Lipid-mediated regulation of SKN-1/Nrf in response to germ cell absence
Figures
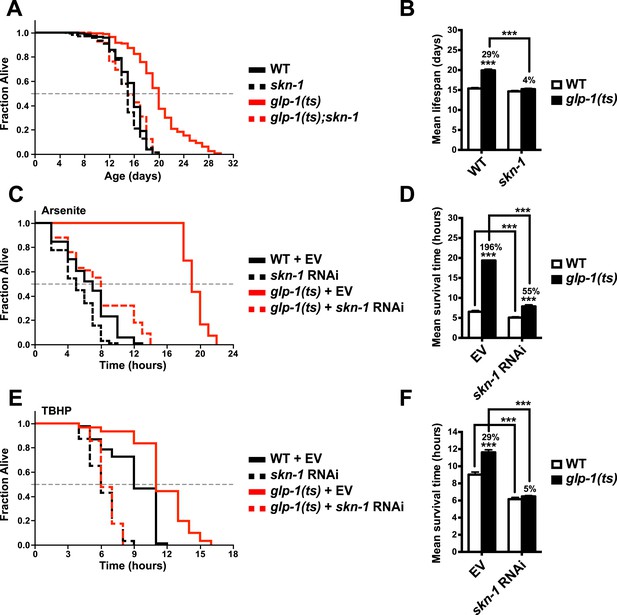
SKN-1 promotes longevity and stress resistance in germline stem cell (GSC)(−) animals.
(A, B) Wild type, skn-1(zu135), glp-1(bn18ts), and glp-1(bn18ts);skn-1(zu135) double mutants were assayed for lifespan at 25°C. skn-1(zu135) is a presumed null that is used throughout the study. Unless otherwise specified, glp-1(ts) refers to glp-1(bn18ts). (A) Composite survival curve. (B) Graph of mean lifespans. (C–F) glp-1(ts) mutants require skn-1 for oxidative stress resistance. Day-3 adult glp-1(ts) and control worms treated with skn-1 RNAi or empty vector were exposed to (C, D) 5 mM sodium arsenite (AS) or (E, F) 15.4 mM tert-butyl hydroperoxide (TBHP). Data are represented as mean ± SEM. p < 0.001***. The interaction between glp-1 and skn-1 was significant for both lifespan and stress resistance (p < 0.001). Statistical analysis and replicates are in Tables 1, 2.
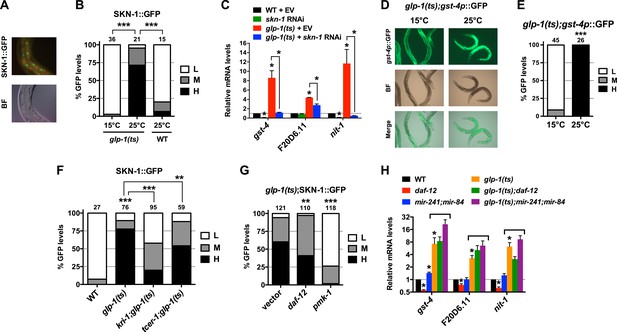
GSCs inhibit SKN-1 activity in the intestine.
(A) Representative images of SKN-1::green fluorescent protein (GFP) in intestinal nuclei; GFP channel (top), bright field (BF; bottom). (B) Accumulation of SKN-1::GFP in intestinal nuclei in GSC(−) animals. (C) skn-1-dependent activation of direct SKN-1 target genes (Robida-Stubbs et al., 2012) in response to GSC absence, detected by qRT-PCR. (D, E) Increased expression of gst-4p::GFP in the intestine of glp-1(ts) animals. Hypodermal gst-4p::GFP expression appeared to be unaffected. (D) Representative 10× images. (E) Intestinal gst-4p::GFP quantification. (F–H) GSCs regulate SKN-1 parallel to DAF-16 and DAF-12. In (H), SKN-1 target genes are assayed by qRT-PCR. glp-1(ts) refers to glp-1(e2141ts), and horizontal black lines indicate strains lacking GSCs. (C, H) Data are represented as mean ± SEM. n = 3 for qRT-PCR samples. (B, E–G) GFP quantification with high, medium, low scoring. Numbers above bars denote sample size. p < 0.05*; p < 0.01**; p < 0.001***.
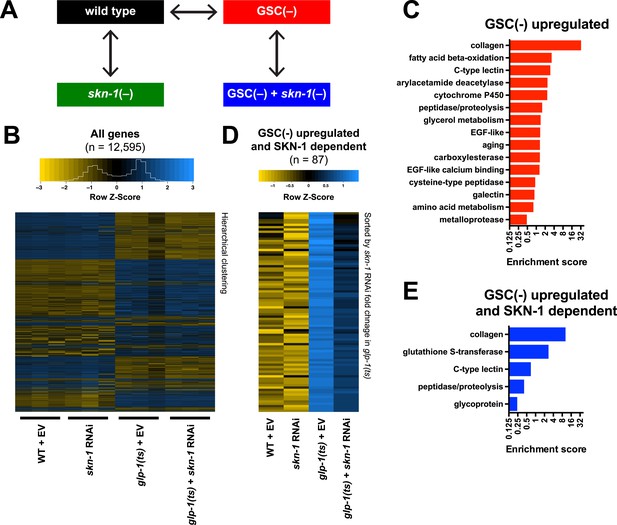
Effects of GSC absence and skn-1 on gene expression.
(A) RNA-seq experiment setup. For each condition, three biological replicates were obtained from synchronized intact day-1 adults at 25°C. Arrows indicate comparisons that were made, and GSC(−) refers to glp-1(ts). (B) Heatmap of all genes evaluated, showing biological replicates. (C) DAVID functional annotation analysis of GSC(−)-upregulated genes. (D) Heatmap of genes upregulated in glp-1(ts) in a skn-1-dependent manner (87 genes; fold change (FC) > 4 in GSC(−); FC < 0.67 with skn-1 RNAi in GSC(−)). (E) Functional annotation of the genes shown in (D). Genes and additional statistics are provided in Supplementary file 1a,d.
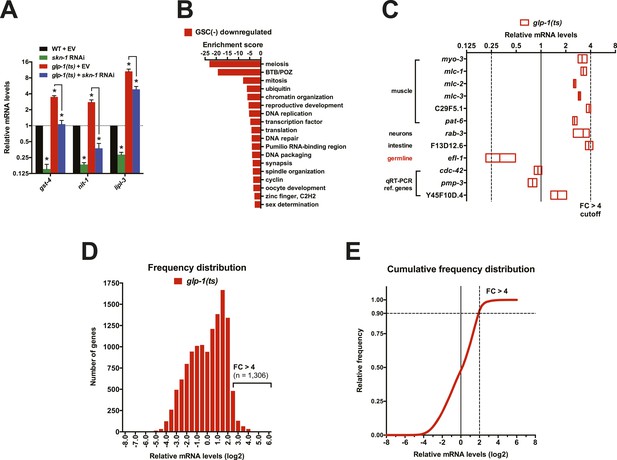
Gene expression changes following GSC inhibition.
(A) qRT-PCR validation of RNA samples used for RNA-seq. gst-4 and nit-1 are direct SKN-1 targets (Robida-Stubbs et al., 2012). Data are represented as mean ± SEM. n = 3; p < 0.05*. Analysis of a different RNA sample set is shown in Figure 2D. (B) DAVID analysis of genes that were downregulated in glp-1(ts). Processes related to GSC maintenance and reproduction were highly represented, as expected. (C) Altered abundance of tissue-specific genes in GSC(−) animals. Adult hermaphrodite worms have 959 somatic cells and ∼2000 GSCs (Kimble and Crittenden, 2005). A representative somatic-specific gene would therefore be predicted to be present at higher relative abundance in GSC(−) samples after normalization to either total RNA or reference housekeeping genes. Accordingly, representative somatic tissue-specific genes (Richmond, 2005; Moerman and Williams, 2006; Chikina et al., 2009) were present at threefold to fourfold higher relative levels in the GSC(−) samples. By contrast, a germline-specific gene (efl-1) was underrepresented (∼4×) in GSC(−) samples. Reference genes that are ubiquitously expressed in all tissues and commonly used for qRT-PCR normalization (Hoogewijs et al., 2008) do not have altered relative abundance in the GSC(−) samples. (D, E) Frequency distribution plots of mRNA levels in glp-1(ts) worms relative to wild type. A cutoff of FC > 4 denotes 1,306 out of 12,595 genes sequenced (10.4%).
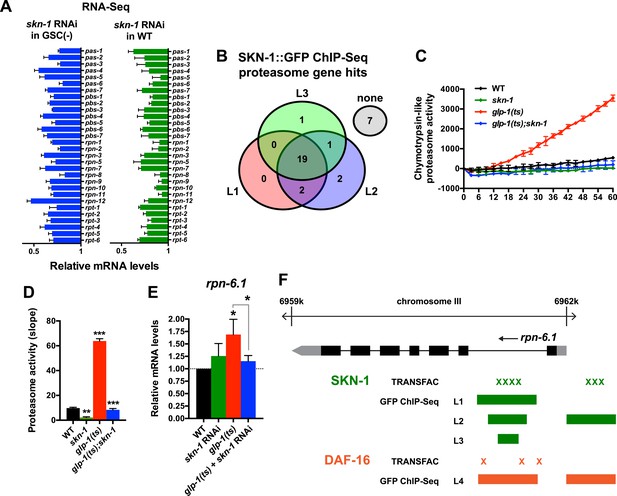
SKN-1 increases proteasome activity in response to GSC absence.
(A) Reduction in relative proteasome gene subunit mRNA levels by skn-1 RNAi, detected by RNA-seq. (B) Venn diagram indicating the number of proteasome subunit genes (pas, pbs, rpn, rpt families) that have SKN-1::GFP ChIP-seq peak hits near the transcription start site at the indicated larval stage (Niu et al., 2011). (C, D) SKN-1-dependence of increased chymotrypsin-related proteasome activity in GSC-ablated worms. The slopes from (C) are graphed in (D). Additional experiments are in Figure 4—figure supplement 1. (E) SKN-1-dependence of rpn-6.1 upregulation. Data are represented as mean ± SEM. n = 3 for all experiments. p < 0.05*; p < 0.01**; p < 0.001***. (F) Direct binding of SKN-1 and DAF-16 to the rpn-6.1 gene, indicated by TRANSFAC transcription factor binding prediction and modENCODE GFP ChIP-seq analyses. Both the predicted promoter and first intron of rpn-6.1 are highly enriched for SKN-1 and DAF-16 binding.
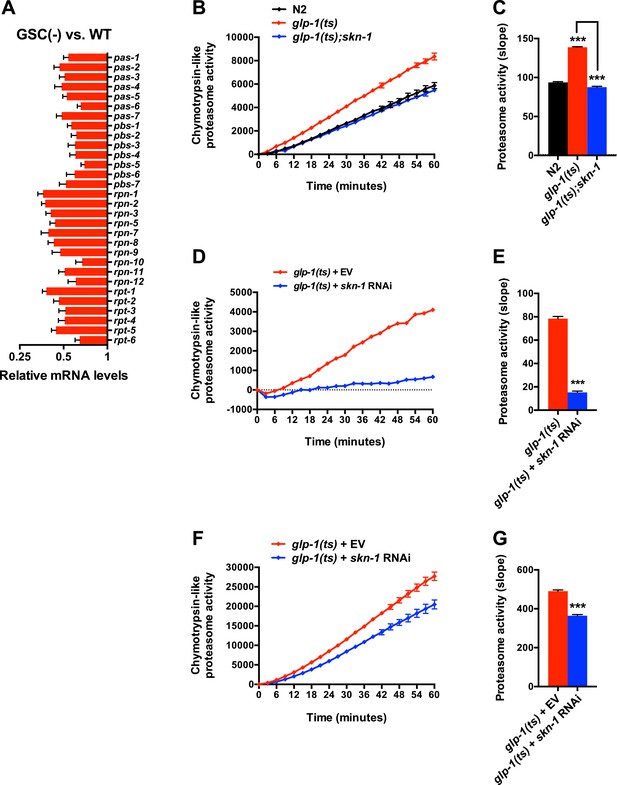
SKN-1-dependence of the increased proteasome activity in GSC(−) animals.
(A) GSC absence reduces relative proteasome subunit gene mRNA abundance, detected by RNA-seq. (B, C) The skn-1(zu135) mutation suppresses the increase in 26S proteasome activity seen in day-1 adult glp-1(ts) animals. (D, E) skn-1 RNAi administered from the egg stage suppresses proteasome activity in day-1 adult glp-1(ts) animals. (F, G) skn-1 RNAi administered post-developmentally, starting at day-1 adulthood, significantly reduces proteasome activity in day-5 adult glp-1(ts) animals. (B, D, F) Kinetic curves of chymotrypsin-like proteasome activity. (C, E, G) Graphs of proteasome activity slopes. Data are represented as mean ± SEM. n = 3; p < 0.001***.
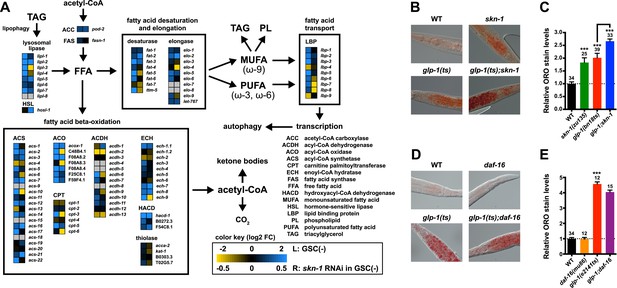
SKN-1 regulates lipid metabolism in GSC(−) animals.
(A) Functional map of lipid metabolism gene expression. Left columns show the effects of GSC absence (GSC(−) vs WT) and right columns the effects of skn-1 RNAi in GSC(−) animals. SKN-1 regulates genes involved in fatty acid (FA) oxidation, breakdown of triacylglycerols (TAG) to free FAs, production of mono- and poly-unsaturated FAs (MUFA, PUFA), and FA transport. Color coding reflects relative representation in RNA-seq data, with blue and yellow indicating increased and decreased expression, respectively. (B–E) Increased fat levels in glp-1(ts) and skn-1 mutants but not daf-16 mutants. (D, E) glp-1(ts) refers to glp-1(e2141ts). Representative 40× differential interference contrast (DIC) images of fixed ORO-stained worms are shown in (B, D), with quantification provided in (C, E). Additional images and quantification are provided in Figure 5—figure supplement 1. Data are represented as mean ± SEM. Numbers above bars denote sample size. p < 0.001***.
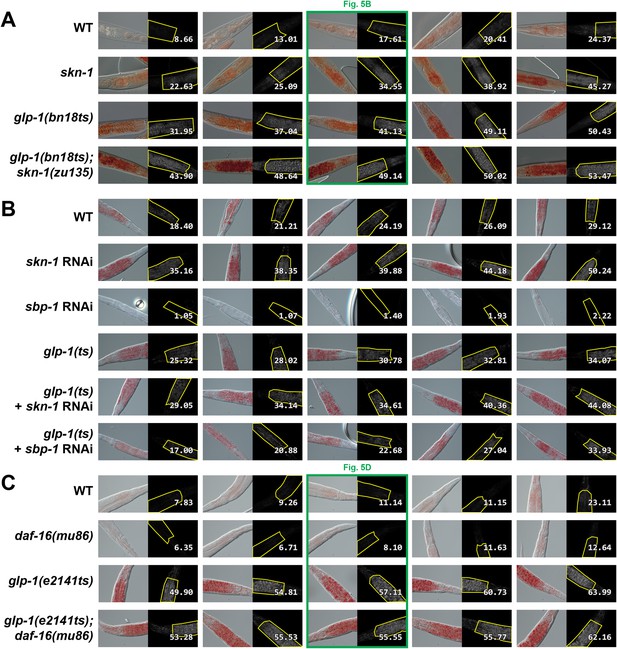

Representative ORO staining images with quantification.
Representative images divided into approximate quintiles, ordered by mean pixel intensity, are shown for each strain. BF images are presented on the left and background subtracted images with quantification on the right. (A) glp-1(bn18ts) and skn-1(zu135) mutants. (B) glp-1(ts) mutants treated with skn-1 and sbp-1 RNAi. Increased staining was observed both with skn-1 mutants and skn-1 RNAi. RNAi against sbp-1, which is required for lipogenesis (Yang et al., 2006), decreases ORO staining in both WT and glp-1(ts) genetic backgrounds. (C) glp-1(e2141ts) and daf-16(mu86) mutants. Numbers indicate mean pixel intensity above background (see ‘Materials and methods’ for additional details).
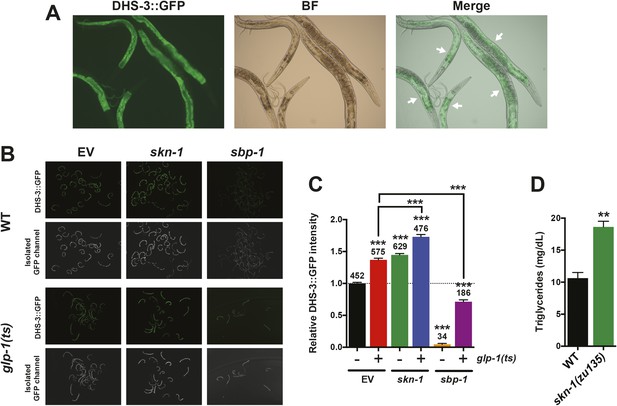
Analysis of the intestinal lipid droplet marker DHS-3::GFP, and TAG levels.
skn-1 RNAi increases DHS-3::GFP intensity in both wild-type and GSC(−) animals, whereas sbp-1 RNAi decreases DHS-3::GFP intensity. (A) 10× slide-mounted DHS-3::GFP images. White arrows indicate intestine-specific expression. (B, C) COPAS Biosorter quantification of DHS-3::GFP in live day-1 adult worms. (B) Representative 10× inverted scope images of worms suspended in M9 buffer used for COPAS scoring. (C) Graph of mean DHS-3::GFP fluorescence, assayed by COPAS. Numbers above bars denote sample size. Asterisks directly above bars indicate p values relative to WT or RNAi control. Asterisks above black lines denote effect of RNAi in glp-1(ts) background. RNAi was started from egg stage, and animals were raised at 25°C. (D) TAG levels are significantly elevated in skn-1 mutants. Data are represented as mean ± SEM. p < 0.01**; p < 0.001***.
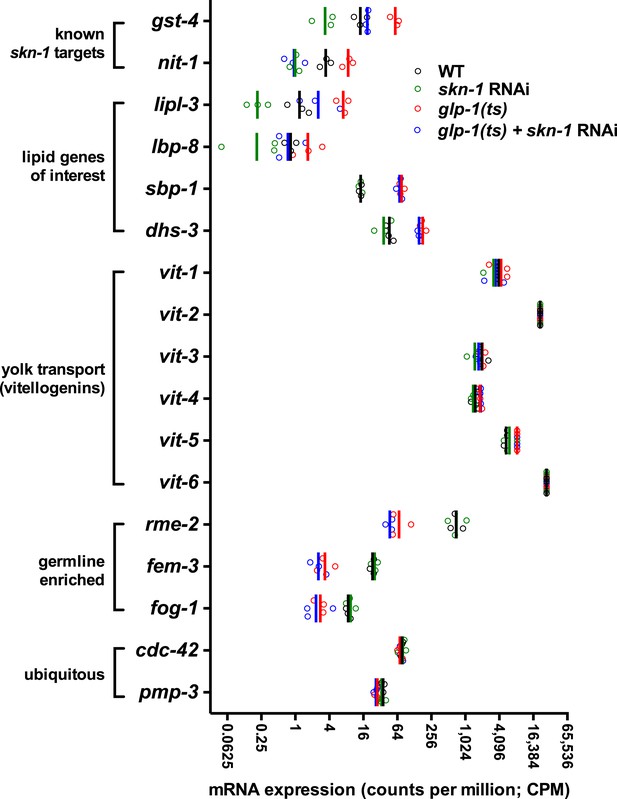
RNA-seq counts of select lipid metabolism and yolk transporter genes.
Expression of the known SKN-1 target genes gst-4 and nit-1 is elevated in GSC(−) animals in a skn-1-dependent manner. The TAG lipase lipl-3 and FABP lbp-8 are similarly elevated in GSC(−) animals in a skn-1-dependent manner, but expression of sbp-1, dhs-3, and yolk protein vitellogenins (VIT genes) are not affected by skn-1 RNAi. rme-2 expression is germline enriched but regulated independently of skn-1. Open circles denote replicates. Vertical lines indicate mean counts per million (CPM).
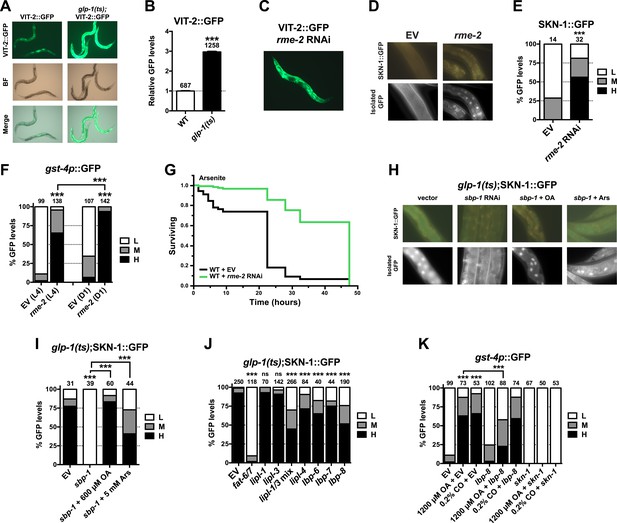
GSC absence activates SKN-1 through FA signaling.
(A) Accumulation of yolk transporter VIT-2::GFP in the soma of GSC(−) animals. Detailed higher magnification images are provided in Figure 6—figure supplement 1. (B) COPAS quantification of VIT-2::GFP. Data are represented as mean ± SEM. (C) Knockdown of the oocyte-specific yolk receptor rme-2 promotes somatic VIT-2 accumulation. (D–F) rme-2 RNAi activates SKN-1 in the intestine. (D, E) SKN-1::GFP accumulates in intestinal nuclei in rme-2 RNAi-treated worms. (F) In rme-2 RNAi-treated worms, gst-4p::GFP levels in the intestine are increased at the L4 stage and increased further by day-1 adulthood. (G) rme-2 RNAi enhances resistance to AS, in a skn-1-dependent manner (see replicates in Table 2). (H, I) An oleic acid (OA)-dependent signal is required for SKN-1 to be activated by GSC inhibition but not oxidative stress. In GSC(−) animals, SKN-1 nuclear accumulation is abolished by sbp-1 RNAi and rescued by OA supplementation. SKN-1 remains capable of responding to oxidative stress (30 min AS exposure) after sbp-1 RNAi in GSC(−) (H, I) or WT (Figure 6—figure supplement 2A) worms. (J) Dependence of SKN-1::GFP accumulation in GSC(−) animals on FAT-6/7-mediated FA desaturation, and proteins that generate free unsaturated FAs (LIPL-1/-3 lipases), or transport them to the nucleus (LBP-6/7/8). (K) OA and coconut oil (CO) increase skn-1-dependent gst-4p::GFP expression in the intestine. lbp-8 RNAi reduces induction by OA. (A, C) Representative 10× GFP images. (D, H) Representative 40× GFP images of day-1 adults. (E, F, I–K) GFP quantification with high, medium, low scoring. Numbers above bars denote sample size. p < 0.001***.
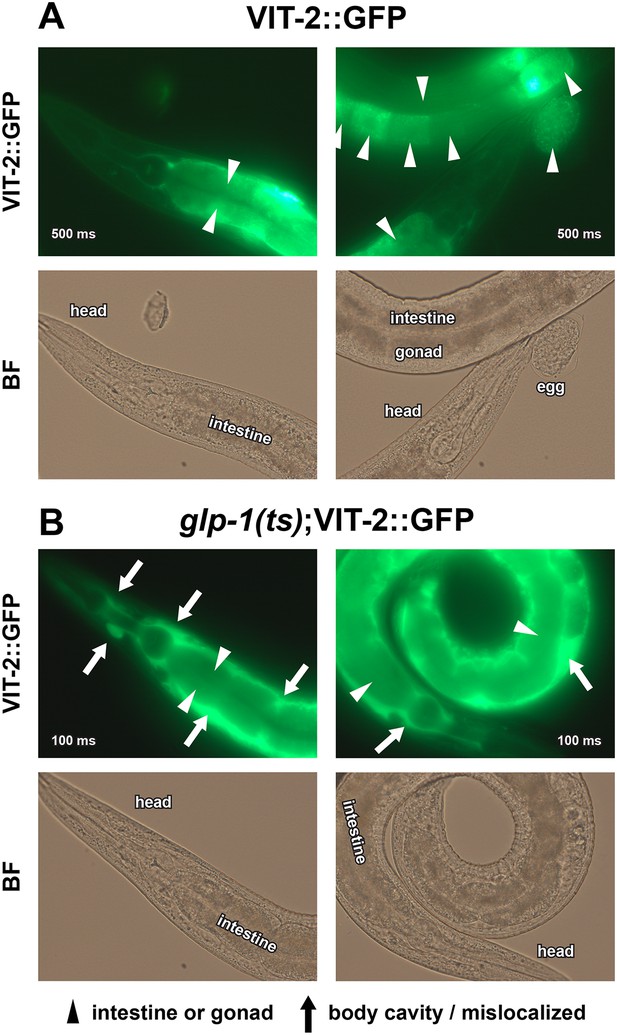
Enlarged VIT-2::GFP images.
40× GFP and BF images of (A) GSC(+) and (B) GSC(−) animals are shown. Note that glp-1(ts);VIT-2::GFP is presented with 5× lower exposure times due to increased VIT-2::GFP intensity in GSC(−) animals. All worms shown are day-1 adults raised at 25°C. Arrowheads indicate the normal distribution of VIT-2::GFP in the intestine and oocytes, and arrows indicate the ectopic accumulation of VIT-2::GFP seen in GSC(−) animals. Numbers superimposed on images denote GFP exposure times.
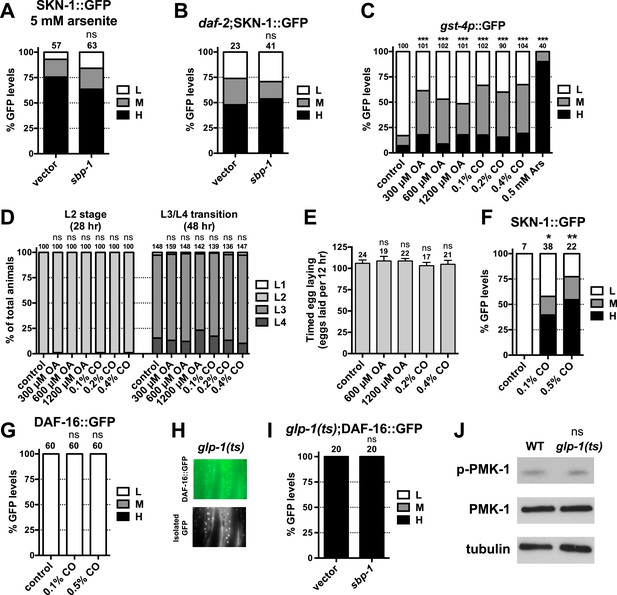
SKN-1 is activated in response to FA signaling.
Accumulation of SKN-1 in intestinal nuclei in (A) response to arsenite exposure for 30 min or (B) daf-2 mutants is not impaired by sbp-1 RNAi. (C) Effects of OA and CO doses on intestinal gst-4p::GFP expression, compared to AS treatment. (D) Larval development and (E) egg laying rate are not affected by OA or CO treatment. CO supplementation induces intestinal (F) SKN-1 nuclear accumulation but not (G) DAF-16 accumulation. (H, I) DAF-16 accumulation in glp-1(e2141ts) is unaffected by sbp-1 RNAi. (H) Representative 40× DIC images of day-1 adults. (J) PMK-1 (p38 kinase) phosphorylation is not affected by GSC removal, consistent with a previous report (Alper et al., 2010). PMK-1 phosphorylation is increased dramatically by AS oxidative stress and reflects activation of its kinase activity (Inoue et al., 2005). (A–C, F, G, I) GFP quantification with high, medium, low scoring. Numbers above bars denote sample size. p < 0.05*; p < 0.01**; p < 0.001***.
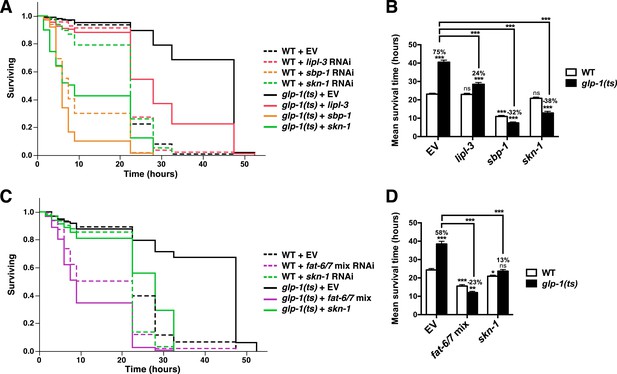
FA desaturation is required for GSC(−) stress resistance.
glp-1(ts) and control day-1 adult worms treated with lipl-3, sbp-1, fat-6/7 mix, skn-1, or empty vector RNAi were exposed to 5 mM AS. Knockdown of lipl-3 (A, B), and either sbp-1 (A, B) or fat-6/7 (C, D) abolished the increase in AS resistance seen in glp-1(ts) animals. (B, D) Data are represented as mean ± SEM. p < 0.05*; p < 0.01**; p < 0.001***. The interaction between glp-1(ts) and fat-6/7, lipl-3, sbp-1, and skn-1 were significant (p < 0.001). Additional information and statistics are provided in Table 2.
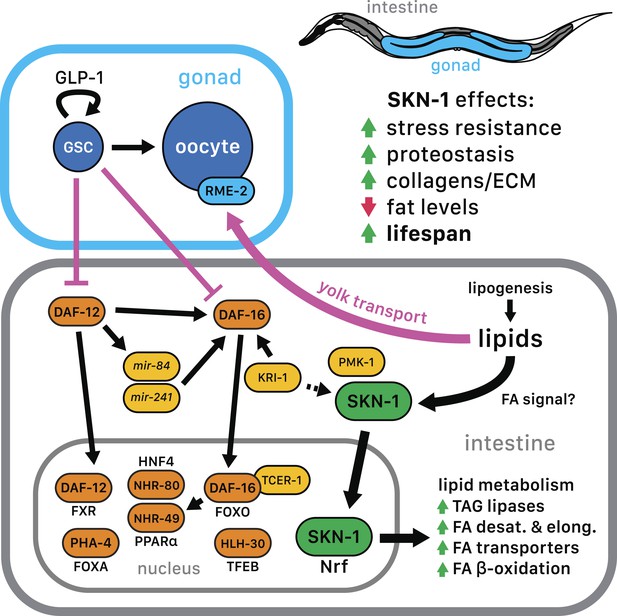
SKN-1 regulation in the GSC longevity pathway.
GSC absence results in activation of transcription factors in the intestine, with SKN-1 being regulated in parallel to DAF-12 and DAF-16. Yolk transport to oocytes is disrupted by GSC loss, resulting in lipid accumulation in the intestine and body cavity. The resulting SKN-1 activation requires OA, the FAT-6/7 FA desaturases, and the lysosomal lipases LIPL-1/3. This lipid-based signaling to SKN-1 depends partially upon LBP-8, which transports FAs from the lysosome to the nucleus. SKN-1 induces transcription of genes involved in stress resistance, detoxification, proteasome maintenance, extracellular matrix, and lipid metabolism, thereby reducing fat storage and increasing stress resistance and lifespan. Magenta denotes processes that are active in the presence of GSCs.
Tables
Lifespans
| Set | Strain | Mean lifespan ± SEM (days) | Median lifespan (days) | 75th % (days) | N | % Mean lifespan Ext. | p value |
|---|---|---|---|---|---|---|---|
| Lifespan at 25°C | |||||||
| Composite lifespan at 25°C with FUdR (from 2 replicates) | |||||||
| C1 | N2 | 15.40 ± 0.2 | 16 | 17 | 133/148 | – | – |
| skn-1(zu135) | 14.66 ± 0.2 | 15 | 16 | 158/167 | – | – | |
| glp-1(bn18ts) | 19.94 ± 0.3 | 20 | 22 | 164/171 | 29.48 | <0.0001* | |
| glp-1(bn18ts);skn-1(zu135) | 15.23 ± 0.2 | 15 | 18 | 149/170 | 3.89 | <0.0001† | |
| two-way ANOVA glp-1(ts) and skn-1 interaction | <0.0001 | ||||||
| Replicate lifespans at 25°C with FUdR | |||||||
| #1 | N2 | 13.99 ± 0.4 | 14 | 17 | 40/55 | – | – |
| skn-1(zu135) | 12.54 ± 0.3 | 13 | 14 | 41/50 | – | – | |
| glp-1(bn18ts) | 22.46 ± 0.8 | 25 | 27 | 50/55 | 60.54 | <0.0001* | |
| glp-1(bn18ts);skn-1(zu135) | 13.29 ± 0.4 | 12 | 14 | 53/70 | 6.00 | <0.0001† | |
| glp-1(ts) and skn-1 interaction | <0.0001 | ||||||
| #2 | N2 | 16.07 ± 0.2 | 16 | 17 | 93/93 | – | – |
| skn-1(zu135) | 15.42 ± 0.2 | 15 | 17 | 117/117 | – | – | |
| glp-1(bn18ts) | 18.86 ± 0.3 | 19 | 21 | 114/116 | 17.36 | <0.0001* | |
| glp-1(bn18ts);skn-1(zu135) | 16.42 ± 0.2 | 17 | 18 | 96/100 | 6.49 | <0.0001† | |
| glp-1(ts) and skn-1 interaction | 0.0002 | ||||||
| Lifespan at 20°C (25°C during development, downshifted to 20°C at D1 adulthood) | |||||||
| Lifespan at 20°C without FUdR | |||||||
| #3 | N2 | 20.36 ± 0.6 | 18 | 21 | 42/50 | – | – |
| skn-1(zu135) | 18.39 ± 0.5 | 18 | 18 | 27/47 | – | – | |
| glp-1(bn18ts) | 25.87 ± 1.3 | 24 | 33 | 35/66 | 27.06 | 0.0002* | |
| glp-1(bn18ts);skn-1(zu135) | 20.02 ± 0.6 | 18 | 24 | 34/55 | 8.86 | <0.0001† | |
| glp-1(ts) and skn-1 interaction | 0.0230 | ||||||
| Composite lifespan at 20°C with FUdR (from 2 replicates) | |||||||
| C2 | N2 | 20.51 ± 0.5 | 21 | 24 | 86/90 | – | – |
| skn-1(zu135) | 17.78 ± 0.4 | 19 | 20 | 88/94 | – | – | |
| glp-1(bn18ts) | 24.52 ± 0.6 | 25 | 28 | 81/104 | 19.55 | <0.0001* | |
| glp-1(bn18ts);skn-1(zu135) | 20.47 ± 0.5 | 20 | 24 | 91/98 | 15.13 | <0.0001† | |
| glp-1(ts) and skn-1 interaction | 0.1892 | ||||||
| Replicate lifespans at 20°C with FUdR | |||||||
| #4 | N2 | 18.09 ± 0.4 | 17 | 19 | 33/37 | – | – |
| skn-1(zu135) | 14.78 ± 0.5 | 14 | 17 | 31/35 | – | – | |
| glp-1(bn18ts) | 21.79 ± 1.0 | 21 | 28 | 28/50 | 20.45 | 0.0002* | |
| glp-1(bn18ts);skn-1(zu135) | 16.83 ± 0.3 | 17 | 18 | 36/43 | 13.87 | <0.0001† | |
| glp-1(ts) and skn-1 interaction | 0.1491 | ||||||
| #5 | N2 | 22.00 ± 0.6 | 23 | 25 | 53/53 | – | – |
| skn-1(zu135) | 19.40 ± 0.4 | 20 | 21 | 57/59 | – | – | |
| glp-1(bn18ts) | 25.91 ± 0.7 | 25 | 30 | 53/54 | 17.77 | <0.0001* | |
| glp-1(bn18ts);skn-1(zu135) | 22.80 ± 0.6 | 24 | 26 | 55/55 | 17.53 | <0.0001† | |
| glp-1(ts) and skn-1 interaction | 0.6607 | ||||||
-
Percent lifespan extension refers to glp-1(ts) vs wild type or skn-1 control. p values were calculated by log-rank test. Symbols denote effect relative to N2* or glp-1(ts)†. The interaction effect of glp-1(ts) and skn-1 was calculated by two-way ANOVA using mean lifespan. The last p value reflects the specific requirement of skn-1 for glp-1(ts) lifespan, as opposed to its effect on lifespan in general. Homozygous skn-1 mutants produce eggs that do not hatch because of a catastrophic defect in developmental patterning but do not exhibit known defects in the germline itself (Bowerman et al., 1992).
Stress resistance assays
| Set | Strain | Mean survival ± SEM (hrs) | Median survival (hrs) | 75th % survival (hrs) | N | % Mean survival Ext. | p value |
|---|---|---|---|---|---|---|---|
| AS (day 1 adulthood) 25°C continuous; RNAi from L1 | |||||||
| #1 | N2 + vector RNAi | 20.20 ± 1.0 | 22.5 | 22.5 | 122/122 | – | – |
| N2 + rme-2 RNAi | 39.51 ± 1.1 | 47.5 | 47.5 | 117/117 | 95.59 | <0.0001* | |
| #2 | N2 + vector RNAi | 26.82 ± 1.2 | 28.0 | 28.0 | 57/57 | – | – |
| N2 + rme-2 RNAi | 53.21 ± 0.9 | 52.5 | 71.5 | 252/252 | 98.44 | <0.0001* | |
| glp-1(bn18ts) + vector RNA | 53.06 ± 1.0 | 52.5 | 71.5 | 216/216 | 97.85 | <0.0001* | |
| #3 | N2 + vector RNAi (20°C) | 38.59 ± 0.6 | 47.5 | 47.5 | 281/281 | – | – |
| N2 + rme-2 RNAi | 63.00 ± 0.7 | 71.5 | 71.5 | 287/287 | 63.25 | <0.0001* | |
| #4 | N2 + vector RNAi (20°C) | 49.10 ± 0.8 | 46.0 | 64.0 | 306/306 | – | – |
| N2 + vector/skn-1 mix RNAi | 32.03 ± 0.6 | 40.0 | 40.0 | 271/271 | – | – | |
| N2 + vector/rme-2 mix RNAi | 65.38 ± 1.1 | 64.0 | 70.0 | 361/361 | 33.16 | <0.0001* | |
| N2 + rme-2/skn-1 mix RNAi | 33.76 ± 0.5 | 40.0 | 40.0 | 409/409 | 5.40 | <0.0001‡ | |
| two-way ANOVA rme-2 and skn-1 interaction | <0.0001 | ||||||
| #5 | N2 + vector RNAi | 23.18 ± 0.5 | 22.5 | 22.5 | 125/125 | – | – |
| N2 + lipl-3 RNAi | 23.04 ± 0.6 | 22.5 | 28.0 | 139/139 | – | – | |
| N2 + sbp-1 RNAi | 11.01 ± 0.6 | 7.5 | 22.5 | 163/163 | – | – | |
| N2 + skn-1 RNAi | 20.88 ± 0.7 | 22.5 | 28.0 | 115/115 | – | – | |
| glp-1(bn18ts) + vector RNAi | 40.58 ± 1.1 | 47.5 | 47.5 | 105/105 | 75.05 | <0.0001* | |
| glp-1(bn18ts) + lipl-3 RNAi | 28.52 ± 1.0 | 28.0 | 32.5 | 178/178 | 23.77 | <0.0001† | |
| glp-1(bn18ts) + sbp-1 RNAi | 7.50 ± 0.5 | 6.0 | 7.5 | 138/138 | −31.88 | <0.0001† | |
| glp-1(bn18ts) + skn-1 RNAi | 12.90 ± 0.9 | 9.0 | 22.5 | 129/129 | −38.21 | <0.0001† | |
| glp-1(ts) and lipl-3 interaction | <0.0001 | ||||||
| glp-1(ts) and sbp-1 interaction | <0.0001 | ||||||
| glp-1(ts) and skn-1 interaction | <0.0001 | ||||||
| #6 | N2 + vector RNAi | 24.36 ± 0.8 | 22.5 | 28.0 | 121/121 | – | – |
| N2 + fat-6/7 mix RNAi | 15.56 ± 0.8 | 22.5 | 22.5 | 159/159 | – | – | |
| N2 + skn-1 RNAi | 20.94 ± 0.6 | 22.5 | 22.5 | 124/124 | – | – | |
| glp-1(bn18ts) + vector RNAi | 38.49 ± 1.5 | 47.5 | 47.5 | 98/98 | 58.03 | <0.0001* | |
| glp-1(bn18ts) + fat-6/7 mix RNAi | 12.04 ± 0.7 | 9.0 | 22.5 | 153/153 | −22.66 | <0.0001† | |
| glp-1(bn18ts) + skn-1 RNAi | 23.69 ± 0.9 | 28.0 | 32.5 | 116/116 | 13.13 | <0.0001† | |
| glp-1(ts) and fat-6/7 interaction | <0.0001 | ||||||
| glp-1(ts) and skn-1 interaction | <0.0001 | ||||||
| AS (day 3 adulthood) 25°C during development, 20°C from D1; RNAi from D1 | |||||||
| #7 | N2 + vector RNAi | 6.55 ± 0.3 | 7.0 | 8.0 | 104/104 | – | – |
| N2 + skn-1 RNAi | 5.08 ± 0.2 | 5.0 | 7.0 | 103/103 | – | – | |
| glp-1(bn18ts) + vector RNAi | 19.36 ± 0.1 | 19.0 | 20.0 | 97/97 | 195.57 | <0.0001* | |
| glp-1(bn18ts) + skn-1 RNAi | 7.86 ± 0.4 | 8.0 | 12.0 | 100/100 | 54.72 | <0.0001† | |
| glp-1(ts) and skn-1 interaction | <0.0001 | ||||||
| #8 | N2 + vector RNAi | 8.31 ± 0.4 | 9.0 | 10.0 | 98/98 | – | – |
| N2 + skn-1 RNAi | 6.89 ± 0.3 | 8.0 | 9.0 | 90/90 | – | – | |
| glp-1(bn18ts) + vector RNAi | 16.95 ± 1.0 | 20.0 | 26.0 | 81/81 | 103.97 | <0.0001* | |
| glp-1(bn18ts) + skn-1 RNAi | 9.90 ± 0.4 | 10.0 | 12.0 | 91/91 | 43.69 | <0.0001† | |
| glp-1(ts) and skn-1 interaction | <0.0001 | ||||||
| TBHP (day 3 adulthood) 25°C during development, 20°C from D1; RNAi from D1 | |||||||
| #9 | N2 + vector RNAi | 9.02 ± 0.3 | 9.0 | 11.0 | 84/108 | – | – |
| N2 + skn-1 RNAi | 6.16 ± 0.2 | 6.0 | 7.0 | 63/98 | – | – | |
| glp-1(bn18ts) + vector RNAi | 11.62 ± 0.3 | 11.0 | 13.0 | 61/61 | 28.82 | <0.0001* | |
| glp-1(bn18ts) + skn-1 RNAi | 6.48 ± 0.1 | 8.0 | 7.0 | 63/65 | 5.19 | <0.0001† | |
| glp-1(ts) and skn-1 interaction | <0.0001 | ||||||
| #10 | N2 | 4.29 ± 0.2 | 4.0 | 4.0 | 65/94 | – | – |
| skn-1(zu135) | 4.51 ± 0.1 | 5.0 | 3.0 | 73/74 | – | – | |
| glp-1(bn18ts) | 6.25 ± 0.2 | 6.0 | 4.0 | 73/74 | 45.69 | <0.0001* | |
| glp-1(bn18ts);skn-1(zu135) | 4.78 ± 0.1 | 5.0 | 4.0 | 73/75 | 5.99 | <0.0001† | |
| glp-1(ts) and skn-1 interaction | <0.0001 | ||||||
-
Survival after sodium arsenite (AS) or tert-butyl hydroperoxide (TBHP) treatment was assayed in adult animals. The increase in oxidative stress resistance of glp-1(ts) germline stem cell (GSC(−)) animals was impaired by loss of fat-6/7, lipl-3, sbp-1, and skn-1. Representative assays are shown. Percent survival extension refers to glp-1(ts) or rme-2 RNAi vs the matching wild type or skn-1 control. p values were calculated by log-rank test. Symbols denote effect relative to N2*, glp-1(ts)†, or rme-2 RNAi‡. The interaction effect of GSC(−) or rme-2 with skn-1, or fat-6/7, lipl-3, and sbp-1 were calculated by two-way ANOVA using mean lifespan. The last p value reflects the specific requirement of each gene for GSC(−) or rme-2 stress resistance as opposed to its effect on stress resistance in general.
C. elegans strains used in this study
| Code | Genetic background | Transgene | Reference |
|---|---|---|---|
| AA003 | daf-12(rh61rh411) X | – | (Shen et al., 2012) |
| AA983 | glp-1(e2141ts) III; daf-12(rh61rh411) X | – | (Shen et al., 2012) |
| AA1049 | mir-241(n4315) V; mir-84(n4037) X | – | (Shen et al., 2012) |
| AA1709 | glp-1(e2141ts) III; mir-241(n4315) V; mir-84(n4037) X | – | (Shen et al., 2012) |
| AA2735 | glp-1(e2141ts) III | – | (Shen et al., 2012) |
| CF1903 | glp-1(e2141ts) III | – | (Berman and Kenyon, 2006) |
| CF1935 | daf-16(mu86) I; glp-1(e2141ts) III | muIs109[daf-16p::GFP:: DAF-16 + odr-1p::RFP] X | (Berman and Kenyon, 2006) |
| CL2166 | N2 | dvIs19[pAF15 (gst-4p::GFP::NLS)] III | (Link and Johnson, 2002) |
| DH1033 | sqt-1(sc103) II | bIs1[vit-2p::VIT-2:: GFP + rol-6(su1006)] X | (Grant and Hirsh, 1999) |
| EU31 | skn-1(zu135) IV | – | (Bowerman et al., 1992) |
| LD001 | N2 | ldIs7[SKN-1b/c:: GFP + rol-6(su1006)] | (An and Blackwell, 2003) |
| LD002 | N2 | ldIs1[SKN-1b/c:: GFP + rol-6(su1006)] | (An and Blackwell, 2003) |
| LD1025 | daf-2(e1370) III | ldIs7[SKN-1b/c:: GFP + rol-6(su1006)] | (Tullet et al., 2008) |
| LD1425 | glp-1(bn18ts) III | ldIs1[SKN-1b/c:: GFP + rol-6(su1006)] | This study |
| LD1434 | glp-1(bn18ts) III; skn-1(zu135) IV | – | This study |
| LD1473 | kri-1(ok1251) I; glp-1(bn18ts) III | ldIs1[SKN-1b/c:: GFP + rol-6(su1006)] | This study |
| LD1474 | tcer-1(tm1452) II; glp-1(bn18ts) III | ldIs1[SKN-1b/c:: GFP + rol-6(su1006)] | This study |
| LD1548 | N2 | Is[dhs-3p::DHS-3:: GFP] I | (Zhang et al., 2012) |
| LD1549 | glp-1(bn18ts) III | Is[dhs-3p::DHS-3:: GFP] I | This study |
| LD1644 | sqt-1(sc103) II; glp-1(bn18ts) III | bIs1[vit-2p::VIT-2:: GFP + rol-6(su1006)] X | This study |
| LD1653 | glp-1(bn18ts) III | – | (Dorsett et al., 2009) Outcrossed from DG2389 |
| LD1744 | glp-1(bn18ts) III | ldEx119[pAF15 (gst-4p::GFP::NLS) + rol-6(su1006)] | This study |
| TJ356 | N2 | zIs356[daf-16p:: DAF-16a/b::GFP + rol-6] IV | (Henderson and Johnson, 2001) |
qRT-PCR primers used in this study
| Gene | Sequence | Annotation | Primer pair |
|---|---|---|---|
| gst-4 | K08F4.7 | Glutathione S-transferase | FWD: CCCATTTTACAAGTCGATGG REV: CTTCCTCTGCAGTTTTTCCA |
| F20D6.11 | F20D6.11 | Flavin-adenine dinucleotide (FAD)-binding oxidoreductase | FWD: GGAAATTCTCGGTAGAATCGAA REV: ACGATCACGAACTTCGAACA |
| nit-1 | ZK1058.6 | Nitrilase | FWD: AATCCTCCGACTATCCCTTG REV: AGCGAATCGTTTCTTTTGTG |
| rpn-6.1 | F57B9.10 | 19S non-ATPase subunit | FWD: AATATTGGAAAAGCACCTGAAATGT REV: TTTGATGTGGAAGTGAAGTCATTGT |
| lipl-3 | R11G11.14 | Lysosomal triglyceride lipase | FWD: ATGGGCAGGCAAATCCACCA REV: AGTTGTTCTGCGCAATTATA |
| *cdc-42 | R07G3.1 | Housekeeping gene | FWD: CTGCTGGACAGGAAGATTACG REV: CTCGGACATTCTCGAATGAAG |
| *Y45F10D.4 | Y45F10D.4 | Housekeeping gene | FWD: GTCGCTTCAAATCAGTTCAG CREV: GTTCTTGTCAAGTGATCCGACA |
-
Select primer sequences were obtained from previous publications (Robida-Stubbs et al., 2012; Vilchez et al., 2012; O'Rourke and Ruvkun, 2013).
Additional files
-
Supplementary file 1
(a) List of genes activated by GSC absence. The gene list is sorted by functional grouping, then by fold change of mRNA expression in glp-1(ts) relative to wild type. A fold change cutoff of 4 (p < 0.05; n = 1306) was used (see Figure 3—figure supplement 1 for rationale). A more conservative fold change cutoff of 5 that captured fewer genes (n = 615) was used for DAVID cluster analysis. FDR denotes false discovery rate. (b) List of genes activated by SKN-1 in wild-type animals. The gene list is sorted by functional grouping, then by fold change of mRNA expression in WT skn-1 RNAi-treated worms relative to WT control vector-treated worms. A fold change cutoff of 0.67 (n = 295) was used to enrich for higher confidence SKN-1 targets. (c) List of genes activated by SKN-1 in GSC(−) animals. The gene list is sorted by functional grouping, then by fold change of mRNA expression in glp-1(ts) skn-1 RNAi-treated worms relative to glp-1(ts) control vector-treated worms. A fold change cutoff of 0.67 (p < 0.05; n = 529) was used to enrich for higher confidence SKN-1 targets. (d) List of genes activated by GSC absence in a SKN-1-dependent manner. The gene list is sorted by functional grouping, then by fold change of mRNA expression in glp-1(ts) skn-1 RNAi-treated worms relative to glp-1(ts) control vector-treated worms. We employed a GSC(−) fold change cutoff of >4 and skn-1 RNAi cutoff of <0.67 to generate the list (see Figure 3—figure supplement 1 for GSC(−) FC cutoff rationale). Statistics were generated by min analysis. Predicted SKN-1 binding sites were determined using 1.5 kb upstream sequences from WBcel235 and the SKN-1 JASPAR matrix (Staab et al., 2013a). SKN-1::GFP ChIP-seq binding analysis was performed using the L1, L2, and L3 data sets available from modENCODE (Niu et al., 2011). Additional details are available in ‘Materials and methods’.
- https://doi.org/10.7554/eLife.07836.022





