Author response:
The following is the authors’ response to the previous reviews.
Public Reviews:
Reviewer #1 (Public Review):
Although this manuscript contains a potentially interesting piece of work that delineates a mechanism of IQCH that associates with spermatogenesis, this reviewer feels that a number of issues require clarification and re-evaluation for a better understanding of the role of IQCH in spermatogenesis. With the shortage of logics and supporting data, causal relationships are still not clear among IQCH, CaM, and HNRPAB. The most serious point in this manuscript could be that the authors try to generalize their interpretations with too simplified model from limited pieces of their data. The way the data and the logic are presented needs to be largely revised, and several interpretations should be supported by direct evidence.
Response: Thank you for the reviewer’s comment. IQCH is a calmodulin-binding protein, and the binding of IQCH and CaM was confirmed by LC-MS/MS analysis and co-IP assay using sperm lysate. We thus speculated that if the interaction of IQCH and CaM might be a prerequisite for IQCH function. To prove that speculation, we took HNRPAB as an example. We knocked down IQCH in cultured cells, and a decrease in the expression of HNRPAB was observed. Similarly, when we knocked down CaM in cultured cells, and a decrease in the expression of HNRPAB was also detected. However, these results cannot exclude that IQCH or CaM could regulate HNRPAB expression alone. To investigate that if IQCH or CaM could regulate HNRPAB expression alone, we overexpressed IQCH in cells that knocked down CaM, while the expression of HNRPAB cannot be rescued, suggesting that IQCH cannot regulate HNRPAB expression when CaM is reduced. In consistent, we overexpressed CaM in cells that knocked down IQCH, while the expression of HNRPAB cannot be rescued, suggesting that CaM cannot regulate HNRPAB expression when IQCH is reduced. Thus, IQCH or CaM cannot regulate HNRPAB expression alone. Moreover, we deleted the IQ motif of IQCH, which is required for binding to CaM. The co-IP results showed that the interaction of IQCH and CaM was disrupted when deleting the IQ motif of IQCH, and the expression of HNRPAB was decreased. Therefore, we suggested that the interaction of IQCH and CaM might be required for IQCH regulating HNRPAB. In future studies, we will further investigate the relationships among IQCH, CaM, and HNRPAB.
Reviewer #3 (Public Review):
(1) More background details are needed regarding the proteins involved, in particular IQ proteins and calmodulin. The authors state that IQ proteins are not well-represented in the literature, but do not state how many IQ proteins are encoded in the genome. They also do not provide specifics regarding which calmodulins are involved, since there are at least 5 family members in mice and humans. This information could help provide more granular details about the mechanism to the reader and help place the findings in context.
Response: Thanks to reviewer’s suggestion. We have provided additional background information regarding IQ-containing protein family members in humans and mice, as well as other IQ-containing proteins implicated in male fertility, in the Introduction section. Furthermore, we have supplemented the Introduction with background information concerning the association between CaM and male infertility.
(2) The mouse fertility tests could be improved with more depth and rigor. There was no data regarding copulatory plug rate; data was unclear regarding how many WT females were used for the male breeding tests and how many litters were generated; the general methodology used for the breeding tests in the Methods section was not very explicitly or clearly described; the sample size of n=3 for the male breeding tests is rather small for that type of assay; and, given that ICHQ appears to be expressed in testicular interstitial cells (Fig. S10) and somewhat in other organs (Fig. S2), another important parameter of male fertility that should be addressed is reproductive hormone levels (e.g., LH, FSH, and testosterone). While normal epididymal size in Fig. S3 suggests that hormone (testosterone) levels are normal, epididymal size and/or weight were not rigorously quantified.
Response: Thanks to reviewer’s comment. We have provided the data regarding copulatory plug rate and the average number of litters for breeding tests in revised Figure 3—figure supplement 2. The methodology used for the breeding tests has been revised to be more detailed and explicit in the revised Method section. Moreover, we have increased the sample size for male breeding tests to n=6. We measured the serum levels of FSH, LH, and Testosterone in the WT (9.3±1.9 ng/ml, 0.93±0.15 ng/ml, and 0.2±0.03 ng/ml) and Iqch KO mice (12±2 ng/ml, 1.17±0.2 ng/ml, and 0.2±0.04 ng/ml). There was no significant difference observed in the serum levels of reproductive hormones between WT and Iqch KO mice; therefore, we did not include the data in the study. Furthermore, we have added quantitative data on epididymal size in the revised Figure 3—figure supplement 2.
(3) The Western blots in Figure 6 should be rigorously quantified from multiple independent experiments so that there is stronger evidence supporting claims based on those assays.
Response: We appreciate the reviewer's comment. As suggested, we have added quantified data in Figure 6—figure supplement 2 from the results of Western blotting in Figure 6.
(4) Some of the mouse testis images could be improved. For example, the PNA and PLCz images in Figure S7 are difficult to interpret in that the tubules do not appear to be stage-matched, and since the authors claimed that testicular histology is unaffected in knockout testes, it should be feasible to stage-match control and knockout samples. Also, the anti-ICHQ and CaM immunofluorescence in Figure S10 would benefit from some cell-type-specific co-stains to more rigorously define their expression patterns, and they should also be stage-matched.
Response: Thanks to reviewer’s suggestions. We have included immunofluorescence images of anti-PLCz, anti-PNA and anti-IQCH and CaM during spermatogenesis development.
Recommendations for the authors:
Reviewer #1 (Recommendations For The Authors):
(1) There are multiple grammatical errors and statements drawn beyond the results. The entire manuscript would benefit from professional editing.
Response: We are sorry for the grammatical errors. We have enlisted professional editing services to refine our manuscript.
(2) Line 40, "Firstly" is not appropriate here.
Response: Thanks to reviewer’s comment. The word "Firstly" has been removed from the revised manuscript.
(3) Line 44, "processes".
Response: Thanks to reviewer’s suggestion. We have changed “process” in to “processes” on line 45.
(4) "spermatocytogenesis (mitosis)" is incorrect.
Response: Thanks to reviewer’s comment. We have changed “spermatocytogenesis (mitosis)” in to “mitosis” on line 47.
(5) Ca and Ca2+ are both used in line 67 - 77. Be consistent.
Response: We appreciate the reviewer's detailed checks. We have maintained consistency by revising instances of "Ca" to "Ca2+" in revised manuscript.
(6) Line 238 to 240, "To elucidate the molecular mechanism by which IQCH regulates male fertility, we performed liquid chromatography tandem mass spectrometry (LC-MS/MS) analysis using mouse sperm lysates and detected 288 interactors of IQCH (Data S1)."It is not clear how LC-MS/MS using mouse sperm lysates could detect "288 interactors of IQCH"? A co-IP experiment for IQCH using sperm lysates prior to LC-MS/MS is needed to detect "interactors of IQCH". However, in the Methods section, consistent with the main text, proteomic quantification was conducted for protein extract from sperm. Figure legend for Fig. 5 did not explain this, either.Thus, it is unable to evaluate Figure 5.
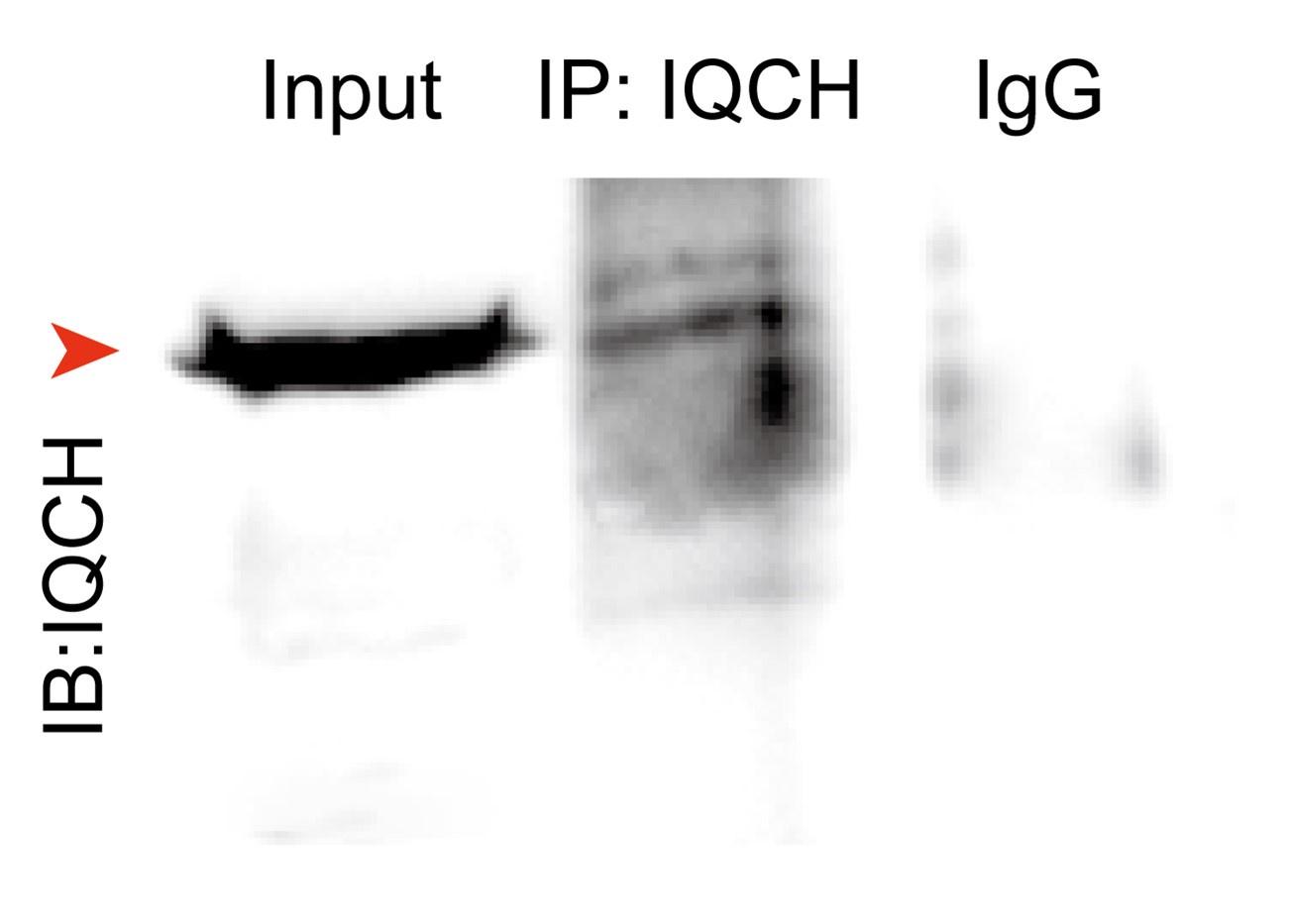
Response: We sincerely apologize for the oversight. Following reviewer’s suggestions, we have supplemented the method details of LC-MS/MS experiment in the Methods section of revised manuscript. Additionally, we conducted a co-IP experiment for IQCH using sperm lysates prior to LC-MS/MS and we did not include the corresponding figure in the manuscript. The results are as follows:
Author response image 1.
The results of a co-IP experiment for IQCH using sperm lysates from WT mice.
(7) Line 246, "... key proteins that might be activated by IQCH". What does "activated" here refer to? Should it be "upregulated"?
Response: We are sorry to our inexact statement. Instead, "upregulated" would better convey the intended meaning. According to reviewer’s suggestions, we have modified "activated" into "upregulated".
(8) Line 252 to 254, "the cross-analysis revealed that 76 proteins were shared between the IQCH-bound proteins and the IQCH-activated proteins (Fig. 5E), implicating this subset of genes as direct targets." This is a confusing statement. Is the author trying to say, IQCH-bound proteins have upregulated expression, suggesting that IQCH enhances their expression?
Response: We appreciate the reviewer's comment regarding the clarity of the statement in Line 252 to 254 of the manuscript. We have modified this sentence into “Importantly, cross-analysis revealed that 76 proteins were shared between the IQCH-bound proteins and the downregulated proteins in Iqch KO mice (Figure 5E), suggesting that IQCH might regulate their expression by the interaction.”
(9) Line 260 to 261, "SYNCRIP, HNRNPK, FUS, EWSR1, ANXA7, SLC25A4, and HNRPAB ... the loss of which showed the greatest influence on the phenotype of the Iqch KO mice." There is no evidence suggesting that the loss of SYNCRIP, HNRNPK, FUS, EWSR1, ANXA7, SLC25A4, and HNRPAB leads to Iqch KO phenotype.
Response: We apologize for our inaccurate statement. According to the literature, Fus KO, Ewsr1 KO, and Hnrnpk KO male mice were infertile, showing the spermatogenic arrest with absence of spermatozoa (Kuroda et al. 2000; Tian et al. 2021; Xu et al. 2022). Syncrip is involved meiotic process in Drosophila by interacting with Doublefault (Sechi et al. 2019). HNRPAB might be associated with mouse spermatogenesis by binding to Protamine 2 and contributing its translational regulation. Specifically, ANXA7 is a calcium-dependent phospholipid-binding protein that is a negative regulator of mitochondrial apoptosis (Du et al. 2015). Loss of SLC25A4 results in mitochondrial energy metabolism defects in mice (Graham et al. 1997). Moreover, RNA immunoprecipitation on formaldehyde cross-linked sperm followed by qPCR detected the interactions between HNRPAB and Catsper1, Catsper2, Catsper3, Ccdc40, Ccdc39, Ccdc65, Dnah8, Irrc6, and Dnhd1, which are essential for sperm development (Fukuda et al. 2013). Our Iqch KO mice showed abnormal sperm count, motility, morphology, and mitochondria, so we inferenced that IQCH might play a role in spermatogenesis by regulating the expression of SYNCRIP, HNRNPK, FUS, EWSR1, ANXA7, SLC25A4, and HNRPAB to some extent. We have changed an appropriate stamen that “We focused on SYNCRIP, HNRNPK, FUS, EWSR1, ANXA7, SLC25A4, and HNRPAB, which play important roles in spermatogenesis.”
(10) Fig. 6C and 6D use different styles of error bars.
Response: We are sorry for our oversight. In accordance with the reviewer's recommendations, we have modified the representation of error bars in the revised Fig. 6C.
(11) Line 296 to 297, "As expected, CaM interacted with IQCH, as indicated by LC-MS/MS analysis". It is not clear how LC-MS/MS detects protein interaction.
Response: As reviewer’s suggestions, we have supplemented the method details of LC-MS/MS experiment in the Methods section of revised manuscript. The results of proteins interacting with IQCH in sperm lysates from the LC-MS/MS experiment analysis were submitted as Figure 5—source data 1.
(12) It is still not clear how the interaction between IQCH, CaM, and HNRPAB is required for the expression of each other.
Response: Thank you for the reviewer’s comment. IQCH is a calmodulin-binding protein, and the binding of IQCH and CaM was confirmed by LC-MS/MS analysis and co-IP assay using sperm lysate. We thus speculated that if the interaction of IQCH and CaM might be a prerequisite for IQCH function. To prove that speculation, we took HNRPAB as an example. We knocked down IQCH in cultured cells, and a decrease in the expression of HNRPAB was observed. Similarly, when we knocked down CaM in cultured cells, and a decrease in the expression of HNRPAB was also detected. However, these results cannot exclude that IQCH or CaM could regulate HNRPAB expression alone. To investigate that if IQCH or CaM could regulate HNRPAB expression alone, we overexpressed IQCH in cells that knocked down CaM, while the expression of HNRPAB cannot be rescued, suggesting that IQCH cannot regulate HNRPAB expression when CaM is reduced. In consistent, we overexpressed CaM in cells that knocked down IQCH, while the expression of HNRPAB cannot be rescued, suggesting that CaM cannot regulate HNRPAB expression when IQCH is reduced. Thus, IQCH or CaM cannot regulate HNRPAB expression alone. Moreover, we deleted the IQ motif of IQCH, which is required for binding to CaM. The co-IP results showed that the interaction of IQCH and CaM was disrupted when deleting the IQ motif of IQCH, and the expression of HNRPAB was decreased. Therefore, we suggested that the interaction of IQCH and CaM might be required for IQCH regulating HNRPAB. In future studies, we will further investigate the relationships among IQCH, CaM, and HNRPAB.
Reviewer #3 (Recommendations For The Authors):
The authors have addressed my minor concerns. However, they neglected to address any of my more significant concerns in the public review. I assume that they simply overlooked these critiques, despite the fact that eLife explicitly states that "...as a general rule, concerns about a claim not being justified by the data should be explained in the public review." Therefore, the authors should have looked more carefully at the public reviews. As a result, my major concerns about the manuscript remain.
Response: We apologize for overlooking the public review process. We have improved our study based on the feedback received during the public review.




