Quantifying the impact of immune history and variant on SARS-CoV-2 viral kinetics and infection rebound: A retrospective cohort study
Figures
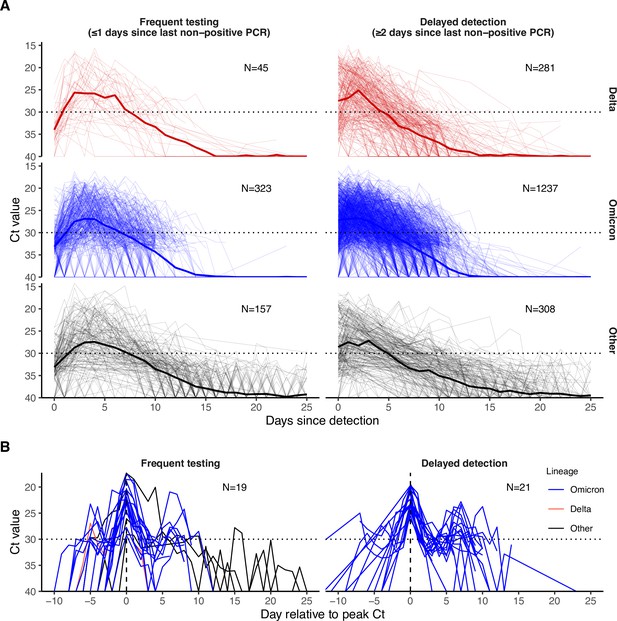
PCR Ct value trajectories for confirmed and suspected infections.
(A) PCR Ct value trajectories for each acute Delta (red), Omicron BA.1 (blue), and other (black) infection. Individuals are grouped by the gap between detection and their most recent negative or inconclusive PCR test (Frequent testing vs. Delayed detection). Thick lines depict the mean Ct value over time, counting negative tests as Ct = 40. Thin lines depict individual level Ct values over time. The horizontal dotted lines mark Ct = 30, which we consider here as a proxy for possible infectiousness and antigen test positivity. (B) Subsets of PCR Ct value trajectories that were classified as rebounds, stratified by testing frequency group. Rebounds are defined here as any trajectory with an initial Ct value <30, followed by a sequence of two or more consecutive negative tests or tests with Ct value ≥30, and subsequently followed by two or more consecutive tests with Ct value <30.
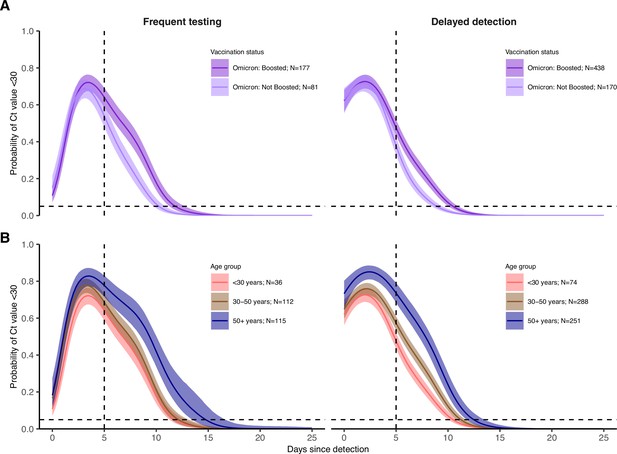
Posterior estimates from a generalized linear model predicting probability of Ct value <30 with spline terms for the interaction between days since detection with age group and the interaction between days since detection with vaccination status and variant.
Shown are the marginal effects of (A) vaccination status and (B) age group on the proportion of individuals with Ct <30 on each day post detection after conditioning on being an Omicron BA.1 infection and (A) <30 years old (B) boosted at the time of infection. Solid colored lines and shaded ribbons show the posterior mean (solid line) and 95% credible intervals (shaded ribbon) of each conditional effect. Dotted horizontal and vertical lines mark 5% probability and day 5 post detection, respectively.
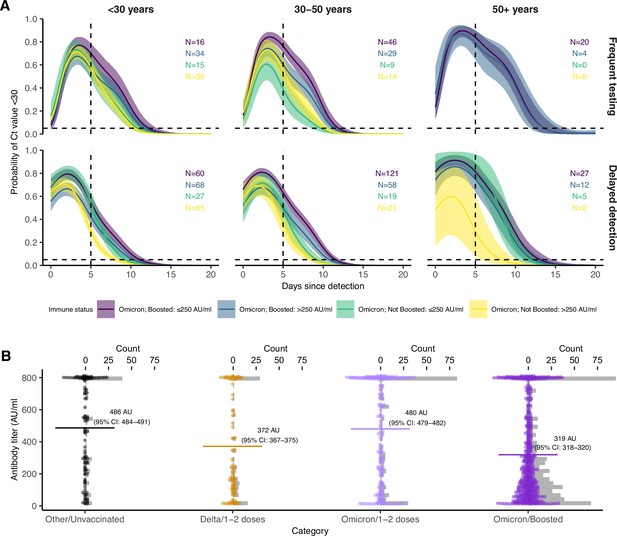
Effect of booster status, antibody titer against the founder SARS-CoV-2 strain and age group on the probability of Ct value <30 over days post detection.
(A) Proportion of Omicron BA.1 infections with Ct value <30 on each day post detection, stratified by age group and the interaction of booster status at the time of infection and antibody titer group. Shown are posterior estimates from a generalized linear model predicting probability of Ct <30 with a spline term on days since detection, conditional on titer/vaccination status category and age group. Solid lines show posterior mean and shaded ribbons show 95% credible intervals of each conditional effect. (B) Distribution of measured antibody titers (colored points) stratified by variant and vaccination status of each detected infection, with mean titers (horizontal lines) and bootstrapped 95% confidence intervals (CIs) shown in text. Note that the 95% CIs are very small relative to the range and are thus not plotted. Grey bars are histograms of antibody titer counts in bins of 10 arbitrary units (AU)/ml. Note also that stratification is by infection event and not individual, and that antibody titers were measured at a single point in time rather than near the time of infection. The Diasorin Trimeric Assay values are truncated between 13 and 800 AU/ml.
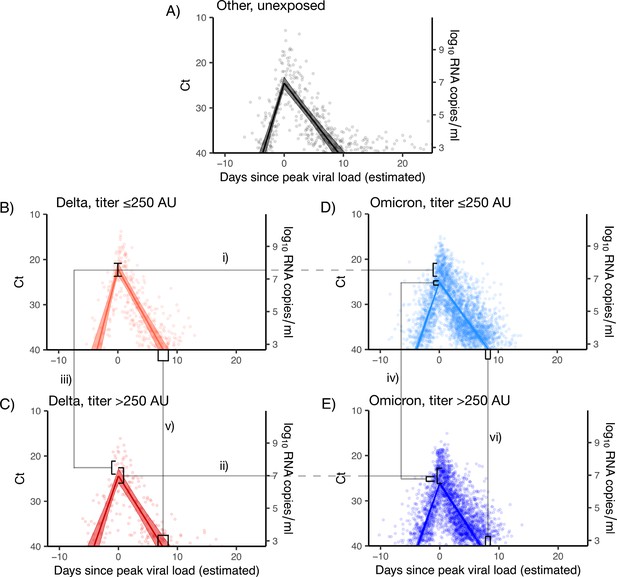
Estimated viral trajectories by variant and titer.
Points depict measured Ct values, lines depict the estimated population mean viral trajectories, and shaded regions depict the 95% credible intervals for the estimated population viral trajectories. (A) Non-Delta and non-Omicron infections in individuals who were previously unexposed (no prior record of vaccination or infection), (B) Delta infections with titer ≤250, (C) Delta infections with titer >250, (D) Omicron BA.1 infections with titer ≤250, (E) Omicron BA.1 infections with titer >250. Peak viral loads were higher for Delta infections than for Omicron BA.1 infections when stratifying by titer (i and ii), and titers ≤250 were associated with higher viral loads when stratifying by variant (iii and iv). Low titers were also associated with longer clearance times (v and vi).
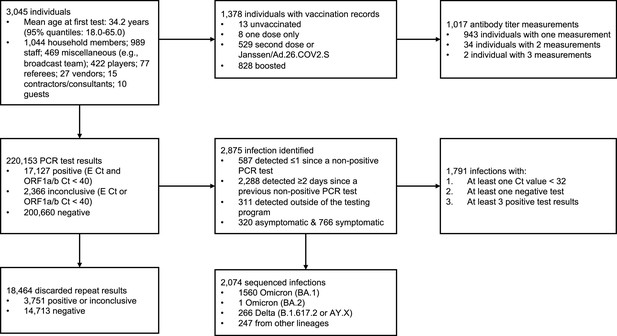
Summary of cohort.
Top row describes cohort demographics and data on immune histories. Middle row describes infection data. Bottom row provides additional information on the infection data.
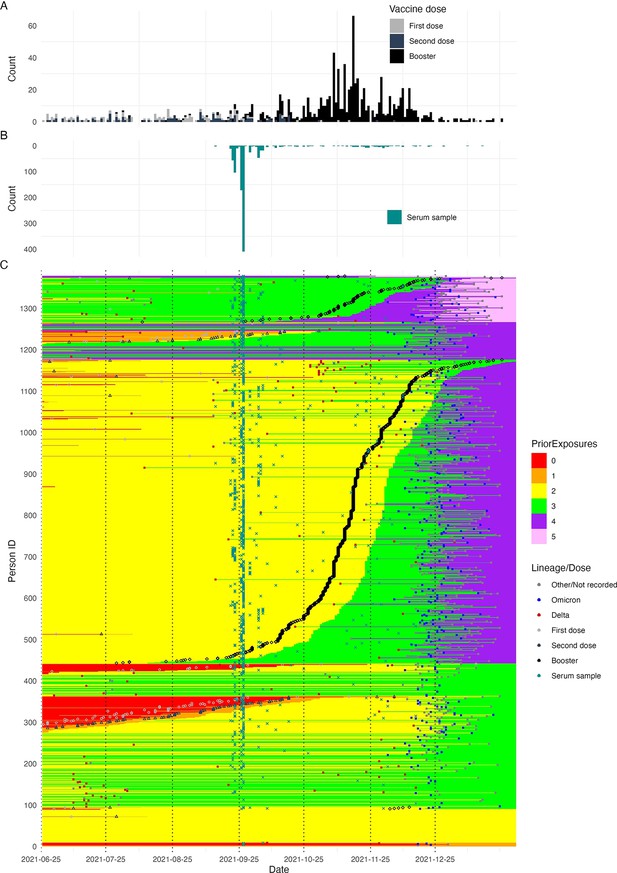
Summary of all infection and vaccination events over time, in addition to serum sample collection dates, included in the dataset.
(A) Histogram showing the distribution of vaccination dates (note that most first doses were administered prior to 2021-06-25), showing when in time the majority of individuals were vaccinated. (B) Histogram showing the timing of serum sample collections in the cohort. (C) Heatmap showing the entire time course of infection and vaccination histories for each individual in the cohort. Each row represents one individual and columns represent date. Each cell is shaded by the number of prior exposures at that date, showing how each individual’s cumulative exposure history increases over time. Points show the timing of each detected infection, recorded vaccination, and serum sample collection date. Points are colored by the variant or vaccination number of that exposure.
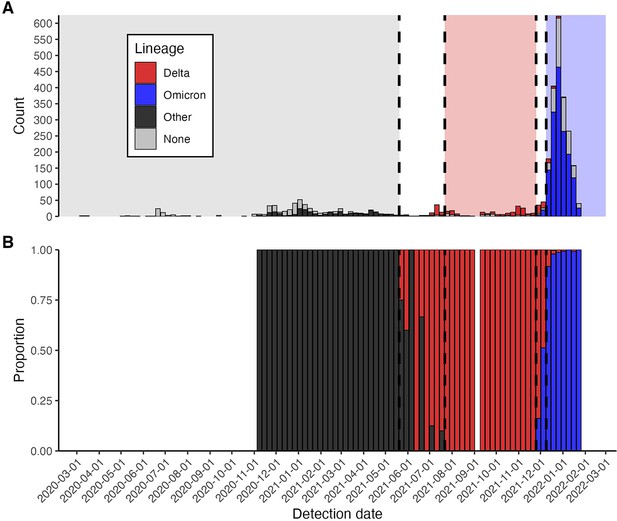
Frequency and proportion of SARS-CoV-2 variant detections.
(A) Frequency of sequenced and unsequenced detected infections over time by week. Vertical dashed lines and shaded backgrounds demarcate periods of variant dominance.
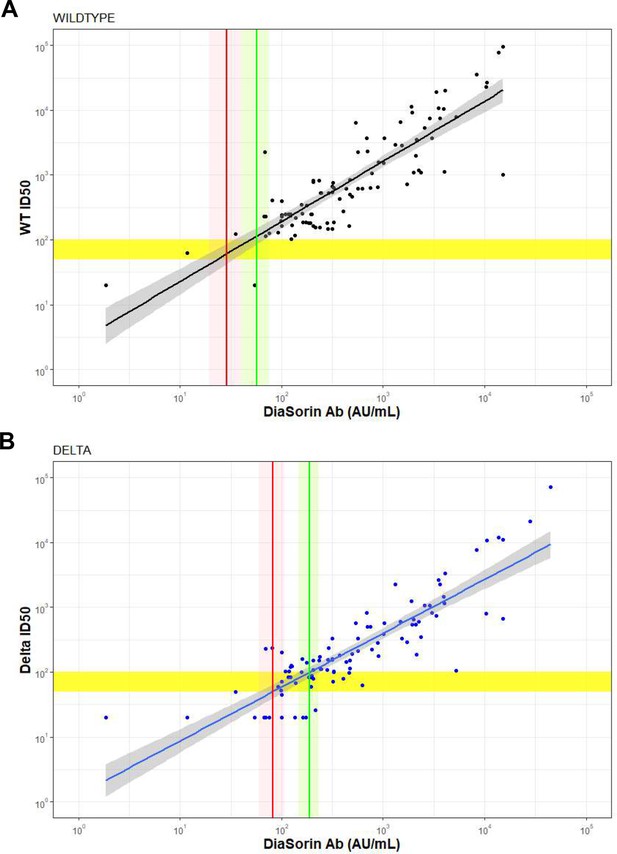
Correlation between authentic virus neutralization assay (ID50) and the Diasorin antibody titer against (A) wildtype and (B) Delta.
Horizontal yellow bar shows an ID50 titer of 50 and 100 respectively, Diagonal lines and shaded regions show mean and 95% confidence intervals (CI) for a linear regression between the Diason antibody titer and ID50 titer. Vertical line and shaded regions show point estimate and 95% CI for the Diasorin antibody titer corresponding to an ID50 titer of 50 (red) and 100 (green).
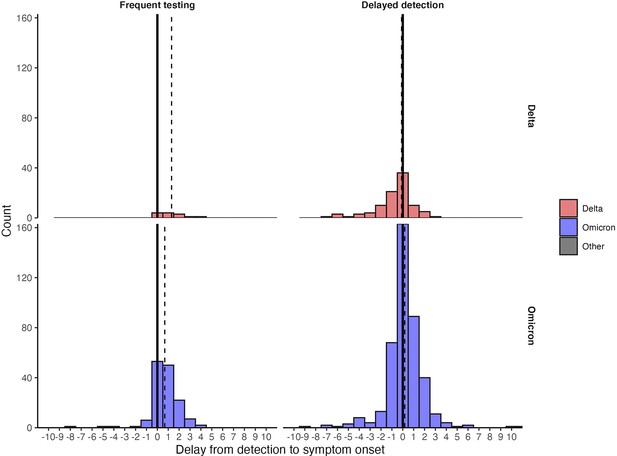
Distribution of delays from detection to symptom onset among individuals with known symptom status.
Dashed lines mark the median delay between detection and symptom onset. Solid lines mark the day of detection (0).
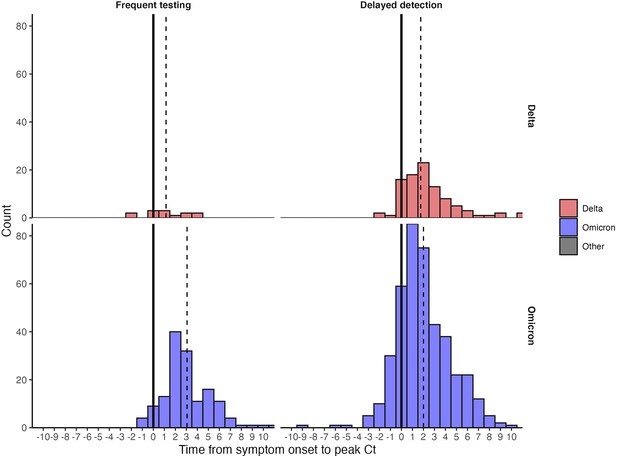
Distribution of delays from symptom onset to peak Ct values among individuals with known symptom status.
Dashed lines mark the median delay between detection and symptom onset. Solid lines mark the day of symptom onset (0).
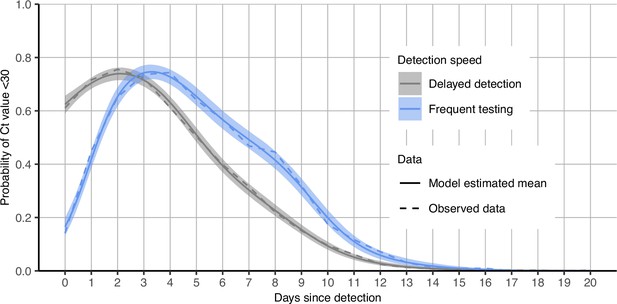
Proportion of infections with Ct <30 on each day post detection stratified by detection speed.
Solid colored lines and shaded ribbons show posterior means and 95% credible intervals from a generalized linear model predicting probability of Ct <30 as a function of days since detection. Dashed lines show proportion with Ct <30 from the observed data.
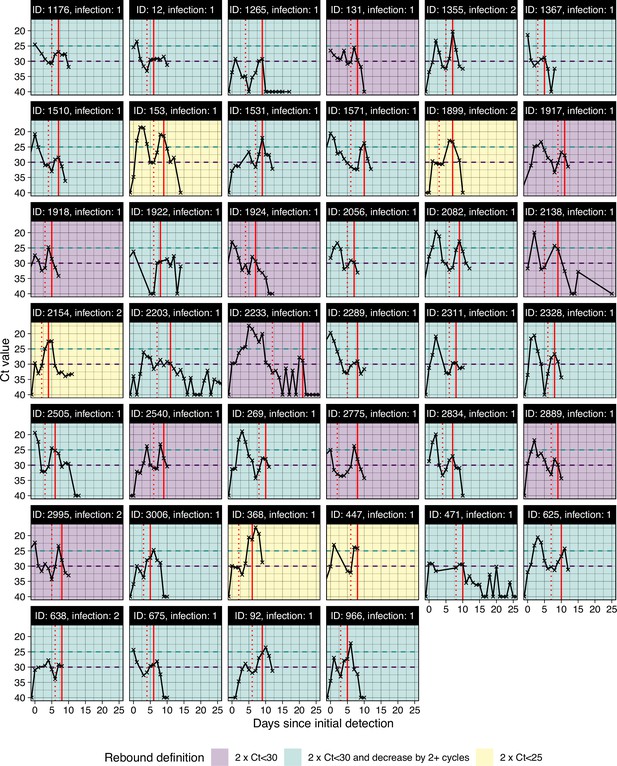
All viral trajectories classified as rebound shown in Figure 1B.
Subplots are colored by the most stringent definition for rebound. To be included here, individuals must have 2+consecutive days of Ct ≥30 after an initial Ct <30. The vertical red dotted line marks this initial clearance time. Trajectories are then classified as rebounds following either two consecutive tests with Ct <30 (purple), two consecutive tests with Ct <30 but with at least a 2 Ct decrease (green), or two consecutive tests with Ct <25 (yellow). The vertical red line marks the timing of rebound detection. The horizontal dashed lines show the different Ct value thresholds for rebound classification. Panels are labeled by arbitrary person ID and infection number. (B) Proportion of sequenced infections attributable to Delta, Omicron BA.1 or other variants.
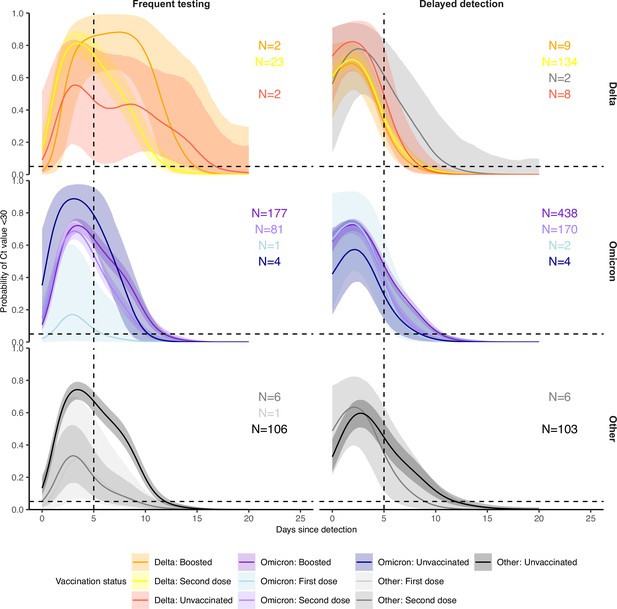
Proportion of infections with Ct <30 on each day post detection by confirmed or suspected variant, vaccination status and detection group, conditional on being in the <30 years age group.
Solid colored lines and shaded ribbons are posterior estimates from a generalized linear model predicting probability of Ct <30 as a function of days since detection and vaccination status, showing the posterior mean (solid line) and 95% credible intervals (shaded ribbon) of each conditional effect. Dashed horizontal and vertical lines show 5% probability and day 5 post detection respectively. Labels show sample size within each group. Note that data is unavailable for some variant and vaccination status combinations.
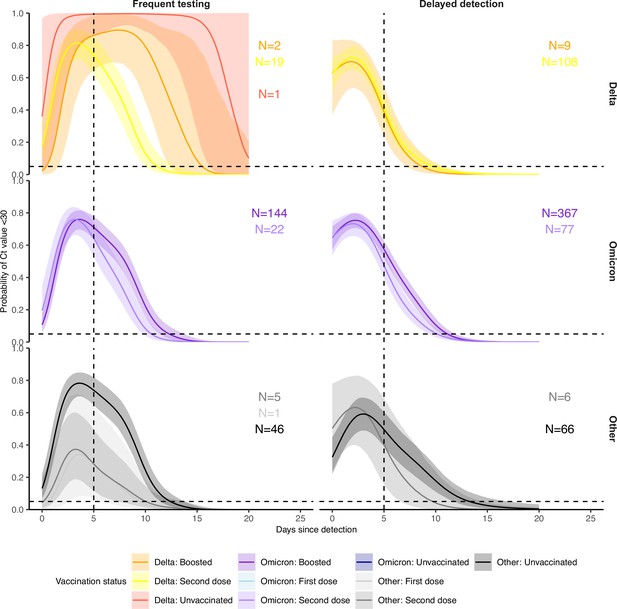
Identical to Appendix 1—figure 9, but after excluding data from all players.
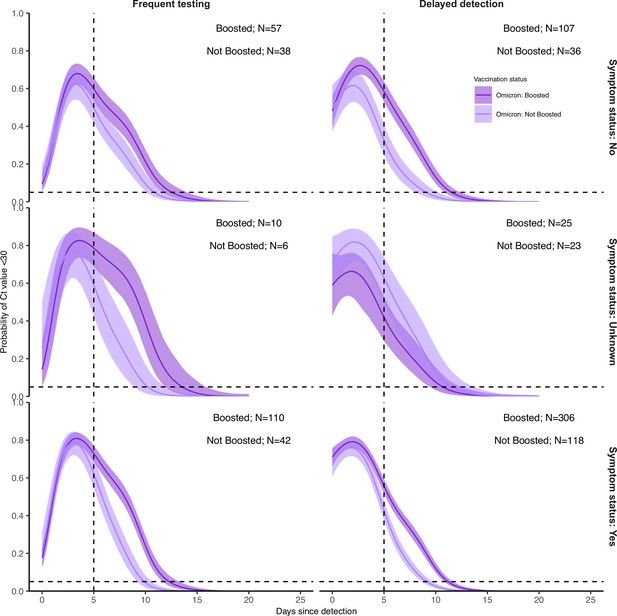
Proportion of Omicron BA.1 infections, stratified by symptom status, with Ct <30 on each day post detection by booster status and detection group.
Solid colored lines and shaded ribbons are posterior estimates from a generalized linear model predicting probability of Ct <30 as a function of days since detection, variant (only BA.1 results shown) and vaccination status, showing the posterior mean (solid line) and 95% credible intervals (shaded ribbon) of each conditional effect. Dashed horizontal and vertical lines show 5% probability and day 5 post detection respectively. Note that age group is not included as an effect in this model.
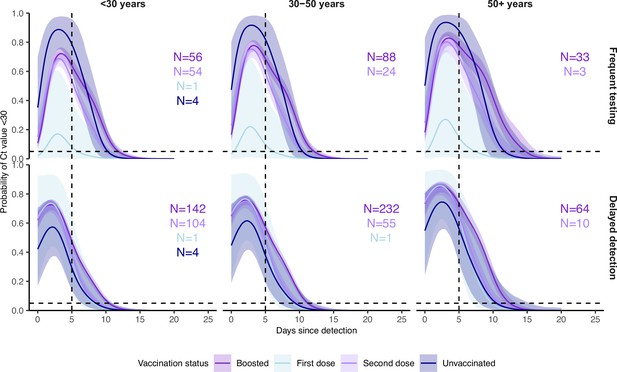
Proportion of Omicron BA.1 infections with Ct <30 on each day post detection stratified by vaccination status, age group and detection group.
Solid colored lines and shaded ribbons are posterior estimates from a generalized linear model predicting probability of Ct <30 as a function of days since detection, vaccination status and variant, showing the posterior mean (solid line) and 95% credible intervals (shaded ribbon) of each conditional effect. Note that only Omicron BA.1 estimates are shown, though the model included data from Delta and pre-Delta infections. Dashed horizontal and vertical lines show 5% probability and day 5 post detection respectively.
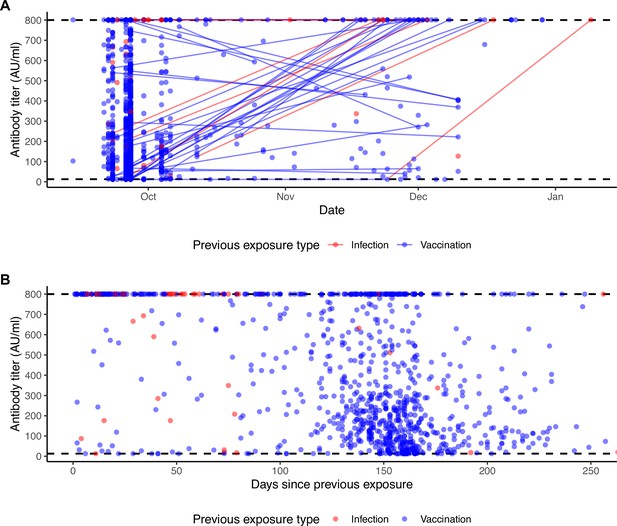
Individual antibody titers to the ancestral SARS-CoV-2 spike over time.
(A) Measured antibody titers by date of sample collection. Lines show longitudinal samples from the same individual, colored by the most recent exposure at the time of sample collection. Lines going up therefore represent antibody boosting events, and lines going down represent waning. (B) Measured antibody titers by days since previous exposure at time of sample collection. Dashed lines show the limit of assay detection.
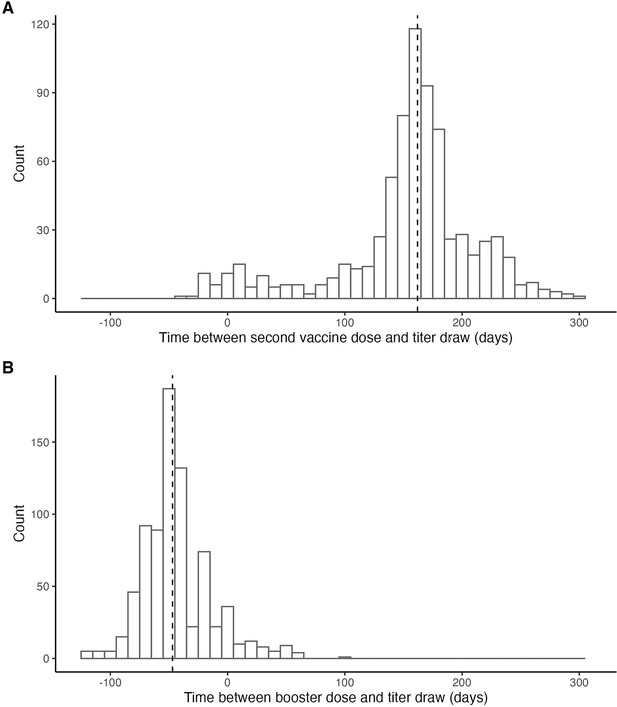
Histogram of time between (A) second vaccine dose and antibody titer measurement and (B) booster dose and antibody titer measurement.
Dashed line marks the median lag (162 days). 1 individual was infected between receiving their second vaccine dose and having a titer measurement taken (Delta infection). 42 individuals were infected between having their titer measurement taken and receiving their booster vaccine dose (32 Delta; 9 unsequenced; 1 confirmed Omicron BA.1).
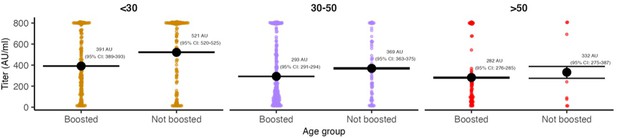
Distribution of antibody titers among Omicron BA.1-infected individuals (colored points) stratified by age group and vaccination status at the time of infection.
Shown are mean titers (large black point) and bootstrapped 95% confidence intervals for the mean (black bars). Note that stratification is by infection and not individual, and that antibody titers were measured at a single point in time rather than near the time of infection. The Diasorin Trimeric Assay values are truncated between 13 and 800 AU/ml.
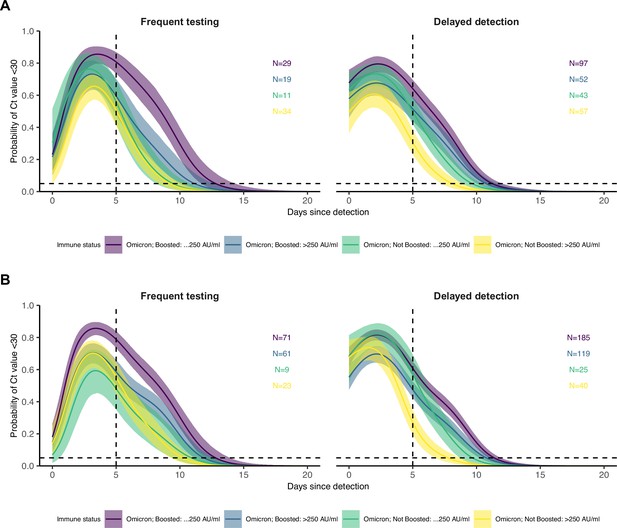
Proportion of BA.1 Omicron infections with Ct <30 on each day post detection stratified by the interaction of booster status at the time of infection and antibody titer group (note, not stratified by age group here due to small subgroup size).
Shown are posterior estimates from a generalized linear model predicting probability of Ct <30 with a spline term on days since detection, conditional on titer/vaccination status category. Solid lines show posterior mean and shaded ribbons show 95% credible intervals of each conditional effect. (A) Model estimates including only individuals who had antibody titers measured between 100 and 200 days following a known previous infection of vaccination. (B) Model estimates including only infections which occurred between 60 and 90 days after an antibody titer measurement.
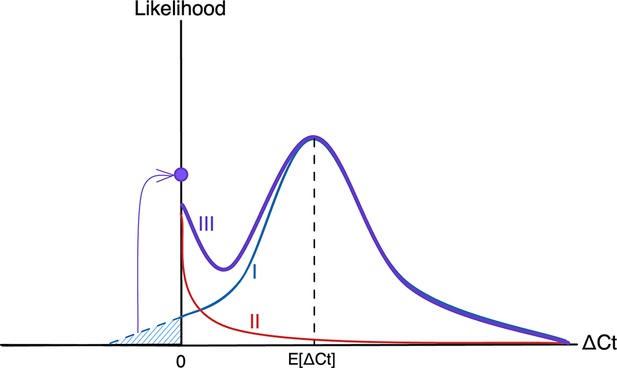
Schematic diagram of the likelihood function for viral kinetic inference.
The plot depicts the likelihood as a function of ΔCt, the difference between the observed Ct value and the limit of detection, so that ΔCt = 0 (the origin) represents observations at the limit of detection, with viral load increasing toward the right-hand side of the plot. The likelihood function (III, purple) is made up of two fundamental components: the process likelihood (I, blue) and the false negative distribution (II, red). The main component of the process likelihood (I) is defined by a normal distribution with mean E[ΔCt], a function of the estimated viral kinetic parameters as defined by the viral kinetic model. Any mass of the process likelihood that extends below the limit of detection (blue hatched region) is instead added to a point probability mass at the origin, since viral loads below the limit of detection register at the limit of detection. The false negative distribution (II) is an exponential distribution with fixed rate to account for a small amount of noise near the limit of detection. Summing the process likelihood (I) and the false negative likelihood (I) using the mixing probability λ (=1 - sensitivity) yields the overall likelihood (III).
Tables
Characteristics of the documented infections.
Counts (N) correspond to numbers of infections.
| Variable | Category | N | Percent |
|---|---|---|---|
| Total | – | 1280 | 100 |
| Variant | Delta | 180 | 14.1 |
| Omicron BA.1 | 878 | 68.6 | |
| Other | 222 | 17.3 | |
| Vaccination Status | Unvaccinated | 228 | 17.8 |
| First dose | 6 | 0.5 | |
| Second dose | 420 | 32.8 | |
| Boosted | 626 | 48.9 | |
| Antibody Titer | 13–250 | 473 | 37.0 |
| 250–800 | 504 | 39.4 | |
| Unknown | 303 | 23.7 | |
| Symptomatic | No | 257 | 20.1 |
| Yes | 664 | 51.9 | |
| Unknown | 359 | 28 | |
| Detection Speed | Delayed detection | 877 | 68.5 |
| Frequent testing | 403 | 31.5 | |
| Age Group | 0–30 | 556 | 43.4 |
| 31–50 | 568 | 44.4 | |
| 50+ | 155 | 12.1 | |
| Unknown | 1 | 0.1 | |
| Cumulative Infection Number | 1 | 1128 | 88.1 |
| 2 | 149 | 11.6 | |
| 3 | 2 | 0.2 | |
| 4 | 1 | 0.1 | |
| Days Since Previous Exposure | Naïve (no prior exposure) | 220 | 17.2 |
| <1 month | 273 | 21.3 | |
| 1–3 months | 403 | 31.5 | |
| >3 months | 384 | 30.0 |
Number of rebound infections classified under different definitions for initial clearance and subsequent rebound.
| Initial clearance duration (consecutive days with Ct ≥30) | Rebound duration (days above Ct value threshold) | Ct value threshold of rebound | Rebounds | Total | Percentage |
|---|---|---|---|---|---|
| ≥4 | ≥4 | Ct <30 | 0 | 749 | 0.00% |
| ≥4 | ≥3 | Ct <30 | 1 | 749 | 0.13% |
| ≥4 | ≥2 | Ct <30 | 4 | 749 | 0.53% |
| ≥3 | ≥4 | Ct <30 | 2 | 999 | 0.20% |
| ≥3 | ≥3 | Ct <30 | 7 | 999 | 0.70% |
| ≥3 | ≥2 | Ct <30 | 16 | 999 | 1.60% |
| ≥2 | ≥4 | Ct <30 | 7 | 1334 | 0.53% |
| ≥2 | ≥3 | Ct <30 | 18 | 1334 | 1.35% |
| ≥2 | ≥2 | Ct <30 | 40 | 1334 | 3.00% |
| ≥4 | ≥4 | Ct <25 | 0 | 749 | 0.00% |
| ≥4 | ≥3 | Ct <25 | 0 | 749 | 0.00% |
| ≥4 | ≥2 | Ct <25 | 0 | 749 | 0.00% |
| ≥3 | ≥4 | Ct <25 | 1 | 999 | 0.10% |
| ≥3 | ≥3 | Ct <25 | 1 | 999 | 0.10% |
| ≥3 | ≥2 | Ct <25 | 2 | 999 | 0.20% |
| ≥2 | ≥4 | Ct <25 | 1 | 1334 | 0.08% |
| ≥2 | ≥3 | Ct <25 | 2 | 1334 | 0.15% |
| ≥2 | ≥2 | Ct <25 | 5 | 1334 | 0.38% |
Number of identified rebounds stratified by variant, either confirmed through sequencing or assumed based on detection date.
Rebounds are defined here as any trajectory with an initial Ct <30, followed by a sequence of two or more consecutive negative tests or tests with Ct ≥30, and subsequently followed by two or more consecutive tests with Ct <30.
| Variant | Rebounds | Total infections | Percentage rebounded |
|---|---|---|---|
| Omicron | 36 | 877 | 4.10% |
| Delta | 1 | 178 | 0.562% |
| Other | 3 | 279 | 1.08% |
Number of identified rebounds stratified by vaccination status.
Rebounds are defined here as any trajectory with an initial Ct <30, followed by a sequence of two or more consecutive negative tests or tests with Ct ≥30, and subsequently followed by two or more consecutive tests with Ct <30.
| Vaccination status | Rebounds | Total infections | Percentage rebounded |
|---|---|---|---|
| Boosted | 32 | 494 | 6.48% |
| No record | 5 | 398 | 1.26% |
| Second dose | 3 | 323 | 0.929% |
| Unvaccinated | 2 | 159 | 1.26% |
Comparison of linear logistic regression models predicting probability of Ct <30 on each day since detection among individuals in the frequent testing group.
Models are ranked based on their expected log pointwise predictive density (ELPD), where a lower ELPD implies better predictive accuracy. Model weight refers to the weight of each model in a Bayesian Model Averaging analysis, where a higher value implies a greater contribution to model prediction when combining multiple models. AUC = area under the curve.
| Model | ELPD difference | SE difference | Weight | AUC | Classification accuracy | Accuracy (≥30) | Accuracy (<30) |
|---|---|---|---|---|---|---|---|
| Cumulative number of exposures, variant and age | 0.000 | 0.000 | 0.157 | 0.897 | 0.828 | 0.890 | 0.647 |
| Days since previous exposure, variant and age | –1.813 | 10.011 | 0.299 | 0.896 | 0.823 | 0.878 | 0.664 |
| Vaccination status, variant and age | –7.440 | 9.828 | 0.089 | 0.896 | 0.824 | 0.881 | 0.656 |
| Cumulative number of exposures and age | –27.935 | 8.823 | 0.104 | 0.894 | 0.823 | 0.887 | 0.637 |
| Cumulative number of exposures and variant | –31.536 | 9.428 | 0.000 | 0.894 | 0.824 | 0.891 | 0.630 |
| Vaccination status and age | –33.933 | 12.389 | 0.000 | 0.893 | 0.822 | 0.882 | 0.646 |
| Vaccination status and variant | –35.274 | 13.693 | 0.156 | 0.893 | 0.824 | 0.886 | 0.645 |
| Days since previous exposure and variant | –39.750 | 14.206 | 0.000 | 0.891 | 0.822 | 0.883 | 0.644 |
| Days since previous exposure and age | –40.929 | 12.503 | 0.000 | 0.892 | 0.820 | 0.877 | 0.654 |
| Variant and age | –52.437 | 11.681 | 0.000 | 0.890 | 0.817 | 0.884 | 0.624 |
| Cumulative number of exposures | –53.661 | 12.344 | 0.143 | 0.892 | 0.822 | 0.895 | 0.610 |
| Vaccination status | –58.503 | 15.162 | 0.000 | 0.890 | 0.822 | 0.888 | 0.632 |
| Age | –60.160 | 12.688 | 0.000 | 0.890 | 0.818 | 0.882 | 0.632 |
| Days since previous exposure | –69.842 | 15.547 | 0.045 | 0.888 | 0.817 | 0.884 | 0.624 |
| Variant | –87.427 | 15.554 | 0.007 | 0.887 | 0.818 | 0.889 | 0.611 |
| Baseline | –93.087 | 15.843 | 0.000 | 0.886 | 0.817 | 0.890 | 0.606 |
Comparison of linear logistic regression models predicting probability of Ct <30 on each day since detection among individuals in the delayed testing group.
Models are ranked based on their expected log pointwise predictive density (ELPD), where a lower ELPD implies better predictive accuracy. Model weight refers to the weight of each model in a Bayesian Model Averaging analysis, where a higher value implies a greater contribution to model prediction when combining multiple models. AUC = area under the curve.
| Model | ELPD difference | SE difference | Weight | AUC | Classification accuracy | Accuracy (≥30) | Accuracy (<30) |
|---|---|---|---|---|---|---|---|
| Cumulative number of exposures, variant and age | 0.000 | 0.000 | 0.154 | 0.914 | 0.850 | 0.893 | 0.712 |
| Vaccination status, variant and age | –1.817 | 7.807 | 0.000 | 0.913 | 0.849 | 0.890 | 0.720 |
| Vaccination status and age | –6.061 | 8.899 | 0.134 | 0.913 | 0.849 | 0.888 | 0.724 |
| Cumulative number of exposures and age | –7.900 | 4.710 | 0.007 | 0.913 | 0.849 | 0.893 | 0.712 |
| Days since previous exposure, variant and age | –8.046 | 8.929 | 0.288 | 0.913 | 0.849 | 0.890 | 0.721 |
| Days since previous exposure and age | –20.190 | 9.360 | 0.000 | 0.912 | 0.850 | 0.890 | 0.725 |
| Variant and age | –22.457 | 8.628 | 0.081 | 0.912 | 0.850 | 0.888 | 0.728 |
| Vaccination status and variant | –59.936 | 14.986 | 0.162 | 0.909 | 0.847 | 0.888 | 0.717 |
| Cumulative number of exposures and variant | –60.139 | 12.098 | 0.007 | 0.910 | 0.846 | 0.891 | 0.703 |
| Age | –63.495 | 12.834 | 0.052 | 0.910 | 0.849 | 0.882 | 0.743 |
| Cumulative number of exposures | –65.893 | 12.851 | 0.114 | 0.910 | 0.846 | 0.891 | 0.707 |
| Vaccination status | –67.520 | 15.418 | 0.000 | 0.909 | 0.847 | 0.888 | 0.717 |
| Days since previous exposure and variant | –82.798 | 15.986 | 0.000 | 0.909 | 0.846 | 0.888 | 0.714 |
| Days since previous exposure | –93.537 | 16.058 | 0.000 | 0.909 | 0.846 | 0.887 | 0.718 |
| Variant | –107.362 | 15.800 | 0.000 | 0.907 | 0.845 | 0.883 | 0.723 |
| Baseline | –148.350 | 18.608 | 0.000 | 0.905 | 0.842 | 0.880 | 0.722 |
Posterior estimates of viral trajectory attributes by variant and vaccination status.
Estimates are posterior means with 95% credible intervals.
| Trajectory value | Variant/vaccination status | Estimate |
|---|---|---|
| Peak viral load (Ct) | Other: Unvaccinated | 25.0 (24.2, 25.9) |
| Delta: 1–2 doses | 22.4 (21.4, 23.5) | |
| Omicron: 1–2 doses | 25.6 (25.0, 26.2) | |
| Omicron: Boosted | 25.7 (25.4, 26.1) | |
| Peak viral load (log10 copies/ml) | Other: Unvaccinated | 6.8 (6.6, 7.0) |
| Delta: 1–2 doses | 7.5 (7.2, 7.8) | |
| Omicron: 1–2 doses | 6.6 (6.5, 6.8) | |
| Omicron: Boosted | 6.6 (6.5, 6.7) | |
| Proliferation time (days) | Other: Unvaccinated | 3.5 (3.1, 3.9) |
| Delta: 1–2 doses | 3.6 (3.1, 4.1) | |
| Omicron: 1–2 doses | 3.6 (3.3, 4.0) | |
| Omicron: Boosted | 4.0 (3.8, 4.3) | |
| Clearance time (days) | Other: Unvaccinated | 9.9 (9.2, 10.6) |
| Delta: 1–2 doses | 7.6 (7.0, 8.3) | |
| Omicron: 1–2 doses | 6.2 (5.8, 6.6) | |
| Omicron: Boosted | 8.4 (8.0, 8.7) |
Posterior estimates of viral trajectory attributes by variant and antibody titer.
Estimates are posterior means with 95% credible intervals.
| Trajectory attribute | Variant/immune status | Estimate |
|---|---|---|
| Peak viral load (Ct) | Other: Unexposed | 24.8 (24.0, 25.6) |
| Delta: Exposed,≤250 AU | 22.1 (20.6, 23.4) | |
| Delta: Exposed,>250 AU | 24.4 (22.7, 26.1) | |
| Omicron: Exposed,≤250 AU | 25.2 (24.7, 25.6) | |
| Omicron: Exposed,>250 AU | 26.2 (25.7, 26.6) | |
| Peak viral load (log10 copies/ml) | Other: Unexposed | 6.9 (6.6, 7.1) |
| Delta: Exposed,≤250 AU | 7.6 (7.3, 8.0) | |
| Delta: Exposed,>250 AU | 7.0 (6.5, 7.4) | |
| Omicron: Exposed,≤250 AU | 6.8 (6.7, 6.9) | |
| Omicron: Exposed,>250 AU | 6.5 (6.4, 6.6) | |
| Proliferation time (days) | Other: Unexposed | 3.5 (3.2, 3.9) |
| Delta: Exposed,≤250 AU | 3.6 (2.8, 4.5) | |
| Delta: Exposed,>250 AU | 4.2 (3.3, 5.3) | |
| Omicron: Exposed,≤250 AU | 3.9 (3.6, 4.2) | |
| Omicron: Exposed,>250 AU | 3.7 (3.5, 4.0) | |
| Clearance time (days) | Other: Unexposed | 9.8 (9.1, 10.6) |
| Delta: Exposed,≤250 AU | 7.7 (6.9, 8.7) | |
| Delta: Exposed,>250 AU | 7.6 (6.4, 8.8) | |
| Omicron: Exposed,≤250 AU | 8.4 (8.0, 8.8) | |
| Omicron: Exposed,>250 AU | 6.9 (6.6, 7.3) |
Posterior estimates of Omicron BA.1 viral trajectory attributes by symptom and vaccination status.
Estimates are posterior means with 95% credible intervals.
| Trajectory attribute | Variant/immune status | Estimate |
|---|---|---|
| Peak viral load (Ct) | Omicron: 1–2 doses, no symptoms | 26.7 (25.7, 27.7) |
| Omicron: 1–2 doses, symptoms | 25.3 (24.5, 26.1) | |
| Omicron: boosted, no symptoms | 26.3 (25.7, 27) | |
| Omicron: boosted, symptoms | 25.4 (25, 25.8) | |
| Peak viral load (log10 copies/ml) | Omicron: 1–2 doses, no symptoms | 6.3 (6.1, 6.6) |
| Omicron: 1–2 doses, symptoms | 6.7 (6.5, 6.9) | |
| Omicron: boosted, no symptoms | 6.4 (6.3, 6.6) | |
| Omicron: boosted, symptoms | 6.7 (6.6, 6.8) | |
| Proliferation time (days) | Omicron: 1–2 doses, no symptoms | 3.7 (3.2, 4.2) |
| Omicron: 1–2 doses, symptoms | 3.7 (3.2, 4.2) | |
| Omicron: boosted, no symptoms | 4.3 (3.9, 4.8) | |
| Omicron: boosted, symptoms | 3.8 (3.5, 4.1) | |
| Clearance time (days) | Omicron: 1–2 doses, no symptoms | 5.9 (5.2, 6.6) |
| Omicron: 1–2 doses, symptoms | 6.3 (5.8, 6.8) | |
| Omicron: boosted, no symptoms | 7.4 (6.8, 8.0) | |
| Omicron: boosted, symptoms | 8.7 (8.3, 9.1) |
Posterior estimates of Omicron BA.1 viral trajectory attributes by symptom and antibody titer.
Estimates are posterior means with 95% credible intervals.
| Trajectory attribute | Variant/immune status | Estimate |
|---|---|---|
| Peak viral load (Ct) | Omicron: Low titer, no symptoms | 26.1 (25.2, 26.9) |
| Omicron: Low titer, symptoms | 24.8 (24.3, 25.4) | |
| Omicron: High titer, no symptoms | 26.7 (25.9, 27.5) | |
| Omicron: High titer, symptoms | 25.9 (25.4, 26.5) | |
| Peak viral load (log10 copies/ml) | Omicron: Low titer, no symptoms | 6.5 (6.3, 6.8) |
| Omicron: Low titer, symptoms | 6.9 (6.7, 7.0) | |
| Omicron: High titer, no symptoms | 6.3 (6.1, 6.6) | |
| Omicron: High titer, symptoms | 6.6 (6.4, 6.7) | |
| Proliferation time (days) | Omicron: Low titer, no symptoms | 4.3 (3.8, 4.9) |
| Omicron: Low titer, symptoms | 3.6 (3.3, 4.0) | |
| Omicron: High titer, no symptoms | 3.8 (3.4, 4.3) | |
| Omicron: High titer, symptoms | 3.8 (3.4, 4.2) | |
| Clearance time (days) | Omicron: Low titer, no symptoms | 7.4 (6.7, 8.1) |
| Omicron: Low titer, symptoms | 8.7 (8.2, 9.3) | |
| Omicron: High titer, no symptoms | 6.4 (5.8, 7.1) | |
| Omicron: High titer, symptoms | 7.1 (6.6, 7.5) |
Posterior estimates of Omicron BA.1 viral RNA clearance times by age and vaccination status.
Estimates are posterior means with 95% credible intervals. Low titer is ≤250 AU, high titer is >250 AU.
| Trajectory attribute | Variant/immune status | Estimate |
|---|---|---|
| Peak viral load (Ct) | Omicron:<30, unboosted | 25.6 (25, 26.3) |
| Omicron:<30, boosted | 25.8 (25.1, 26.4) | |
| Omicron: 30–50, unboosted | 25.3 (24.2, 26.5) | |
| Omicron: 30–50, boosted | 25.7 (25.2, 26.2) | |
| Omicron: 50+, unboosted | 25.5 (22.8, 27.9) | |
| Omicron: 50+, boosted | 25.5 (24.6, 26.3) | |
| Peak viral load (log10 copies/ml) | Omicron:<30, unboosted | 6.6 (6.4, 6.8) |
| Omicron:<30, boosted | 6.6 (6.4, 6.8) | |
| Omicron: 30–50, unboosted | 6.7 (6.4, 7) | |
| Omicron: 30–50, boosted | 6.6 (6.5, 6.7) | |
| Omicron: 50+, unboosted | 6.7 (6, 7.4) | |
| Omicron: 50+, boosted | 6.7 (6.5, 6.9) | |
| Proliferation time (days) | Omicron:<30, unboosted | 3.7 (3.4, 4.1) |
| Omicron:<30, boosted | 4.2 (3.8, 4.7) | |
| Omicron: 30–50, unboosted | 3.6 (3, 4.4) | |
| Omicron: 30–50, boosted | 4 (3.6, 4.3) | |
| Omicron: 50+, unboosted | 4.4 (3, 6.3) | |
| Omicron: 50+, boosted | 3.9 (3.4, 4.4) | |
| Clearance time (days) | Omicron:<30, unboosted | 6 (5.6, 6.5) |
| Omicron:<30, boosted | 7.2 (6.7, 7.7) | |
| Omicron: 30–50, unboosted | 6.4 (5.7, 7.2) | |
| Omicron: 30–50, boosted | 8.6 (8.1, 9.1) | |
| Omicron: 50+, unboosted | 8.7 (6.6, 11.2) | |
| Omicron: 50+, boosted | 9.6 (8.8, 10.6) |
Posterior estimates of Omicron BA.1 viral RNA clearance times by age and titer.
Estimates are posterior means with 95% credible intervals. Low titer is ≤250 AU, high titer is >250 AU.
| Trajectory attribute | Variant/immune status | Estimate |
|---|---|---|
| Peak viral load (Ct) | Omicron:<30, low titer | 24.8 (24, 25.5) |
| Omicron:<30, high titer | 26.2 (25.6, 26.8) | |
| Omicron: 30–50, low titer | 25.4 (24.8, 26) | |
| Omicron: 30–50, high titer | 26.1 (25.4, 26.8) | |
| Omicron: 50+, low titer | 24.9 (23.8, 25.9) | |
| Omicron: 50+, high titer | 26.3 (24.8, 27.6) | |
| Peak viral load (log10 copies/ml) | Omicron:<30, low titer | 6.9 (6.7, 7.1) |
| Omicron:<30, high titer | 6.5 (6.3, 6.7) | |
| Omicron: 30–50, low titer | 6.7 (6.5, 6.9) | |
| Omicron: 30–50, high titer | 6.5 (6.3, 6.7) | |
| Omicron: 50+, low titer | 6.8 (6.6, 7.1) | |
| Omicron: 50+, high titer | 6.5 (6.1, 6.9) | |
| Proliferation time (days) | Omicron:<30, low titer | 3.9 (3.4, 4.3) |
| Omicron:<30, high titer | 4 (3.6, 4.4) | |
| Omicron: 30–50, low titer | 4 (3.6, 4.4) | |
| Omicron: 30–50, high titer | 3.6 (3.2, 4) | |
| Omicron: 50+, low titer | 3.8 (3.2, 4.5) | |
| Omicron: 50+, high titer | 3.7 (2.9, 4.7) | |
| Clearance time (days) | Omicron:<30, low titer | 7.2 (6.7, 7.8) |
| Omicron:<30, high titer | 6.2 (5.8, 6.7) | |
| Omicron: 30–50, low titer | 8.7 (8.1, 9.3) | |
| Omicron: 30–50, high titer | 7.4 (6.8, 8) | |
| Omicron: 50+, low titer | 9.9 (8.7, 11.1) | |
| Omicron: 50+, high titer | 9.5 (8, 11.1) |





