Autophagosome membrane expansion is mediated by the N-terminus and cis-membrane association of human ATG8s
Figures
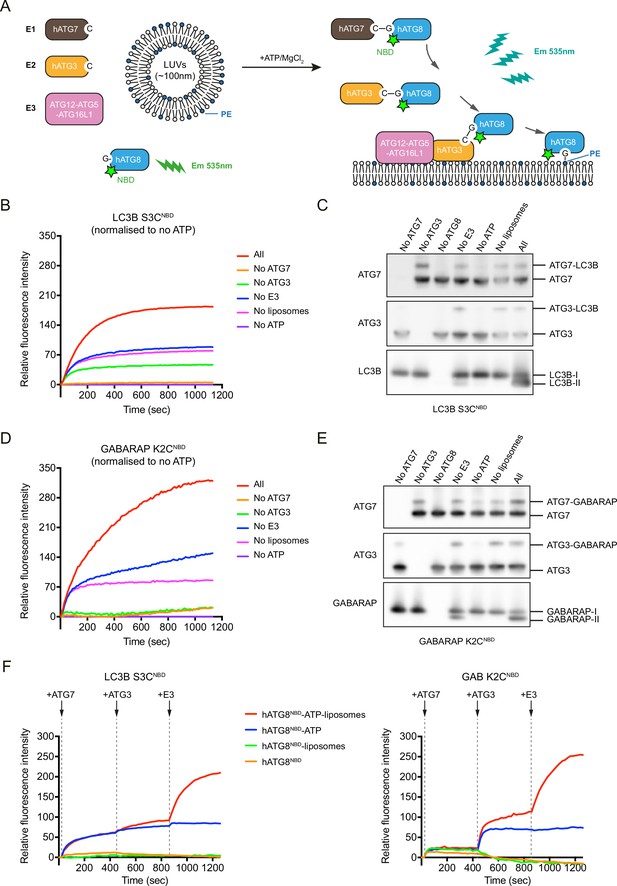
A real-time assay to track LC3B and GABARAP N-termini dynamics during lipidation.
(A) A schematic diagram of the real-time lipidation assay using N-terminal NBD-labelled hATG8. (B) The NBD fluorescence changes of LC3B S3CNBD during lipidation reaction. ‘All’ condition contains 0.2 μM ATG7, 0.2 μM ATG3, 0.05 μM ATG12–ATG5-ATG16L1, 1 mM ATP, 1 mM large unilameller vesicles (LUVs) (50% DOPE/50% POPC) and 1 μM LC3B S3CNBD, at 37°C. The NBD fluorescence was traced immediately once ATP was added (time point at 0 s). The rest conditions were performed in the absence of one component from the ‘All’ condition. The relative NBD fluorescence was normalised to ‘no ATP’ condition. Data represent mean values (n = 3). (C) Western blots of samples from (B). After 20 min reaction, the reactions from the real-time assay were stopped by adding sample buffer and immunoblot for ATG7, ATG3, and LC3B. (D) The NBD fluorescence changes of GABARAP K2CNBD during lipidation reaction. Experimental conditions are the same as described in (B). Data represent mean values (n = 3). (E) Western blots of samples from (D). (F) Step-by-step real-time lipidation assay with LC3B S3CNBD and GABARAP K2CNBD. The NBD fluorescence was recorded continuously while adding ATG7 (between 60–80 s), adding ATG3 (between 480–500 s), and adding E3 complex (between 900–920 s). Data represent mean values (n = 3).
-
Figure 1—source data 1
Uncropped blot and gel images of Figure 1C and E and source data file (Excel) for Figure 1B, D and F.
- https://cdn.elifesciences.org/articles/89185/elife-89185-fig1-data1-v2.zip

The increase of LC3B/GABARAP N-terminal NBD fluorescence reflects the conjugation with ATG7, ATG3 and lipidation of LC3B/GABARAP.
(A) Control experiments for the real-time assay with LC3B S3CNBD and GABARAP K2CNBD in Figure 1. Instead of removing one component from the ‘All’ conditions, the assays were performed with components indicated in the legend. The NBD fluorescence was recorded once ATP was added (time point at 0 s). The relative NBD fluorescence was normalised to that of LC3B S3CNBD and GABARAP K2CNBD protein only, respectively. (B) Western blots of sample from (A). (C) Sequence alignment of six human ATG8 proteins using ESPript 3. The identical and similar residues are indicated in red and yellow, respectively. N-terminus (orange), loop 3 (green), basic patch RRR/RKR (blue), and loop 6 (light blue) are shown.
-
Figure 1—figure supplement 1—source data 1
Uncropped blot images of Figure 1—figure supplement 1B and source data file (Excel) for Figure 1—figure supplement 1A.
- https://cdn.elifesciences.org/articles/89185/elife-89185-fig1-figsupp1-data1-v2.zip
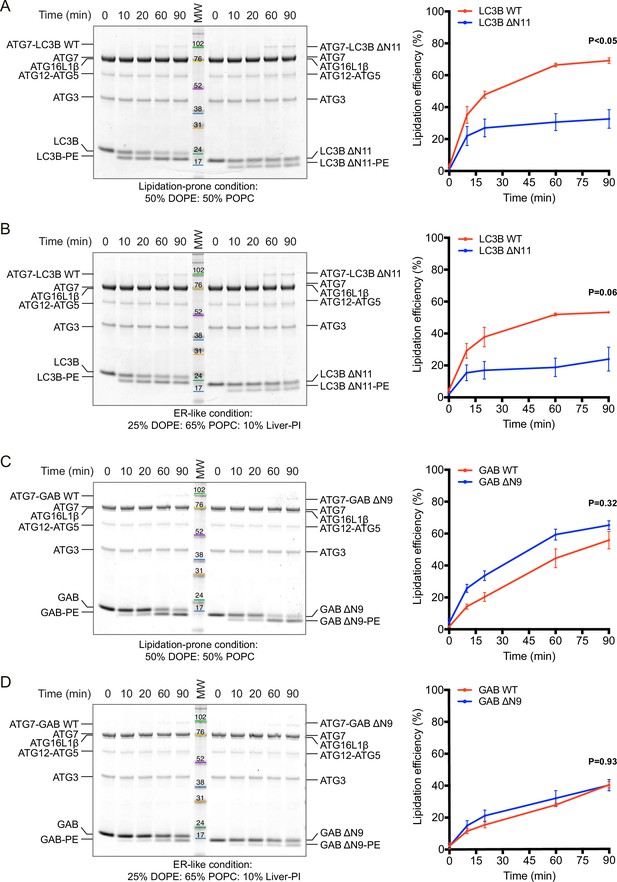
LC3B N-terminus is required for in vitro lipidation reaction, while GABARAP N-terminus is dispensable for this process.
(A, B) In vitro lipidation of LC3B WT and ΔN11. (C, D) In vitro lipidation of GABARAP WT and ΔN9. Lipidation reaction contains 1 μM ATG7, 1 μM ATG3, 0.25 μM ATG12-ATG5-ATG16L1, 5 μM ATG8 (LC3B WT or ΔN11, GABARAP WT, or ΔN9) and 1 mM large unilameller vesicles (LUVs) (lipidation-prone condition: 50% DOPE/50% POPC or ER-like condition: 25% DOPE/65% POPC/10% Liver-PI). The reactions were initiated by adding 5 mM MgATP and then incubated at 37°C for the indicated time points. The lipidated form of the ATG8 proteins were detected by SDS-PAGE followed by Coomassie blue stain. MW: molecular weight in kDa. Lipidation efficiency (%) was calculated in which the amount of lipidated ATG8 was divided by that of total ATG8 (n = 3, mean ± SEM).
-
Figure 1—figure supplement 2—source data 1
Uncropped gel images of Figure 1—figure supplement 2A–D and source data file (Excel) for Figure 1—figure supplement 2.
- https://cdn.elifesciences.org/articles/89185/elife-89185-fig1-figsupp2-data1-v2.zip
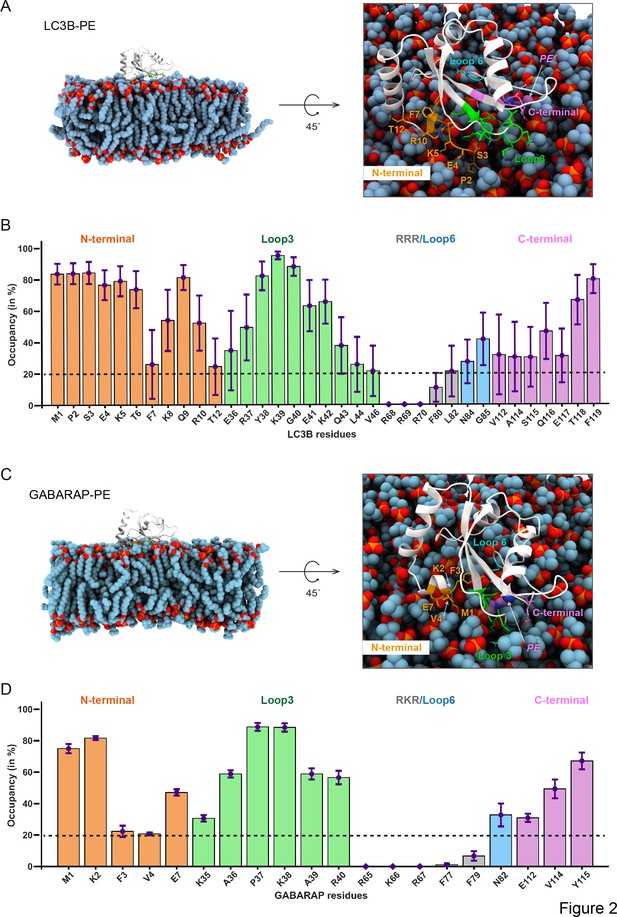
Molecular dynamics simulations of lipidated LC3B and GABARAP in POPC membrane elicit cis-membrane association of LC3B/GABARAP N-termini.
(A, C) The representative structure of LC3B-PE (300 ns) and GABARAP-PE (840 ns). (B, D) The percentage of time (occupancy) LC3B/GABARAP residues are in contact with POPC membrane during 1 μs of the trajectory. The bars are coloured according to the region which is N-terminal as orange, loop 3 as green, loop 6 as light blue, and C-terminal as purple.
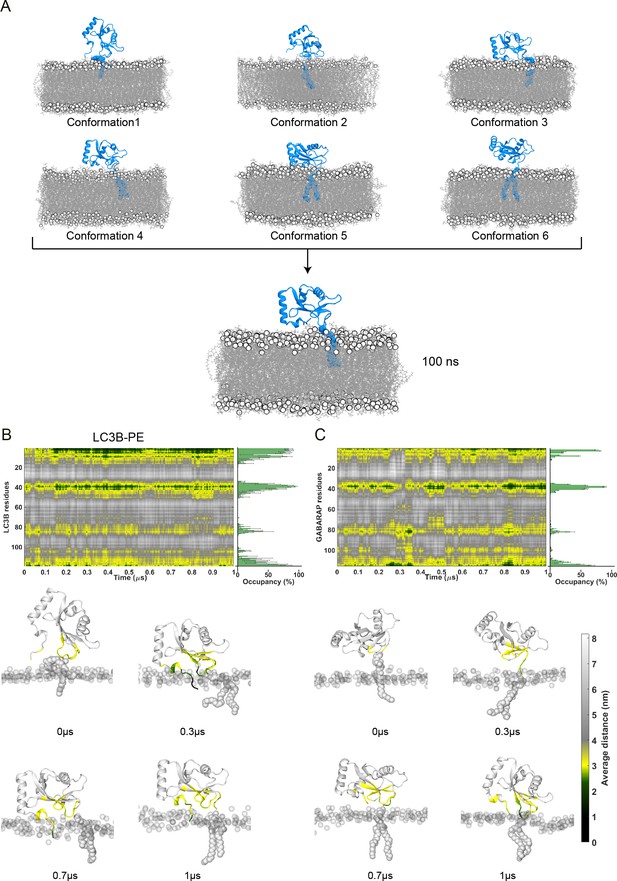
Molecular dynamics (MD) simulation set-ups of lipidated LC3B and GABARAP in POPC membrane.
(A) Pool of six Initial conformations with varying lipid-contacting orientations. After an initial refinement of 100 ns, all conformations converged to a single orientation with N-terminal moving closer to lipids. (B, C) Time evolution of distance between LC3B/GABARAP residues respectively and POPC membrane highlighting four distinct membrane interacting regions mainly, N-terminal, loop 3, loop 6, and C-terminal. Horizontal bar graph in the right shows the contact occupancy (in percentage) of all replicates. The contacts were categorised into four parts depending upon the residues’ side-chain orientation on the membrane: Inserted at ≤2 nm coloured in black, at membrane surface at ≤2.8 nm coloured in green, in proximity at <3 nm coloured in yellow, and not interacting at >3.5 nm coloured in grey. Lower panel depicts the snapshots at different time points, wherein protein residues are coloured according to their distance with the membrane. The headgroup of POPC membrane is highlighted in transparent silver colour, whereas the protein and lipid anchor are highlighted in silver colour.
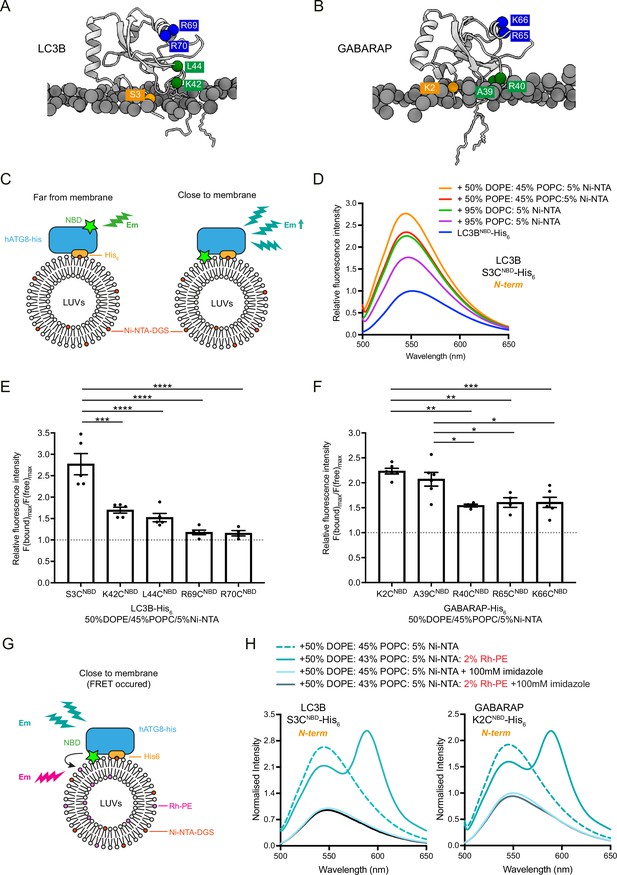
Analysis of membrane association interface of liposome-conjugated LC3B and GABARAP.
(A, B) Representative structures of lipidated LC3B (300 ns) and GABARAP (840 ns) from Molecular dynamics (MD) simulation. The individual amino acids of interest are labelled with NBD. (C) Scheme for the NBD spectra assay to characterise the membrane association residues in LC3B and GABARAP, using LC3B-His6/GABARAP- His6 and large unilameller vesicles (LUVs) containing nickel lipids to mimic the lipidated status. If the residue is embedded in the membrane, the NBD fluorescence gets increased. (D) Example of fluorescence spectra of LC3B S3CNBD- His6 in the absence of LUVs (protein only, 1 μM) and in the presence of LUVs containing 5% Ni-NTA (1 mM). Spectra represent mean values (n = 5) (E, F) Quantification of NBD fluorescence increase of the screened residues in LC3B-His6 and GABARAP-His6. F(bound)max/F(free)max ratio represents the maximum emission intensity of NBD-labelled LC3B-His6/GABARAP-His6 bound to liposomes containing 50% DOPE/45%POPC/5% Ni-NTA, normalised to the maximum emission intensity of NBD-labelled LC3B-His6/GABARAP-His6 in the absence of liposomes (n = 4–5, mean ± SEM). (G) Schematic diagram for the FRET assay to confirm the interaction between NBD-labelled LC3B-His6/GABARAP-His6 and rhodamine labelled liposomes. If the residue interacts with membrane, the emission of NBD would excite the rhodamine on the liposomes. (H) FRET assay with LC3B S3CNBD-His6 and GABARAP K2CNBD-His6. Each NBD-labelled LC3B-His6/GABARAP-His6 (1 μM) was mixed with 1 mM blank liposomes (50% DOPE/45% POPC/5% Ni-NTA) (dashed cyan) or rhodamine liposomes (50% DOPE/43%POPC/5% Ni-NTA/2% Rh-PE) (cyan). In parallel, addition of 100 mM imidazole to remove NBD-labelled LC3B-His6/GABARAP-His6 from liposomes was performed as negative controls. Spectra represent mean values (n = 3). Differences were statistically analysed by one-way ANOVA and Tukey multiple-comparison test. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001.
-
Figure 3—source data 1
Source data file (Excel) for Figure 3D, E, F and H.
- https://cdn.elifesciences.org/articles/89185/elife-89185-fig3-data1-v2.zip

Relative fluorescence spectra of NBD-labelled and liposome conjugated LC3B/GABARAP.
(A) LC3B loop 3 (green) and RRR region (blue) were tested for membrane binding with selected residues. (B) GABARAP N-terminal (orange), loop 3 (green), and RKR region (blue) were tested for membrane association activity with selected residues. Spectra represent mean values (n = 4–5).
-
Figure 3—figure supplement 1—source data 1
Source data file (Excel) for Figure 3—figure supplement 1.
- https://cdn.elifesciences.org/articles/89185/elife-89185-fig3-figsupp1-data1-v2.zip

FRET between NBD-labelled LC3B-His6/GABARAP-His6 and rhodamine-labelled liposomes.
(A–C) FRET assay with NBD-labelled residues in LC3B-His6. (D, E) FRET assay with NBD-labelled residues in GABARAP-His6. Spectra represent mean values (n = 3).
-
Figure 3—figure supplement 2—source data 1
Source data file (Excel) for Figure 3—figure supplement 2.
- https://cdn.elifesciences.org/articles/89185/elife-89185-fig3-figsupp2-data1-v2.zip
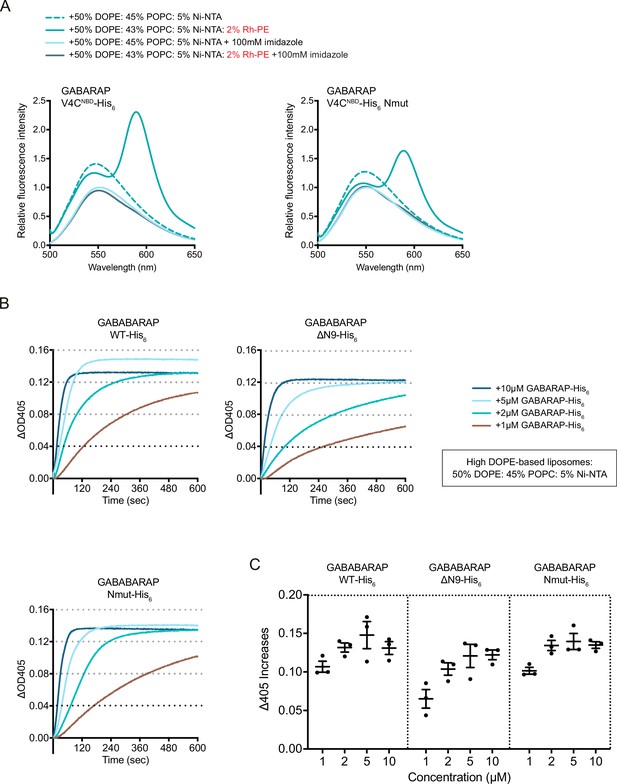
Point mutations in GABARAP N-terminus partially impair its N-terminal cis-membrane association but do not affect membrane tethering activity of GABARAP.
(A) FRET assay with GABARAP V4CNBD-His6 or V4CNBD Nmut-His6. Each NBD-labelled GABARAP-His6 (1 μM) was mixed with 1 mM blank liposomes or rhodamine liposomes: 50% DOPE/45% POPC/5% Ni-NTA (dashed cyan) or 50% DOPE/43% POPC/5% Ni-NTA/2% Rh-PE (cyan). Addition of 100 mM imidazole to remove NBD-labelled LC3B-His6/GABARAP-His6 from liposomes was performed as negative controls. Spectra represent mean values (n = 3). (B) Liposome tethering assays with GABARAP-His6, N-terminal deletion ΔN9-His6 and Nmut-His6 with 1 mM high DOPE-base liposomes containing 50% DOPE/45% POPC/5% Ni-NTA. Various amount of C-terminal His6-tagged GABARAP proteins were used. Liposome tethering of each condition (ΔOD405) was measured and normalised to that of liposomes only (n = 3). (C) Membrane tethering capacity after 10 min (ΔOD405 increase) of each condition shown in (B) was plotted (n = 3, mean ± SEM).
-
Figure 4—source data 1
Source data file (Excel) for Figure 4A and B.
- https://cdn.elifesciences.org/articles/89185/elife-89185-fig4-data1-v2.zip
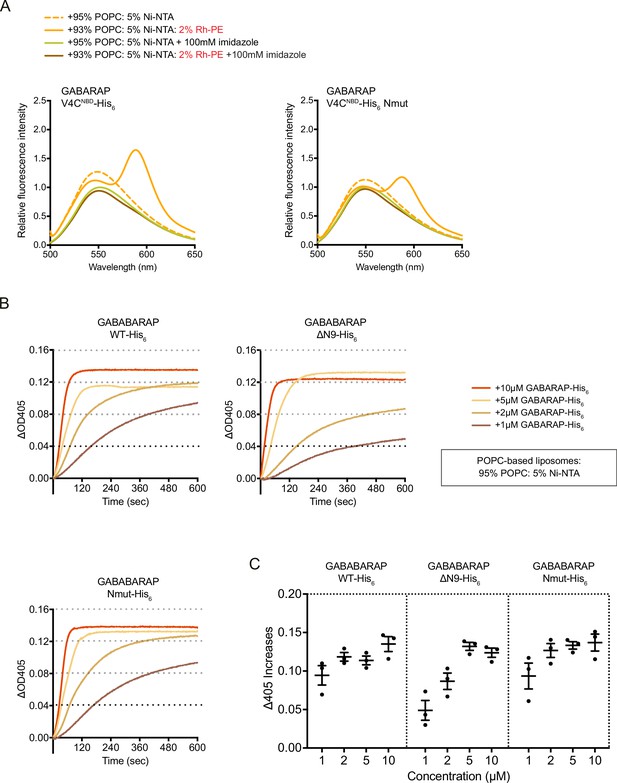
The same experiments have been done with POPC-based liposomes.
(A) FRET assay with GABARAP V4CNBD-His6 or V4CNBD Nmut-His6. Each NBD-labelled GABARAP-His6 (1 μM) was mixed with 1 mM blank liposomes or rhodamine liposomes: 95% POPC/5% Ni-NTA (dashed orange) or 93% POPC/5% Ni-NTA/2% Rh-PE (orange). Addition of 100 mM imidazole to remove NBD-labelled LC3B-His6/GABARAP-His6 from liposomes was performed as negative controls. Spectra represent mean values (n = 3). (B) Liposome tethering assays with GABARAP-His6, N-terminal deletion ΔN9-His6 and Nmut-His6 with 1 mM POPC-based liposomes containing 95% POPC/5% Ni-NTA. Various amount of C-terminal His6-tagged GABARAP proteins were used. Liposome tethering of each condition (ΔOD405) was measured and normalised to that of liposomes only (n = 3). (C) Membrane tethering capacity after 10 min (ΔOD405 increase) of each condition shown in (B) was plotted (n = 3, mean ± SEM).
-
Figure 4—figure supplement 1—source data 1
Source data file (Excel) for Figure 4—figure supplement 1A and B.
- https://cdn.elifesciences.org/articles/89185/elife-89185-fig4-figsupp1-data1-v2.zip
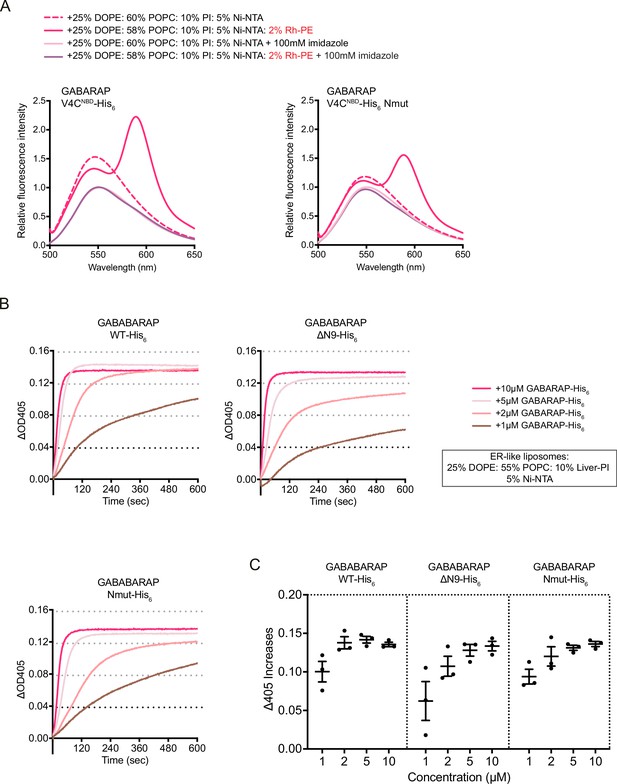
The same experiments have been done with ER-like liposomes.
(A) FRET assay with GABARAP V4CNBD-His6 or V4CNBD Nmut-His6. Each NBD-labelled GABARAP-His6 (1 μM) was mixed with 1 mM blank liposomes or rhodamine liposomes: 25% DOPE/60% POPC/10% Liver-PI/5% Ni-NTA (dashed magenta) or 25% DOPE/58% POPC/10% Liver-PI/5% Ni-NTA (magenta). Addition of 100 mM imidazole to remove NBD-labelled LC3B-His6/GABARAP-His6 from liposomes was performed as negative controls. Spectra represent mean values (n = 3). (B) Liposome tethering assays with GABARAP-His6, N-terminal deletion ΔN9-His6, and Nmut-His6 with 1 mM ER-like liposomes containing 25% DOPE/60% POPC/10% Liver-PI/5% Ni-NTA. Various amount of C-terminal His6-tagged GABARAP proteins were used. Liposome tethering of each condition (ΔOD405) was measured and normalised to that of liposomes only (n = 3). (C) Membrane tethering capacity after 10 min (ΔOD405 increase) of each condition shown in (B) was plotted (n = 3, mean ± SEM).
-
Figure 4—figure supplement 2—source data 1
Source data file (Excel) for Figure 4—figure supplement 2A and B.
- https://cdn.elifesciences.org/articles/89185/elife-89185-fig4-figsupp2-data1-v2.zip
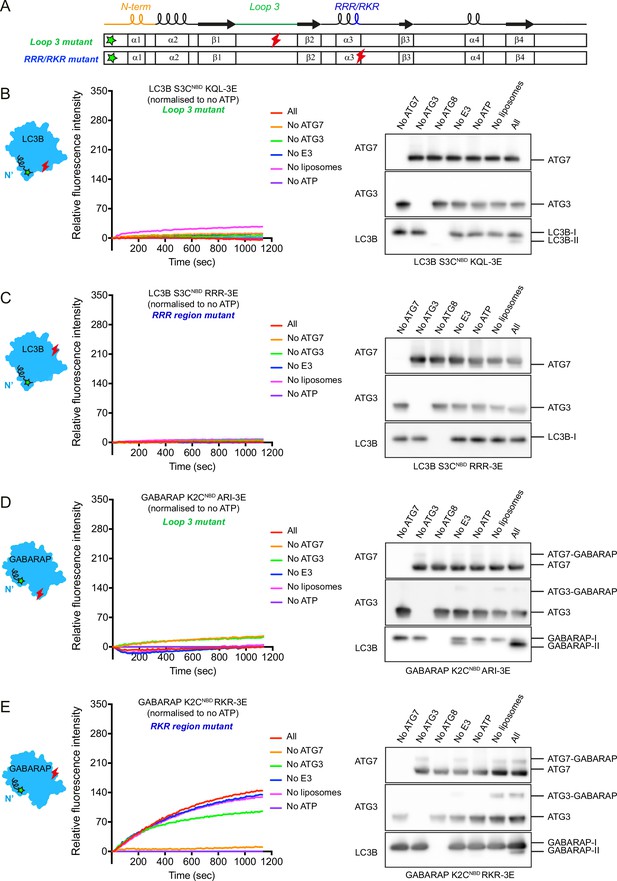
Membrane insertion of LC3B/GABARAP N-termini is hindered by alteration of residues in loop 3 and RRR/RKR regions.
(A) Scheme of N-terminal NBD-labelled LC3B (S3CNBD) or GABARAP (K2CNBD) with less hydrophobic and charge-inverting mutations (3E) in loop 3 and RRR/RKR regions. The green and red marks indicate the position of NBD labelling and 3E mutations, respectively. (B) The real-time lipidation assay with LC3B S3CNBD KQL-3E (loop 3 mutant). (C) The real-time lipidation assay with LC3B S3CNBD RRR-3E (RRR region mutant). (D) The real-time lipidation assay with GABARAP K2CNBD ARI-3E (loop 3 mutant). (E) The real-time lipidation assay with GABARAP K2CNBD RKR-3E (RKR region mutant). All spectra represent mean value (n = 3).
-
Figure 5—source data 1
Uncropped blot images of Figure 5B–E and source data file (Excel) for Figure 5B–E.
- https://cdn.elifesciences.org/articles/89185/elife-89185-fig5-data1-v2.zip
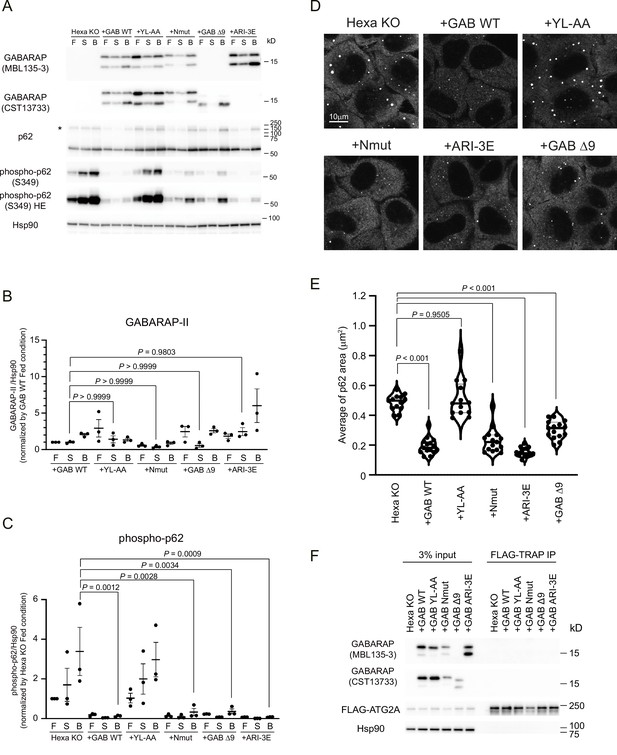
GABARAP N-terminus can be dispensable for the degradation of p62 body.
(A) Hexa KO cells stably expressing non-tagged GABARAP WT, YL-AA, Nmut, Δ9, or ARI-3E were starved for 8 hr with (B) or without 100 nM Bafilomycin A1 (S), or cultured in full media (F). Cell lysates were analysed by immunoblotting using the indicated antibodies. The asterisk indicates the position of high molecular weight forms of p62. (B, C) Band intensity quantification of GABARAP-II (B), and phosphorylated p62 (Ser349) (C). All data were normalised with those of HSP90. Data represent the mean ± SEM of three independent experiments. (D) The cells were starved for 2 hr before p62 (red) was visualised. Scale bar, 10 μm. (E) The violin plot of average p62 area. The thick and thin lines in the violin plot represent the medians and quartiles, respectively. n = 13 for Hexa KO and GAB YL-AA; n = 15 for GAB WT, Nmut, Δ9, and ARI-3E. (F) Co-immunoprecipitation experiments of FLAG-ATG2A with GABARAPs. Cell lysates were subjected to IP using anti-FLAG magnetic beads. The resulting precipitates were examined by immunoblot analysis with the indicated antibodies. Differences were statistically analysed by one-way ANOVA and Tukey multiple-comparison test. See also Figure 6—figure supplement 1.
-
Figure 6—source data 1
Uncropped blot images of Figure 6A and F.
- https://cdn.elifesciences.org/articles/89185/elife-89185-fig6-data1-v2.zip
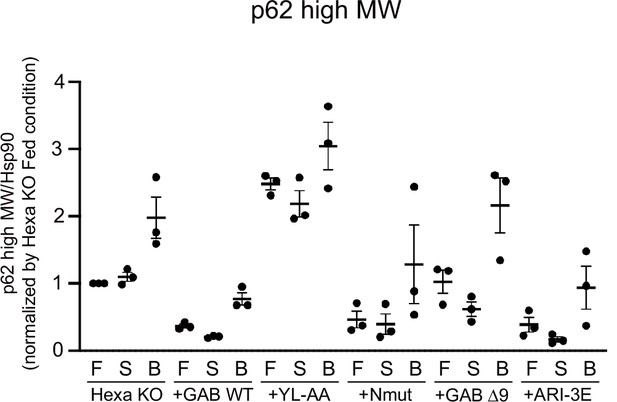
Band intensity quantification of high molecular weight forms of p62.
All data were normalised with those of HSP90. Data represent the mean ± SEM of three independent experiments.
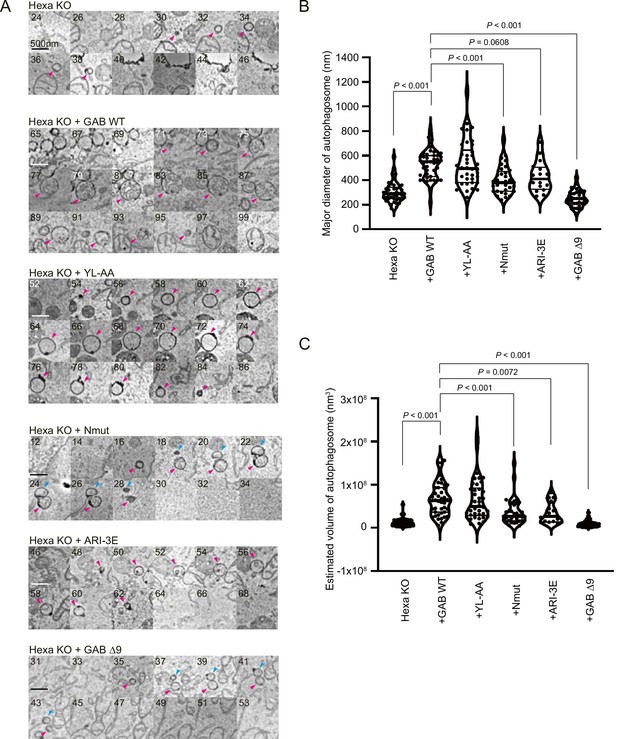
Autophagosome size is regulated by the N-terminus and cis-membrane association of GABARAP protein.
Three-dimensional correlative light and electron microscopy (3D-CLEM) analysis. The starved cells were fixed, imaged by confocal microscopy, and subsequently relocated and imaged by scanning electron microscopy. (A) Consecutive 25 nm SEM slice images are shown. The number in each image indicates the slice number. The arrowheads show autophagosomal structures. Scale bars, 500 nm. (B, C) Size distributions of autophagosomes. Length, width, and height of each autophagosome are measured. The major diameter (B) and estimated volume (C) of each autophagosome are shown. The thick and thin lines in the violin plot represents the medians and quartiles, respectively. n = 42 for Hexa KO; n = 43 for GAB WT; n = 38 for YL-AA; n = 33 for Nmut; n = 16 for ARI-3E; n = 33 for Δ9. See also Figure 7—figure supplement 1.
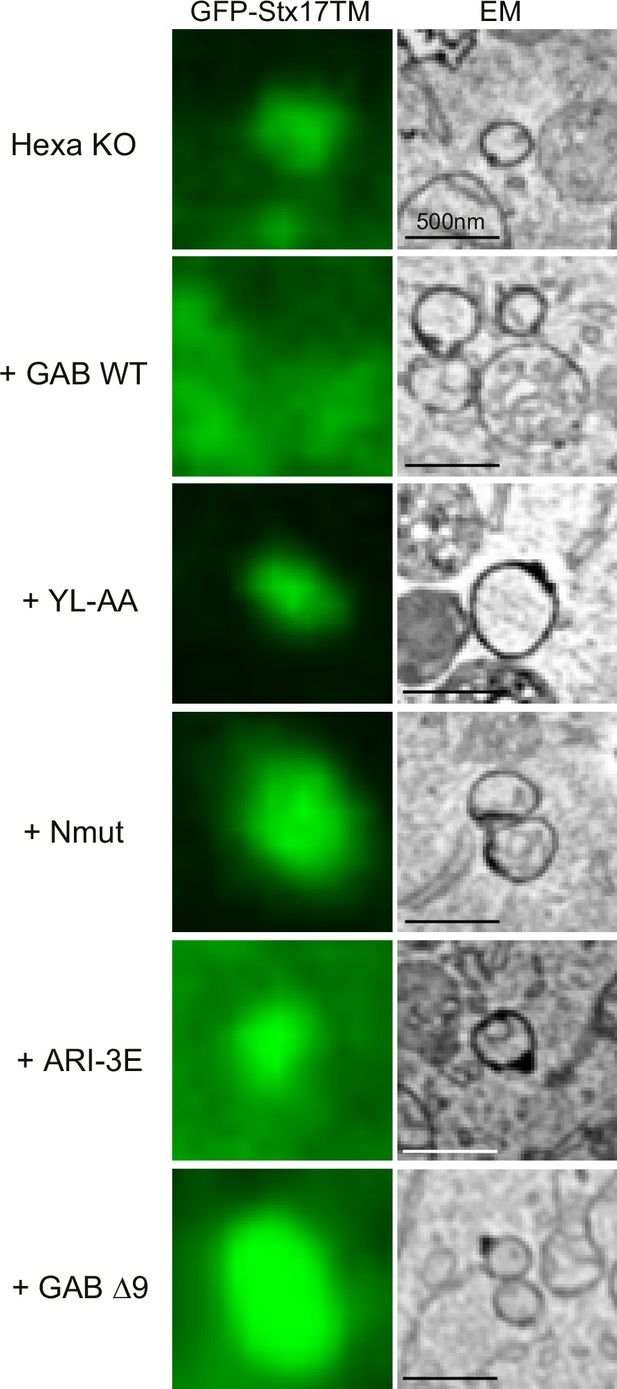
Correlative light and electron microscopy (CLEM) images of STX17TM-positive autophagosomes.
CLEM images of the area demonstrated in Figure 7 for serial images are shown. Scale bars, 500 nm.
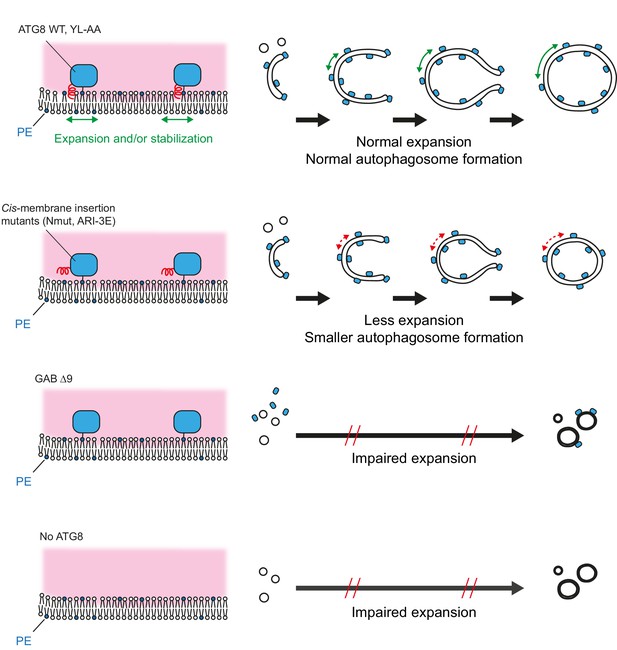
Model of autophagic membrane expansion mediated by lipidated mammalian ATG8 proteins.
After ATG8 lipidation, their N-termini are inserted into autophagic membranes. The cis-membrane association of the N-termini promotes autophagic membrane expansion by inducing membrane area expansion or stabilising the membrane. Reduced N-terminal cis-membrane association results in less autophagic membrane expansion, and consequently smaller autophagosomes form. Deletion of ATG8 N-terminus or lack of ATG8 proteins impairs autophagosome expansion.
Additional files
-
MDAR checklist
- https://cdn.elifesciences.org/articles/89185/elife-89185-mdarchecklist1-v2.docx
-
Supplementary file 1
Plasmids used in this study.
- https://cdn.elifesciences.org/articles/89185/elife-89185-supp1-v2.docx





