Machine learning-assisted elucidation of CD81–CD44 interactions in promoting cancer stemness and extracellular vesicle integrity
Figures
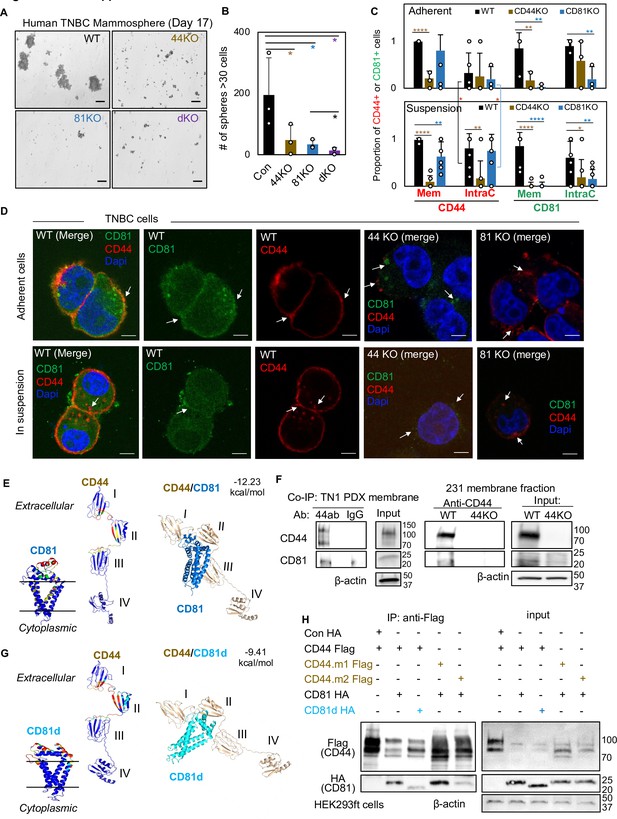
CD81 interacts with CD44 on the membrane and promotes mammosphere formation of triple negative breast cancer (TNBC) cells.
Representative images (A) and bar graphs (B) of the mammospheres of MDA-MB-231 cell groups (WT, CD44KO, CD81KO, dKO pool populations), 17 days after seeded at 2000 cells per well (6-well plate) in serum-free mammosphere formation media. N = 4 replicates. Error bars represent standard deviation. Repeated three times. Scale bar = 100 µm. One-tailed Student T-test *p < 0.05. Bar graphs (C) and representative images (D) of membrane and intracellular CD44 and CD81 localization in WT, CD44KO, and CD81KO MDA-MB-231 cells (adherent and in suspension), detected by immunofluorescence staining with anti-CD44-Texas Red, anti-CD81-Alexa488, and DAPI (4’, 6-diamidino-2-phenylindole) for DNA staining. Sample size N = 20 (WT), 14 (CD44KO), and 20 (CD81KO) adherent cells and N = 31 (WT), 31 (CD44KO), and 28 CD81KO cells in suspension. Error bars represent standard deviation. Scale bar = 5 µm. Two-tailed Student T-test p = 0.03 for intracellular CD44 levels between adherent and suspension cells (both WT and CD81KO cells). Compared to WT cells, T-test p values for adherent CD44KO/CD81KO cells are 9.52e−06/0.22 (membrane CD44), 0.81/0.65 (intracellular CD44), 0.009/0.006 (membrane CD81), 0.15/0.002 (intracellular CD81); and p values for cells in suspension are 1.21e−11/0.003 (membrane CD44), 0.002/0.72 (intracellular CD44), 3.08e−06/2.64e−07 (membrane CD81), and 0.04/0.004 (intracellular CD81). Significant pair comparison differences are marked in the graph. ANOVA analyses among three groups with p values for adherent and in-suspension cells, respectively: 0.0004 and 7.357E−08 (membrane CD44), 0.9119 and 0.0024 (intracellular CD44), 0.0022 and 1.079E−09 (membrane CD81), and 0.0255 and 0.0103 (intracellular CD81). (E) Predictive modeling of CD44 and CD81 interaction with hot spots shown in red and yellow. Top-ranked structural models of predictive interactions between CD44/CD81 (estimated binding energy: −12.23 kcal/mol) and between CD44/CD81d (deletion mutant) (estimated binding energy: −9.41 Kcal/mol). (F) Immunoblots of endogenous CD44 and CD81 immunoprecipitated by anti-CD44 from the lysates (membrane fraction) of WT/44KO MDA-MB-231 and (total) WT TN1 patient-derived xenograft (PDX) cells. (G) Predictive modeling of CD44 and CD81d (deletion mutant) interaction with hot spots shown in red and yellow. Top-ranked structural models of predictive interactions between CD44/CD81d (deletion mutant) (estimated binding energy: −9.41 kcal/mol). (H) Representative immunoblots of CD81-HA (CD81d) immunoprecipitated by anti-CD44-Flag (CD44 mutants CD44.1 and CD44.2) from the lysates of HEK293ft cells (N = 3 biological replicates; Con, control). Cells were transfected with either Flag-CD44 or mutants (Flag-CD44.1, Flag-CD44.2) and Con HA, HA-CD81, or HA-CD81d with deletion at amino acids 159–187 (CD81d). Forty-eight hours after transfection the cells were harvested for Flag IP.
-
Figure 1—source data 1
Uncropped blots associated with Figure 1F.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig1-data1-v2.pdf
-
Figure 1—source data 2
Uncropped blots associated with Figure 1H.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig1-data2-v2.pdf
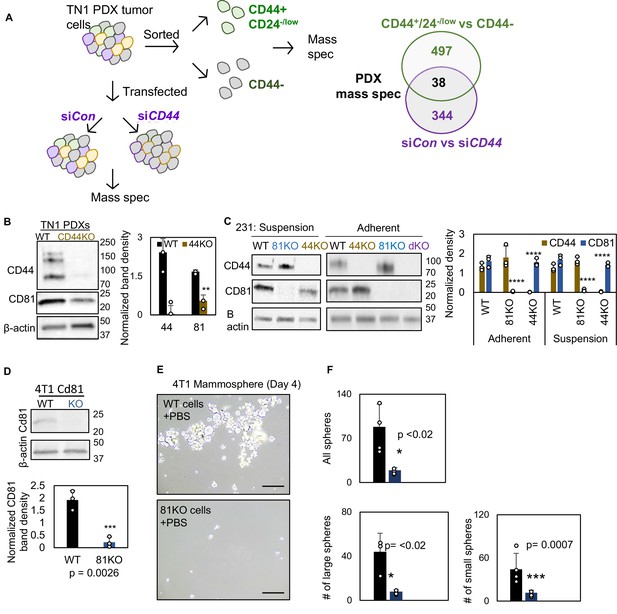
Characterization of CD44KO and CD81KO breast cancer cell lines.
(A) Schematic of mass spectrometry workflow. TN1 patient-derived xenograft (PDX) tumor was dissociated and sorted based on CD44 and CD24 expression and analyzed by mass spec. Four hundred and ninety-seven differential proteins were identified. TN1 PDX tumor was dissociated and transfected with siCD44 and siCon and analyzed by mass spec. Three hundred and forty-four differential proteins were identified. Thirty-eight differential proteins overlapped in both screens, one of which was CD81. (B, C) Immunoblots and quantification of CD44 and CD81 in dissociated PDX (WT and CD44KO) tumor cells in suspension and MDA-MB-231 cells, in suspension and adherent (WT, CD44KO, CD81KO, and dKO). N= 3. Error bars represent standard deviation. *p < 0.05, **p = 0.018, ***p = 0.0026, ****p = <0.0008. (D) Mammosphere formation and immunoblot of mouse 4T1 cells (WT and Cd81 KO via CRISPR/Cas9). Cells were seeded 2000 cells/well in 6-well plate in four replicate.Two-tailed Student T-test was used. Repeated twice. Representative images (E) and quantifications (F) of mammospheres derived from 4T1 cells in suspension, including WT cells and Cd81KO cells. 2000 cells were seeded in 6-cm plates in mammosphere media, and the images were captured on day 4. N= 3. Error bars represent standard deviation. Two-tailed Student T-test was used to calculate p values.
-
Figure 1—figure supplement 1—source data 1
Uncropped blots associated with Figure 1—figure supplement 1B.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig1-figsupp1-data1-v2.pdf
-
Figure 1—figure supplement 1—source data 2
Uncropped blots associated with Figure 1—figure supplement 1C.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig1-figsupp1-data2-v2.pdf
-
Figure 1—figure supplement 1—source data 3
Uncropped blots associated with Figure 1—figure supplement 1D.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig1-figsupp1-data3-v2.pdf
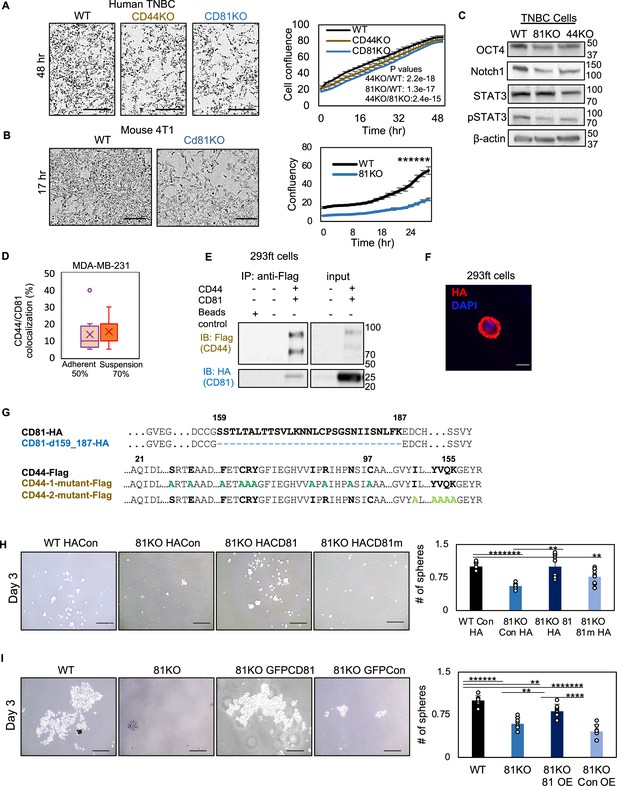
CD81 colocalizes with CD44 on cellular membrane in breast cancer cell line.
(A, B) IncuCyte images and curve analyses of cell confluence of human MDA-MB-231 WT, CD44KO, and CD81KO cells (C) and mouse 4T1 WT and Cd81KO cells (D) over time. Scale bar is 300 µm. N= 3. Error bars represent standard deviation. Two-tailed Student T-test ******p = 1.97E−08. (C) Immunoblot analysis of OCT4, Notch1, STAT3, and pSTAT3 expression in WT, CD81KO, and CD44KO MDA-MB-231 cells (pooled KO cells). (D) % of CD44 and CD81 colocalization shown in 50% of adherent (14 out of 26 cells) and 70% of suspension (22 out of 31 cells) MDA-MB-231 cells (WT). Pearson coefficient r = 0.57 (average from three experimental replicates r1 = 0.52, r2 = 0.54, r3 = 0.64). (E) Immunoblots of exogenously expressed CD44-Flag and CD81-HA immunoprecipitated by anti-Flag (CD44) from the lysates of transfected HEK293ft cells. Repeated three times. (F) Immunofluorescence of HA tagged CD81d in HEK293ft cells showing its membrane localization. (G) Alignment of CD44 and CD81 amino acids showing deletion in CD81 and point mutations in CD44 domains I and II. (H) Representative images and bar graphs of the mammospheres of MDA-MB-231 cell groups (WT HACon, CD81KO HACon, CD81KO HACD81, CD81KO HACD81m), 3 days after seeded at 1000 cells per well (12-well plate) in serum-free mammosphere formation media. N = 4 replicates. Repeated two times. Error bars represent standard deviation. Scale bar = 100 µm. Two-tailed Student T-test *******p = 0.00000001, ****p = 0.0004, **p = 0.005. (I) Representative images and bar graphs of the mammospheres of MDA-MB-231 cell groups (WT, CD81KO, CD81KO GFPCD81) overexpression (OE), CD81KO GFPCon, 3 days after seeded at 1000 cells per well (12-well plate) in serum-free mammosphere formation media. N = 4 replicates. Repeated two times. Error bars represent standard deviation. Scale bar = 100 µm. Two-tailed Student T-test *******p = 0.00000005, ******p = 0.0000009, ****p = 0.00004, ***p < 0.004, **p = 0.005.
-
Figure 1—figure supplement 2—source data 1
Uncropped blots associated with Figure 1—figure supplement 2C.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig1-figsupp2-data1-v2.pdf
-
Figure 1—figure supplement 2—source data 2
Uncropped blots associated with Figure 1—figure supplement 2E.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig1-figsupp2-data2-v2.pdf
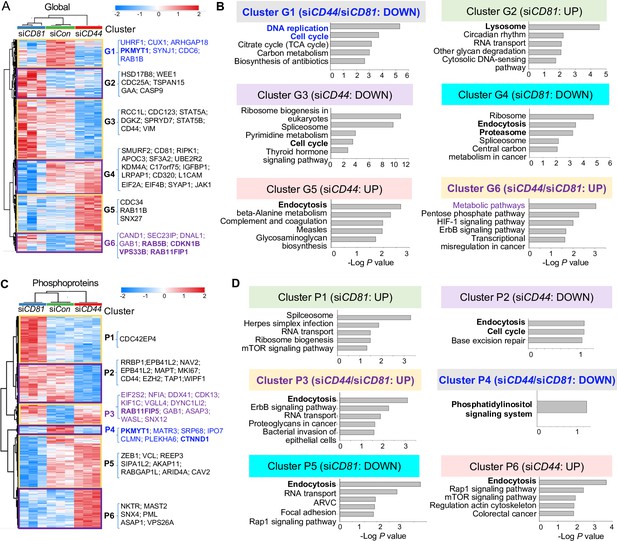
Global mass spectrometry and phosphoproteomic profiling of MDA-MB-231 cells with siCD81 and siCD44 KDs.
Global mass spectrometry heatmap (A) and KEGG pathway analysis (B) of altered top pathways with significantly expressed proteome within six different clusters (G1–G6) in siControl, siCD81, and siCD44 cells at 3 hr in suspension (N = 3 replicates, ANOVA T-test false discovery rate (FDR) <0.01, p = <0.05). Phosphoproteomic mass spectrometry heatmap (C) and KEGG pathway analysis (D) of altered top pathways with significantly changed phosphoproteome within six different clusters (P1–P6) in siControl, siCD81, and siCD44 cells at 3 hr in suspension (N = 3 replicates, ANOVA T-test FDR <0.01, p = <0.05).
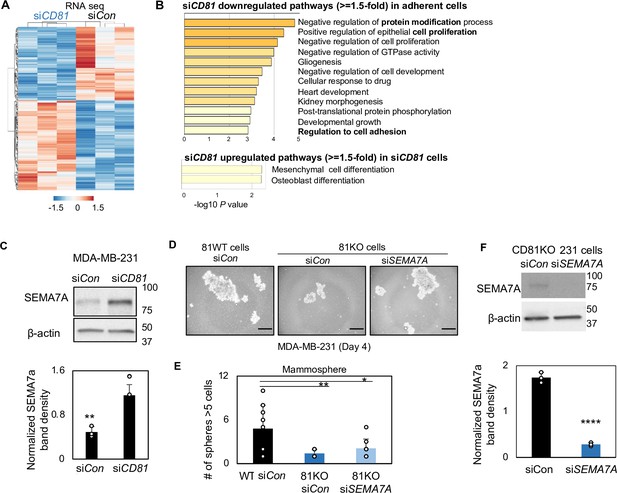
SEMA7A increased in CD81-depleted cells.
RNA-sequencing-based transcriptome analysis shows siCD81-altered pathways (A) and heatmap (B), including down- and upregulated genes with ≥1.5-fold changes in MDA-MB-231 cells compared to the siRNA control (N = 3 replicates). (C) Immunoblots of top target SEMA7A altered by siCD81 in adherent cells. **p = 0.001. Error bars represent standard deviation. Representative images of formed mammospheres (D), quantification (E) of mammospheres, and immunoblots of SEMA7A after siSEMA7A gene KD in CD81KO cells (F) showing downregulation of SEMA7A rescues mammosphere formation of CD81KO MDA-MB-231 cells, SEMA7A was knocked down via siRNA transfection and seeded at 2000 cells/well in 12-well plate. After 4 days, SEMA7A KD in CD81KO cells increased mammosphere number and size compared to the control. Scale bar = 50 µm. N= 4. Error bars represent standard deviation. ****p = 0.00003, **p = 0.007, *p = 0.03.
-
Figure 2—figure supplement 1—source data 1
Uncropped blots associated with Figure 2—figure supplement 1C.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig2-figsupp1-data1-v2.pdf
-
Figure 2—figure supplement 1—source data 2
Uncropped blots associated with Figure 2—figure supplement 1F.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig2-figsupp1-data2-v2.pdf
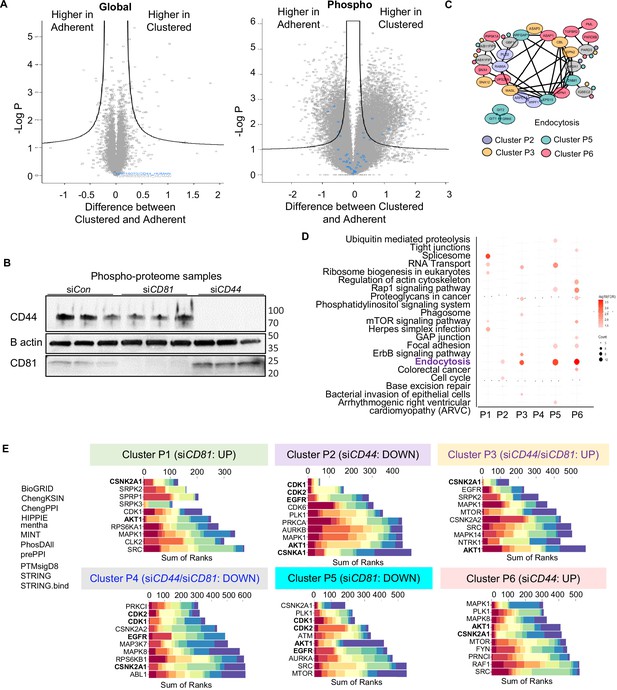
Global and phosphoproteomic analyses in CD81- and CD44-depleted cells.
(A) The Volcano plots of different abundance in global proteome and phosphoproteome between adherent and clustered tumor cells. (B) Immunoblots and bar graphs of CD44 and CD81 in MDA-MB-231 cells after transient knockdowns after siCD81and siCD44 transfections. (C) The protein–protein interactions network of altered phosphoproteome in the endocytosis pathway across four phosphoproteome clusters P2, P3, P5, and P6. The enrichment of KEGG pathway (D) and Kinase Enrichment Analysis (KEA) (E) from expressed phosphopeptides in the different clusters P1–P6 comparing siControl with siCD81 and siCD44 groups.
-
Figure 2—figure supplement 2—source data 1
Uncropped blots associated with Figure 2—figure supplement 2B.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig2-figsupp2-data1-v2.pdf
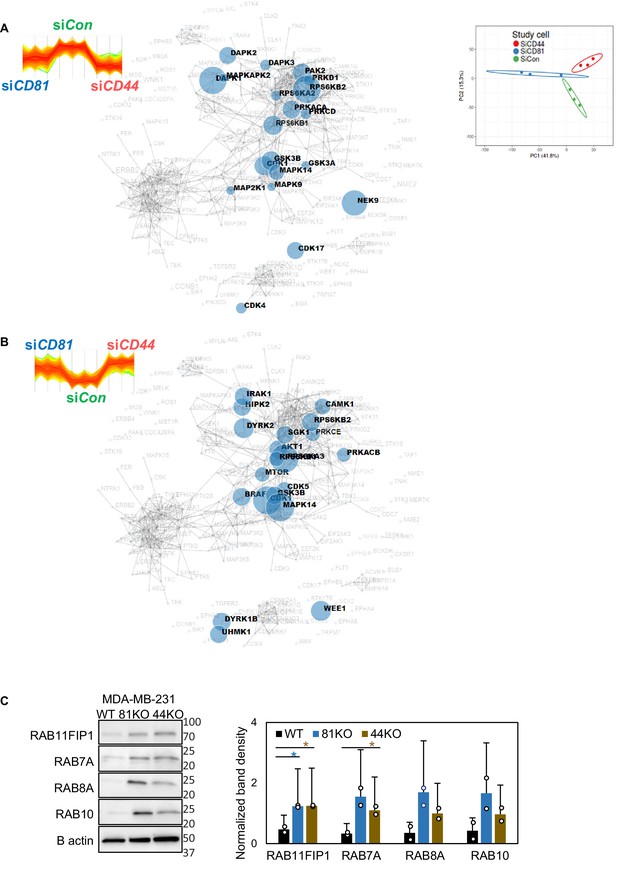
The kinase reactome networks in CD81- and CD44-depleted cells.
The MaxQuant kinase reactome networks based on the downregulated (A) and upregulated (B) phosphorylation sites in both siCD81- and siCD44-transfected cells and known curation of kinase–substrate interactions in the literature. (C) Immunoblot validation of mass spectrometry analysis targets including Rab11FIP1, Rab7a, Rab8a, and Rab10. N= 3. Error bars represent standard deviation. *p = <0.05.
-
Figure 2—figure supplement 3—source data 1
Uncropped blots associated with Figure 2—figure supplement 3C.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig2-figsupp3-data1-v2.pdf
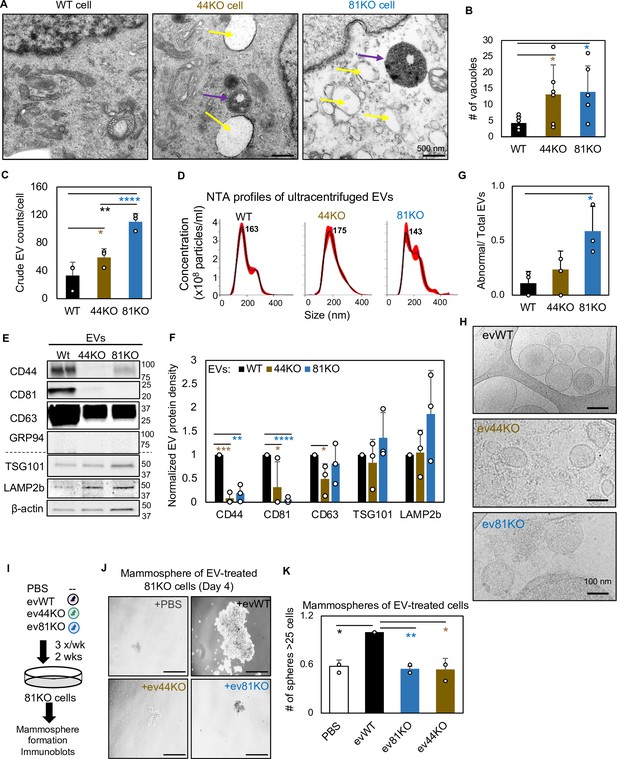
CD81 and CD44 are required for exosome-induced cancer stemness.
Transmission electron microscopy images (A) of WT, 44KO, and 81KO MDA-MB-231 cells and number of vacuoles observed per cell (B). Yellow arrows point to vacuoles or early endosomes/endocytic vesicles, purple arrows to multivesicular bodies. Scale bar = 500 nm. N= 6. Error bars represent standard deviation. Student two-tailed T-test *p = 0.017. (C) Counts of extracellular vesicles (EVs) per cell in crude culture supernatants of WT, 44KO, and 81KO cells, measured by Apogee (N = 3 or 5). Error bars represent standard deviation. Two-tailed Student T-test ****p = 0.0001, **p = 0.003, one-tailed Student T-test *p = 0.036. Nanoparticle tracking analysis (NTA)-based size distributions (repeated three times) (D) and (E, F) representative immunoblots (n = 3) and quantification for EV proteins in ultracentrifuge-isolated EV particles from the culture media of WT, 44KO, and 81KO MDA-MB-231 cells. N= 3. Error bars represent standard deviation. T-test *p = 0.04/0.02 (one tailed), **p = 0.001 (two tailed), ***p = 0.0002 (two tailed), ****p = 8.5e−6 (two tailed). Cryo-EM images (repeated twice) (G) and quantification (H) of membrane integrity in evWT, ev44KO, and ev81KO, taken at ×8000 nominal magnification (a pixel size of 4.125 Å). Vesicles assessed included 18 WT, 17 CD44KO, and 63 CD81KO. Scale bar = 200 nm. (I) Schematic of 81KO MDA-MB-231 cells educated with phosphate-buffered saline (PBS) or evWT, ev44KO, and ev81KO every 2 days for 2 weeks and seeded at low density to evaluate mammosphere formation. Representative images (J) and bar graph quantification (K) of mammospheres 4 days after seeding of 1000 cells per well (24-well plate) after education with PBS, WT EVs, CD44KO EVs, and CD81KO EVs. Scale bar = 100 µm. N= 4. Error bars represent standard deviation. One-tailed Student T-test *p = 0.02, **p = 0.01. Repeated twice times.
-
Figure 3—source data 1
Uncropped blots associated with Figure 3E.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig3-data1-v2.pdf
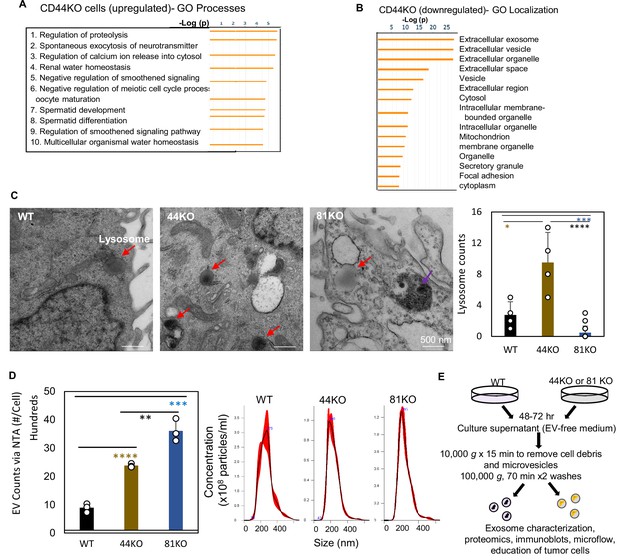
Effects of CD44/CD81 depletion on cellular pathways related to extracellular vesicles (EVs).
(A) GO Localization analysis of downregulated proteins in pooled CD44KO cells compared to WT MDA-MB-231 cells (N = 3 replicates, p < 0.05). (B) GO Processes analyses of global mass spectrometry-based upregulated proteins in the CD44KO cells in comparison to CD44 WT MDA-MB-231 cells. (C) Transmission electron microscopy image and quantification of lysosomes (red arrows) in MB-MDA-231 cells. Scale bar = 500 nm. N= 4. Error bars represent standard deviation. Student two-tailed T-test *p = 0.02, ***p = 5e−04, ****p = 2e−09. The purple arrow points to a multivesicular body (MVB) which is not significant different among three types of cells (1 or 0 observed per cell). (D) Left panel: nanoparticle tracking analysis (NTA) counts of EVs per cell in purified EVs (100,000 × g x 70 min) from WT, 44KO, and 81KO cells, measured by NanoSight (N = 3). Error bars represent standard deviation. Two-tailed Student T-test ****p = 0.0001, ***p = 0.0003, **p = 0.006. Right panel: NTA-based size distributions (repeated three times) of the three types of EVs. (E) Schematic of EV isolation by ultracentrifugation steps and characterization.
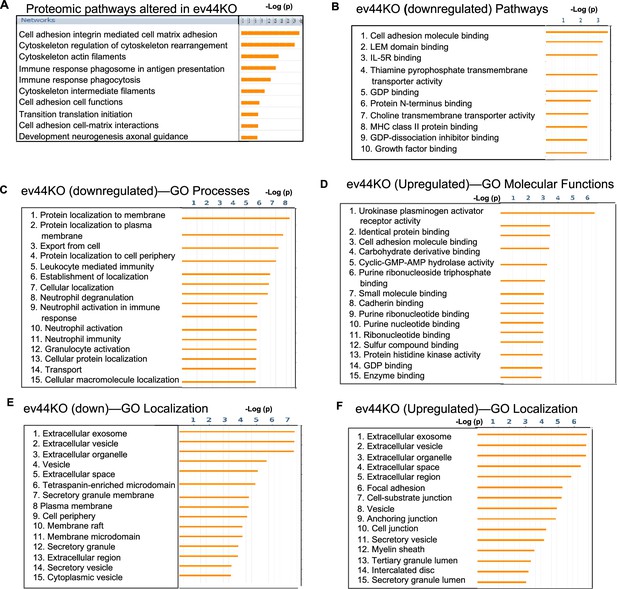
Proteomic pathway analysis of evCD44 derived from CD44KO cells.
(A) Proteomic pathways altered in the extracellular vesicles (EVs) derived from CD44KO cells (ev44KO) compared to evWT from WT cells. (B–F) GO analyses of proteins down- or upregulated in the ev44KO versus evWT.
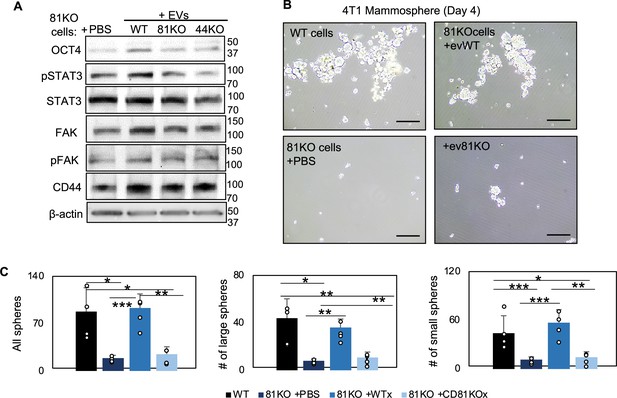
CD81 is required for exosome-induced effects on stemness phenotype.
(A) Immunoblot analyses for OCT 4, STAT3, phosphoSTAT3 (pSTAT3), FAK, pFAK, CD44, and β-actin using CD81KO MDA-MB-231 cells educated with phosphate-buffered saline (PBS) or extracellular vesicles (EVs) derived from WT, CD81KO, and CD44KO, and CD81KO cells. (B, C) Representative images (B) and quantifications of mammospheres derived from 4T1 cells in suspension, including WT cells and Cd81KO cells, the latter of which were educated with PBS or exosomes (10 µg for 1 week). 2000 cells were seeded in 6-cm plates in mammosphere media, and the images were captured on day 4. N= 4. Error bars represent standard deviation. Two-tailed Student T-test was used *p<0.05, **p<0.005, ***p<0.001.
-
Figure 3—figure supplement 3—source data 1
Uncropped blots associated with Figure 3—figure supplement 3.
- https://cdn.elifesciences.org/articles/82669/elife-82669-fig3-figsupp3-data1-v2.pdf
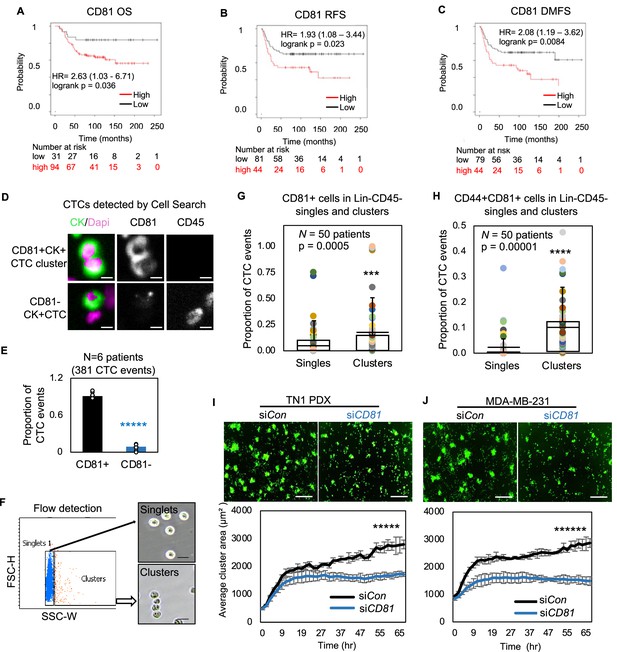
CD81 is associated with patient survival and enriched in circulating tumor cells (CTCs) promoting tumor cell aggregation.
Kaplan–Meier plots of CD81 protein expression in patients with triple negative breast cancer (TNBC) correlate with an unfavorable overall survival (A), relapse-free survival (B), and distant-metastasis-free survival (C). Representative images (D) and quantified % (E) of CD81+ and CD81− CTC events in the blood of six patients with metastatic breast cancer, analyzed on CellSearch. Scale bar = 5 µm. N= 3. Error bars represent standard deviation. Two-tailed Student T-test *****p = 9E−11. Representative images of flow cytometry gated singlets and clusters (F, scale bar = 25 µm) and bar graphs of proportion of CD81+ (G) and CD44+CD81+ (H) in putative Lin-CD45- CTC events (622,509) in the blood (N = 50 patients). N= 50. Error bars represent standard deviation. Two-tailed Student T-test ***p = 0.005, ****p = 0.00001. IncuCyte images (top panels) and quantified tumor cell aggregation curves of TN1 patient-derived xenograft (PDX) (I), MDA-MB-231 (J) cells upon CD81 KD (repeated at least three times). Scale bar = 300 µm. N= 5. Error bars represent standard deviation. Two-tailed Student T-test *****p = 1E−12, ******p = 8E−19.
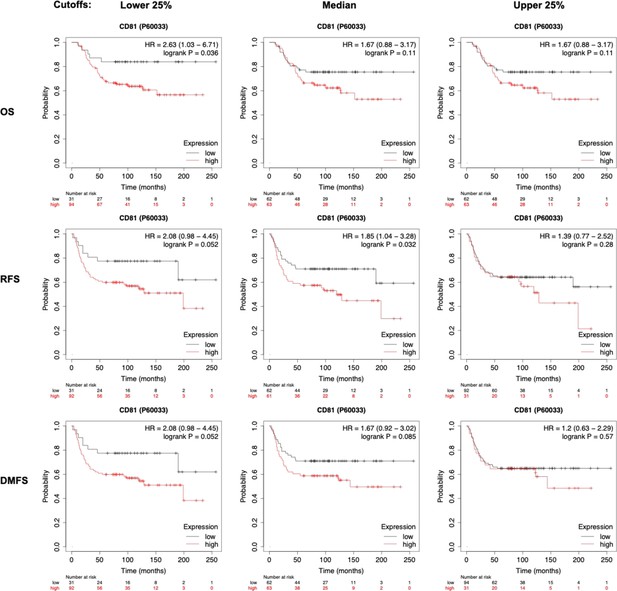
Kaplan–Meier plots of patient survival based on CD81 protein expression.
Kaplan–Meier plots of patient outcomes with triple negative breast cancer (TNBC), stratified by CD81 expression levels that correlate with an unfavorable overall survival (OS), relapse-free survival (RFS), and distant-metastasis-free survival (DMFS). Various cutoffs were made at lower 25%, median, or upper 25%.
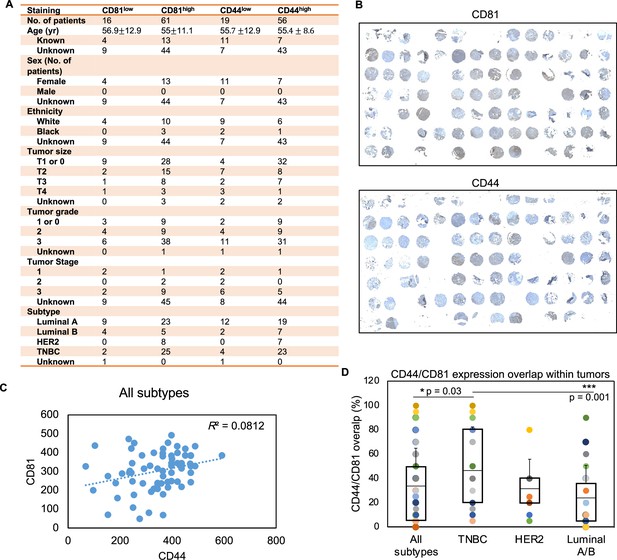
Tissue microarray (TMA) clinical characteristics.
(A) Table of tumor TMA breast cancer patient characteristics. (B) Immunohistochemical staining of CD81 and CD44 expression (in brown) and hemotoxylin (in blue) in TMA of breast tumors (N = 77 with CD81 IHC and N = 75 with CD44 IHC). (C) A scatter plot of CD44 and CD81 expression on tumor microarray. CD81 and CD44 expression levels were quantified using ImageJ and assessed in all subtypes of breast tumors (N = 77, N = 75, patient tumors, respectively). (D) Percentage (%) of tumor regions with overlapped CD81 and CD44 expression within each tumor (N = 75 with both CD81 and CD44 IHC). Error bars represent standard deviation. Two-tailed Student T-test p = 0.03 and 0.001 showing triple negative breast cancer (TNBC) with a higher overlap between CD81/CD44 than the average level of all subtypes as well as that of the Luminal A/B subtype.
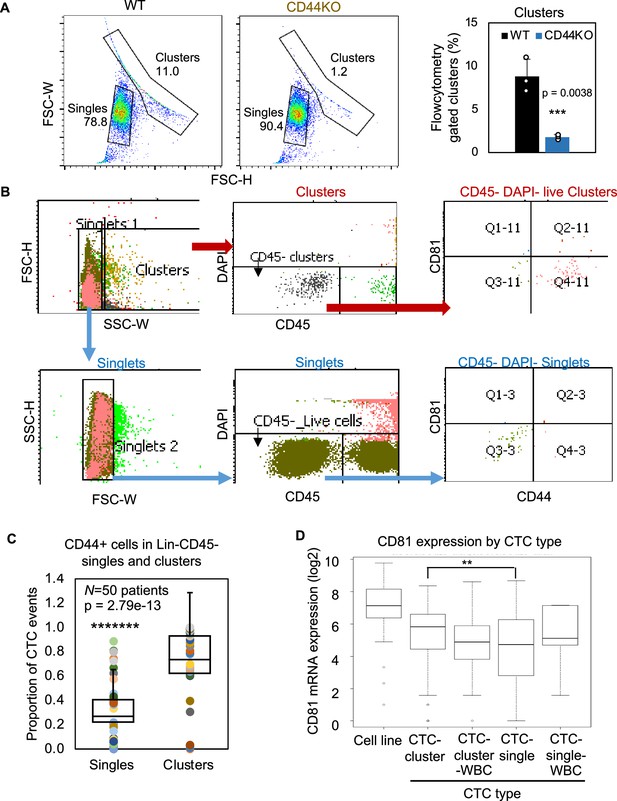
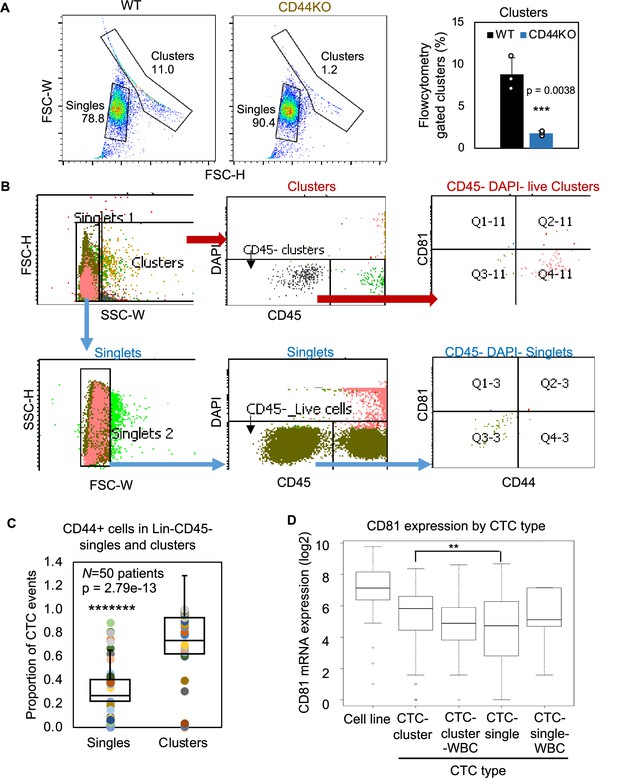
Gating strategy of circulating tumor cells (CTCs) by flow cytometry and the RNA expression level of CD81 in single versus clustered CTCs.
(A) Forward scatter channel (FSC)-gated singles and clusters of WT and CD44KO MDA-MB-231 cells in suspension (n = 3 replicates). N= 3. Error bars represent standard deviation. T-Test p = 0.0038. (B) Gating strategies for patient blood-isolated CD45− single cells and clusters for CD81 and CD44 analyses. (C) Plots of proportion of putative CD44 + CTC events (EpCAM+/− clusters and singles in the blood of 50 patients with metastatic breast cancer, analyzed on flow cytometer). N= 50. Error bars represent standard deviation. Two-tailed Student T-test *******p = 2.79e-13. (D) mRNA expression of CD81 in single/clusters of CTCs from a publicly available dataset (AlQuraishi, 2021 and Fernandez et al., 2014). Error bars represent standard deviation. Wilcoxon **p for CTC-single versus CTC-cluster = 0.0034.
Cluster formation of tumor cells (MDA-MB-231, 4T1, and patient-derived xenograft [PDX] models) in CD81 and CD44 modified cells.
IncuCyte images (right) and quantified tumor cell aggregation curves (left) of MDA-MB-231 WT/CD81KO cells (*p = 0.02) (A), 4T1 WT/Cd81KO cells (******p < 5.8E−08) (B), and TN2 PDX siCon/siCD81 (****p = 0.0004) (C). Scale bar = 300 µm. N= 5. Error bars represent standard deviation. Two-tailed Student T-test was used. Repeated at least twice. (D) IncuCyte images and quantified tumor cell aggregation curves of WT/81KO MDA-MB-231 cells upon overexpression with either HACon, HACD81, or HACD81m. Scale bar = 300 µm. (E) Clustering analysis of MDA-MB-231 WT/CD44KO/CD81KO cells treated with 10 µg/µl IgG or anti-CD81 activating antibody. N= 5. Error bars represent standard deviation. Two-tailed Student T-test was used. p value comparing WT aCD81 to WT IgG *p = 0.022, to CD44KO IgG *p = 0.013, to CD44KO aCD81 *p = 0.011, to CD81KO IgG **p = 0.008, and to CD81KO aCD81 **p = 0.006.
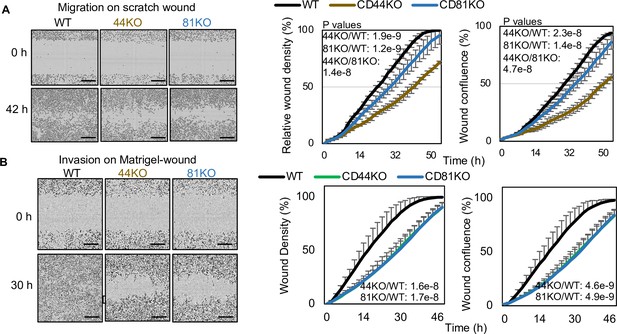
CD81 KD inhibits breast cancer cell migration and invasion.
(A) Cell migration of MDA-MB-231 WT, CD44KO, and CD81KO cells to close scratch wound, analyzed by Incucyte time-lapse imaging in every 2 hr. CD44KO and CD81KO groups took longer time to fill the wound gap. N= 3. Error bars represent standard deviation. Two-tailed Student T-test was used. Repeated twice. (B) Cell invasion of MDA-MB-231 WT, CD44KO, and CD81KO cells on Matrigel-covered scratch wound was evaluated by Incucyte time-lapse imaging software which tracks wound closure every 2 hr. After 30 hr the control cell wounds began to close while the CD44KO and CD81KO groups took longer. N= 3. Error bars represent standard deviation. Two-tailed Student T-test was used.
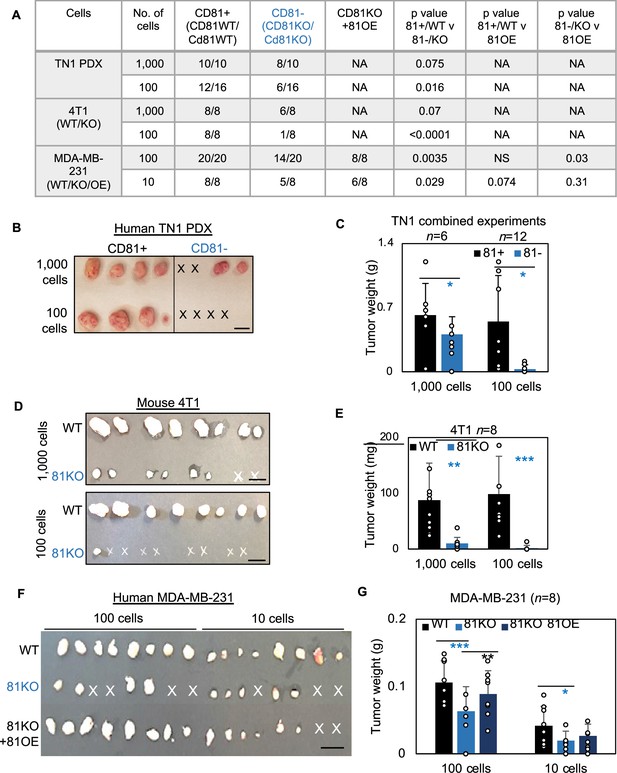
CD81 promotes tumorigenesis of triple negative breast cancer (TNBC) cells.
(A) Table of serial dilutions of tumorigenic results with CD81+ and CD81−TN1 PDX, CD81 WT/KO 4T1 cells, and CD81 WT/KO/KO + CD81 overexpression (OE) MDA-MB-231. One-tailed Student T-test. Pictures of harvested tumors (B) and graphs of tumor weight comparisons (C) with CD81+ and CD81−TN1 PDX tumor implants. Scale bar = 1.3 cm. n = 10–16 injections per group. N= 6-12. Error bars represent standard deviation. Two-tailed Student T-test comparisons between CD81+ and CD81− cells: *p = 0.026 (1000 cells), 0.019 (100 cells). Pictures of harvested tumors (D) graphs of tumor weight comparisons (E). Scale bar = 1 cm. n = 8 injections (2 injections/mouse). N= 8. Error bars represent standard deviation. Two-tailed Student T-test **p = 0.002, ***p = 0.001. Pictures of harvested tumors (F) and graphs of tumor weight comparisons (G) with CD81 WT/KO/KO + CD81 OE MDA-MB-231 cells. Scale bar = 1 cm. n = 8 injections (4 injections/mouse). N= 8. Error bars represent standard deviation. Two-tailed Student T-test ***p = 0.0007, **p = 0.006, *p = 0.026.
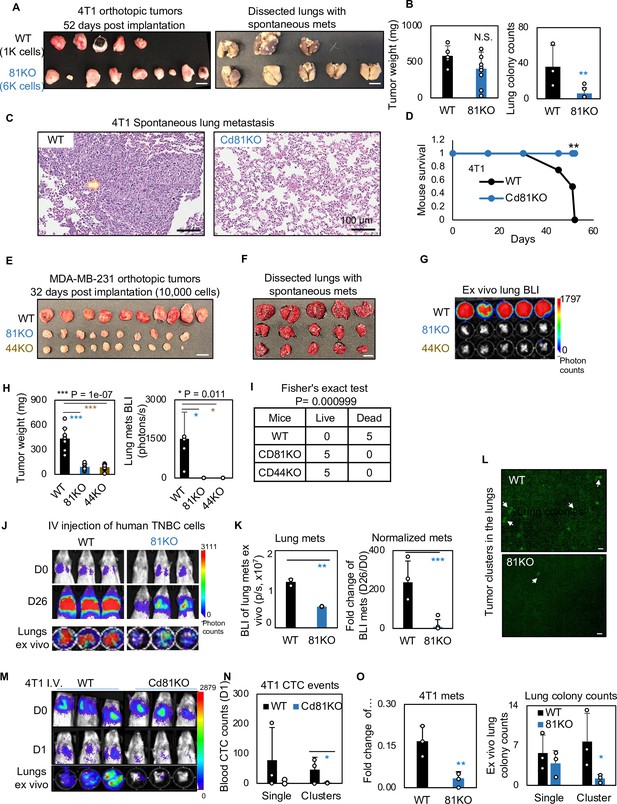
CD81 deficiency abrogates lung metastasis in breast cancer.
(A) Photos of 4T1 orthotopic tumors (left panels) grown from implantations of comparable (1000) WT and (6000) Cd81KO cells into the L4/R4 mammary fat pads (N = 5 Balb/c mice with 10 injections) and the fixed lungs with overt metastatic colonies. By the terminal day 52, two mice from the WT group died and the left three were sick and sacrificed. Scale bar = 1 cm. (B) Bar graph of the tumor weights and lung colonies count between the WT tumors and KO tumors. (C) IHC (HE) images of lung colonies from the WT group mice (a higher number of visible lung metastases at a larger size) as compared to the Cd81 KO group. Scale bar = 100 µm. N.S. = not significant, two-tailed Student T-test **p = 0.01 (n = 5 mice). Error bars represent standard deviation. . (D) Distinct mouse survival between 4T1 WT and Cd81 KO tumor bearing mice with spontaneous lung metastases. Two-tailed Student T-test **p = 0.01 (n = 5 mice). Photos of WT and 81KO MDA-MB-231 orthotopic tumors (E) grown from 10,000 cell implantations and dissected mouse lungs on day 32 (F) (N = 5 NSG mice with 10 injections) Scale bar = 1 cm. Error bars represent standard deviation. (G) BLI images and of spontaneous metastases in the lungs ex vivo following after orthotopic implantation of WT and 81KO MDA-MB-231 cells into NSG mice. Error bars represent standard deviation. *p = 0.011 (n = 5 mice). (H) Quantification of tumor weights, lung metastases, and relative metastatic burden normalized by tumor weight. Error bars represent standard deviation. Two-tailed Student T-test was used. ***p = 1e−07, *p = 0.011. (I) Table of mouse mortality or survival by day 32 after orthotopic implantation. Three mice from the WT group died and the left two were sick and sacrificed whereas the 81KO tumor-bearing mice would have survived. Fisher’s test was used **p = 0.01 (n = 5). (J) BLI images of lung colonization following the tail vein injections of MDA-MB-231 WT and 81KO cells into NSG mice on days 0 and 26. The bottom row shows dissected lungs ex vivo. (K) Quantified BLI signals (left panel) and normalized metastasis intensity (relative to the day 0 signals) of MDA-MB-231 cells in the dissected lungs ex vivo on day 26 after tail vein injections. Error bars represent standard deviation. Two-tailed Student T-test **p < 0.007. ***p = 0.001 (N = 3 mice). (L) Representative fluorescence images (top two panels) of mouse lungs and quantified metastatic colonies (singles and clustered, bottom panel) of WT and CD81KO L2G+ MDA-MB-231 cells (D26). Scale bar = 100 mm. Error bars represent standard deviation. Two-tailed Student T-test was used. *p = 0.044. BLI images of mice (days 0 and 1) and dissected lungs on day 1 (M) lood circulating tumor cell (CTC) counts (L2G+ singles and clusters) measured via flow cytometry on day 1 (N), and relative lung metastasis via BLI on day 1 (O, relative to day 0) following the tail vein injections of 4T1-WT and Cd81KO cells into Babl/c mice (N = 3), *p = 0.047, **p = 0.007. Error bars represent standard deviation. Two-tailed Student T-test was used. These experiments were repeated at least twice to show consistent conclusions.
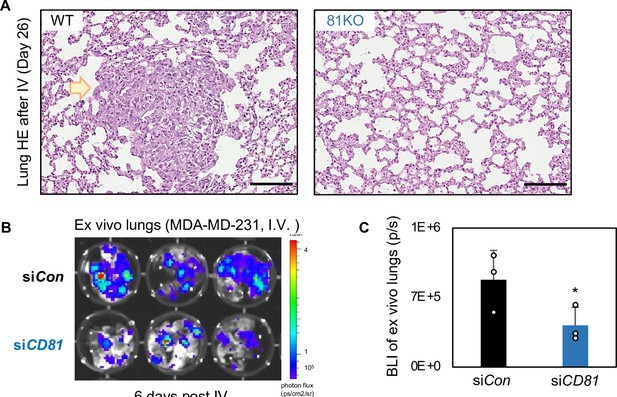
CD81 KD inhibits breast cancer cell metastasis.
(A) HE images of lung metastasis colonies (arrow pointed region) observed in the mice injected with WT cells and absent with 81KO MDA-MB-231 cells, harvested on day 26 after tail vein injection. Scale bar = 200 µm. (B, C) CD81 control and CD81KD MDA-MB-231 cells were evaluated for their metastatic potential. 100 K cells were injected into mice by tail vein injection to observe lung colonization. Six days post tail vein injection lung colonization was observed lower levels in the CD81KD group compared to the control (n = 3) (error bars represent standard deviation,one-tailed Student T-test *p = 0.04).
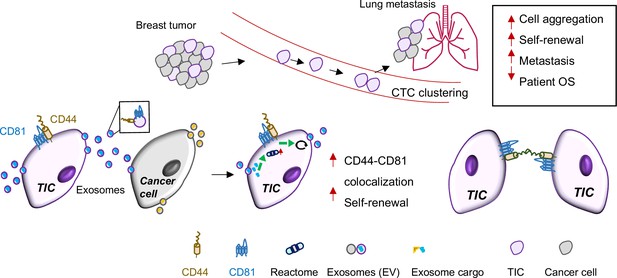
Schematic summary.
Schematic summary of CD81 in interacting with CD44 on the cytoplasmic membrane of tumor-initiating cells (TIC), facilitating the exosome cargo packaging with CD44 and CD81, promoting exosome-induced self-renewal in recipient cells via phosphoreactome pathways, and strengthening CD44-mediated circulating tumor cell (CTC) cluster formation and lung metastasis.
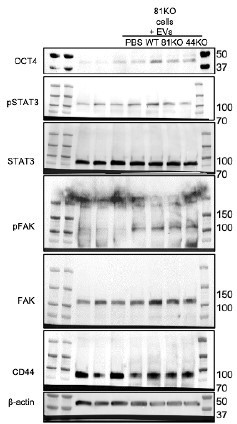
Full blots of Suppl.
Figure 8A. Full Immunoblot analyses for OCT4, STAT3, phosphoSTAT3 (pSTAT3), FAK, pFAK, CD44 and β Actin using CD81KO MDA-MB-231 cells educated with PBS or EVs derived from WT, CD81KO, and CD44KO, and CD81KO cells.
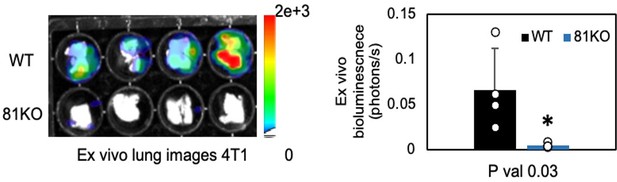
Repeated Figure 6M.
BLI images of dissected lungs on day 1 following the tail vein injections of 4T1-WT and Cd81KO cells into Babl/c mice (n=4) * P = 0.03. Two-tailed student T-test was used.

CD81 is required for EV-induced effects on stemness signature proteins and mammosphere formation.
Representative images (left two panels) and quantifications (right panel) of mammospheres (Day 5) derived from 500 MDA-MB-231 cells in suspension after transfections with scrambled (Scr) RNAs and human siRab27a.
Tables
| EV-producing cell | EV counts/mL | |
|---|---|---|
| NTA | MFV | |
| WT | 1.24e+09 +/- 9.44e+07 | 0.96e+09 +/- 2e+08 |
| CD44KO | 1.57e+09 +/- 2.80e+07 | 1.56e+09 +/- 2e+08 |
| CD81KO | 1.62e+09 +/- 1.68e+07 | 1.75e+09 +/- 9e+07 |
Additional files
-
Supplementary file 1
Overlapping list of 38 proteins in CD44 and siCD44.
- https://cdn.elifesciences.org/articles/82669/elife-82669-supp1-v2.xlsx
-
Supplementary file 2
Down or up-regulated genes without cutoffs.
- https://cdn.elifesciences.org/articles/82669/elife-82669-supp2-v2.xlsx
-
Supplementary file 3
RNAseq data: down- and up-regulated pathways with 1.5 fold change.
- https://cdn.elifesciences.org/articles/82669/elife-82669-supp3-v2.xlsx
-
Supplementary file 4
Global and phosphoproteomics data in cluster versus adherent cells.
- https://cdn.elifesciences.org/articles/82669/elife-82669-supp4-v2.xlsx
-
Supplementary file 5
KEGG Pathways of global proteome clusters (G1 to G6) and phosphoproteome cluster (P1 to P6).
- https://cdn.elifesciences.org/articles/82669/elife-82669-supp5-v2.xlsx
-
Supplementary file 6
Heatmap for siCD81 and siCD44 global proteomics in clusters.
- https://cdn.elifesciences.org/articles/82669/elife-82669-supp6-v2.xlsx
-
Supplementary file 7
Phosphoproteomics data for heatmap in siCD81 and siCD44.
- https://cdn.elifesciences.org/articles/82669/elife-82669-supp7-v2.xlsx
-
Supplementary file 8
Differentially Expressed Proteins in MDA-MB-231 WT versus CD44KO cell by mass spectrometry.
- https://cdn.elifesciences.org/articles/82669/elife-82669-supp8-v2.xlsx
-
Supplementary file 9
Altered proteins in EVs of MDA-MB-231 WT versus CD44KO by mass spectrometry.
- https://cdn.elifesciences.org/articles/82669/elife-82669-supp9-v2.xlsx





