LXRs link metabolism to inflammation through Abca1-dependent regulation of membrane composition and TLR signaling
Figures
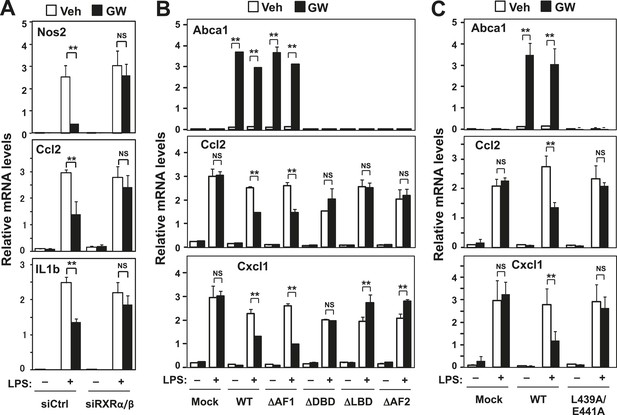
RXR and transactivation are required for LXR-dependent inflammatory repression.
(A) Bone marrow-derived macrophages from wild-type mice were transfected with siRNA targeting RXRα and RXRβ (siRXRαβ or control (siCtrl) for 48 hr, pretreated with GW3965 (1 µM) overnight, and then stimulated with LPS (10 ng/ml) for 4 hr. (B) Immortalized MEFs from Lxrα−/−Lxrβ−/− mice reconstituted with wild-type human LXRα, AF1-deletion mutant (∆AF1), DNA-binding domain deletion mutant (∆DBD), ligand-binging domain deletion mutant (∆LBD), AF2-deletion mutant (∆AF2) or control mock were pretreated with the LXR agonist GW3965 (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 4 hr. (C) Immortalized MEFs from Lxrα−/−Lxrβ−/− mice reconstituted with wild-type human LXRα, L439A/E441A mutant or control mock were pretreated with the LXR agonist GW3965 (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 4 hr. Gene expression was analyzed by real-time PCR. N = 4 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM.
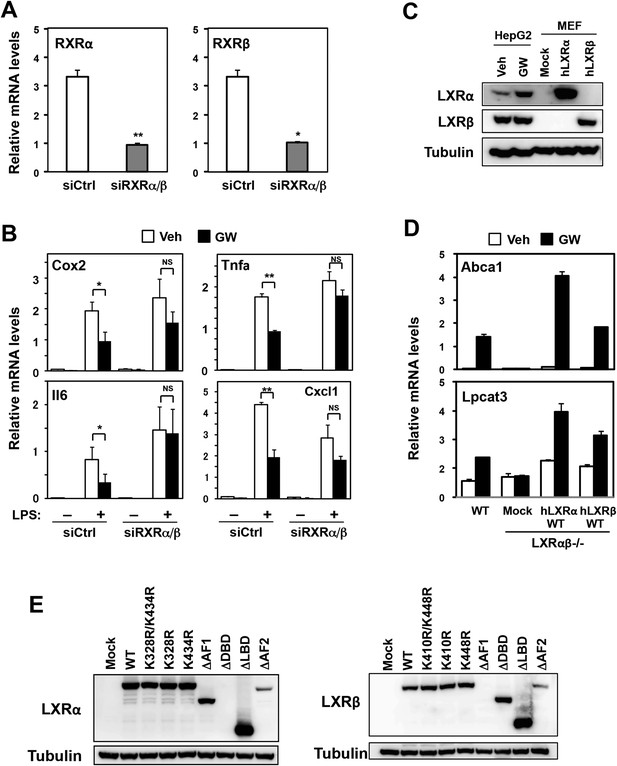
Effects of RXR knockdown on LXR-mediated inflammatory repression.
(A, B) Bone marrow-derived macrophages from wild-type mice were transfected with siRNA targeting RXRα and RXRβ (siRXRαβ or control (siCtrl) for 48 hr, pretreated with GW3965 (1 µM) overnight, and then stimulated with vehicle or LPS (10 ng/ml) for 4 hr as indicated. Gene expression was analyzed by real-time PCR. N = 4 per group. (C) Immunoblot analysis of LXRα and LXRβ protein in HepG2 cells treated with vehicle or GW3965 (2 µM) overnight and immortalized MEFs from Lxrα−/−Lxrβ−/− mice reconstituted with wild-type human LXRα, human LXRβ or control mock. (D) Immortalized MEFs from Lxrα−/−Lxrβ−/− mice reconstituted with wild-type human LXRα and LXRβ were treated with the LXR agonist GW3965 (1 µM) overnight. Gene expression was analyzed by real-time PCR. N = 4 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM. (E) Immunoblot analysis of LXR proteins in immortalized MEFs from Lxrα−/−Lxrβ−/− mice reconstituted with wild-type human LXRs or indicated mutants. Note, the epitope recognized by the LXRα antibody is in the DBD and that recognized by the LXRβ antibody is in the AF-1 domain. Therefore, these respective deletion mutants are not detected.
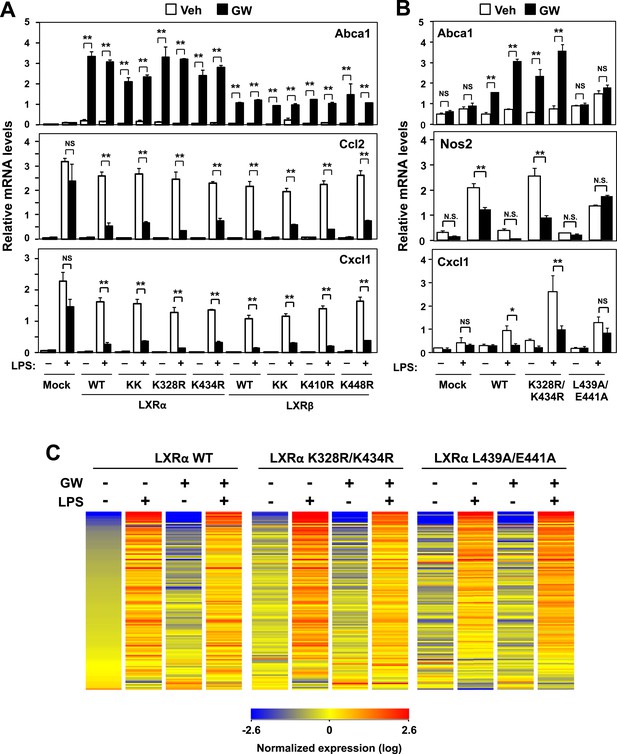
Transactivation but not sumoylation is required for LXR-mediated inflammatory repression.
(A) Immortalized MEFs from Lxrα−/−Lxrβ−/− mice reconstituted with wild-type human LXRα, sumoylation site mutants (K328R/K434R (KK), K328R, K434R), wild-type human LXRβ, sumoylation site mutants (K410R/K448R (KK), K410R, K448R), or mock control were pretreated with GW3965 (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 4 hr. (B, C) Immortalized bone marrow-derived macrophages from Lxrα−/−Lxrβ−/− mice reconstituted with wild-type human LXRα, K328R/K434R (KK) mutant, L439A/E441A mutant, or mock control were pretreated with GW3965 (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 4 hr. Gene expression was analyzed by real-time PCR (B) and Agilent microarrays (C). Selected genes that are annotated with the ‘Immune system process’ GO term from the array studies are presented as a heatmap (≧ twofold change). N = 4 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM.
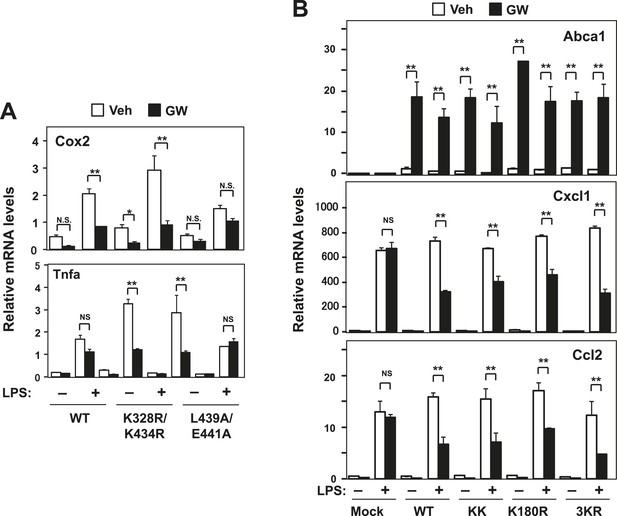
Effect of lysine mutants on LXR-mediated inflammatory repression.
(A) Immortalized bone marrow-derived macrophages from Lxrα−/−Lxrβ−/− mice reconstituted with wild-type human LXRα K328R/K434R mutant (KK), LXRα L439A/E441A mutant, or mock control were pretreated with GW3965 (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 4 hr. Gene expression was analyzed by real-time PCR. N = 4 per group. (B) Immortalized MEFs from Lxrα−/−Lxrβ−/− mice reconstituted with wild-type human LXRα, K328R/K434R mutant (KK), K180R mutant, K177R/K178R/K180R mutant (3 KR) or control mock were pretreated with the LXR agonist GW3965 (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 4 hr. Gene expression was analyzed by real-time PCR. N = 4 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM.
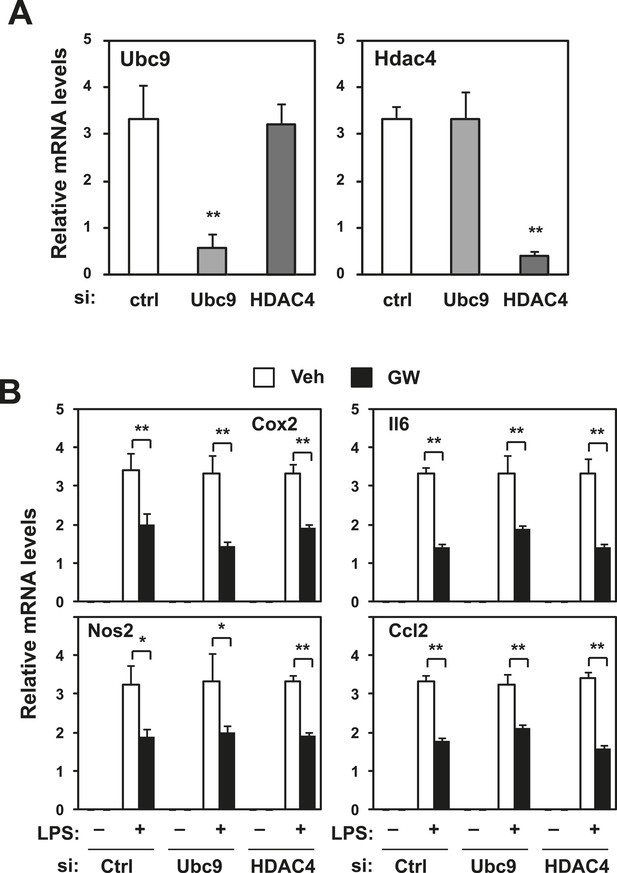
Repression does not require Ubc9 or Hdac4.
Immortalized bone marrow-derived macrophages were transduced with control siRNA or siRNA targeting Ubc9 or Hdac4 as indicated. (A) Validation of mRNA knockdown for Ubc9 or Hdac4. (B) Regulation of inflammatory gene expression by LXR agonists in presence or absence of Ubc9 or Hdac4. N = 4 per group. *p < 0.05, **p < 0.01. Error bars represent means ± SEM.
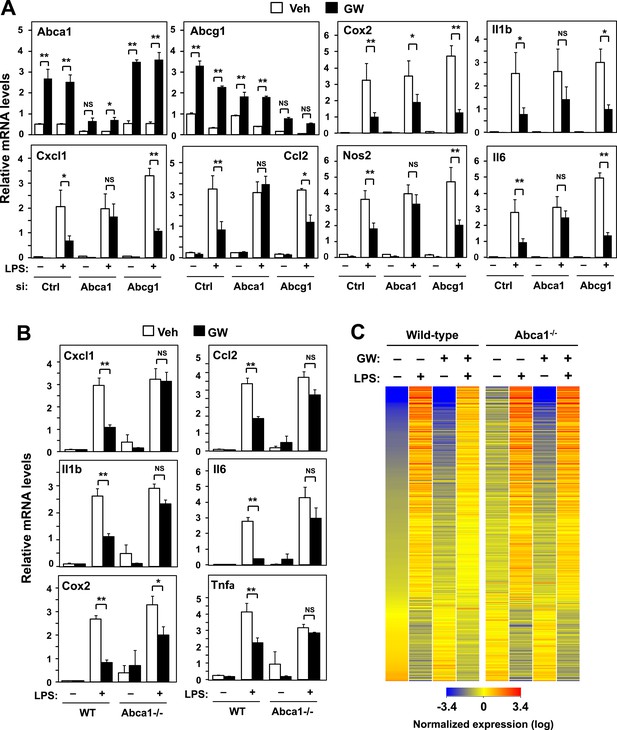
ABCA1 induction is critical for LXR-mediated repression.
(A) Bone marrow-derived macrophages from wild-type mice were transfected with siRNA targeting Abca1, Abcg1 or control (Ctrl) for 48 hr, pretreated with GW3965 (1 µM) overnight, and then stimulated with LPS (10 ng/ml) for 4 hr. (B, C) Bone marrow-derived macrophages from myeloid-specific Abca1−/− and control wild-type mice were pretreated with GW3965 (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 4 hr. Gene expression was analyzed by real-time PCR (B) and Agilent microarrays (C). Selected genes from the array studies that are annotated with the ‘Immune system process’ GO term are presented as a heatmap (≥twofold changes shown). N = 4 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM.
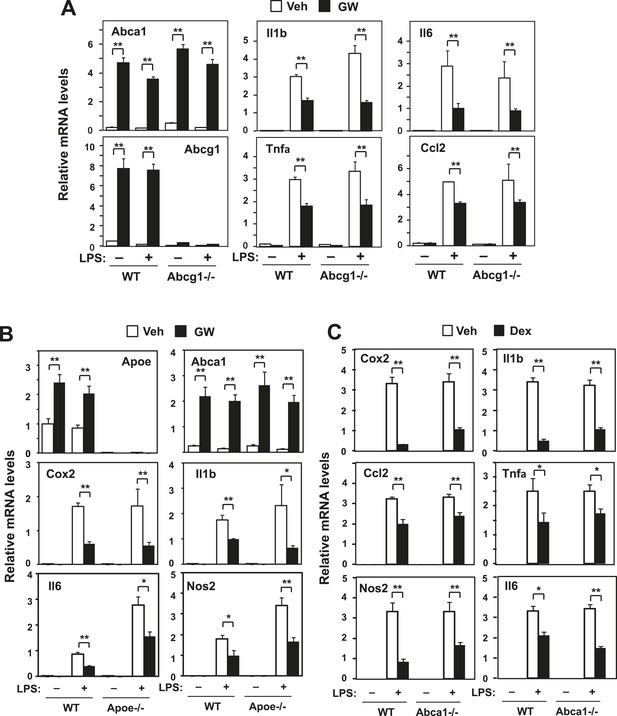
Loss of Abcg1 or ApoE does not compromise LXR-mediated repression.
(A) Bone marrow-derived macrophages from wild-type mice were transfected with siRNA targeting Abcg1 or control (Ctrl) for 48 hr, pretreated with GW3965 (1 µM) overnight, and then stimulated with LPS (10 ng/ml) for 4 hr. (B) Bone marrow-derived macrophages from wild-type or Apoe−/− mice were pretreated with GW3965 (1 µM) overnight, and then stimulated with LPS (10 ng/ml) for 4 hr. (C) Bone marrow-derived macrophages from myeloid-specific Abca1−/− and control wild-type mice were treated with dexamethasone (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 4 hr. Gene expression was analyzed by real-time PCR. N = 4 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM.
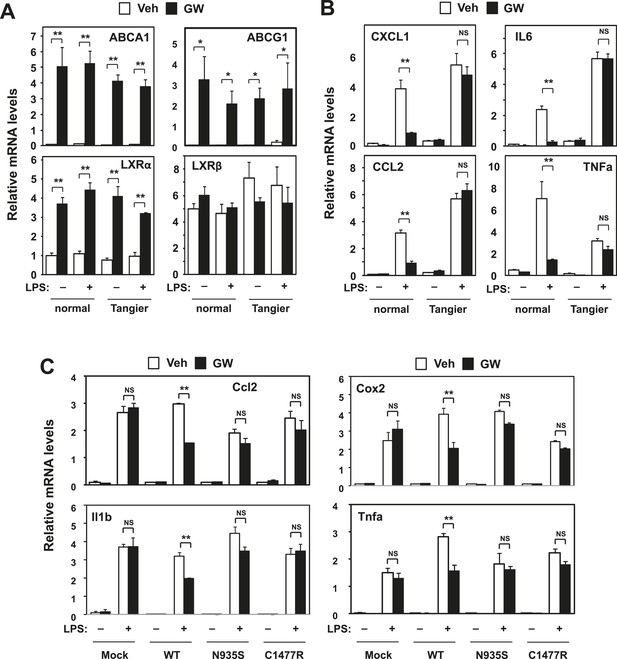
Intracellular cholesterol content affects LXR-mediated repression.
(A, B) Skin fibroblasts from a healthy donor (normal) and a Tangier disease patient (Tangier) were pretreated with GW3965 (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 4 hr. (C) Immortalized bone marrow-derived macrophages from Abca1−/− mice reconstituted with wild-type Abca1, N935S mutant, C1447R mutant or mock control were pretreated with GW3965 (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 4 hr. Gene expression was analyzed by real-time PCR. N = 4 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM.
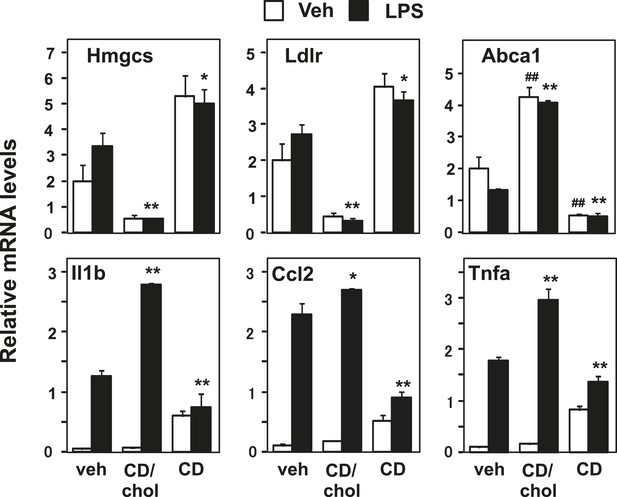
Manipulation of membrane cholesterol content affects inflammatory responses.
Bone marrow-derived macrophages were incubated with cyclodextrin cholesterol (CD-Chol, 100 µM) or hydroxypropyl-β-cyclodextrin (CD, 10 mM) for 1 hr and stimulated with LPS (10 ng/ml) for 4 hr. Gene expression was analyzed by real-time PCR. N = 4 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM.
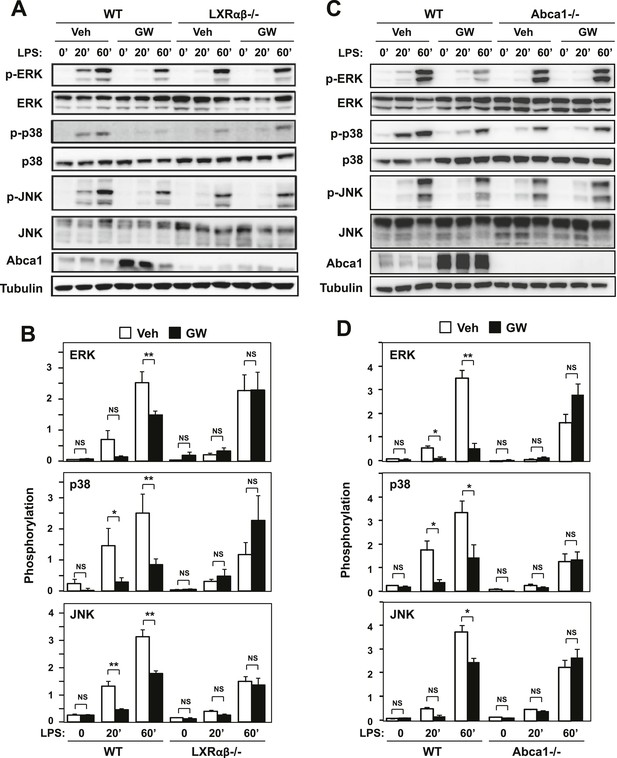
Ligand activation of LXR inhibits LPS-induced MAP kinase activation through Abca1 induction.
(A–D) Bone marrow-derived macrophages from Lxrα−/−Lxrβ−/− and control wild-type mice (A, B), or bone marrow-derived macrophages from myeloid-specific Abca1−/− and control wild-type mice (C, D) were pretreated with GW3965 (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 20 min or 1 hr. Whole cell lysates were harvested and protein expression was analyzed by immunoblotting with the indicated antibodies (A, C). Protein expression was quantified by Image Quant TL7.0 (B, D). N = 4–6 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM.
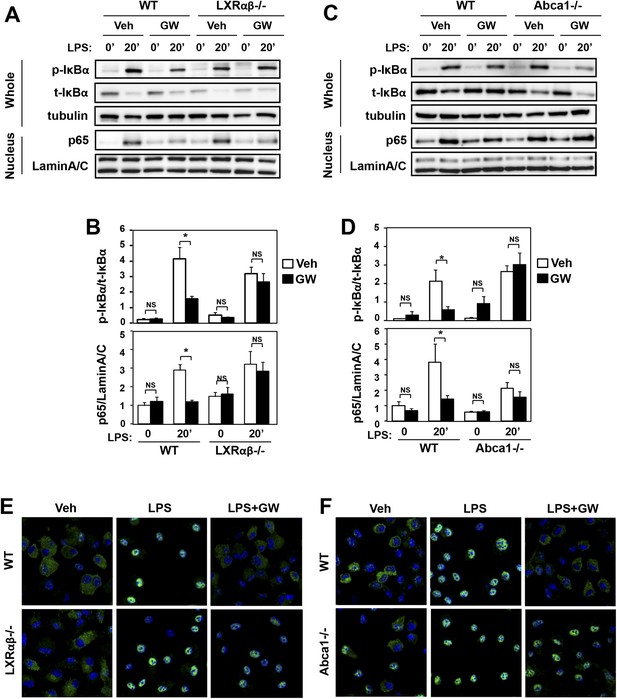
Ligand activation of LXR inhibits LPS-induced NF-κB activation through Abca1 induction.
(A–F) Bone marrow-derived macrophages from Lxrα−/−Lxrβ−/− and control wild-type mice (A, B, E), or bone marrow-derived macrophages from myeloid-specific Abca1−/− and control wild-type mice (C, D, F) were pretreated with GW3965 (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 20 min. Whole cell lysates and nuclear lysates were harvested and protein expression was analyzed by immunoblotting with the indicated antibodies (A, C). Protein expression was quantified by Image Quant TL7.0 (B, D). N = 4–6 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM. Nuclear translocation of p65 was assessed by staining of p65 (green) and DAPI (blue) (E, F).
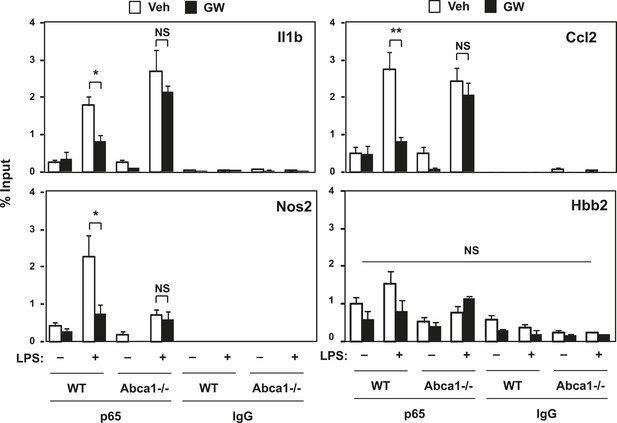
LXR activation decreases p65 occupancy on inflammatory gene promoters.
Recruitment of p65 to inflammatory gene promoters was assessed by ChIP-qPCR assays. Chromatin from wild-type or Abca1−/− cells was precipitated with p65 antibody or control IgG. N = 4 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM.
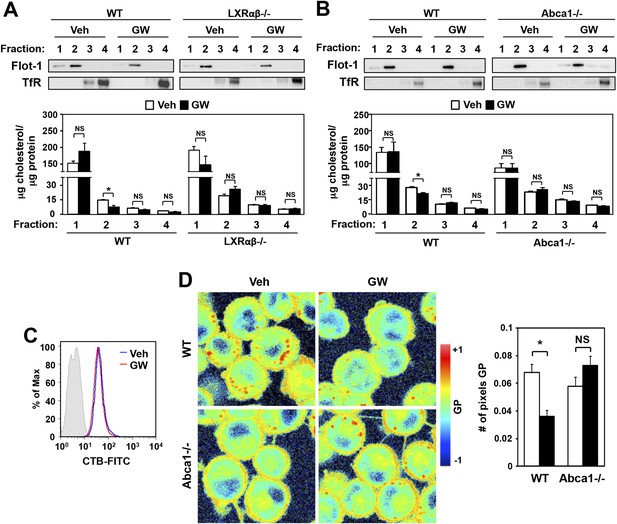
Ligand activation of LXR decreases cholesterol content in lipid rafts and increases membrane lipid mobility.
(A, B) Bone marrow-derived macrophages from Lxrα−/−Lxrβ−/− and control wild-type mice (A), or bone marrow-derived macrophages from myeloid-specific Abca1−/− and control wild-type mice (B) were pretreated with GW3965 (1 µM) overnight. Lipid raft and non-raft fractions were isolated using the detergent method and each fraction was analyzed by Western blotting. The free cholesterol concentration in each fraction was determined and normalized to protein concentration. N = 5 per group. (C) The abundance of lipid rafts in the plasma membrane was analyzed by flow cytometry after staining with cholera toxin B (CTB). (D) Wild-type or Abca1−/− iBMDM were treated with oxidized LDL (50 µg/ml) for 48 hr, and GW3965 (1 µM) overnight. GP images (left) were obtained from fluorescence-lifetime imaging microscopy (FLIM). The GP scale used to pseudocolor the intensity image is shown at right. Plasma membrane fluidity was determined with GP and number of pixel in plasma membrane (right). *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM.
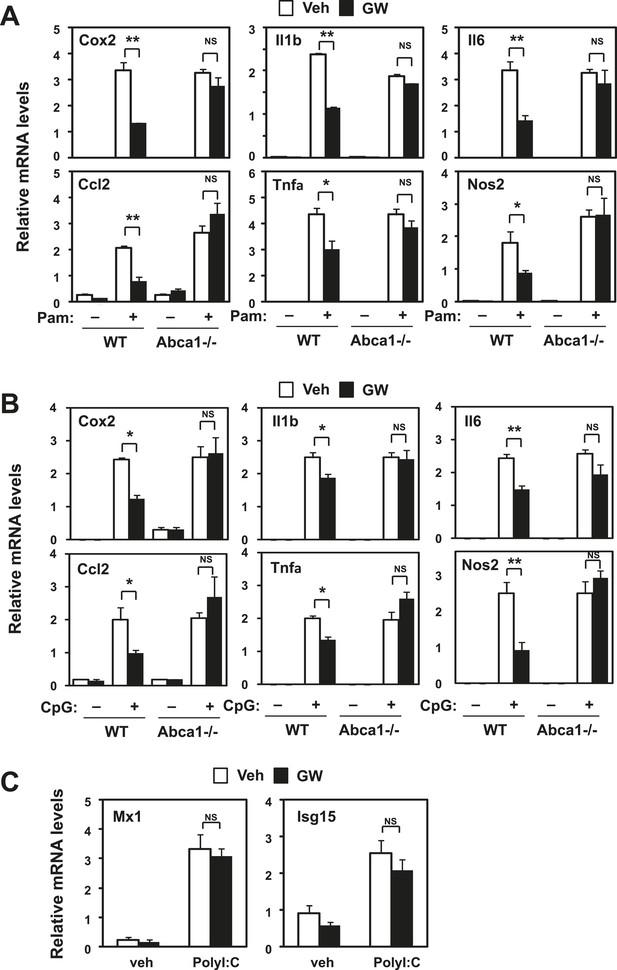
Inhibition of TLR2 and TLR9 signaling by LXR requires Abca1.
Bone marrow-derived macrophages from myeloid-specific Abca1−/− and control wild-type mice (A, B, C) were pretreated with GW3965 (1 µM) overnight, followed by stimulation with Pam3CSK4 (100 ng/ml), LPS (10 ng/ml), CpG (1 µM) or polyI:C (10 µg/ml) for 4 hr as indicated. Gene expression was analyzed by real-time PCR. N = 4 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM.
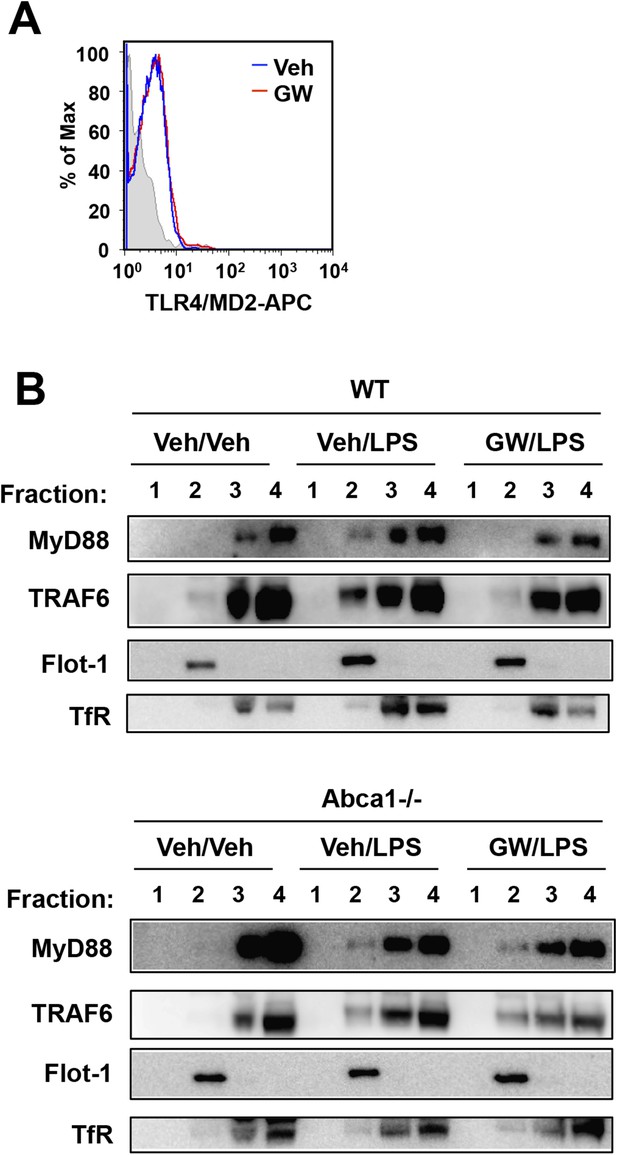
LXR activation inhibits recruitment of MyD88 and TRAF6 to lipid rafts.
(A) Wild-type iBMDM were pretreated with GW3965 (1 µM) overnight and TLR4 expression was analyzed by flow cytometry. (B) Wild-type or Abca1−/− iBMDM were pretreated with GW3965 (1 µM) overnight, followed by stimulation with LPS (10 ng/ml) for 10 min. Lipid raft and non-raft fractions were isolated using detergent method and each fraction was analyzed by Western blotting. Results are representative of two independent experiments.
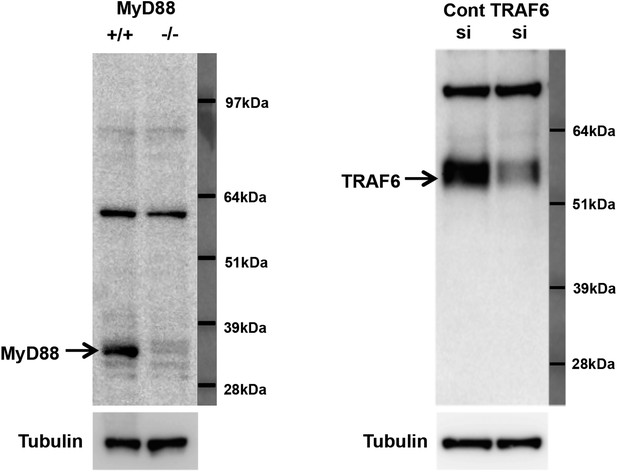
Specificity of MyD88 and TRAF6 antibodies.
(A) Immunoblot analysis of wild-type and MyD88−/− bone marrow macrophages. (B) Immunoblot analysis of wild-type bone marrow macrophages transduced with siRNA targeting TRAF6 or control siRNA.
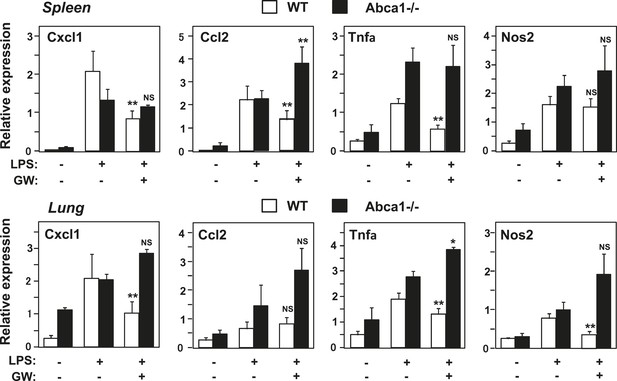
Abca1 contributes to LXR anti-inflammatory effects in vivo.
Myeloid-specific Abca1−/− and control wild-type mice were gavaged with GW3965 (40 mg/kg) for 3 days, followed by challenge with LPS (1 mg/kg) by intraperitoneal injection. 2 hr later, spleens (upper) and lungs (lower) were harvested, RNA isolated and gene expression analyzed by real-time PCR. N = 3–4 per group. *p < 0.05, **p < 0.01, NS, not significant. Error bars represent means ± SEM.
Tables
qPCR primers
| Murine qPCR primers | Forward | Reverse |
|---|---|---|
| 36B4 | GGCCCTGCACTCTCGCTTTC | TGCCAGGACGCGCTTGT |
| Abca1 | CGTTTCCGGGAAGTGTCCTA | GCTAGAGATGACAAGGAGGATGGA |
| Abcg1 | TCACCCAGTTCTGCATCCTCT | GCAGATGTGTCAGGACCGAGT |
| Apoe | GACTTGTTTCGGAAGGAGCTG | CCACTCGAGCTGATCTGTCA |
| Cox-2 | CAGGTCATTGGTGGAGAGGTG | GGATGTGAGGAGGGTAGATCA |
| Cxcl1 | CACTGCACCCAAACCGAAGT | GGACAATTTTCTGAACCAAGGG |
| Hmgcs | GCCGTGAACTGGGTCGAA | GCATATATAGCAATGTCTCCT |
| Il-1b | AGAAGCTGTGGCAGCTACCTG | GGAAAAGAAGGTGCTCATGTCC |
| Il-6 | GCTACCAAACTGGATATAATCAGGA | CCAGGTAGCTATGGTACTCCAGAA |
| iNos | GCAGCTGGGCTGTACAAA | AGCGTTTCGGGATCTGAAT |
| Isg15 | CAGGACGGTCTTACCCTTTC | CGCTGCAGTTCTGTACCACT |
| Ldlr | AGGCTGTGGGCTCCATAGG | TGCGGTCCAGGGTCATCT |
| Mcp-1 | CATCCACGTGTTGGCTCA | GATCATCTTGCTGGTGAATGA |
| Mx-1 | AAACCTGATCCGACTTCACTTCC | TGATCGTCTTCAAGGTTTCCTTGT |
| Rxra | ACCGCTCCATAGCTGTGAAAG | TGAGCGCTGTTCCGGTGTA |
| Rxrb | GCCACTGGCATGAAAAGG | ATCTCCATCCCCGTCTTTGT |
| Srebp-1c | GGAGCCATGGATTGCACATT | GGCCCGGGAAGTCACTGT |
| Tnfa | TCTTCTCATTCCTGCTTGTGG | GGTCTGGGCCATAGAACTGA |
| Human qPCR primers | Forward | Reverse |
|---|---|---|
| 36B4 | CCACGCTGCTGAACATGCT | TCGAACACCTGCTGGATGAC |
| Abca1 | GCCTGCTAGTGGTCATCCTG | CCACGCTGGGATCACTGTA |
| Abcg1 | ATGTCAGGTATGGGTTCGAAG | TCTGGTCGATGTCACAGTGC |
| Cxcl1 | TCAAGAATGGGCGGAAAGC | CAGCATCTTTTCGATGATTTTC |
| IL-6 | CCAGGAGCCCAGCTATGAAC | CCCAGGGAGAAGGCAACTG |
| Lxra | AAGCCCTGCATGCCTACGT | TGCAGACGCAGTGCAAACA |
| Lxrb | TCGTGGACTTCGCTAAGCAA | GCAGCATGATCTCGATAGTGGA |
| Mcp-1 | AGAAGCTGTGATCTTCAAGACCATT | TGCTTGTCCAGGTGGTCCAT |
| Tnfa | TCTTCTCGAACCCCGAGTGA | CCTCTGATGGCACCACCAG |
ChIP primers
| Murine ChIP primers | Forward | Reverse |
|---|---|---|
| Hbb2 | AGGTGCACCATGATGTCTGT | AGCAGGGTCAGTTGCTTCTT |
| Il-1b | GGACAATTGTGCAGATGGTG | CCTACCTTTGTTCCGCACAT |
| iNos | GGAGTGTCCATCATGAATGAG | CAACTCCCTGTAAAGTTGTGACC |
| Mcp-1 | TCCAGGGTGATGCTACTCCT | AGTGAGAGTTGGCTGGTGCT |





