two odorants with 2 min intervals. Memory persistency, a mean of PIs for 3rd-6th tests divided by PI of 1st test, was significantly smaller for second-order memory. **; p<0.0022; Mann-Whitney test; N=6. Means and SEMs are displayed with individual data points. (H) Learning curves by first-order, second-order, or second-order without omission of optogenetic reward. Flies expressing CsChrimson in sugar sensory neurons with Gr64f-GAL4 were trained by pairing S2+ odor with activation of LED (First) or S1 odor that was previously paired with LED (Second). In the no omission protocol, sugar sensory neurons were activated immediately after S1 by repeating 1 s red LED illumination with 1 s intervals for three times. Preference between S2+ and S2− odors was tested after 1st, 3rd, 5th, 7th, and 9th training sessions. After 9th training, memory by second-order protocol was lower than other protocols and its peak at 3rd training (p<0.05); Dunn’s multiple comparison tests following Kruskal-Wallis test; N=8. (I) Learning of S2 odors was compromised when S1 odor paired with Gr64f>CsChrimson precedes S2+ odor. *, p<0.05 by Dunn’s tests following Kruskal-Wallis test; N=12.
-
Figure 1—source data 1
Numerical data for Figure 1.
- https://cdn.elifesciences.org/articles/79042/elife-79042-fig1-data1-v2.xlsx
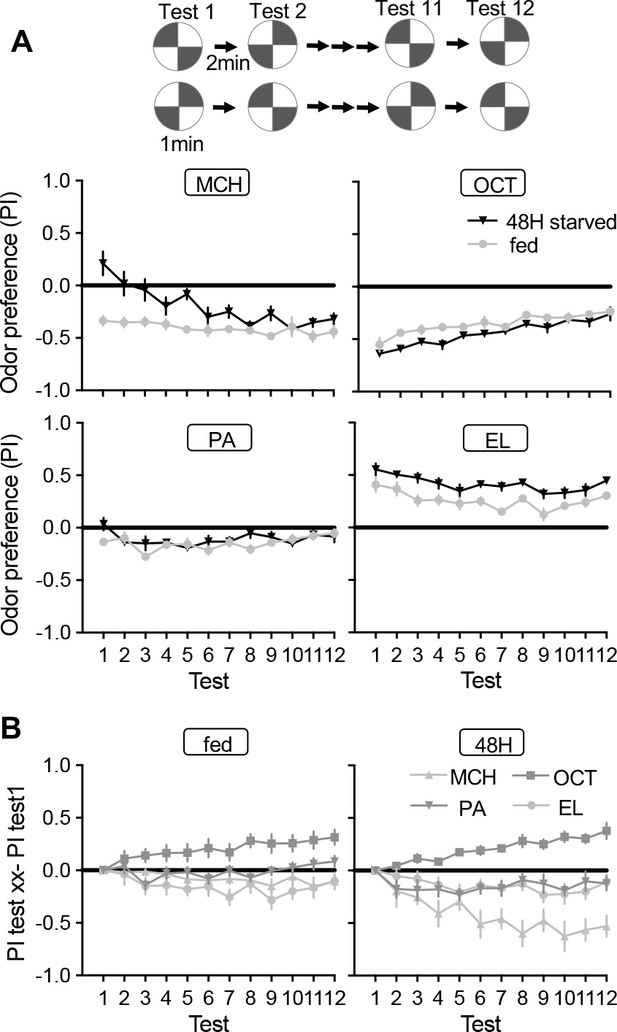
Dynamics of odor preference.
(A) Twelve repetition of odor preference of fed and 40–48 hr starved flies. PIs represent results of reciprocal experiments. The half groups of flies went through the identical tests but with alternating positions of odors and air quadrants to cancel out potential positional preference for each rigs. The mean odor preferences during 12 tests were significantly different between fed and starved flies for MCH and EL. p<0.01; unpaired t-test; N=8. Means and SEM are shown. (B) Delta between the first test and subsequent tests in fed (left) and 40–48 hr starved flies (right). Areas under curve for OCT was significantly higher than that for other odors in both fed and starve flies, whereases the area under the curve for MCH was lower than other odors only in starved flies. p<0.01; unpaired t-test; N=8.
-
Figure 1—figure supplement 1—source data 1
Numerical data for Figure 1.
- https://cdn.elifesciences.org/articles/79042/elife-79042-fig1-figsupp1-data1-v2.xlsx
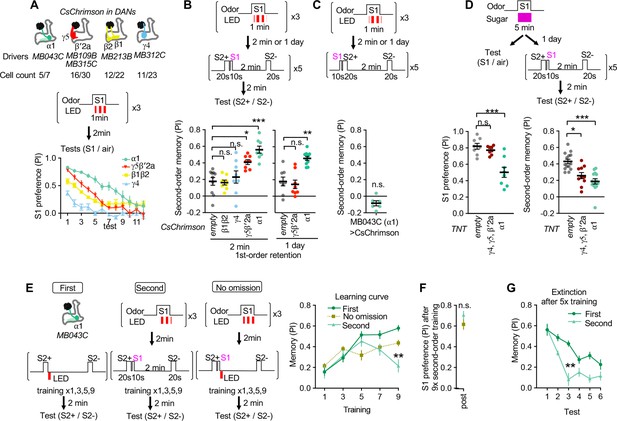
Identification of the teacher compartment(s).
(A) Dynamics of S1 odor (MCH) preference after pairing 1 min of S1 odor with activation of different PAM-cluster DANs for three times. Numbers of CsChrimson-mVenus in each driver per hemisphere and total number of corresponding DAN cell types in EM hemibrain data are indicated. At 3rd-7th tests, MCH preference of MB043C>CsChrimson flies was higher than all other genotypes. p<0.05; Dunn’s multiple comparison tests following Kruskal-Wallis test; N=6. (B) The second-order conditioning 2 min or 1 day after the first-order conditioning with optogenetic activation of various DAN types. Second-order memory was tested immediately after pairing S2+ odor with S1 odor (MCH) five times. n.s., not significant; *, p=0.0330; **, p=0.0046 ***, p<0.001; Dunn’s multiple comparison tests following Kruskal-Wallis test; N=8–10. (C) The second-order memory immediately after backward second-order conditioning. Flies expressing CsChrimon-mVenus by MB043C split-GAL4 were trained with identical protocol as in B, except that the onset of S1 odor was shifted to the 10 second before the onset of the first S2 odor. n.s., not significant from zero; Wilcocxon signed-rank test; N=6. (D) Preference to the S1 odor (left) and second-order memory (right) by flies expressing TNT with empty, MB196B or MB043C split-GAL4. MB196B labels ~ 27 cells per hemisphere, including PAM-γ4, PAM-γ4<γ1γ2, γ5 and β′2a. *, p=0.0126; ***, p<0.001; Dunn’s multiple comparison tests following Kruskal-Wallis test; N=8 for S1 preference; N=10–14 for second-order. (F) Learning curves by first-order, second-order, or second-order without omission of optogenetic reward. Flies expressing CsChrimson with MB043C split-GAL4 were trained by pairing S2+ odor directly with optogenetic activation of DANs (First) or S1 odor that was previously paired with DAN activation (Second). In the no omission protocol, DANs were activated immediately after S1 by repeating 1 s red LED illumination with 1 s intervals for three times. Preference between S2+ and S2− odors was tested after 1st, 3rd, 5th, 7th, and 9th training sessions. After 9th training, memory by second-order protocol was lower than other protocols and its peak at 5th training. **, p<0.01; Dunn’s multiple comparison tests following Kruskal-Wallis test; N=8–10. (G) The preference for the S1 odor (MCH) after the 9th session of second-order conditioning as in F. n.s., not significant; Mann-Whitney test; N=8. (H) Comparison of memory decay after repetitive tests. Flies were trained five times with first or second-order conditioning protocol as in F but without tests. Immediately after the 5th training, preference between two S2 odors was measured repeatedly without training. At third test, second-order memory was significantly lower than first-order memory. **, p=0.0036; Dunn’s multiple comparison tests following Kruskal-Wallis test; N=8.
-
Figure 2—source data 1
Numerical Data for Figure 2.
- https://cdn.elifesciences.org/articles/79042/elife-79042-fig2-data1-v2.xlsx
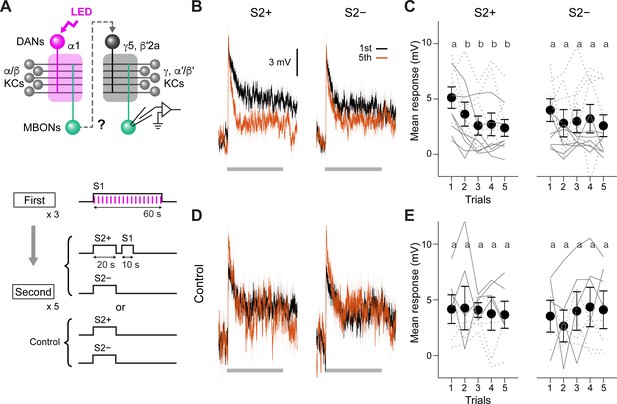
Second-order conditioning induces cross-compartmental plasticity.
(A) Experimental design and protocol. ChrimsonR-mVenus was selectively expressed in PAM-α1 using MB043-split-LexA (58E02-ZpLexADBD in JK22C; 32D11-p65ADZp in JK73A; see Figure 3—figure supplement 1 for expression pattern), and in vivo whole-cell recordings were made from MBON-γ5β′2a, which was labeled by mScarlet using a split-GAL4 driver SS01308. For the first-order conditioning, 1 min presentation of S1 (MCH) was paired with LED stimulation (1ms, 2 Hz, 120 times), which caused odor-specific suppression of responses in MBON-α1 (Figure 3—figure supplement 2). After repeating first-order conditioning three times with 2 min intervals, second-order conditioning was performed by presenting S2+ (either PA or EL) for 20 s, and then S1 for 10 s with 5 s delay. S2− was presented alone 2 min later. Second-order conditioning was repeated five times, and the responses to S2 were recorded. In control experiments, first-order conditioning was performed in the same manner, but the presentation of S1 was omitted during second-order conditioning. Reciprocal experiments were performed by swapping S2+ and S2− in separate flies. (B) Mean responses ( ± SEM in light colors) to S2+ and S2− in the first (black) and fifth trials (red) during second-order conditioning (n=14, including reciprocal experiments). Horizontal gray bars indicate 20 s odor presentation period. (C) Mean response magnitudes ( ± SEM) evoked by S2+ and S2−. The response magnitude was calculated by averaging the depolarization during the response window (0–20.6 s from odor onset). Each solid (PA used as S2+; n=7) and dashed line (EL as S2+; n=7) indicates data from a single fly. Responses to S2+ underwent depression after the first trial, while those to S2− did not change. Different letters indicate significant differences detected by Tukey’s post hoc multiple comparisons test (p<0.05) following repeated-measures two-way ANOVA (p=0.003). There was no significant change in the peak amplitude (p=0.87). (D, E) Same as (B) and (C) except that the data are from control experiments (n=4 each with PA or EL used as S2+, respectively). Neither responses to S2+ nor S2− changed (p=0.28; repeated-measures two-way ANOVA). The peak response did not change either (p=0.22).
-
Figure 3—source data 1
Numerical data for Figure 3.
- https://cdn.elifesciences.org/articles/79042/elife-79042-fig3-data1-v2.zip
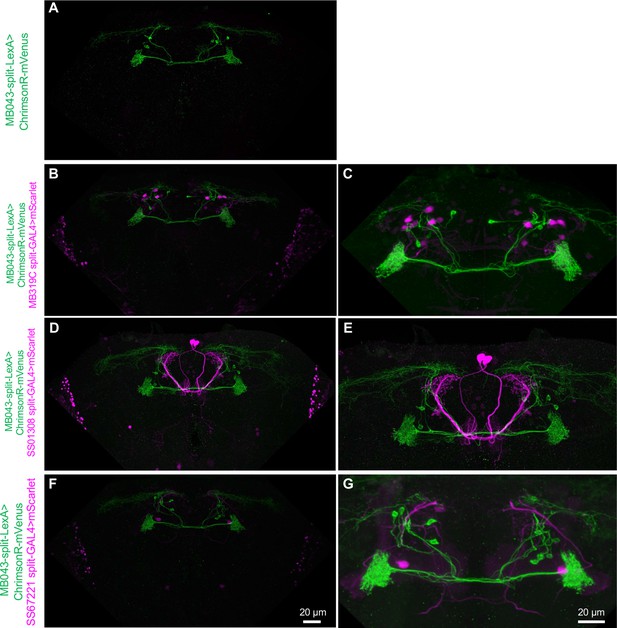
Expression patterns of MB043-split-LexA, MB319C and SS67221-split-GAL4.
(A) MB043-split-LexA drove expression ChrimsonR-mVenus in 4.7 PAM-α1 neurons on average: (4, 4, 5, 4, 5, 6) cells per hemisphere were observed in three brain samples. (B–C) The expression pattern of MB043-split-LexA was unaffected in the presence of MB319C-split-GAL4. We observed a few additional mScarlet-positive cells in addition to the two MBON-α1, presumably because interference between the AD hemi-driver of the MB043-split-LexA and DBD hemi-driver of MB319C-split-GAL4. For electrophysiology, MBON-α1 was found based on their soma location, brightness of mScarelt signals and odor response. (D–E) Expression patterns of MB043-split-LexA and SS67221-GAL4-split-GAL4. Additional cells expressed mScarlet in optic lobes, but MBON-γ5β′2 a was unambiguously labeled by mScarlet in the central brain. (F–G) Expression patterns of MB043-split-LexA and SS67221-GAL4-split-GAL4 did not interfere.
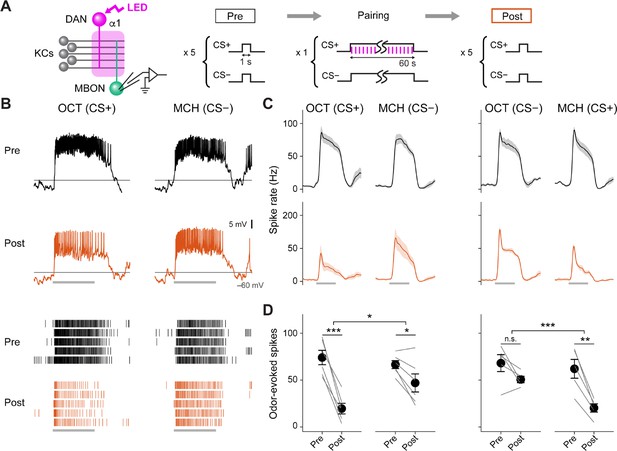
Optogenetic conditioning in α1 compartment induces depression in MBON-α1.
(A) Experimental design and protocol. ChrimsonR-mVenus was selectively expressed in PAM-α1 using MB043-split-LexA, and in vivo whole-cell recordings were made from MBON-α1, which was labeled by mScarlet using a split-GAL4 driver MB319C. 1 min presentation of conditioned stimulus (CS+; OCT or MCH) was paired with LED stimulation (1ms, 2 Hz, 120 times), followed by 1 min presentation of CS− alone. Reciprocal experiments were performed by swapping CS+ and CS− in a separate set of flies. (B) Membrane voltage (upper panels) and spike data (lower panels) from a single representative fly, in which OCT was used as CS+. Gray bars indicate 1 s odor presentation. (C) Time courses of instantaneous spike rate (mean ± SEM; n=6 and 5 for each set of experiment). (D) Summary data of mean odor-evoked spike counts ( SEM). Gray lines indicate data from individual neurons. After each pairing, responses to CS+ were suppressed, while those to CS− were either showed less suppression than CS+ or no change (repeated-measures two-way ANOVA followed by Tukey’s post hoc multiple comparisons test; *p<0.05, **p<0.005, ***p<0.001).
-
Figure 3—figure supplement 2—source data 1
Numerical data for Figure 3—figure supplement 2.
- https://cdn.elifesciences.org/articles/79042/elife-79042-fig3-figsupp2-data1-v2.zip
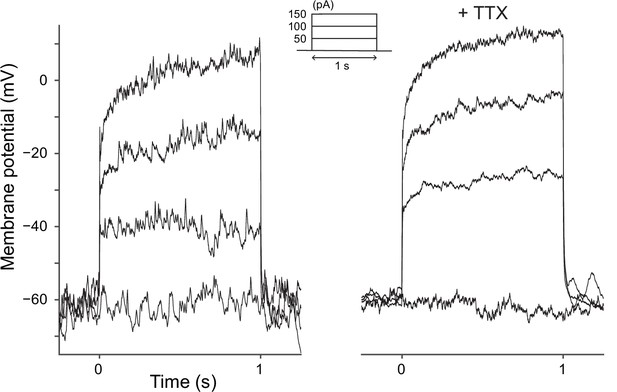
Response to current injection in MBON-γ5β′2a.
Representative somatic voltage responses to current injection in MBON-γ5β′2a with (right) or without (left) tetrodotoxin (TTX; 1 mM). The current injection waveforms are shown in the inset in the middle. Depolarization typically increased the frequency of small, fast membrane potential fluctuations, which were partially suppressed by TTX. However, those events were not readily distinguishable from the putative synaptic potentials that were remaining in the presence of TTX. Therefore, it remains inconclusive whether MBON-γ5β′2a elicits action potentials.
-
Figure 3—figure supplement 3—source data 1
Numerical data for Figure 3—figure supplement 3.
- https://cdn.elifesciences.org/articles/79042/elife-79042-fig3-figsupp3-data1-v2.xlsx
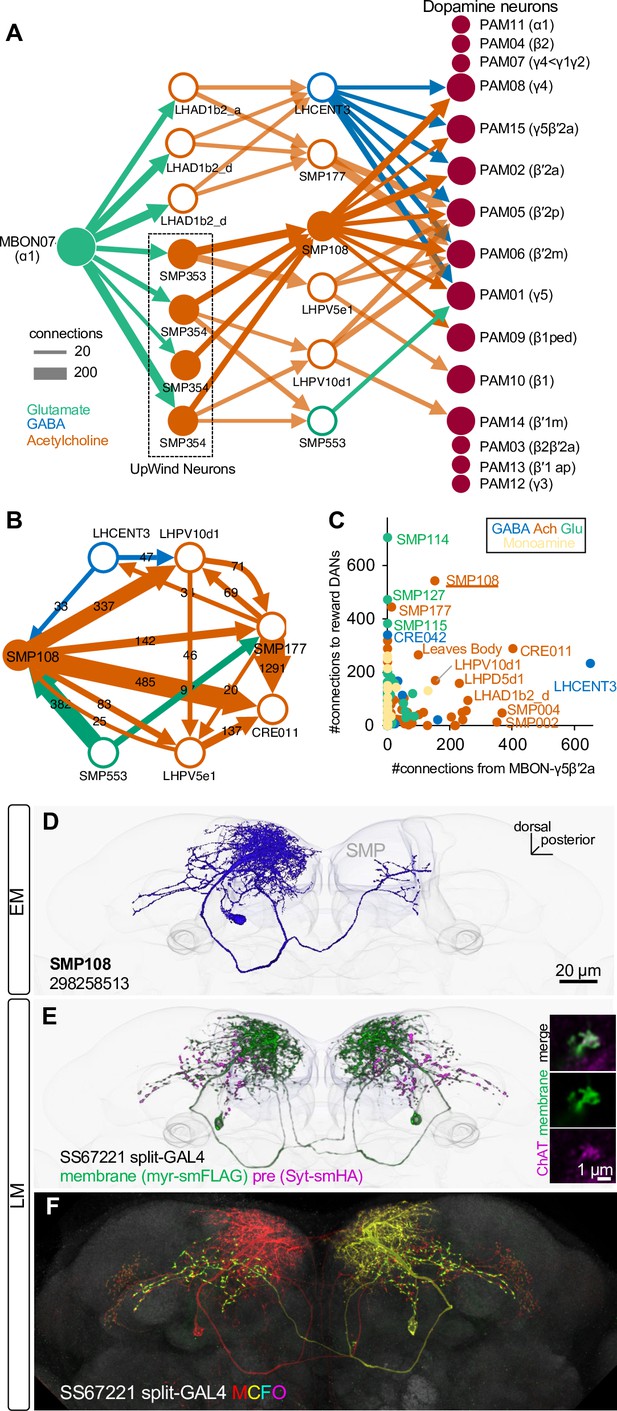
SMP108 is a key interneuron between MBON-ɑ1 and DANs.
(A) The connections from MBON-ɑ1 to PAM cluster DANs with two interneurons identified in the hemibrain EM data (Scheffer et al., 2020). The width of arrows indicate number of connections. The colors of circles and arrows indicate type of putative neurotransmitter. Single SMP353 and three SMP354s have similar morphology and projection patterns and converge on to SMP108. Cholinergic interneurons SMP353/SMP354 and SMP108 are shown as filled orange circles and arrows. Other cholinergic connections are shown in transparent orange. See Supplementary file 1 for a full connectivity matrix and neurotransmitter predictions. See Figure 4—figure supplement 2 for the SMP108’s connections with subtypes of DANs. (B) Connections between the six neurons in the second layer in A and CRE011. SMP108 outputs to all three other putative cholinergic interneurons. LHPV10d1 is the top target of SMP108. SMP553 send its first and second strongest outputs to SMP108 and SMP177. (C) Total number of connections to reward DANs (PAM01, 02, 04, 06, 07, 08, 10, 11,15) which can induce appetitive memory with optogenetic activation, plotted against number of inputs from MBON-γ5β′2a. Each circle represents one of 396 interneuron cell types that have at least 100 total connections with MBONs and DANs. Similar to SMP108, CRE011 is an outlier cell type in terms of the high number of direct inputs from MBON-γ5β′2a and outputs to reward DANs. See Figure 4—figure supplement 3 for other kinds of connections between these interneurons and DANs/MBONs. (D) A projection of a reconstructed SMP108 neuron in the hemibrain EM images aligned to a standard brain with outline of the brain and the MB lobes. (E) Confocal microscope images of SS67221 split-GAL4 driver with membrane-targeted reporter myr-smFLAG and presynaptic reporter Syt-smHA. Inset shows anti-ChAT immunoreactivity of SMP108’s axon terminals. (F) Morphology of individual SMP108 visualized by multi-color flip out of SS67221 split-GAL4.
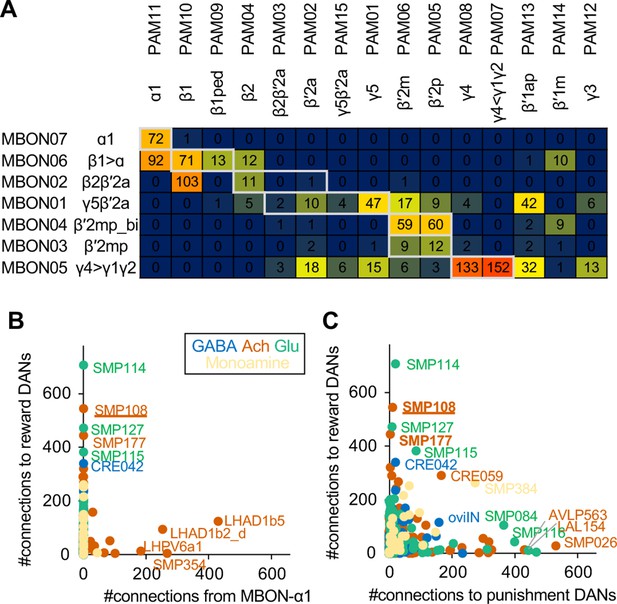
Connections of MBON-ɑ1 and SMP108.
(A) Numbers of direct MBON-to-DANs synaptic connections. White boxes indicate within-a-compartment connection such as connection from MBON-ɑ1 to PAM-ɑ1. (B) Total number of outputs to reward DANs (i.e. PAM01, 02, 04, 06, 07, 08, 10, 11,15) which can induce appetitive memory upon optogenetic activation, plotted against total number of connection from MBON-ɑ1 for 396 cell types that have at least 100 total connections with MBONs and DANs. Colors indicate predicted neurotransmitters. One of outstanding cell type, LHAD1b5, cannot mediate cross-compartmental pathways because it is exclusively connected with PAM-ɑ1 but not other reward DANs. Another outstanding cell type LHAD1b2_d is a part of two-hop pathways from MBON-ɑ1 to PAM-ɑ1 (Figure 3A). (C) Total number of outputs to reward DANs plotted against total number of outputs to punishment DANs (i.e. PPL101, 103, 105, 106 and PAM12).
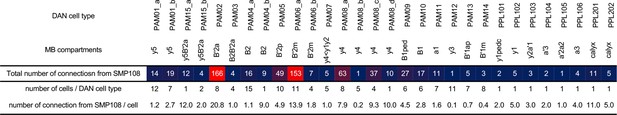
Connections from SMP108 to DAN subtypes.
The number in the top row is the total number of connections from the SMP108 to subtypes of DANs that were defined by their projection and connectivity patterns (Li et al., 2020). The number of cells per subtype and number of connections per cell are shown in the middle and bottom row, respectively.
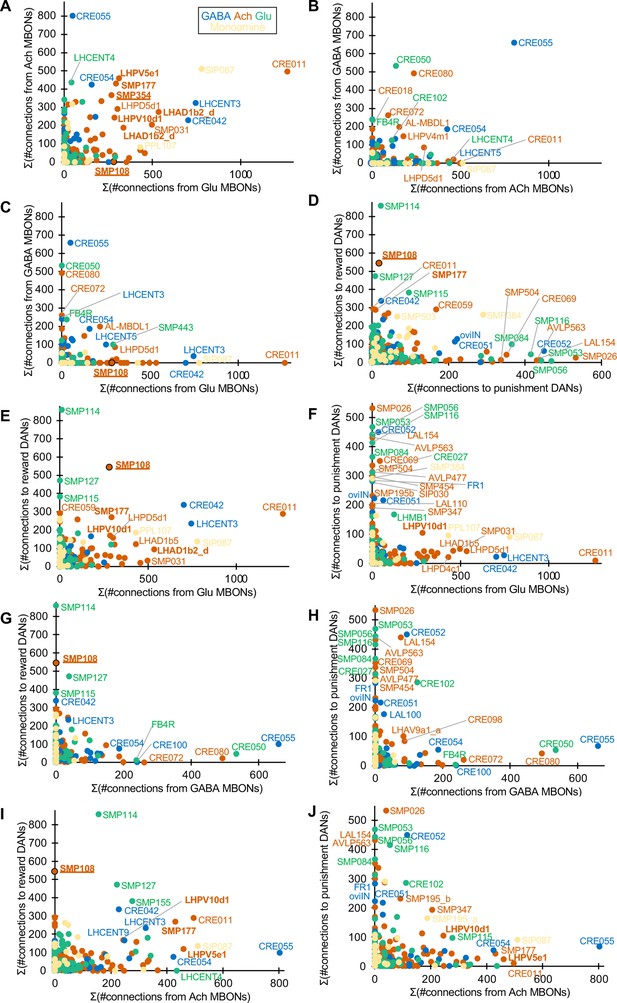
Connections of interneurons with DANs and MBONs.
Scatter plots display designated pairs of connections for 396 cell types that have at least 100 total connections with MBONs and DANs in EM hemibrain data. Colors indicate predicted neurotransmitters. Names of outlier cell types are labeled. See Supplementary file 1 for full data. Note that SMP354 and other cholinergic interneurons in 2-hop pathway (Figure 4A), SMP177, LHPV5e1, LHAD1b2_d, LHPV10d1 showed a shared property of receiving converging inputs from glutamatergic MBONs and cholinergic MBONs, whose activity represent appetitive or aversive memories, respectively. In addition to input from MBON-ɑ1, SMP354 receive converging inputs from MBON-ɑ3, a compartment of long-term aversive memory.
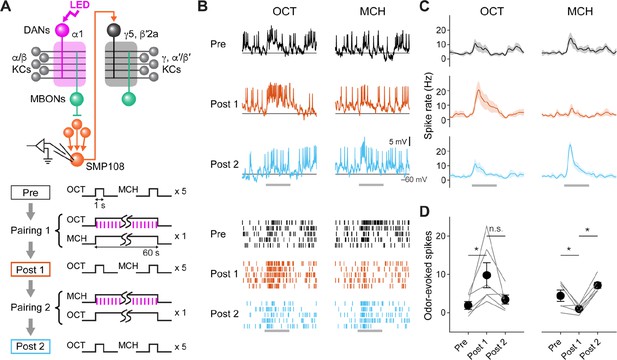
SMP108 acquires enhanced responses to reward-predicting odors.
(A) Experimental design and protocol. ChrimsonR-mVenus was expressed in PAM-cluster DANs, which include PAM-α1, using R58E02-LexA. In vivo whole-cell recordings were made from SMP108, which was labeled by GFP using a split-GAL4 driver SS45234. In the first pairing (Pairing 1), 1 min presentation of OCT was paired with LED stimulation (1ms, 2 Hz, 120 times), followed by 1 min presentation of MCH alone. Odors were flipped in the second round of pairing (Pairing 2). Responses to each odor (1 s presentation) were measured before (Pre) and after pairing 1 (Post 1), and after pairing 2 (Post 2). (B) Membrane voltage (upper panels) and spike data (lower panels) from a single representative neuron. Gray bars indicate 1 s odor presentation. (C) Time courses of instantaneous spike rate (mean ± SEM; n=6). (D) Summary data of mean odor-evoked spike counts ( ± SEM). Gray lines indicate data from individual neurons. After each pairing, responses to paired odors were potentiated, while those to unpaired odors tended to decrease. Repeated-measures two-way ANOVA (p=0.0001) followed by Tukey’s post hoc multiple comparisons test. *p<0.05.
-
Figure 5—source data 1
Numerical data for Figure 5.
- https://cdn.elifesciences.org/articles/79042/elife-79042-fig5-data1-v2.zip
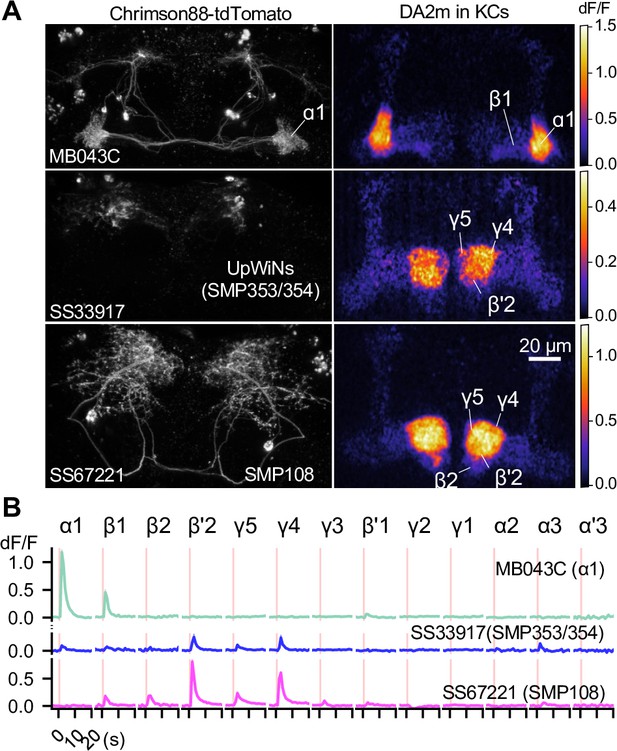
SMP108 promotes dopamine release in multiple compartments.
(A) Representative images of Chrimson88-tdTtomato expression patterns (left) and maximum intensity projections of DA2m dF/F in the MB lobes (right). Release of dopamine upon activation of DANs or SMP108 pathways, measured with dopamine sensor DA2m expressed in Kenyon cells. 10XUAS-Syn21-Chrimson88-tdTtomato-3.1 in attP18 was driven with designated split-GAL4 driver lines. Fluorescence of DA2m in response to one second of 660 nm LED light was measured in dissected brains with two-photon imaging of volume containing MB lobes (see Materials and methods). (B) Mean DA2m dF/F in ROIs defined for each MB compartment. SEMs are shown as shading, although they are often within width of lines representing means. N=8–12. See Figure 6—figure supplement 1 for quantification and the data with direct simulation of DANs.
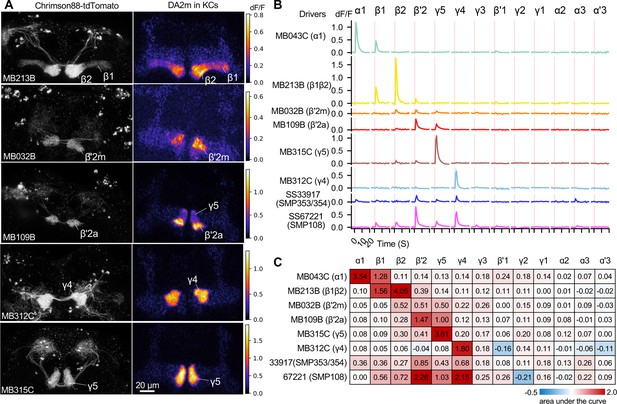
Patterns of dopamine release by different driver lines.
Representative images of neurons expressing Chrimson88-tdTtomato by designated driver lines (left) and maximum intensity projection of DA2m dF/F in the MB lobes (right). (A) Representative images of Chrimson88-tdTtomato expression patterns (left) and max intensity projections of DA2m dF/F in the MB lobes (right) as in Figure 6A. (B) Mean DA2m dF/F in ROIs defined for each MB compartment. N=8–12. (C) Area under the curve during the 10 s period after activation.
-
Figure 6—figure supplement 1—source data 1
Numerical Data for Figure 6.
- https://cdn.elifesciences.org/articles/79042/elife-79042-fig6-figsupp1-data1-v2.xlsx
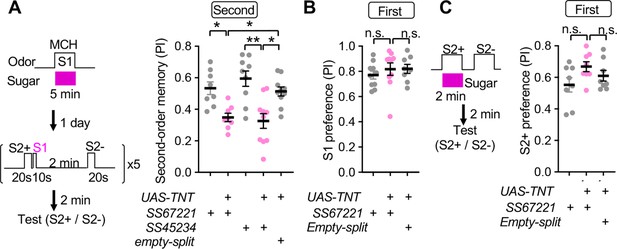
SMP108 is required for second-order memory.
(A) Second-order memory immediately after five training sessions as in Figure 1D following 5-min first-order conditioning a day before. Blocking SMP108 by expressing TNT with SS67221 or SS45234 impaired the second-order memory compared to genetic controls. N=10–12. (B) Preference to the S1 (MCH) odor over the air one day after pairing with sugar for 5 min. N=8–10. (C) First-order memory immediately after pairing S2+ odor with sugar for 2 min. N=8. *, p<0.05; **, p<0.01; Dunn’s multiple comparison tests following Kruskal-Wallis test.
-
Figure 7—source data 1
Numerical Data for Figure 7.
- https://cdn.elifesciences.org/articles/79042/elife-79042-fig7-data1-v2.xlsx
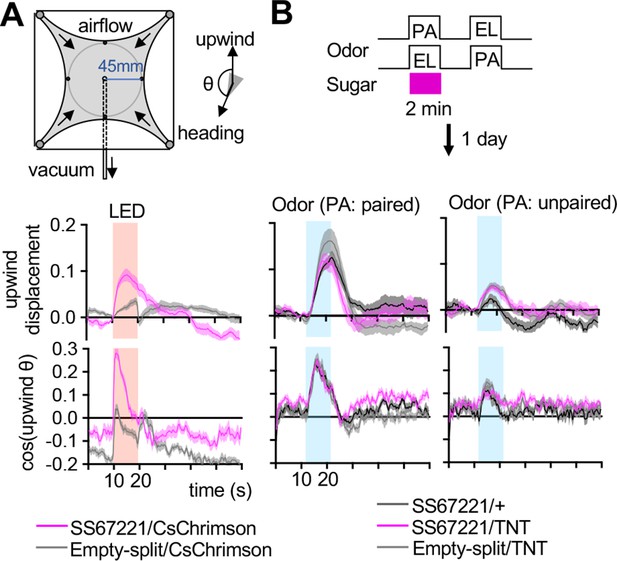
SMP108 can drive upwind steering but dispensable for the conditioned responses.
(A) Diagram of the circular arena (top). Airflow was constantly set at 400 mL/min throughout the experiments, and 10 s of 627 nm LED stimulations was applied for six times with 2 min intervals. Six trial averages of upwind displacement from the onset of LED (middle) and cosine of angle to upwind direction (bottom) are shown for flies expressing CsChrimson SS67221 (SMP108) or empty-split-GAL4. SS67221 >CsChrimson flies showed enhanced upwind displacement (P<0.05) and orientation toward upwind during LED ON period (p<0.01); See the method for the calculation of upwind displacement. N=12. (B) Groups of flies were trained by pairing either PA or EL with sugar for 2 min, and their response to airflow in the presence of odors were examined 20–24 hr later. Flies showed enhanced upwind displacement and orientation to upwind in the presence of reward-predicting odor. Upwind steering of flies with blocked SMP108 (SS67221/UAS-TNT) was indistinguishable with control genotypes. N=15–16.
-
Figure 7—figure supplement 1—source data 1
Numerical Data for Figure 7—figure supplement 1.
- https://cdn.elifesciences.org/articles/79042/elife-79042-fig7-figsupp1-data1-v2.xlsx
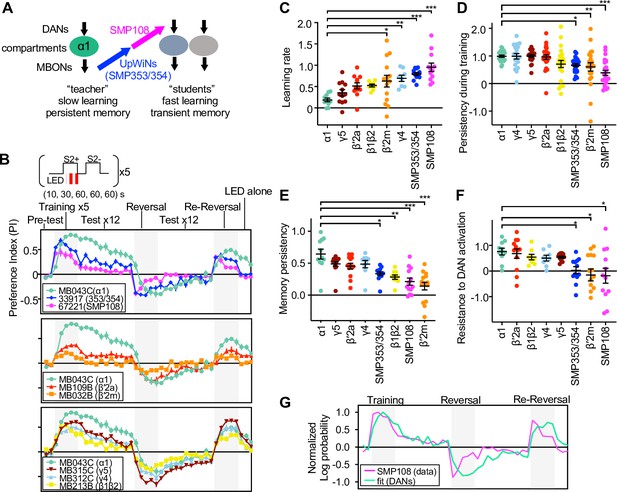
SMP108 pathway induces transient memory.
(A) Teacher-student compartments model of second-order conditioning hypothesizes that ‘teacher’ compartment with slow learning rate and persistent memory instructs other compartments with faster learning rate and transient memory dynamics via SMP353/SMP354 and SMP108. (B) Dynamics of memory with optogenetic activation of SMP108 (SS67221), SMP353/354 (SS33917) or various types of DANs. See texts and Materials and methods for explanation of the protocol, and Figure 8—figure supplement 1 for specificity of expression pattern in the central brain and the ventral nerve cord. Means and SEM are displayed. N=8–14. (C) Learning rate defined as a (PI after first 10 s training)/(peak PI during the first 5 training trials) for each driver line. (D) Persistency during training defined as (PI after 5th training)/(peak PI during the first 5 training trials). (E) Persistency of memory defined as (mean of PIs during 12 tests after first training trials)/(peak PI during the first 5 x training trials). (F) Resistance to DAN activation defined as (mean of last three tests following activation LED without odors)/(PI after 5th conditioning in re-reversal phase), which measures both transiency during training and extinction during 12 tests. p<0.05; **, p<0.01; ***, p<0.01; Dunn’s multiple comparison tests following Kruskal-Wallis test; N=8–14. (G) The log-probability ratio of choosing the S2+ against S2− for SS67221 (SMP108) data were fitted best with weights of (0.57, 0.46,0.157,0,0,0) for data of DAN driver lines (MB032B, MB213B, MB312C, MB043C, MB109B, and MB315C).
-
Figure 8—source data 1
Numerical Data for Figure 8.
- https://cdn.elifesciences.org/articles/79042/elife-79042-fig8-data1-v2.xlsx
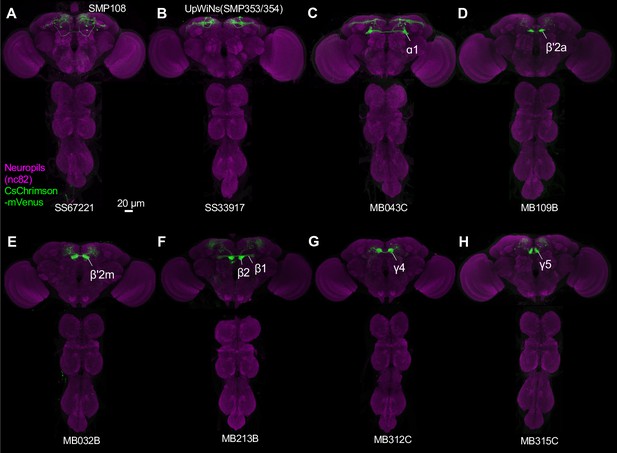
Expression patterns of drivers.
(A–H) Projection of confocal microscopy stacks for expression patterns of CsChrimson-mVenus driven by designated split-GAL4 driver lines in brains and ventral nerve cords. Confocal stacks are available at https://splitgal4.janelia.org.
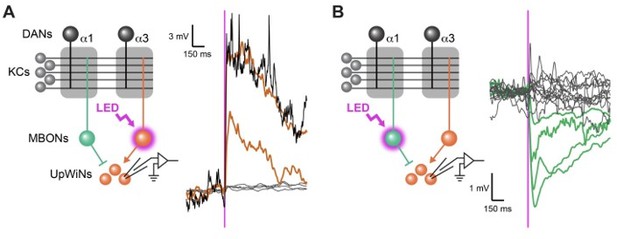
Functional connectivity between MBON-α1 and UpWiNs.
(A) Functional connectivity between MBON-α3 and UpWINs. Chrimson88-tdTomato was expressed in MBON-α3 by MB082C split-GAL4, and the photostimulation responses were measured by whole-cell current-clamp recording in randomly selected SMP353/354 labeled by R64A11-LexA. 2 out of 6 neurons (4 flies) showed excitatory response. Mean voltage traces from individual connected (orange) and unconnected neurons (gray) are overlaid. Connection was strong enough to elicit spikes (black; single-trial response in one of the connected neurons). Magenta vertical line indicates photostimulation (10 msec). (B) Functional connectivity between MBON-α1 and SMP353/354. Chrimson88-tdTomato expression in MBON-α1 was driven by MB310C split-GAL4. 4 out of 17 neurons (12 flies) showed inhibitory response. Mean voltage traces from individual connected (green) and unconnected neurons (gray) are overlaid. Figure 4A and B of Aso et al 2022 available at https://doi.org/10.1101/2022.12.21.521497.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| strain, strain background (Drosophila melanogaster) | Canton S | Martin Heisenberg | N.A. | |
| strain, strain background (Drosophila melanogaster) | 20xUAS-CsChrimson- mVenus attP18 | Klapoetke et al., 2014; PMID: 24509633 | N.A. | |
| strain, strain background (Drosophila melanogaster) | 10XUAS-Chrimson88- tdTomato attP1 | Klapoetke et al., 2014; PMID: 24509633 | N.A. | |
| strain, strain background (Drosophila melanogaster) | 13XLexAop2-IVS- ChrimsonR-mVenus- p10 attP18 | Vivek Jayaraman | N.A. | |
| strain, strain background (Drosophila melanogaster) | 20XUAS-syn21-mScarlet- opt-p10 su(Hw)attp8 | Glenn Turner | N.A. | |
| strain, strain background (Drosophila melanogaster) | pJFRC200-10xUAS- IVS-myr::smGFP-HA in attP18 | Nern et al., 2015; PMID: 25964354 | N.A. | |
| strain, strain background (Drosophila melanogaster) | pJFRC225-5xUAS- IVS-myr::smGFP-FLAG in VK00005 | Nern et al., 2015; PMID: 25964354 | N.A. | |
| strain, strain background (Drosophila melanogaster) | pBPhsFlp2::PEST in attP3 | Nern et al., 2015; PMID: 25964354 | N.A. | |
| strain, strain background (Drosophila melanogaster) | pJFRC201-10XUAS-FRT>STOP > FRT-myr::smGFP-HA in VK0005 | Nern et al., 2015; PMID: 25964354 | N.A. | |
| strain, strain background (Drosophila melanogaster) | pJFRC240-10XUAS-FRT>STOP > FRT-myr::smGFP-V5-THS-10XUAS-FRT>STOP > FRT-myr::smGFP-FLAG_in_su(Hw)attP1 | Nern et al., 2015; PMID: 25964354 | N.A. | |
| strain, strain background (Drosophila melanogaster) | LexAop2-DA2m VK00005 | Sun et al., 2020; PMID: 33087905 | N.A. | |
| strain, strain background (Drosophila melanogaster) | MB043-split-LexA | This paper | N.A. | Available from Aso lab |
| strain, strain background (Drosophila melanogaster) | empty-split-GAL4 (p65ADZp attP40, ZpGAL4DBD attP2) | Seeds et al., 2014; PMID: 25139955 | N.A. | |
| strain, strain background (Drosophila melanogaster) | MB032B split-GAL4 | Aso et al., 2014; PMID: 25535793 | N.A. | |
| strain, strain background (Drosophila melanogaster) | MB043C split-GAL4 | Aso et al., 2014; PMID: 25535793 | N.A. | |
| strain, strain background (Drosophila melanogaster) | MB109B split-GAL4 | Aso et al., 2014; PMID: 25535793 | N.A. | |
| strain, strain background (Drosophila melanogaster) | MB213B split-GAL4 | Aso et al., 2014; PMID: 25535793 | N.A. | |
| strain, strain background (Drosophila melanogaster) | MB315C split-GAL4 | Aso et al., 2014; PMID: 25535793 | N.A. | |
| strain, strain background (Drosophila melanogaster) | SS33917 split-GAL4 | This paper | N.A. | Available from Aso lab |
| strain, strain background (Drosophila melanogaster) | SS45234 split-GAL4 | This paper | N.A. | Available from Aso lab |
| strain, strain background (Drosophila melanogaster) | SS67221 split-GAL4 | This paper | N.A. | Available from Aso lab |
| strain, strain background (Drosophila melanogaster) | UAS-TeNT | Keller et al., 2002: PMID: 11810637 | N.A. | |
| antibody | anti-GFP (rabbit polyclonal) | Invitrogen | A11122 RRID:AB_221569 | 1:1000 |
| antibody | anti-Brp (mouse monoclonal) | Developmental Studies Hybridoma Bank | nc82 RRID:AB_2341866 | 1:30 |
| antibody | anti-ChAT (mouse monoclonal) | Developmental Studies Hybridoma Bank | ChAT4B1 RRID:AB_528122 | 1:50 |
| antibody | anti-HA-Tag (mouse monoclonal) | Cell Signaling Technology | C29F4; #3724 RRID:AB_10693385 | 1:300 |
| antibody | anti-FLAG (rat monoclonal) | Novus Biologicals | NBP1-06712 RRID:AB_1625981 | 1:200 |
| antibody | anti-V5-TAG Dylight-549 (mouse monoclonal) | Bio-Rad | MCA2894D549GA RRID:AB_10845946 | 1:500 |
| antibody | anti-mous IgG(H&L) AlexaFluor- 568 (goat polyclonal) | Invitrogen | A11031 RRID:AB_144696 | 1:400 |
| antibody | anti-rabbit IgG(H&L) AlexaFluor-488 (goat polyclonal) | Invitrogen | A11034 RRID:AB_2576217 | 1:800 |
| antibody | anti-mouse IgG(H&L) AlexaFluor-488 conjugated (donkey polyclonal) | Jackson Immuno Research Labs | 715-545-151 RRID:AB_2341099 | 1:400 |
| antibody | anti-rabbit IgG(H&L) AlexaFluor-594 (donkey polyclonal) | Jackson Immuno Research Labs | 711-585-152 RRID:AB_2340621 | 1:500 |
| antibody | anti-rat IgG(H&L) AlexaFluor-647 (donkey polyclonal) | Jackson Immuno Research Labs | 712-605-153 RRID:AB_2340694 | 1:300 |
| antibody | anti-Mouse IgG (H&L) ATTO 647 N (goat polyclonal) | ROCKLAND | 610-156-121 RRID:AB_10894200 | 1:100 |
| antibody | anti-rabbit IgG (H+L) Alexa Fluor 568 (goat polyclonal) | Invitrogen | A-11036 RRID:AB_10563566 | 1:1000 |
| chemical compound, drug | 3-Octanol | Sigma-Aldrich | 218405 | |
| chemical compound, drug | 4-Methylcyclohexanol | VWR | AAA16734-AD | |
| chemical compound, drug | Pentyl acetate | Sigma-Aldrich | 109584 | |
| chemical compound, drug | Ethyl lactate | Sigma-Aldrich | W244015 | |
| chemical compound, drug | Paraffin oil | Sigma-Aldrich | 18512 | |
| software, algorithm | ImageJ and Fiji | NIH Schindelin et al., 2012 | https://imagej.nih.gov/ij/ http://fiji.sc/ | |
| software, algorithm | MATLAB | MathWorks | https://www.mathworks.com/ | |
| software, algorithm | Adobe Illustrator CC | Adobe Systems | https://www.adobe.com/products/illustrator.html | |
| software, algorithm | GraphPad Prism 9 | GraphPad Software | https://www.graphpad.com/scientific-software/prism/ | |
| software, algorithm | Python | Python Software Foundation | https://www.python.org/ | |
| software, algorithm | neuPrint | HHMI Janelia | https://doi.org/10.25378/janelia.12818645.v1 | |
| software, algorithm | Cytoscape | Shannon et al., 2003 | https://cytoscape.org/ | |
| software, algorithm | NeuTu | Zhao et al., 2018 | https://github.com/janelia-flyem/NeuTu; janelia-flyem, 2021 | |
| software, algorithm | ScanImage | Vidrio Technologies | https://vidriotechnologies.com/ | |
| software, algorithm | VVDveiwer | HHMI Janelia | https://github.com/takashi310/VVD_Viewer | |
| other | Grade 3 MM Chr Blotting Paper | Whatmann | 3030–335 | Used in glass vials with paraffin-oil diluted odours |
| other | mass flow controller | Alicat | MCW-200SCCM-D | Mass flow controller used for the olfactory arena |
Additional files
-
Supplementary file 1
Neurotransmitter prediction and a full connection matrix for MBONs, DANs and 396 interneurons cell types.
Numbers in column B-G are numbers of presynaptic sites that are predicted to be designated neurotransmitters. EM id in column K is an identification number in EM hembrain data. The other columns are the connection matrix. Top row indicates the direction of connections. For instance, 153 in the raw 5 of column M indicate the number of connections from MBON01 to SMP108, while 166 in the raw5 of column BD indicate the number of connections from SMP108 to PAM02. For the cell type consisting of multiple cells, a summed number of connections are shown.
- https://cdn.elifesciences.org/articles/79042/elife-79042-supp1-v2.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/79042/elife-79042-mdarchecklist1-v2.docx





