A recurrent regulatory change underlying altered expression and Wnt response of the stickleback armor plates gene EDA
Figures
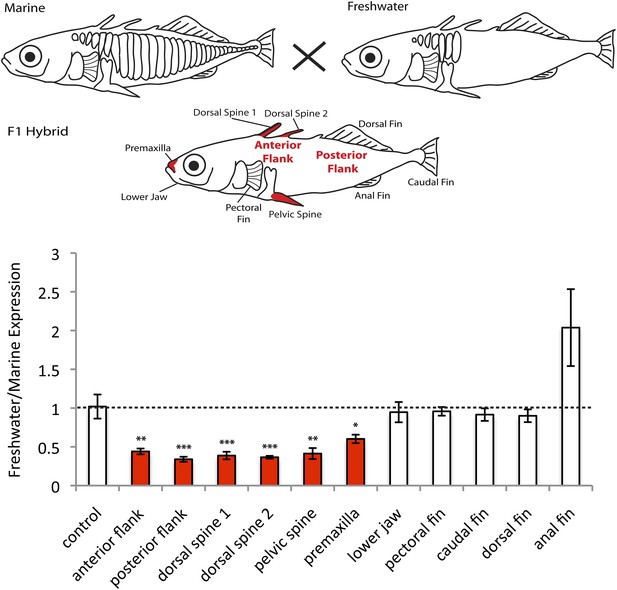
EDA shows allele-specific expression differences in several tissues, indicating cis-regulatory divergence.
Allele-specific expression in F1 freshwater-marine heterozygous larvae reveals significant differential expression of the marine and freshwater alleles in dorsal spines 1 and 2, the pelvic spine, the premaxilla, and the presumptive armor plates (anterior and posterior flanks). In all of these bony tissues the marine allele of EDA is expressed more highly than the freshwater allele, suggesting that there are differences in the cis-regulatory sequences controlling EDA expression. Several other tissues, however, do not show significant allelic imbalance in EDA expression; their allelic ratios are close to 1 (dashed line). The control shows results from a 1:1 mixture of plasmids containing the freshwater and marine alleles. Red-shaded structures and bars indicate tissues with significant allelic-imbalance compared to control (***p < 0.001, **p < 0.01, *p < 0.05 by two-tailed t-test).
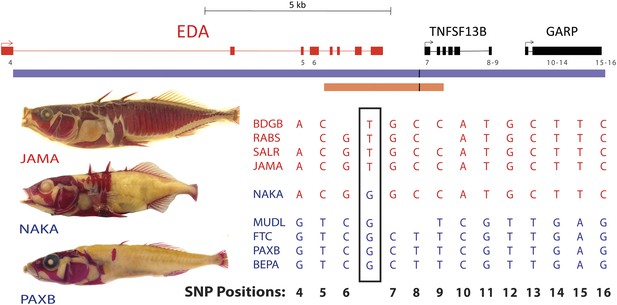
All low-plated populations share a single base pair change in the genetic region controlling armor plates.
Genome-wide comparisons of low- and high-plated fish reveal a T → G base pair change (black box) that is shared between all low-plated populations tested, including the low-plated Japanese NAKA fish that otherwise shows a primarily marine-like haplotype in the EDA region. Geographic population codes and DNA sequences from marine high-plated populations and freshwater low-plated populations are shown in red and blue, respectively, along with representative Alizarin Red stained fish showing typical armor plate patterns in different fish. The 16 kb candidate interval controlling armor plate number (blue bar, Colosimo et al., 2005) is shown beneath predicated genes in the region. Also shown are the numbered positions (4–16) of previously identified SNPs that differentiate most low- and high-plated sticklebacks other than NAKA (Colosimo et al., 2005). These numbered SNPs correspond to positions chrIV: 12800508, 12808303, 12808630, 12811933, 12813328, 12813394, 12815024, 12815027, 12816201, 12816202, 12816360, 12816402, and 12816464 in the stickleback genome assembly (Jones et al., 2012). Blank positions represent occasional gaps in sequence coverage for individual fish from large population surveys (Colosimo et al., 2005; Jones et al., 2012). The position of the shared T → G change (chrIV:12811481) is indicated with a short black vertical line in the overall genomic interval, and in a 3.2 kb region that was used to test for possible regulatory enhancers in the EDA region (orange bar, chrIV:12808949–12812120).
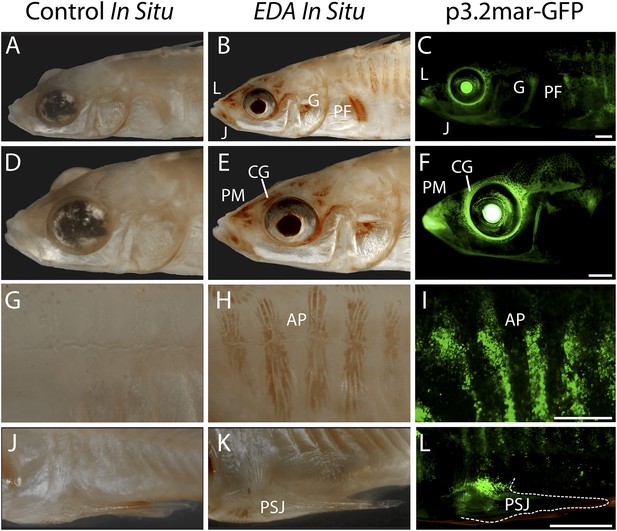
Reporter expression driven by an EDA enhancer matches several regions of endogenous EDA expression.
(A, D, G, J) Negative control DapB RNAscope in situ staining shows no positive brown signal appearing around the face (A and D), the plates (G), or the pelvic junction (J). The slight brown color in the pelvic spine is due to natural pigmentation at this site. (B, E, H, K) Endogenous EDA expression is localized to the premaxilla, lips, lower jaw, cranial ganglia, gill and pectoral fin base (B and E); armor plates (H); and the junction between the pelvic spine and the pelvic girdle (K). (C, F, I, L) The p3.2mar-GFP construct drives reporter expression at several corresponding sites, including the lips, premaxilla, lower jaw and cranial ganglia surrounding the eyes (C and F); in the armor plates; (I) and at the pelvic junction (L). Anatomical abbreviations as in other figures, including: lips (L), premaxilla (PM), lower jaw (J), cranial ganglia (CG), gills (G), pectoral fin base (PF), anterior plates (AP), and pelvic spine junction (PSJ). Scale bars are 1 mm long.
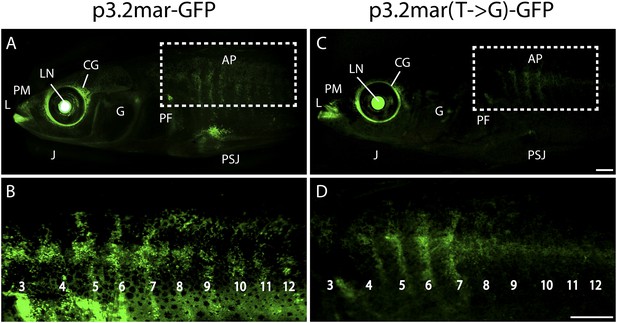
Enhancer expression in plates and other structures is reduced by a single base pair change.
(A, B) A 3.2 kb enhancer region from high-plated fish drives GFP expression in all armor plates (AP) of 2-month-old (20 mm long) marine stickleback larvae, with expression preceding plate ossification, and stronger expression in the first 7 armor plates. The p3.2mar-GFP construct also drives expression in the lips (L), premaxilla (PM), lower jaw (J), cranial ganglia (CG), the base of the pectoral fins (PF), and the pelvic spine-girdle junction (PSJ). Panel B is a higher magnification view of the area boxed in panel A. (C, D) The single base pair change in the p3.2mar(T → G)-GFP construct results in greatly reduced enhancer activity in the posterior plates, and reduced but detectable expression in plates 4–7 (D). This stable line also retains expression in the cranial ganglia and lips, reduced expression in the pelvic junction and the pectoral fin base, and novel strong expression in the spinal cord. Panel D is a higher magnification view of the area boxed in panel B. The hsp70 promoter in the GFP vector drives strong expression in the lens (LN) of all transgenic fish, helping to identify carriers following microinjection experiments (Chan et al., 2010). Scale bars are 1 mm long.
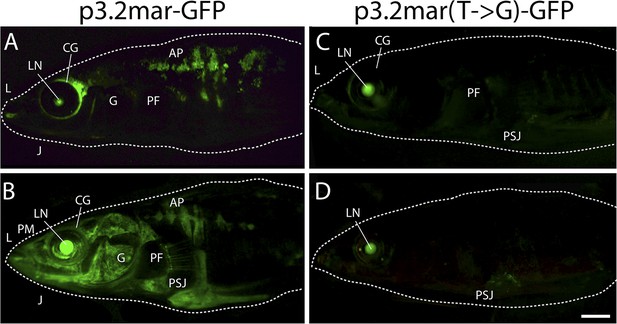
Plate enhancer activity is altered by a single base pair change (additional examples from independent transgenic fish).
(A, B) Examples of transient transgenics with mosaic GFP expression under the control of the marine high-plated marine EDA enhancer (p3.2mar-GFP). Multiple transgenic founders share expression in the cranial ganglia (CG) surrounding the eyes and the lips (L), the premaxilla (PM), under the jaw (J), and in armor plates (AP). (C, D) Site-directed mutagenesis of the p3.2mar-GFP construct generating p3.2mar(T → G)-GFP results in loss of armor plate expression in transient transgenics. However, expression in the cranial ganglia (CG) around the eyes and lips (L), as well as some expression surrounding the base of the pelvic spine-girdle junction (PSJ) remains in several fish. Copy number, integration sites, and mosaicism can vary in injected sticklebacks, giving rise to a range of expression levels. Despite this variability, consistent expression patterns can still be detected by comparing results from multiple injected fish. Overall, posterior plate expression was seen in 9 of 20 transgenic larvae with green eyes following injection of p3.2mar-GFP, vs of 0 of 27 transgenic larvae following injection of p3.2mar(T → G)-GFP. Scale bar in D is 2 mm long.
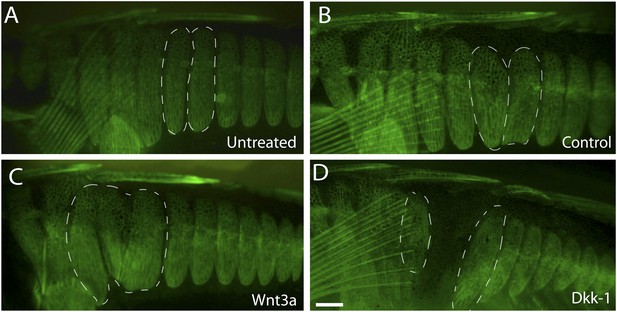
Wnt signaling regulates armor plate development.
Live Calcein staining of 6-month-old fish marks newly ossified bones in green. (A) Armor plates in an untreated high-plated adult marine fish. The normal morphologies of two individual plates are outlined with dashed lines. (B) Control beads soaked in PBS were implanted between the two outlined plates at two months of age. After bead implantation, fish continued to develop a full set of armor plates, with minimal changes in plate morphology (n = 8). (C) Implantation of Wnt-3a beads results in hypermorphic growth and armor plate fusion in the regions surrounding the exogenous Wnt-3a signal (n = 11). (D) Conversely, beads soaked in the Wnt inhibitor Dkk-1 inhibit plate formation surrounding the site of bead implantation (n = 10). Scale bar in D is 2 mm long.
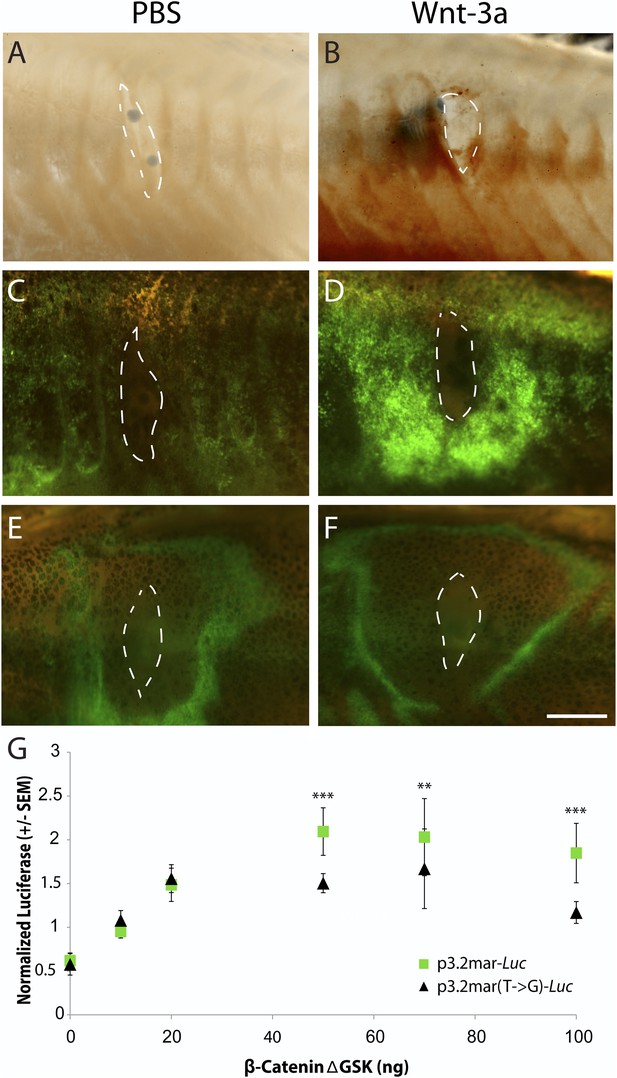
Single point mutation alters Wnt responsiveness of the EDA plate enhancer.
Beads soaked in either PBS or Wnt-3a protein were implanted in the flanks of 2-month-old (24 mm long) marine fish. All images were taken at 48 hr post bead implantation. (A, B) RNAscope in situ hybridization for EDA expression induced by control bead placement (A) or Wnt-3a protein (B). The addition of Wnt-3a beads induces a ring of EDA expression (brown color in B) directly surrounding the implantation site. (C, D) Bead implantation into the stable p3.2mar-GFP transgenic fish line. Control beads fail to induce GFP activity (C), whereas Wnt-3a beads induce a strong GFP response, seen in a ring surrounding the bead implantation site (D). (E, F) Bead implantation into the stable p3.2mar(T → G)-GFP line of transgenic fish. A ring of GFP expression is only seen at a distance from the implantation site of either control (E) or Wnt-3a (F) beads, corresponding to the location where cyanoacrylate glue was placed following implantation. Strong expression immediately surrounding the bead is not seen with Wnt-3a beads, in contrast to the result seen with p3.2mar-GFP transgenic fish (compare panels F and D). Scale bar in F is 1 mm long. (G) In vitro analysis of enhancer response to Wnt signaling via β-catenin co-transfection shows a strong induction of p3.2mar-Luc (green squares) with 50 ng or more of β-catenin in human HaCaT keratinocyte cells. The β-catenin-responsiveness of the p3.2mar(T → G)-Luc is significantly lower (black triangles). Combined p-values were calculated using Meta-P (***p < 0.001, **p < 0.01).
Additional files
-
Supplementary file 1
EDA haplotypes found in typical marine and freshwater populations, and in rare heterozygous carrier fish from Alaska. Previous studies have identified characteristic base pair positions that differentiate the most abundant EDA haplotypes present in marine and freshwater populations. The position of these SNPs in the EDA region are shown, along with the exon number in which they occur, the nature of the mutation (S: synonymous change; N: non-synonymous change; Reg: regulatory change), the SNP number assigned in previous studies, and the characteristic alleles that are found in 10 different completely-plated marine (red) and 14 different low-plated freshwater (blue) populations (top two haplotype rows, see Colosimo et al., 2005). We amplified the genomic region surrounding the new T → G regulatory mutation identified in the current study, and found that 12 of 263 completely plated migratory fish from Rabbit Slough, Alaska (RABS) were heterozygous for the T → G change (minor allele frequency: 2.3%). Analysis of flanking positions suggests that these carrier fish are heterozygous for larger or smaller haplotypes at the EDA locus, or are marine-like at all positions except for the T → G change like NAKA. The six indicated SNPs correspond to positions chrIV: 12808303, 12808630, 12811481, 12811933, 12813328, 12813394 in the stickleback genome assembly (Jones et al., 2012).
- https://doi.org/10.7554/eLife.05290.010





