Substrate stiffness impacts early biofilm formation by modulating Pseudomonas aeruginosa twitching motility
Figures
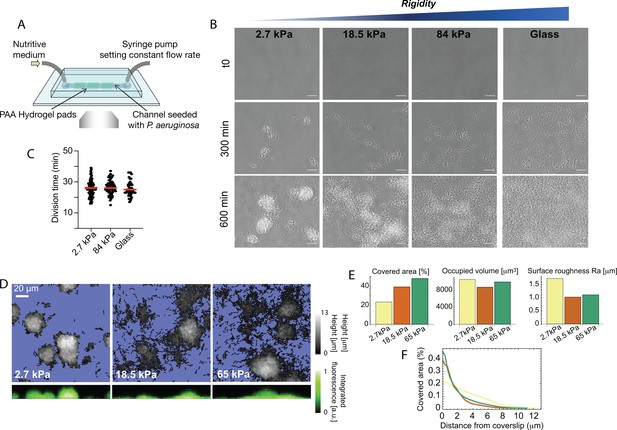
Bacterial microcolony formation depends on substrate rigidity.
(A) Experimental setup: bacteria (P. aeruginosa strain PAO1) are imaged in a flow cell under constant flow of minimal medium. (B) After 10 hr, dense, isolated colonies form on soft PAA (2.7 kPa) while bacteria are more evenly distributed on stiff PAA (84 kPa), closer to what is observed on glass. Scale bars, 20 μm. (C) Bacterial growth is not impacted by substrate rigidity. (D) 3D reconstruction of colonies confirms their hemispherical shape on soft substrates. (E) Surface coverage is lower on soft substrates, but total volume of colonies is conserved, with a higher roughness value. (F) Fraction of area occupied by the bacteria as a function of the distance from the coverslip, showing flatter colonies on rigid substrates.
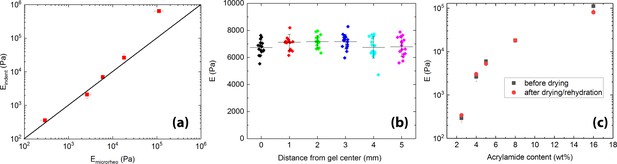
Mechanical characterization of hydrogels by AFM (a) comparison of elastic moduli measured by indentation and by microrheology.
Both techniques yield quantitatively similar results for gels with Young’s moduli kPa. (b) Spatial homogeneity of the gels characterized by indentation measurements. For each position, separated by 1 mm, a 4x4 force spectroscopy map is taken, with a spacing between “pixels” of 3 μm. (c) comparison of elastic moduli measured before and after drying/rehydration of the gels.
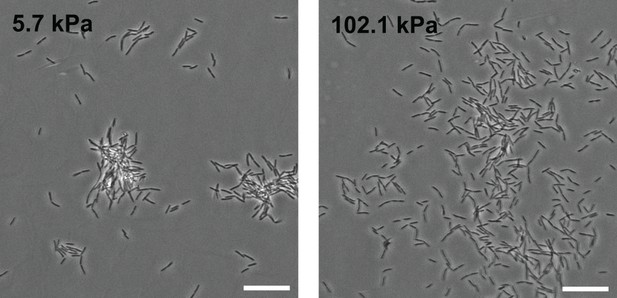
Morphology of microcolonies is strongly impacted by surface rigidity on PEG hydrogels.
Phase contrast images of WT PAO1 bacteria on PEG hydrogels 5h after the onset of surface colonization. Scale bars, 20 μm.
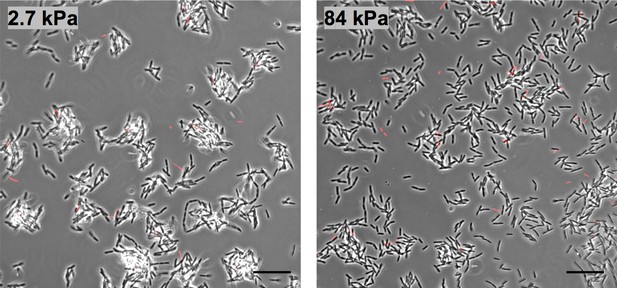
Substrate rigidity impacts early microcolony morphology in the absence of shear flow.
Phase contrast images of WT PAO1 bacteria on PAA hydrogels at the bottom of a 6-well plate, after 150 min of incubation at 37°C without agitation. In red: initial adhering bacteria at t0. (Scale bars, 20 μm.).

In the T4P-deficient mutant PAO1 , substrate rigidity does not significantly impact colony morphology.
Colonies imaged after 5 h. Scale bar 20 μm.
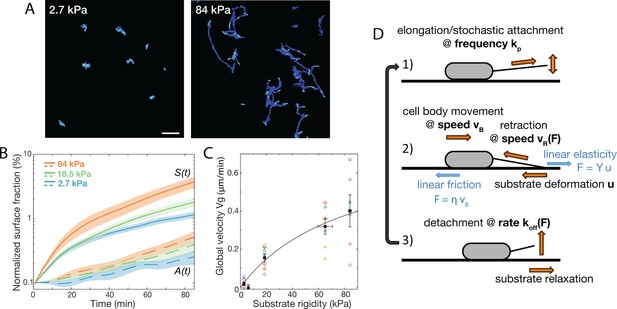
Bacterial surface motility is impaired on soft hydrogels.
(A) Surface explored (dark blue) and current surface coverage (cyan) after 100 min on soft and stiff PAA surfaces. Scale bar: 20 μm. (B) Surface coverage A(t) (broken lines) and cumulative explored area S(t) (full lines) for all tested rigidities (the initial surface coverage was normalized to 0.1%). Shaded areas are standard errors of the mean (84 kPa: 6 data sets from independent experiments, 18.5 kPa: 10 data sets from five independent experiments, 2.7 kPa: 8 data sets from independent experiments).(C) Global bacterial motility averaged over the first 100 min, inferred from the difference between A(t) and S(t) (16 different surfaces, 6 independent experiments). The black line is the fit with the kinetic model using equation A11. with values m.min-1 and kPa. (D) Ingredients of the minimal 1D model for bacterial T4P-powered displacement.
-
Figure 2—source data 1
Surface coverage and explored area vs time.
- https://cdn.elifesciences.org/articles/81112/elife-81112-fig2-data1-v2.xlsx

Behavior of mutants and on soft (2.7 kPa) and stiff (84 kPa) PAA hydrogels.
Left: typical images showing initially attached bacteria (red) and total explored area after 100 minutes (white). Right: Average bacterial velocity Vg calculated using equation 2 of the main text, over the first 100 min of acquisition (1 experiment for each strain, 3 different positions for each gel. Error bars are SEM).
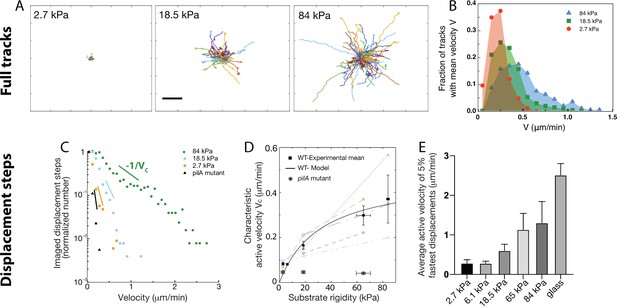
Twitching motility depends on substrate rigidity and is highly distributed in the bacterial population.
Analysis of full tracks (top row) and 1 min displacement steps (bottom row). (A) Individual bacterial tracks on soft (2.7 kPa), intermediate (18.5 kPa) and stiff (84 kPa) PAA during the first 3 hours after bacterial inoculation (total number of tracks is respectively 60, 123, and 175). Scale bar: 10 μm (B) Mean track velocity distribution for different values of the substrate rigidity. Only full tracks were considered (corresponding to the right peak in Figure 3—figure supplement 1). 84 kPa: 330 tracks from 2 independent experiments, 18.5 kPa: 394 tracks from 3 independent experiments, 2.7 kPa: 83 tracks from 2 independent experiments. (C) Normalized velocity distributions for the whole bacterial population on different PAA surfaces. The exponential decrease yields a characteristic active velocity on each substrate. Displacement steps were measured every minute for 100 min, and two positions were acquired on each rigidity. The average of T4P-defective mutant on all surfaces is shown as a reference. (D) Characteristic active velocity values obtained by fitting velocity distributions (6 independent experiments, 16 different surfaces). Values measured on different surfaces in a single experiment (same channel) are shown with the same symbols and connected. Black squares(circled stars) are mean values obtained for the WT(pilA mutant), and error bars show the SEM. The black line is the fit of WT values with the kinetic model-derived equation A11, with μm.min-1 and kPa. (E) Average velocity values of the top 5% fastest displacement steps for different substrates. Error bars are standard errors.
-
Figure 3—source data 1
Instant velocity values from bacterial tracking.
- https://cdn.elifesciences.org/articles/81112/elife-81112-fig3-data1-v2.xlsx

Tracking result reveals bacterial sub-populations on hydrogel substrates.
(A) Distribution of the path duration of bacterial tracks measured on different PAA substrates (WT PAO1). One track starts after a division event, and finishes either at the next division, or when the bacterium leaves the surface. Full lines are bimodal gaussian fits, evidencing a sub-population of bacteria that detach from the surface before dividing. (B) Mean track velocity is similar for short tracks (below the cutoff value tc = 16 min) and long tracks, for all tested substrate rigidities. (total number of analyzed tracks: 60 (2.7 kPa), 124 (18.5 kPa), 175 (84 kPa)).
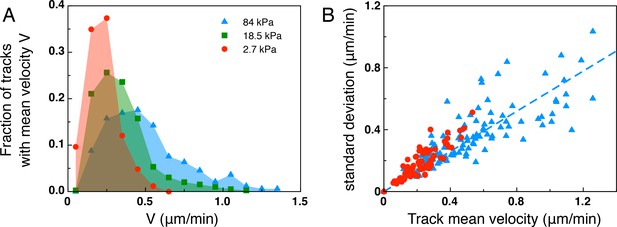
Anaysis of the mean track velocity.
(A) Mean track velocity distribution for different values of the substrate rigidity. Only full tracks were considered (corresponding to the right peak in Figure 3—figure supplement 1). Considering all tracks does not significantly modify the distributions (data not shown). 84 kPa: 330 tracks from 2 independent experiments, 18.5 kPa: 394 tracks from 3 independent experiments, 2.7 kPa: 83 tracks from 2 independent experiments. (B) Standard deviation of the mean velocity for individual tracks, shown for the 2.7 kPa and 84 kPa data sets. Dotted line is a linear fit for the 84 kPa data (y=0.65x).
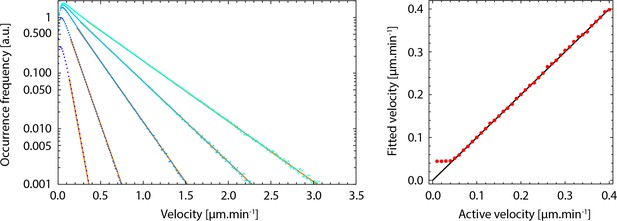
Validation of the fitting of displacement steps distributions with single exponentials.
Left, calculated distributions (dots) of the sum of two uncorrelated random vectors each following a decreasing exponential distribution with characteristic velocities and , at an angle following a random distribution. is equal to the experimentally measured value on the mutant. spans a range similar to experimental measurements, (0.05, 0.1, 0.2, 0.3, and 0.4 μm/min, from dark blue to cyan). Orange lines are exponential functions with the corresponding. Right, characteristic velocities extracted by fitting the exponential part of the simulated distributions, as a function V_C, with. The black line corresponds to y=x.
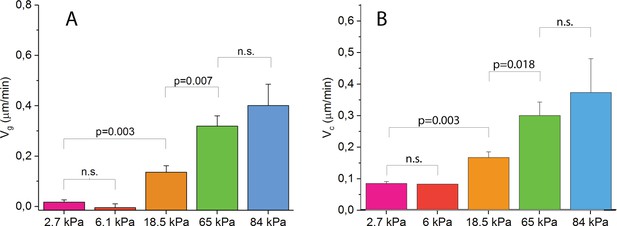
Statistical analysis of (A) and (B) for various gel rigidities.
Indicated are p-values from paired two-sided Mann-Whitney non-parametric tests. n.s. denotes datasets that are not significantly different at a threshold of 0.05.

Characteristic twitching velocity measured on PEG hydrogels is similar to the one measured on PAA hydrogels under identical experimental conditions.
obtained by fitting the distribution of displacement steps on PAA or PEG hydrogels in a similar range of substrate rigidity. Error bars are SEM. Each point is the average of 2 positions from 1 experiment. (Note that for technical reasons these experiments were carried out with a wall shear stress of 6.4 mPa, about 3x higher than all other experiments on PAA).
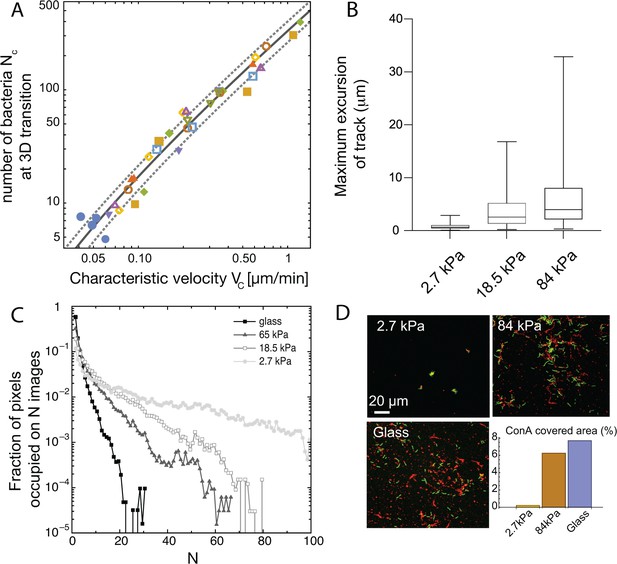
Spatial structuring of surface colonization is impacted by substrate rigidity through twitching velocity.
(A) Size of microcolony (in number of bacteria ) at the 2D to 3D transition as a function of the center-of-mass characteristic velocity defined in Figure 3A. Markers are experimental data from 9 different experiments, each with different substrates including glass (hence leading to higher values that in Figure 3B). Blue dots are data obtained with the pili-deficient mutant pilA:: Tn5. Lines, kinetic model for (solid line) ± its standard deviation (dotted lines). (B) Distribution of track lengths of full trajectories as a function of the substrate rigidity. (C) Distribution of occupation occurrence on each image pixel as a function of rigidity, showing a much more heterogeneous occupancy on soft substrates. (D) ConA staining (red) of EPS deposition during cell (green) exploration of the surface.
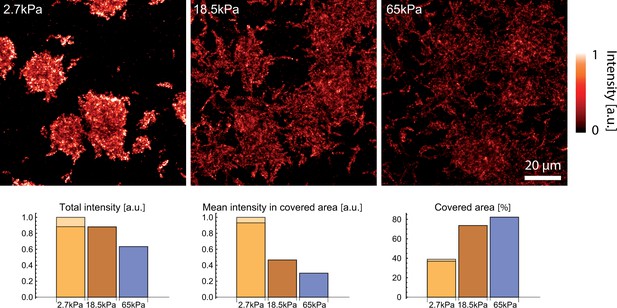
EPS staining with concanavalin A highlights matrix deposits on the surface of substrates of different stiffnesses after 8h of surface colonization.
Matrix deposits are more compact on soft substrates with large areas (>60%) devoid of EPS, while stiffer substrates are almost fully decorated (>% covered area). The two colours for the softer surface corresponds to two areas taken 30 min apart after staining, before and after acquisition of the two other datasets on larger rigidities, to rule out any significant time evolution of the staining.
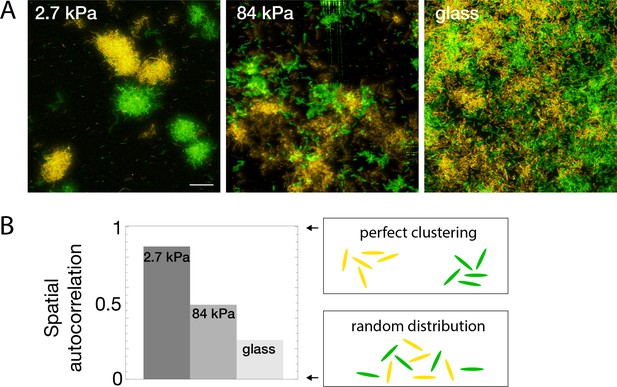
Bacterial spatial distribution is impacted by substrate rigidity.
(A) Images of surfaces seeded with a 1:1 mixture of constitutively fluorescent bacteria expressing GFP or YFP show mostly monoclonal colonies on soft hydrogels, and mixed bacteria on rigid substrates (3D-rendering obtained by stacking images). Scale bar, 20 μm.(B) Spatial correlations quantified via Moran’s I index.
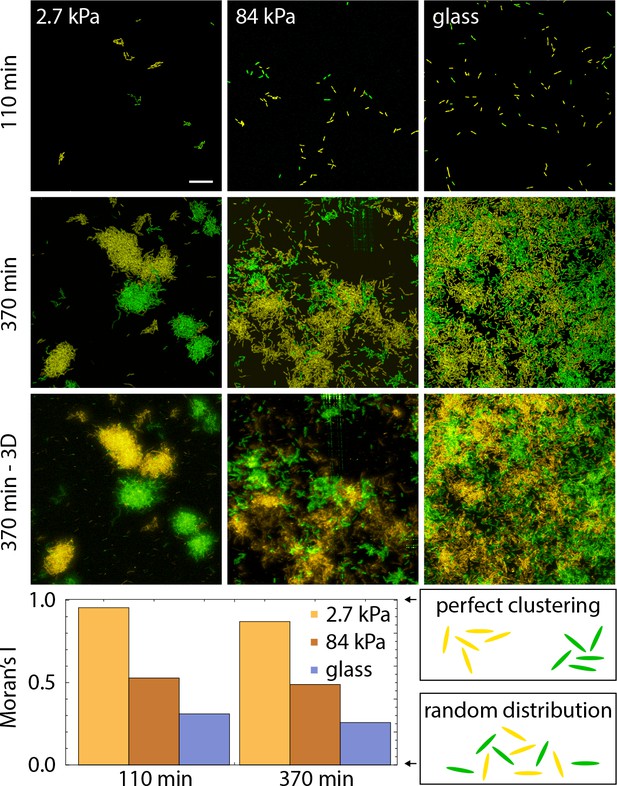
Bacterial spatial distribution as a function of substrate rigidity.
Top, images of surfaces seeded with a 1:1 mixture of constitutively fluorescent bacteria expressing GFP or YFP at two different times after the start of surface colonization (bottom : 3D rendering of a volumetric image). Bottom, Spatial autocorrelation quantified via Moran’s I index at the two time points il lustrated above. The values should not be directly compared between the two time points (see SI I-C) but illustrate that the rigidity modulation of mixing is maintained over time from very sparse to large coverage of the surface.
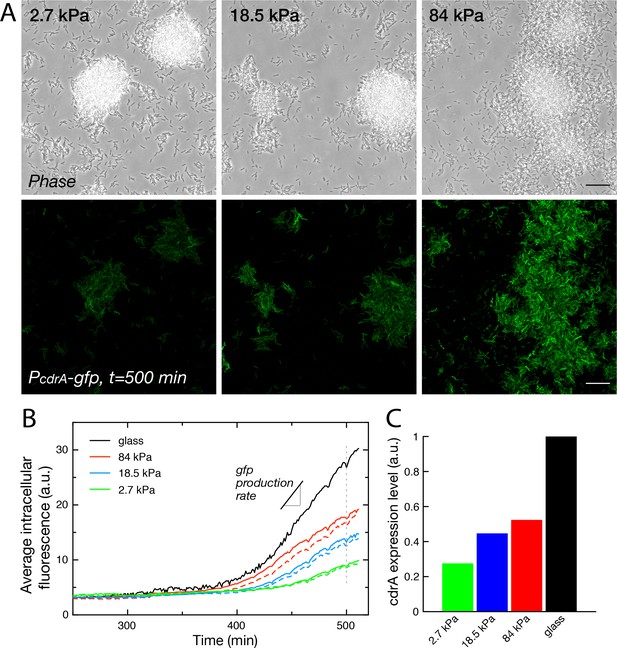
Substrate rigidity influences gene expression levels (A) Colonies grown from a modified PAO1 strain bearing the plasmidic reporter imaged in phase contrast (top) and fluorescence (bottom), after 500 min under flow.
Scale bars, 20 μm. (B) Average intracellular gfp fluorescence as a function of time, on different PAA surfaces and on glass. Broken lines are duplicate positions on a given surface. (C) Average cdrA expression level, obtained from a linear fit of (B).
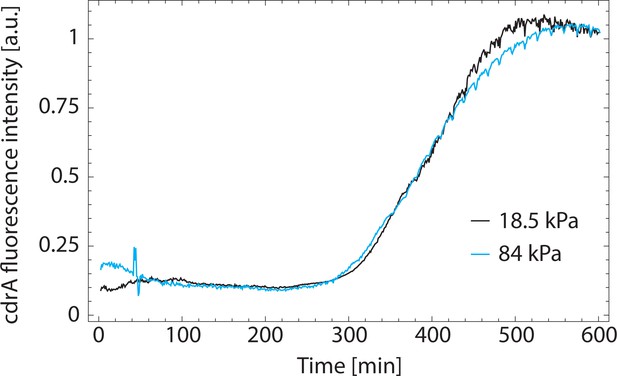
Fluorescence intensity from a PAO1 strain expressing a plasmid over time, showing a plateau of fluorescence expression after ≈500 min.
In our experiments, this change in behavior might be due to a lack of oxygen in the flow upon growth of the biofilm in the microfluidic channel and occurs. Two rigidities are shown and scaled to the same final fluorescence value.
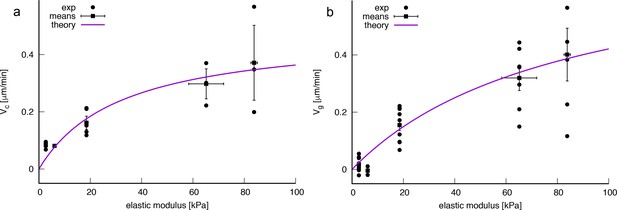
Experimentally measured velocity vs rigidity data and least squared fits of Equation A11 w.r.t. to the experimental values as indicated in the legends.
(a) Local velocity measures. Parameters obtained by a least-square fit: kPa, μm.min-1. (b) Global velocity measures. Parameters obtained by a least square fit: kPa, μm.min-1. Errorbars indicate SEM.
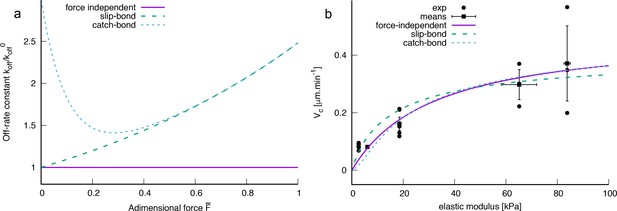
Comparison of various force dependencies of the pilus detachment rate constant koff as indicated in the legend.
(b) Comparison of bacterial velocities obtained by models with various complexity with experimentally measured values (local velocity analysis) as indicated in the legend (parameters were fit w.r.t experimental mean values). The model parameters for the force-independent model are as in Appendix 1—figure 1a. For the slip and catch bond model the fixed parameters are kPa,, (catch bond), (catch bond). For the slip and catch bond model was estimated from least square fits: μm.min-1 (slip) and μm.min-1 (catch). Error estimates are expected to be of the same order of magnitude as for the force independent model (see Appendix 1—figure 1a).
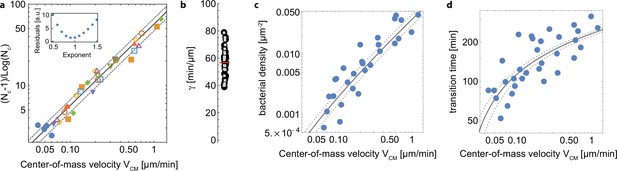
Experimental results and kinetic model for the transtion to 3D microcolonies.
(a) Comparison between the experimental data (markers) and the kinetic model (lines). Blue dots are data obtained with the pili-deficient mutant pilA: Tn5. Inset, residual of the fit of all the experimental data points as a function of the exponent value for , indicating that the best fit is achieved for a value of or close to 1. (b) values for all data points (black disks) and their average (red line). Gray squares are data points with μm.min-1, showing a similar distribution and thus ruling out a significant bias at high velocities. (c) Same dataset as in a but the surface density at the transition is plotted; (d) same dataset as in a but the time at the transition is plotted. The same model is used to describe the data, but converted into the proper quantities.
Videos
surface colonization on 2.7 kPa (left) and 65 kPa (right) PAA hydrogels, imaged with phase contrast microscopy with one image/min over 6 hours.
The two gels were included in the same microfluidic channel and imaged quasi-simultaneously. Scale bar, 20 m.
Principle of the image processing for the tracking of individual bacteria.
Glass surface colonization under controlled shear flow followed over 90 min (1 frame/minute) by phase contrast microscopy. Left, the registered phase contrast image is superimposed with the center of mass of the detected cells after segmentation (green dots). Right, tracks of the detected cells (obtained with the simple LAP tracker of the Trackmate ImageJ plugin), color-coded as a function of the track final length. Scale bar, 20 m.
Tables
Hydrogel compositions and associated Young’s moduli.
| Acrylamide (wt%) | Bis-acryl. (wt%) | PEGDA (wt%) | Modulus (kPa) |
|---|---|---|---|
| 4 | 0.225 | 0.0 | 2.7 ± 0.3 |
| 5 | 0.225 | 0.0 | 6.1 ± 0.2 |
| 8 | 0.264 | 0.0 | 18.5 ± 0.7 |
| 20 | 0.47 | 0.0 | 65 ± 5.6 |
| 15 | 0.65 | 0.0 | 84 ± 1.1 |
| 20 | 0.7 | 0.0 | 103 ± 3.8 |
| 0 | 0 | 5 | 5.7 ± 0.3 |
| 0 | 0 | 20 | 102 ± 8.4 |