Author Response
The following is the authors’ response to the original reviews.
Reviewer #1 (Recommendations For The Authors):
1) Line 99-100 The authors claimed that IQCH is a novel IQ motif-containing protein, which is essential for spermiogenesis and fertilization. However, it is not clear if the currently published paper named an ancient testis-specific IQ motif containing H gene that regulates specific transcript isoform expression during spermatogenesis.
Response: Thanks to the reviewer’s comment. Yes, IQCH is the ancient testis-specific IQ motif containing H gene. According to the reviewer’s suggestion, we have revised the statement “Here, we revealed a testis-specific IQ motif containing H gene, IQCH, which is essential for spermiogenesis and fertilization” in Introduction part of revised manuscript.
2) Line 154-159 Immunofluorescence staining for the marker of the acrosome (peanut agglutinin: PNA) as well as the mitochondrial marker (Transcription Factor A, Mitochondrial: TFAM) was performed to confirm the deficiency of the acrosomes and mitochondria in the proband's spermatozoa. It seems that the spermatozoa acrosomes and mitochondria were severely defective in the proband. The authors should indicate IQCH's role in mitochondrial and acrosome function and IQCH's role in mitochondrial and acrosome function these points by explaining how IQCH is related to mitochondrial and acrosome deficiency. In addition to staining, other functional analyses should be performed to strengthen the claim of acrosome and mitochondrial defects.
Response: We appreciate the reviewer's valuable suggestion. Indeed, in our study, the results of multiomics analysis on WT and Iqch KO testes, including LC-MS/MS analysis, proteomic analysis, and RNA-seq analysis, found a potential role of IQCH in mitochondrial and acrosome function. GO analysis of these analysis indicated a significant enrichment in mitochondrial and acrosomal functions, including acrosomal vesicle, acrosome assembly, vesicle fusion with Golgi apparatus, mitochondrion organization, mitochondrial matrix, and so on.
Among the enriched molecules, in particular, HNRNPK mainly expresses at Golgi phase and Cap phase (Biggiogera et al. 1993). ANXA7 is a calcium-dependent phospholipid-binding protein that is a negative regulator of mitochondrial apoptosis (Du et al. 2015). Loss of SLC25A4 results in mitochondrial energy metabolism defects in mice (Graham et al. 1997). Furthermore, we confirmed that IQCH interacted with HNRNPK, ANXA7, and SLC25A4 through Co-IP, and exhibited downregulation in the sperm of the Iqch KO mice by immunofluorescence and western blotting. Moreover, IQCH can bind to HNRPAB, which could influence the mRNAs level of Catsper-family, such as Catsper1, Catsper2, and Catsper3, which are crucial for acrosome development (Jin ZR et al). In addition, we also detected HNRPAB binding to Dnhd1, which affects mitochondria development (Tan C et al). Therefore, in addition to staining, the other functional analyses also have provided the evidence of acrosome and mitochondrial defects caused by IQCH absence.
3) Line 180-182 IQCH knockout mice were generated. It is not clear why Mut-IQCH mice were not generated to be consistent with the human sequencing data.
Response: Thanks for reviewer’s comments. To understand IQCH's impact on fecundity in mice, we employed CRISPR-Cas9 to generate mice encoding the orthologous variant of IQCH387+1_387+10del detected in humans. Regrettably, due to sequence complexity, the designed sgRNA's specificity and efficiency were low, hindering successful Iqch knock-in mouse construction. Considering IQCH387+1_387+10del results in absent expression, we pursued Iqch knockout mice to explore IQCH's role in spermatogenesis.
4) Line 241.Figure 5A Gene Ontology (GO) analysis of the IQCH-bound proteins revealed a particular enrichment in fertilization, sperm axoneme assembly, mitochondrial organization, calcium channel, and RNA processing. But these GO functions are not shown in Figure 5A. The entire Figure 5 should be revised to enhance readability.
Response: We sincerely apologize for the oversight. These GO functions were indeed identified during the analysis of IQCH-bound proteins. Regrettably, we unintentionally omitted these GO functions when creating the plots. We have revised the plots in Figure 5 in revised manuscript to enhance readability.
5) Line 242 "33 ribosomal proteins were identified (Fig. 5B), indicating that IQCH might be involved in protein synthesis". The authors should perform an analysis to support the claim of protein synthesis defects.
Response: Thanks to reviewer’s suggestions. Initially, we have supplemented Co-IP experiments to confirm the interaction between IQCH and three ribosomal proteins (RPL4, RPS3, and RPS7), chosen from a pool of 33 ribosomal proteins based on different protein scores (Figure R1). In addition, the proteomic analysis revealed 807 upregulated proteins and 1,186 downregulated proteins in KO mice compared to WT mice. We confirmed the key downregulated proteins by western blotting and immunofluorescence staining in the previous manuscript. These results indicated that IQCH might interact with ribosomal proteins to regulate protein expression. Naturally, the regulation of protein synthesis by IQCH requires further experiments for confirmation in future studies.
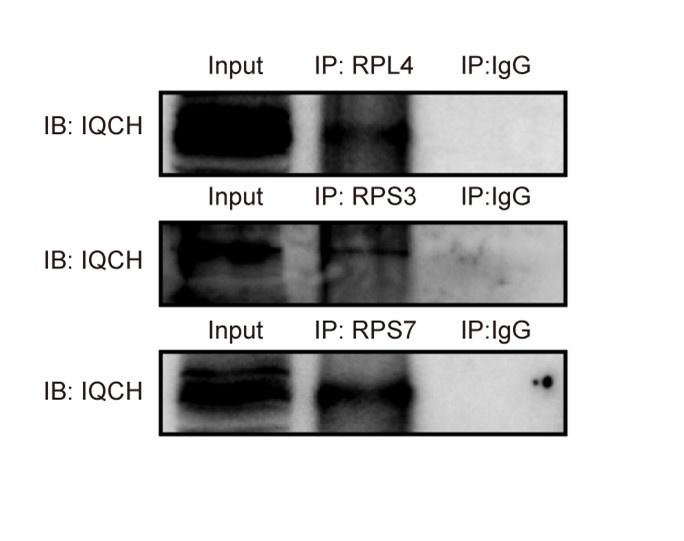
Author response image 1.
The interaction between IQCH and ribosomal proteins. Co-IP assays confirmed that IQCH interacted with RPL4, RPS3, and RPS7 in WT mouse sperm.
6) Line 244 The authors mentioned too many GO functions without focus.
Response: Following reviewer’s suggestions, we have simplified IQCH-associated GO functions in the revised manuscript.
7) Figure 6, there are no negative controls in all co-IP experiments. Band sizes are not marked. Thus, all data can't be evaluated. This also raises concern about whether the LC-MS/MS experiment to identify IQCH interacting protein was well-controlled? All co-IP experiments were poorly designed to draw any conclusion.
Response: Thanks to reviewer’s comments. We have supplemented negative controls in all Co-IP experiments and provided band sizes in Figure 6 in revised manuscript.
8) The authors mentioned that IQCH can bind to CaM. But they didn't detect CaM protein in Figure 5. Did the LC-MS/MS experiment really work?
Response: Thanks to reviewer’s comments. We detected the interaction of CaM protein with IQCH in the LC-MS/MS experiment analysis, which has been submitted as new Data S1 in the revised manuscript. We also confirmed their binding in mouse sperm by Co-IP experiment and immunofluorescence staining, which results were shown in Figure 6 and Figure S10 in the previous study.
9) Figure 6D. Because IQCH is lost in Iqch KO sperm, what is the point of showing in the Co-IP assay that CaM does not bind to IQCH in Iqch KO sperm?
Response: Following reviewer’s suggestions, we have deleted the results of Co-IP assay that CaM could not bind to IQCH in Iqch KO sperm.
10) Figure 6E. The Co-IP assay does not support the authors' claim that the decreased expression of HNRPAB was due to the reduced binding of IQCH and CaM by the knockout of IQCH or CaM.
Response: Thanks to reviewer’s expert comments. Indeed, the results of Figure 6E confirmed the interaction of IQCH and CaM in K562 cells, and also showed that the expression of HNRPAB was reduced when IQCH or CaM was knocked down, suggesting that IQCH or CaM might regulate HNRPAB expression. While in Figure 6F, the downregulation of HNRPAB caused by knocking down IQCH (or CaM) cannot be rescued when overexpressed CaM (or IQCH), indicating that CaM (or IQCH) cannot mediate HNRPAB expression alone. Therefore, the reduced expression of HNRPAB in Figure 6E might result from the weakened interaction between IQCH and CaM, but not a superficial downregulation of IQCH or CaM expression. To avoid the confusion, we have modified the relevant description in the revied manuscript.
Reviewer #2 (Recommendations For The Authors):
Major comments:
1) Lines 117 and 129: Please provide the reference number (NM_xxx.x) for the IQCH isoform that was used to interpret this variant. This is key information. Also, please provide the predicted truncation consequence caused by this splicing variant to IQCH protein.
Response: Thanks to reviewer’s suggestions. We have added reference number (NM_0010317152) of IQCH in manuscript. We employed splice site prediction tools, such as SpliceAI, RDDC, and varSEAK, to assess the expression consequences of this IQCH splicing variant. These tools couldn't anticipate the outcome of this splicing variant. However, the results of minigene splicing assay showed that the IQCH c.387+1_387+10del resulted in degradation of IQCH.
2) Figure 1A: The deleted sequence indicated by the red box does not match IQCH c.387+1_387+10del. Please show a plot of the exon-intron boundary under the Sanger sequencing results of the WT allele.
Response: Thanks to reviewer’s suggestions. We are sorry for the use of non-standard descriptions about the results of Sanger sequencing. According to the HGVS nomenclature (Figure R2), we have modified the red box to match IQCH c.387+1_387+10del and have added the exon-intron boundary in Figure 1A accordingly.
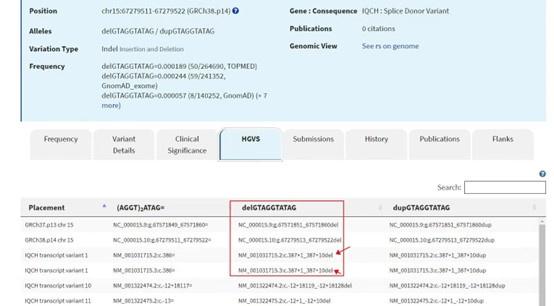
Author response image 2.
HGVS nomenclature description of the IQCH variant. The picture showed a detailed HGVS nomenclature description of IQCH c.387+1_387+10del.
Minor comments:
a) Manuscript title: It is suggested to change the title to "IQCH regulates spermatogenesis by interacting with CaM to promote the expression of RNA-binding proteins".
Response: According to reviewer’s suggestions, we have modified the title as “IQCH regulates spermatogenesis by interacting with CaM to promote the expression of RNA-binding proteins”.
b) Line 116: Please introduce the abbreviation WES. Also, please introduce the other abbreviations (such as WT, SEM, TEM, etc.) the first time they appear.
Response: Thanks to reviewer’s suggestions. We have provided the full explanations for all abbreviations upon their initial appearance.
c) Line 140, "Nonfunctional IQCH": Due to "the lack of IQCH expression" in Line 137, should "Nonfunctional IQCH" be changed into "IQCH deficiency"?
Response: Thanks for reviewer’s the detailed review. We have modified this title in Results part of the revised manuscript as followed: “IQCH deficiency leads to sperm with cracked axoneme structures accompanied by defects in the acrosome and mitochondria”
d) The information on the following references is incomplete: Sechi et al., Tian et al., Wang et al., and Xu et al. Please provide issue/page/article numbers.
Response: We are sorry for our oversight. We have provided the missing issue/page/article numbers for the references.
e) The title of Figure 1: Please emphasize that the male infertile-associated variant is "homozygous".
Response: Thanks to reviewer’s suggestions. We have revised the title of Figure 1 to emphasize the homozygous variant as follows: “Identification of a homozygous splicing mutation in IQCH in a consanguineous family with male infertility”.
f) Table 1: Please provide the reference paper for the normal values.
Response: We appreciate the reviewer's detailed checks. We have provided the reference paper for the normal
values in Table 1.
g) Figure 5F is distorted. Please make sure that it is a perfect circle.
Response: Thanks to reviewer’s suggestions. We have revised both the graphical representation and layout of Figure 5 in revised manuscript to make sure the readability.
Reviewer #3 (Recommendations For The Authors):
While the writing is generally clear, there are multiple examples of where the writing could be improved for clarity.
1) While some terms are defined throughout the manuscript, many abbreviations are not defined upon their first mention, such as WES, RT-PCR, TYH, HTF, KSOM, KEGG, RIPA, PMSE, SDS-PAGE, H&L, and HRP.
Response: Thanks to reviewer’s suggestions. We have provided the full explanations for all abbreviations upon their initial appearance.
2) On line 44, the claim that spermatogenesis is the "most complex biological process" is rather subjective and hard to support with concrete data.
Response: Thanks to reviewer’s suggestions. We have modified this description in the Introduction section as follow: “Spermatogenesis is one of the most complex biological process in male organisms and functions to produce mature spermatozoa from spermatogonia in three phases: (i) spermatocytogenesis (mitosis), (ii) meiosis, and (iii) spermiogenesis.”
3) On line 54, I think the authors meant "heterogeneous," not "heterologous."
Response: Thanks to reviewer’s comment. We have changed “heterologous” into “heterogeneous”.
4) On line 156, I think the authors meant "deficiency," not "deficient."
Response: Thanks to reviewer’s comment. We are sorry to make this mistake. We have made the correction in the revised version of the manuscript.
5) On line 300, K562 cells are mentioned, but neither in the Methods nor the Results are any details about the biological origin of these cells (or rationale for their use other than co-expression of IQCH and CaM) provided.
Response: Thanks to reviewer’s suggestion. K562 cell line is a human leukemia cell line and is enriched in the expression of IQCH and CaM, we thus opted to use this cell line for an easier knockdown of IQCH and CaM. We have supplemented the details about the biological origin of these cells in Method section of revised manuscript.
6) For the Results section describing Figure 6H, it would be nice to provide some explanation of the results of ICHQ overexpression alone relative to control situations and not just relative to the delta-IQ version or relative to simultaneous CaM manipulation.
Response: According to the reviewer’s suggestion, we have supplemented the co-transfection of control and CaM plasmids in HEK293T cells, and the results showed that the expression of HNRPAB in cells co-transfected with control and CaM plasmids was similar to that of co-transfected with IQCH (△IQ) /CaM plasmids, but was lower than that in the cells overexpressing the WT-IQCH and CaM plasmids, confirming the nonfunction of IQCH (△IQ) plasmids. We have shown the results in Figure 6H in the revised manuscript.
7) The sentence on lines 352-354 is confusing.
Response: We apologize for any confusion caused by the sentence in question. We have revisited the sentence and made appropriate revisions to enhance its clarity as follows: “Our findings suggest that the fertilization function is the main action of IQ motif-containing proteins, while each specific IQ motif-containing protein also has its own distinct role in spermatogenesis.”
8) The use of "employee" on line 371 is awkward and not very scientific.
Response: Thanks to reviewer’s comment. We have changed “employee” in to “downstream effector protein” on line 376





