Competition between lysogenic and sensitive bacteria is determined by the fitness costs of the different emerging phage-resistance strategies
Figures
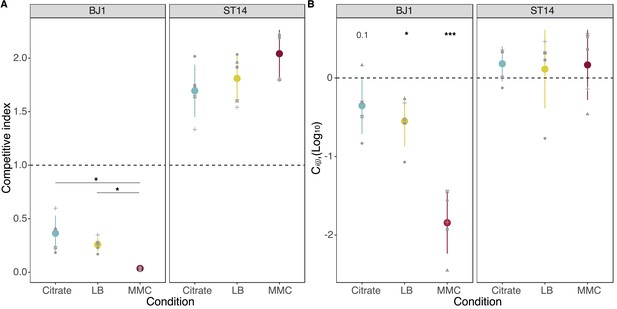
Fitness of strains during competition.
(A) The competitive index is calculated as the final frequency of each strain divided by the initial frequency in the mixed cocultures. * P<0.05, Wilcoxon rank sum test adjusted by Benjamini-Hochberg correction. (B) The effect of mixing two strains during growth in coculture is given as Ci(j), expressed in log10, with i representing either strain BJ1 or strain ST14. Positive values represent increased cell numbers during coculture than those expected from the pure cultures. p-values correspond to a one-sample t-test for a difference of 0. *p<0.05, ***p<0.001. Each dot shape represents an independent biological replicate, N=5. Error bars indicate the standard deviation.
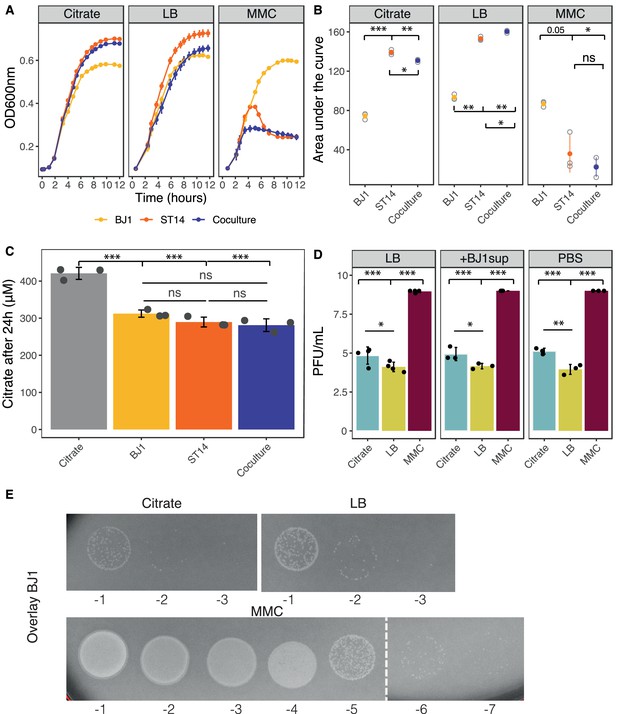
Growth and phage production in experimental conditions.
(A) Growth of each strain and their coculture in each experimental condition. Three independent clones and cocultures were measured. (B) Area under the growth curves, calculated with trapz function from pracma package for R. (C) Citrate amount in 24 hr cultures of strain BJ1, ST14, 1:1 coculture, and control cultures of plain LB with citrate. Each dot represents an independent experiment. Error bars indicate standard deviation. (D) Strain ST14 was grown in LB, in LB diluted (1:1) in spent supernatant from BJ1, or in LB diluted (1:1) in PBS. Phage lysates were prepared and PFU analyzed on a lawn of strain BJ1. Each dot represents an independent biological replicate. Error bars indicate standard deviation. (E) Lysis plaques on an overlay of strain BJ1 after the addition of 10 µl of supernatant of strain ST14 grown in the different experimental conditions. *p<0.05,**p<0.01,***p<0.001 for ANOVA with Tukey post hoc corrections.
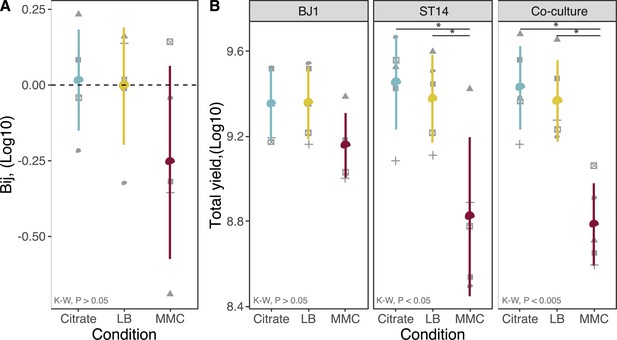
Total growth in the different media.
(A) The effect of mixing on total growth or population yield is expressed as Bi(j), in which positive values represent more growth during coculture than expected from the pure cultures. Each shape represents an independent experiment, N=5. Error bars indicate standard deviation. (B) The total yield of each culture is expressed as the log10-transformed of CFU/mL. Each shape represents an independent biological replicate, N=5. p-values correspond to Kruskal-Wallis(K-W) rank sum test, followed by a pairwise Wilcoxon test for differences across conditions, with Benjamini-Hochberg correction. *p<0.05.

Population yield and proportion of capsulated clones of the two strains during the coevolution experiment.
(A) Total CFU per mL of each strain was estimated every day on selective media. Each line represents an independent coevolving population. (B) Emergence of non-capsulated mutants in each strain. The proportion of capsulated clones in the population is depicted. The insert shows the area under the curve (AUC) during the first nine days of evolution, as calculated by the function trapz from the R package pracma. *p<0.05,**p<0.01,***p<0.001 for ANOVA with Tukey post hoc corrections.
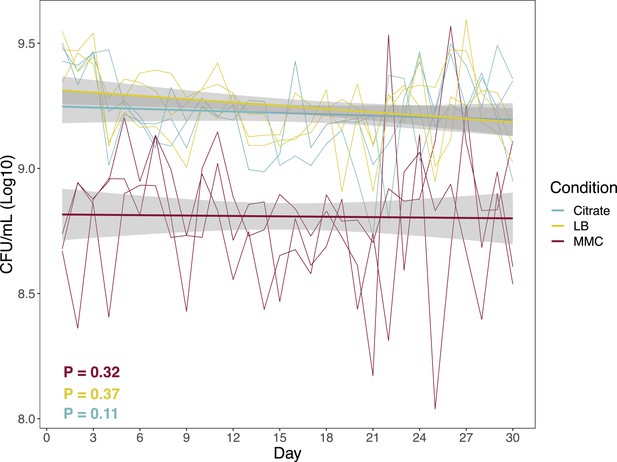
Total number of cells across evolving populations.
Each independent line represents an independently evolving population. Bold lines represent regression lines from a linear model. The slopes of the regression model were calculated for each independent population, and tested with a one-sample t-test for a significant difference from 0.
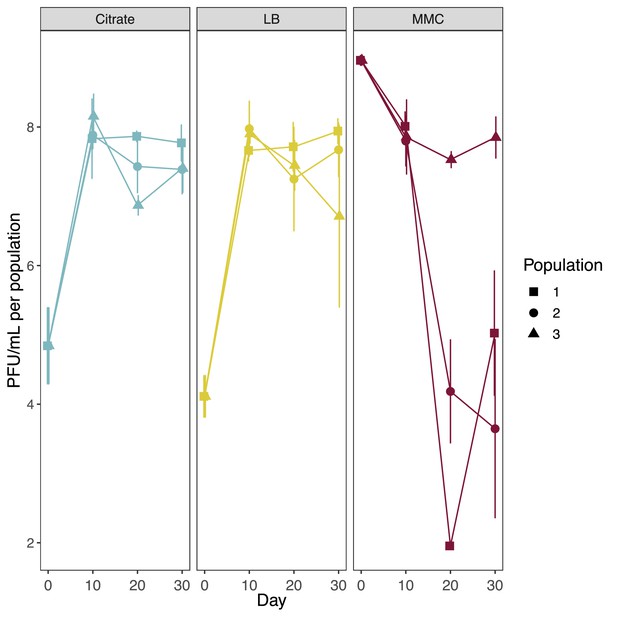
Estimated phage titers in each population throughout the coevolution experiment.
Phages were estimated by plaque counting on ancestral phage susceptible BJ1. Total phage production was inferred for each population by quantifying the efficiency of plaquing (EOP) of one capsulated and one non-capsulated randomly selected ST14 clone, relative to their presence in the population. Different shapes represent independently evolving populations.
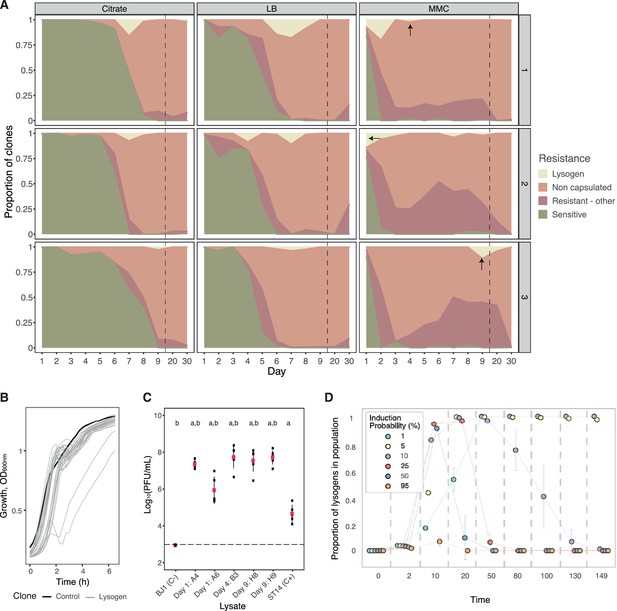
Evolution of resistance mechanisms in strain BJ1.
(A) Ratio of clones from each coevolving population that are susceptible (green), non-capsulated (light pink), capsulated lysogens (beige), or capsulated but resistant by other undefined mechanisms (dark pink). N.B. Dashed line indicates when the x-axis, no longer follows a linear scale. Dark arrows indicate the time points from which the lysogens tested in panel C were retrieved. (B) Growth of newly lysogenized clones reveals significant death during the exponential phase (in the absence of induction), as measured by the optical density. Black line corresponds to the control, BJ1 ancestor. (C) PFU/mL produced without induction by five selected new lysogens derived from BJ1 and isolated at day one for A4 and A6 (Population #2, MMC), at day four for B3 (Population #3, MMC), and day nine for H8 and H9 (Population #3, MMC). Dashed line indicates the limit of detection of the essay. Each black dot represents an independent biological replicate (independent strain lysate) and large red dots represent the mean. Error bars correspond to the standard deviation. Two-sided t-test ‘a’, p<0.001 compared to ancestor BJ1 (negative control, C-) and ‘b’, p<0.05 compared to ST14 (phage producer, positive control, C+). (D) Simulated temporal dynamics of the proportion of lysogens in the populations, as calculated by eVIVALDI. Each circle corresponds to the central tendency of replicate simulations, with the different colors indicating a given probability of spontaneous prophage induction (shown in the legend, values approximated to the nearest major integer). The error bars correspond to the standard deviation across the replicate simulations. In the represented simulations, the probability of acquisition of a phage resistance mutation (capsule loss) is 0.001, and the fitness cost of this mutation is 10% of the bacterial growth rate, as calculated in Buffet et al., 2021.
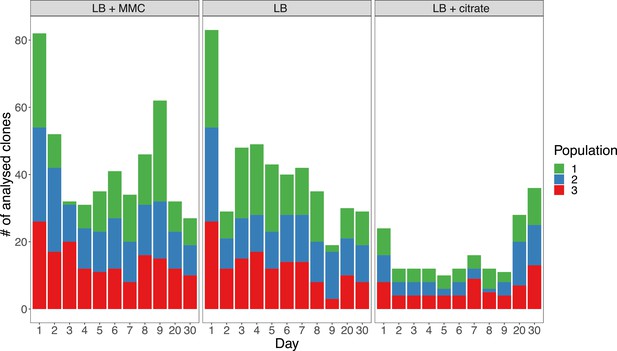
Number of clones from strain BJ1 analyzed each day from each population.
Each day a total of 93 capsulated clones from every population in each evolutionary treatment were isolated, except for day one, when 186 capsulated clones (93 × 2) were isolated. The number of 93 was chosen for it to conveniently fit in a 96-well plate and allow three wells for internal controls (ancestors and blank).
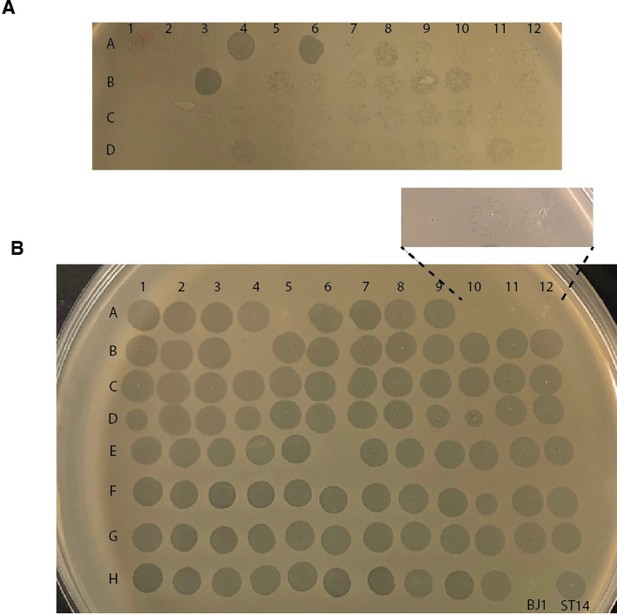
Lysogens can release viable phages that infect the ancestral BJ1 strain.
93 lysogens were grown overnight in 100 µL of LB in a 96-well microtiter plate. Cultures were then diluted 1:100, and allowed to grow for 6 hr (A, non-induced lysogens), or 2 hr, after which, MMC was added to a final concentration of 5 µg/mL (B, induced lysogens). Cultures were allowed to grow for 4 hr. After a total of 6 hr of everyday growth, the microtiter plate was centrifuged to allow cells to pellet, and 4 µL of the supernatant was spotted on an overlay of 0.7% Lennox agar with the ancestral BJ1 strain. Two controls are included; ancestral BJ1 (well H11) and ancestral ST14 (well H12).
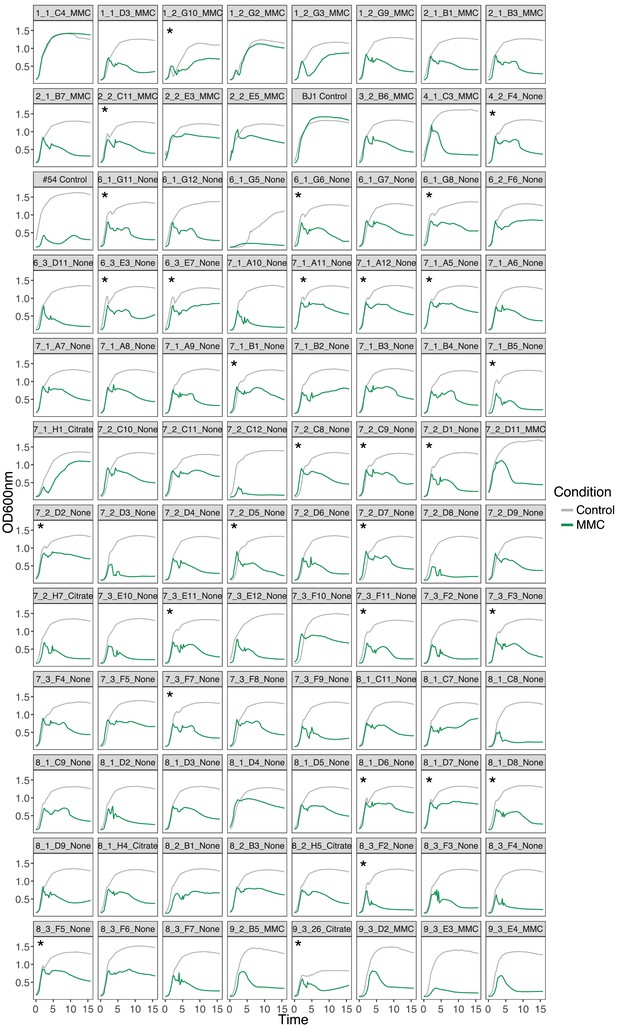
Growth of newly lysogenized clones.
Clones were grown in either LB (control, gray line) for 16 hr or in LB to which MMC (1 µg/ml) was added after 1.5 hr (green). Labels on top of each graph correspond to the isolation day, population number, clone ID, and condition in which it evolved. Star (*) indicates clones in which death is observed at the end of the exponential phase, suggestive of an unusually large phage outburst. Only one replicate per curve is shown for clarity purposes.
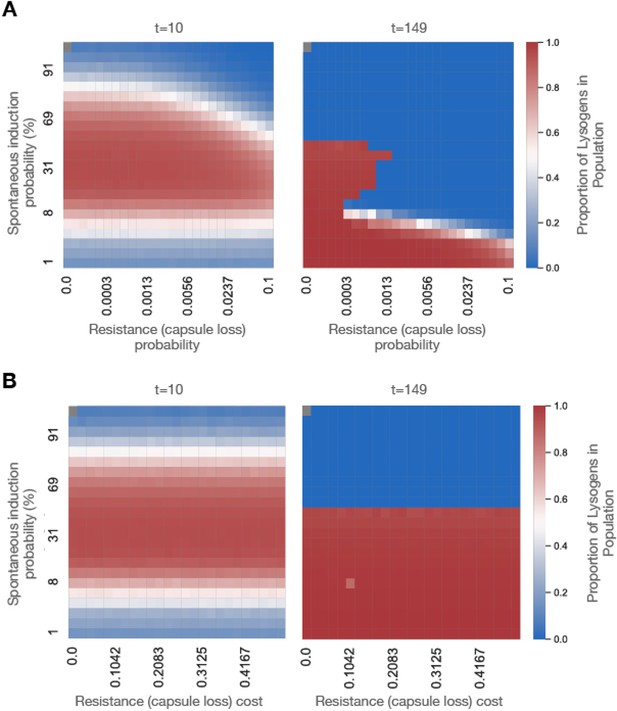
Simulated temporal dynamics of the competition between different phage resistance mechanisms.
eVIVALDI was used to simulate a scenario where an initial population of phage-sensitive bacteria (n=#) is co-inoculated with # temperate phages. Upon infection, bacteria either die from the phage’s lytic cycle or become lysogens if the phage follows the lysogenic cycle. Integrated phage (prophages) can subsequently spontaneously induce and restart the lytic cycle, leading to bacterial death. Bacteria can also become phage-resistant (before or after becoming lysogens) by a random mutation that represents the loss of the capsule. (A) Heatmap of the proportion of lysogens in the populations at intermediate (t=10) and final (t=149) timepoints, as a function of the spontaneous induction probability (y-axis) and the probability of phage-resistant (capsule-less) mutants to emerge. The fitness cost of the resistant mutants is fixed at 10%. (B) Heatmap of the proportion of lysogens in the populations at intermediate (t=10) and final (t=149) timepoints, as a function of the spontaneous induction probability (y-axis) and the fitness cost of a phage-resistant mutation (loss of capsule). The probability of these mutations to emerge is fixed at 0.001. For B, C, and D, dark red colors indicate a prevalence of lysogens in the population, while deep blue colors indicate their absence. Each square in the heatmaps represents the median of the 20 independent replicate simulations for each combination of parameters.

Growth of BJ1-resistant clones.
All isolated clones that were capsulated and tested for resistance (Figure 3A and Figure 3—figure supplement 1) were grown in the absence of phage (control) or with phage. One or two non-capsulated clones per day were randomly selected and included in the analyses. The area under the growth curve of each clone was estimated, and compared to that of the BJ1 ancestor (dashed line). K-W, Kruskal-Wallis test. Post hoc tests for significant differences across groups were calculated and p-values adjusted for multiple testing with Bonferroni correction. a, p<0.05 difference from non-capsulated; b, p<0.05 difference from resistant by other mechanisms; c, p<0.05 difference from lysogens; d, p<0.05 difference from sensitive strains.

Characteristics of phage-resistant clones.
(A) Schematic organization of the capsule operon of strain BJ1. Black triangles represent mutations observed in the capsule operon. Blue arrows indicate core genes common to all K. pneumoniae capsule serotypes. Gray arrows correspond to serotype-specific genes. GT stands for glycosyltransferase. The diagram was generated with genoplotR package. (B) For each clone, we evaluated the amount of capsule produced, the ability of the phage to be adsorbed, and phage the sensitivity on overlay by EOP (PFU/mL) and on liquid culture by AUC. The area under the curve (AUC) difference accounts for the surplus of growth of the control compared to growth with phage. If the clone is resistant, the difference in AUC is ~0. The average of three independent replicates is shown. The experiments were performed with three independently generated lysates, when applicable. ND: none detected.
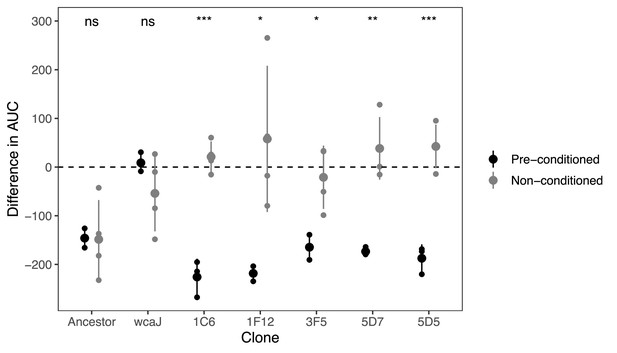
Transient resistance to phages.
The difference in the area under the curve (AUC) represents the effect of adding phage to a growing culture as estimated by the difference in the growth curve in the absence of phage and with phage (Figure 6—figure supplement 1). Values below 0 indicate strains are sensitive to phage, and values close to 0 indicate that there is no effect of adding phage to the culture. Non-conditioned clones are those directly grown from glycerol stock, whereas pre-conditioned clones, had been reisolated twice, and grown overnight prior to performing the growth analyses. The ancestor, BJ1, sensitive, and the non-capsulated mutant (∆wcaJ), are included as controls for the difference in culture conditions. Each small dot represents an independent biological replicate. Statistics represent t-tests to check for differences between clones directly from stock and those sequenced (after two passages in LB). ns means non-significative, *p<0.05, **p<0.01 and ***p<0.001.
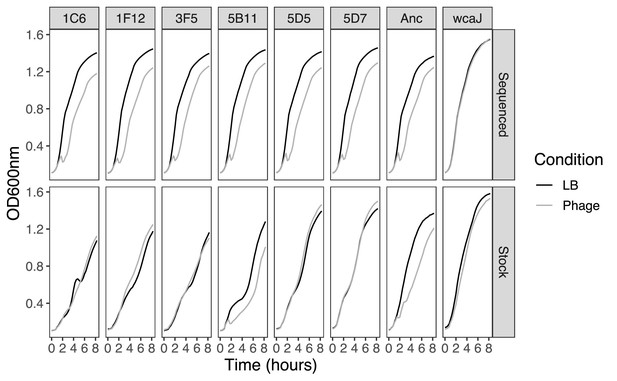
Growth curves of resistant clones with no fixed mutations.
Growth of resistant clones from the original stock was directly initiated from the glycerol stock without performing a preconditioning culture. Sequenced clones underwent two extra rounds of culture since the original stock; restreaked on agar and an overnight LB culture was used to extract the genome and to generate a new glycerol stock, from which all other experiments were initiated. N=4. No error bars are shown for clarity purposes.
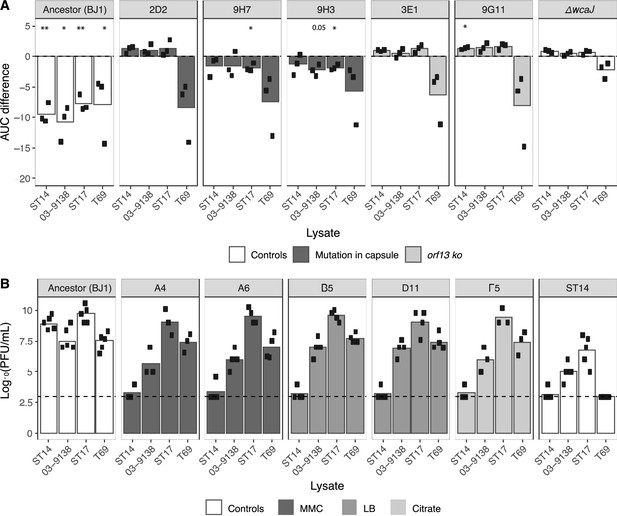
Cross-resistance to phages from other lysates.
(A) The area under the curve (AUC) for capsulated (and non-lysogenized) resistant clones was calculated. The AUC of control cultures with LB was subtracted from those that were challenged with phage lysates. (Growth curves are shown in Figure 7—figure supplement 1A). Each dot represents an independent assay. One-sample t-tests were performed to test the difference from 0 (growth in LB). *p<0.05; **p<0.01 (B) PFU/mL of three independently generated lysates on lawns of new BJ1 lysogens from different evolutionary treatments. Lysogens A4 and A6 evolved in MMC and were isolated on day 1, lysogens B5 and D111 evolved in LB and were isolated on day 6 and 7, respectively, and clone F5 evolved in citrate and was isolated on day 7. Each black dot represents an independent biological replicate. The dashed line represents the limit of detection of the assay.
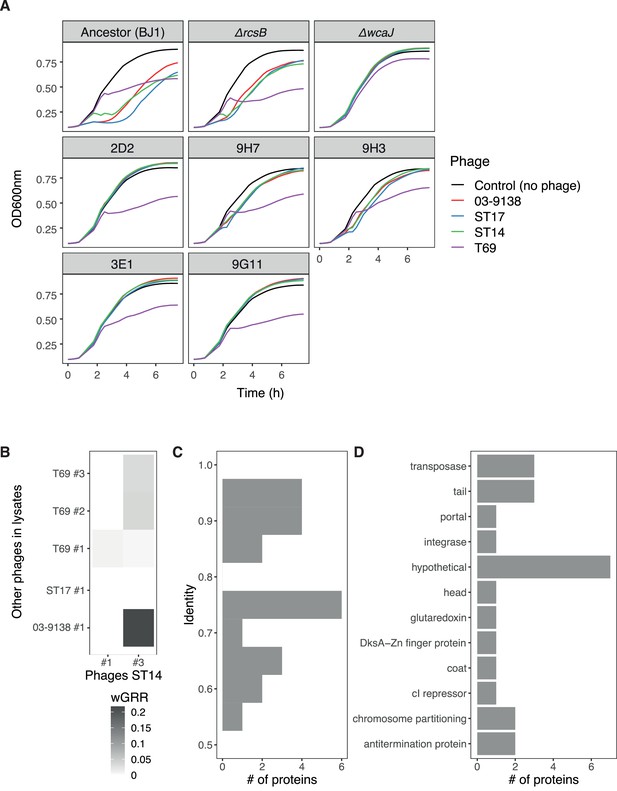
Characterization of lysates produced by other K2 strains.
(A) Growth curves of resistant mutants and control strains in the presence of lysates produced by other K2 strains. Error bars are not shown to improve visibility. (B). Phage similarity as expressed by wGRR of intact phages in lysates. (C). Identity of proteins in intact phages present in other lysates against proteins in phages of strain ST14. (Only values >0.5 as determined by BLASTP are displayed). (D) Predicted function of proteins with an identity >0.5. The functional characterization was performed by annotating the intact phages with prokka v1.14.0 (Seemann, 2014), and pVOG profiles (Grazziotin et al., 2017) searched for using HMMER v3.2 (Eddy, 2011). For each protein, the pVOG with the lowest p-value was kept, and keywords were analyzed.
Tables
List of mutations identified in the resistant clones sequenced.
Location indicates if the mutation is found on the chromosome (C) or plasmid (P). Pop stands for the population. The number of mapped sequences is also reported. Clones with less than 98% of mapped sequences are displayed in italics.
| Clone | Pop | Condition | Day | Location | Position | Change | Mutation type | Annotation | Function | Mapped sequences |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 C6 | 1 | MMC | 1 | 98.3 | ||||||
| 1 F12 | 2 | MMC | 1 | 98.4 | ||||||
| 2D2 | 2 | MMC | 2 | C | 3,994,990 | T→A | I110F (ATT→TTT) | orf13 | K2 capsule gene; Polysialic acid O-acetyltransferase | 98.2 |
| 3E1 | 1 | MMC | 3 | C | 3,995,203 | (T)8→7 | coding (115/669 nt) | orf13 | K2 capsule gene; Polysialic acid O-acetyltransferase | 98.4 |
| 3 F5 | 3 | MMC | 3 | C | 98.6 | |||||
| 5B11 | 1 | LB | 5 | P | 34,553 | A→G | intergenic (+401/–389) | hypothetical protein/ hypothetical protein | 98.2 | |
| P | 34,561 | A→G | intergenic (+409/–381) | hypothetical protein/ hypothetical protein | ||||||
| 5D5 | 2 | LB | 5 | 98.5 | ||||||
| 5D7 | 2 | LB | 5 | 98.6 | ||||||
| 9 G11 | 1 | LB | 9 | C | 3,995,203 | (T)8→9 | coding (115/669 nt) | orf13 | K2 capsule gene; Polysialic acid O-acetyltransferase | 98.3 |
| 9 H7 | 2 | Citrate | 9 | C | 2,890,708 | C→T | R121R (CGG→CGA) | dcuR | Transcriptional regulatory protein DcuR | 98.5 |
| C | 4,009,965 | Δ11 bp | coding (245-255/630 nt) | wzi | K2 regulatory capsule gene | |||||
| 9 H3 | 1 | Citrate | 9 | C | 3,994,389 | C→T | M61I (ATG→ATA) | wcaJ | UDP-glucose: undecaprenyl-phosphate glucose-1-phosphate transferase | 98.5 |
| 9 H12 | 3 | Citrate | 9 | C | 2,575,329 | C→T | D615N (GAC→AAC) | yeaG | Protein kinase YeaG | 98.6 |
| P | 23,432 | G→A | R34Q (CGG→CAG) | soj | Phage cox protein (PF10743), annotated as plasmid-partitioning |
Additional files
-
Supplementary file 1
PHASTER prophage prediction in strain ST14.
The genome was analyzed with PHASTER (Arndt et al., 2016) in April 2021.
- https://cdn.elifesciences.org/articles/83479/elife-83479-supp1-v2.docx
-
Supplementary file 2
Mutations accumulated in wcaJ gene resulting in non-capsulated mutants.
ND; none detected. On the day in which at least 50% of the clones of each independently evolving population were non-capsulated, two of such clones were isolated (# Clone), the wcaJ gene amplified by PCR and sequenced by Sanger. For BJ1, this was repeated twice independently (# Seq) (N=36 for BJ1 and N=18 for ST14). Independently evolving populations (Pop) are identified with a number from 1 to 3. Sequencing of the ancestor revealed that no mutations were present in the wcaJ genes.
- https://cdn.elifesciences.org/articles/83479/elife-83479-supp2-v2.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/83479/elife-83479-mdarchecklist1-v2.docx





