An effector of the Irish potato famine pathogen antagonizes a host autophagy cargo receptor
Figures
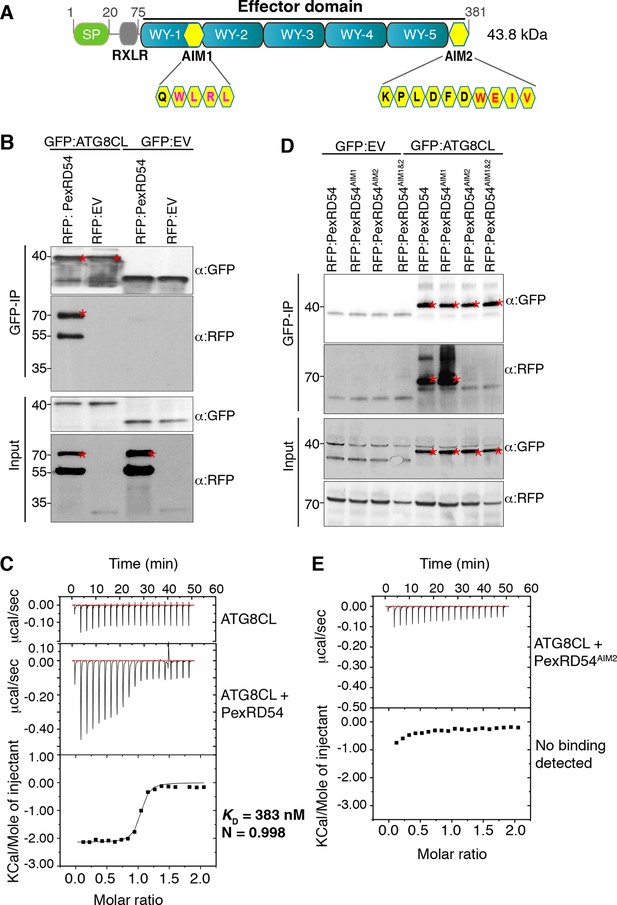
PexRD54 binds to ATG8CL via a C-terminal ATG8 interacting motif (AIM).
(A) Domain organization of PexRD54. PexRD54 is a canonical RXLR effector with five WY folds (Win et al., 2012). The amino acid sequences of candidate AIMs are highlighted in yellow color and indicated in brackets. (B) Validation of PexRD54-ATG8CL association in planta. RFP:PexRD54 or RFP:EV (Empty vector) were transiently co-expressed with GFP:ATG8CL or GFP:EV in N. benthamiana leaves. Immunoprecipitates (IPs) obtained with anti-GFP antiserum and total protein extracts were immunoblotted with appropriate antisera. Stars indicate the expected band sizes. (C) PexRD54 binds ATG8CL in vitro. The binding affinity of PexRD54 to ATG8CL was determined using isothermal titration calorimetry (ITC). Upper panel shows heat differences upon injection of ATG8CL into buffer or PexRD54 and the bottom panel show integrated heats of injection (■) and the best fit (solid line) to a single site binding model using MicroCal Origin. KD=383 nM, N=0.998, ΔH= −8.966 kJ.mol-1 and ΔS = 0.092 J.mol-1.K-1. The values of KD, N, ΔH and ΔS are representative of two independent ITC experiments. (D) ATG8 Interacting Motif 2 (AIM2) mediates ATG8CL binding in planta. RFP:PexRD54, RFP:PexRD54AIM1, RFP:PexRD54AIM2 or RFP:PexRD54AIM1&AIM2 were transiently co-expressed with GFP:ATG8CL or GFP:EV in N. benthamiana leaves. IPs obtained with anti-GFP antiserum and total protein extracts were immunoblotted with appropriate antisera. Stars indicate the expected band sizes. (E) AIM2 mediates ATG8CL binding in vitro. The binding affinity of PexRD54AIM2 to ATG8CL was determined using ITC. Upper panel shows heat differences upon injection of PexRD54AIM2 and the bottom panel show integrated heats of injection (■). No binding was detected between PexRD54AIM2 and ATG8CL.
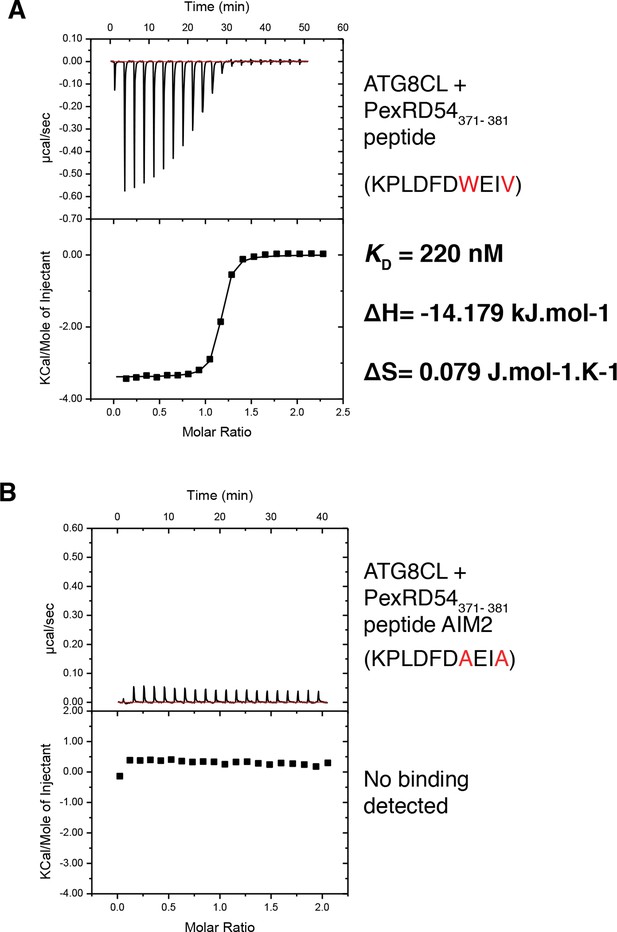
A synthetic peptide composed of the last ten amino acids of PexRD54 is sufficient for binding to ATG8CL.
(A) A synthetic peptide (KPLDFDWEIV) containing the AIM of PexRD54 binds ATG8CL as documented by ITC. (B) A mutant AIM peptide (KPLDFDAEIA) was used as negative control and did not bind ATG8CL. Heat differences upon injection of peptides and integrated heats of injection are shown for both peptides. The values of KD, ΔH and ΔS are representative of two independent experiments.
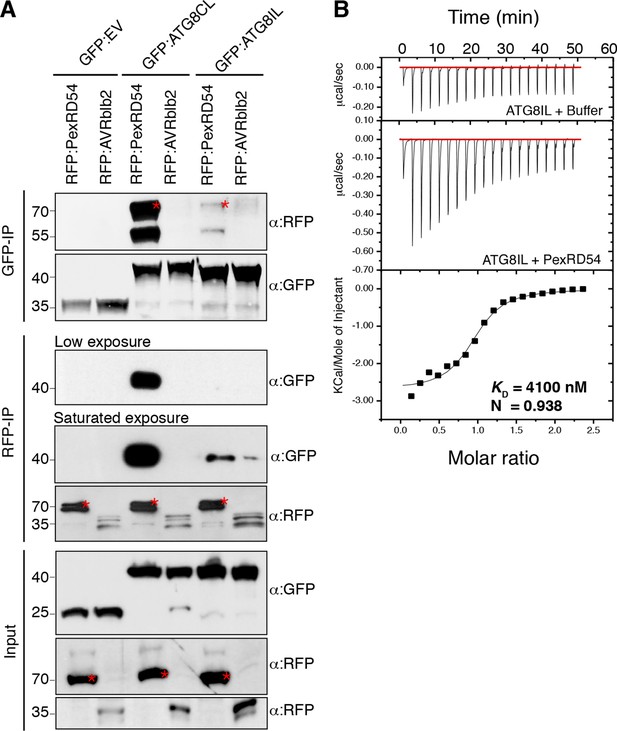
PexRD54 has higher binding affinity to ATG8CL than ATG8IL.
(A) RFP:PexRD54, RFP:AVRblb2 or RFP:EV were transiently co-expressed with GFP:ATG8CL, GFP:ATG8IL or GFP:EV in N. benthamiana leaves and proteins were extracted two days after infiltration and used in immunoprecipitation experiments (IPs). IPs obtained with anti-GFP or anti-RFP antisera and total protein extracts were immunoblotted with appropriate antisera. RFP:AVRblb2 (Bozkurt et al., 2011), an RFP fusion to a different P. infestans RXLR effector, did not associate with ATG8CL or ATG8IL. Both the GFP and RFP IPs indicate higher binding affinity of PexRD54 to ATG8CL than ATG8IL. Stars indicate the expected band sizes. (B) PexRD54 has lower binding affinity to ATG8IL in vitro. The binding affinity of PexRD54 to ATG8IL was determined using isothermal titration calorimetry (ITC). Upper panel shows heat generated upon injection of ATG8IL into buffer or PexRD54 and lower panel shows integrated heats of injection (■) and the best fit (solid line) to a single site binding model using MicroCal Origin. The values of KD = 4100 nM, N = 0.938, ΔH = −11.305 kJ.mol-1 and ΔS= 0.064 J.mol-1K-1 are representative values of two independent ITC experiments. The data show that PexRD54 binds ATG8CL with ~10 times higher affinity than ATGIL.

Amino acid sequence alignment of ATG8 proteins from Arabidopsis thaliana (At), Solanum tuberosum (St), Solanum lycopersicum (Sl) and Nicotiana benthamiana (Nb).
Sequences were aligned using ClustalW and visualized using Boxshade v3.21. Identical amino acids are highlighted in black and amino acids with similar biochemical properties are shown in grey. Red arrowhead points to ATG8CL and blue arrowhead points to ATG8IL.
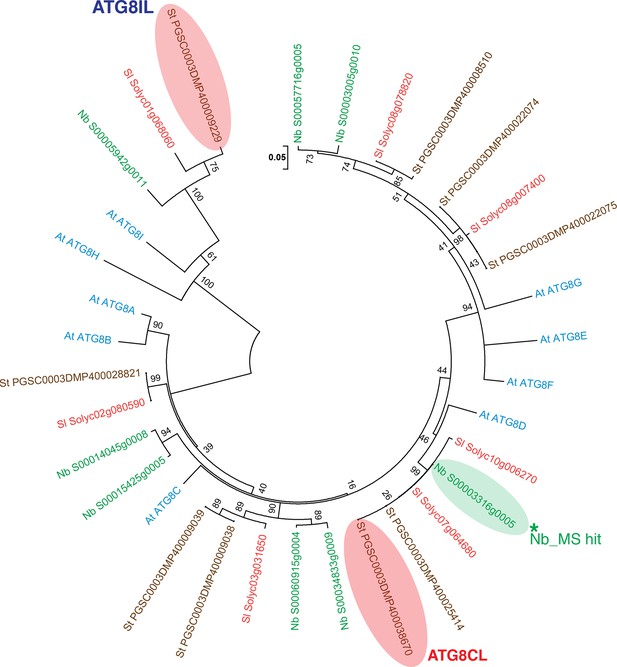
Phylogenetic analysis of ATG8 proteins.
A neighbor-joining tree was generated with ATG8 sequences obtained from genomes of A. thaliana, S. tuberosum, S. lycopersium, and N. benthamiana, using MEGA5 with a 1000 bootstrap value. ATG8CL and ATG8IL are highlighted in pink and the ATG8CL variant that was identified in N. benthamiana is highlighted in green.
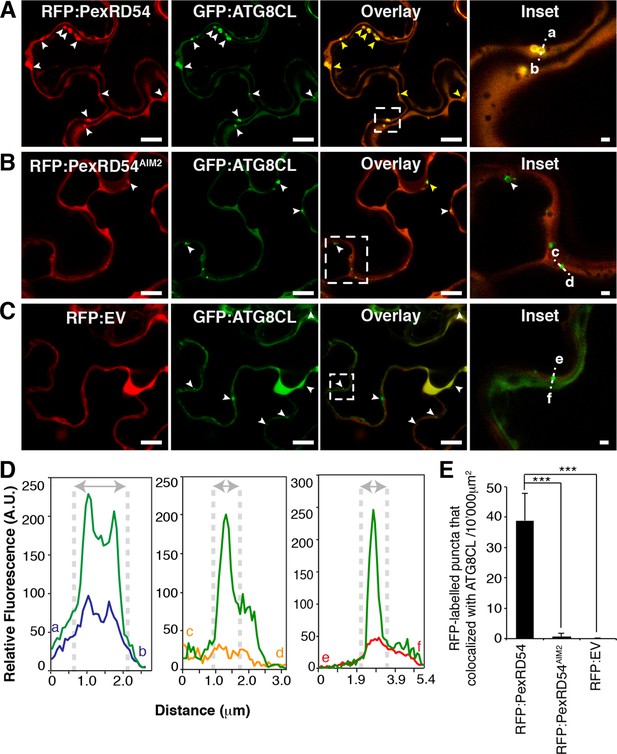
PexRD54 localizes to ATG8CL labelled autophagosomes.
Transient co-expression of GFP:ATG8CL with (A) RFP:PexRD54, (B) RFP:PexRD54AIM2 and (C) RFP control (RFP:EV) in N. benthamiana leaves. Confocal micrographs show single optical sections of RFP:PexRD54, RFP:PexRD54AIM2 and RFP:EV in red, GFP:ATG8CL in green and the overlay indicating colocalization in yellow. White arrowheads point to punctate structures and yellow arrowheads point to puncta where GFP and RFP signals overlap. Far right panels highlight the dotted square region focusing on GFP:ATG8CL labelled puncta in overlaid GFP/RFP channels. Scale bar = 10 μm; scale bar in inset = 1 μm. (D) The intensity plots represent relative GFP and RFP fluorescence signals along the dotted line connecting points a-b, c-d and e-f that span GFP:ATG8CL marked puncta at far right panels. GFP:ATG8CL fluorescence intensity peak overlapped with fluorescence intensity peak of RFP:PexRD54 (left panel) but not with RFP:PexRD54AIM2 (mid panel) or RFP:EV (right panel) validating the localization of RFP:PexRD54 at GFP:ATG8CL labelled autophagosomes. (E) Bar charts showing colocalization of GFP:ATG8CL puncta with RFP:PexRD54, RFP:PexRD54AIM2 or RFP:EV punctate structures. Data are representative of 500 individual images from two biological replicates. Each replicate consists of five independent Z stacks with 50 images each, acquired from five independent leaf areas. (***p<0.001). Error bars represent ± SD.
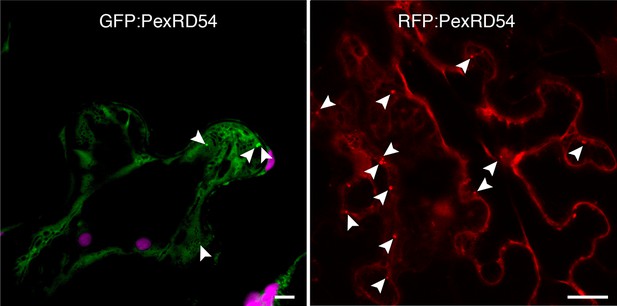
PexRD54 localizes to vesicle like structures.
GFP:PexRD54 or RFP:PexRD54 were transiently expressed in N. benthamiana and examined by confocal laser scanning microscopy 3 days post infiltration. Confocal micrographs show single optical sections of GFP:PexRD54 (left panel) and RFP:PexRD54 (right panel). With both tags, PexRD54 localized to vesicular structures pointed with arrowheads. Scale bar=10 μm.
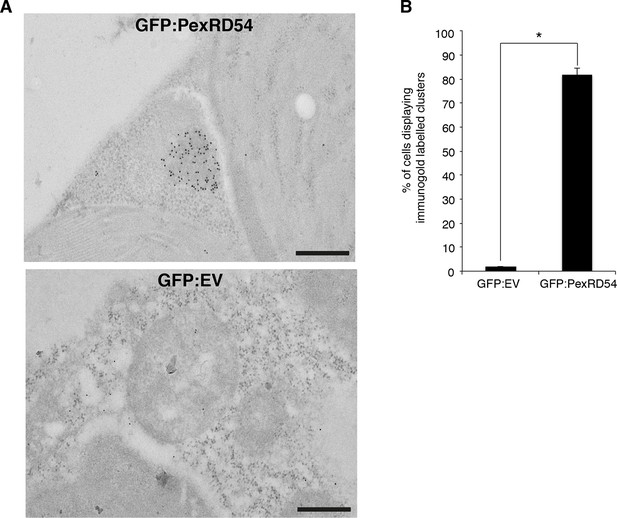
PexRD54 localizes to high electron dense structures.
(A) Transmission electron microscopy (TEM) imaging of PexRD54. N. benthamiana cells transiently expressing GFP:PexRD54 or GFP:EV were collected 3 days post infiltration and probed with anti-GFP antibodies conjugated to gold particles. GFP:PexRD54 labelling were mainly at high electron dense structures, whereas GFP:EV showed cytosolic distribution. Scale bar=200 nm (B) Bar charts showing electron dense labelling in GFP:EV and GFP:PexRD54 expressing samples (*p<0.05).
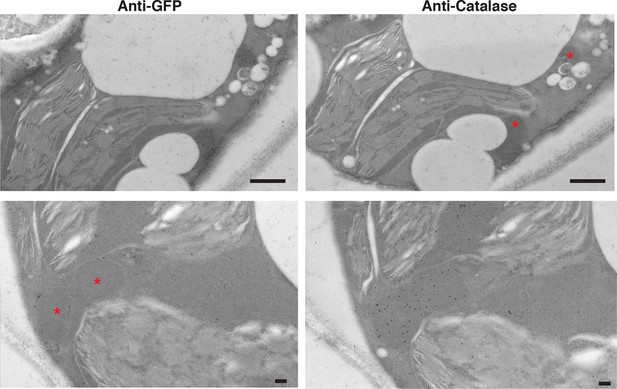
PexRD54 labelled high electron dense structures that are not peroxisomes.
Serial sections of N. benthamiana leaves transiently expressing GFP:PexRD54 were collected 3 days post infiltration and probed with anti-GFP and anti-catalase antibodies conjugated to gold particles. High electron dense structures labelled with GFP antibody were different than the regions labelled with catalase antibody, confirming that PexRD54 labelled structures are not peroxisomes. Stars indicate regions that are labelled by gold particles in the other image. Scale bar=500 nm
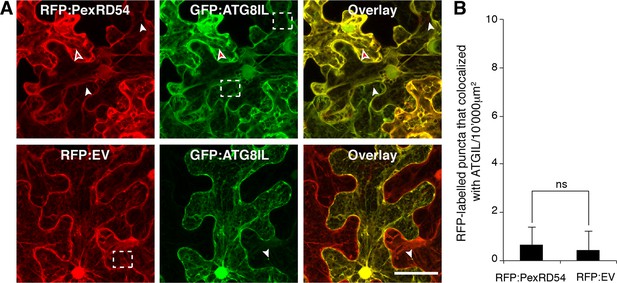
PexRD54 labelled puncta rarely include GFP:ATGIL signal.
GFP:ATG8IL was transiently co-expressed with RFP:PexRD54 or RFP:EV in N. benthamiana leaves and examined by confocal laser scanning microscopy 3 days post infiltration. Maximum intensity projection of confocal micrographs shows that RFP:PexRD54 only partially colocalizes with GFP:ATG8IL. White arrowheads point to vesicles labelled with only GFP:ATG8IL or RFP:PexRD54, whereas the red-filled arrowheads point to puncta where colocalization occurs. Regions where GFP or RFP labelled puncta do not overlap are indicated with dotted squares. Scale bar=50 μm (B) Bar charts showing the number of RFP-labelled puncta that colocalized with GFP:ATG8IL puncta. The data are representative of 500 individual images from two biological replicates. Each replicate consists of five independent Z stack images with 50 images each, acquired from five independent leaf areas. (ns=statistically not significant). Error bars represent ± SD.
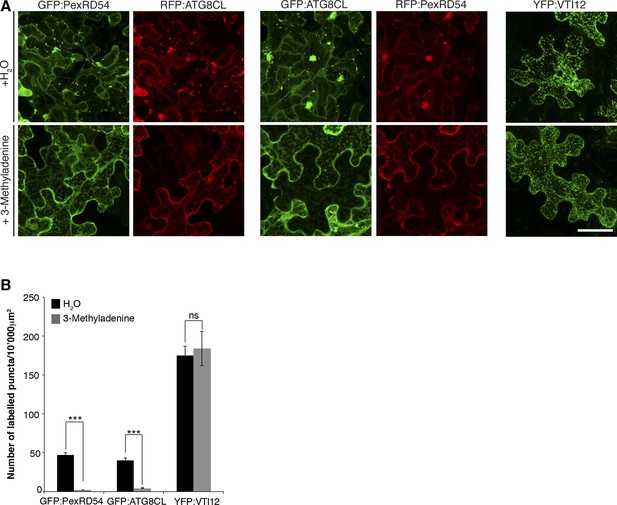
Inhibition of autophagy hinders PexRD54 vesicular distribution.
N. benthamiana leaves transiently expressing GFP:ATG8CL/RFP:PexRD54, GFP:PexRD54/RFP:ATG8CL or trans-golgi network marker YFP:VTI12 were infiltrated with autophagy inhibitor 3-methyl adenine (3-MA) at a concentration of 5 mM or water and imaged 6 hr post infiltration. 3-MA prevented formation of punctate structures of PexRD54 and ATG8CL but not of VTI12. Scale bar= 50 μm (B) Bar charts showing the number of punctate structures in 3-MA or water-treated samples. The data are representative of 500 individual images from two biological replicates. Each replicate consists of five independent Z stacks with 50 images each, acquired from five independent leaf areas. (***p<0.001, ns=statistically not significant). Error bars represent ± SD.
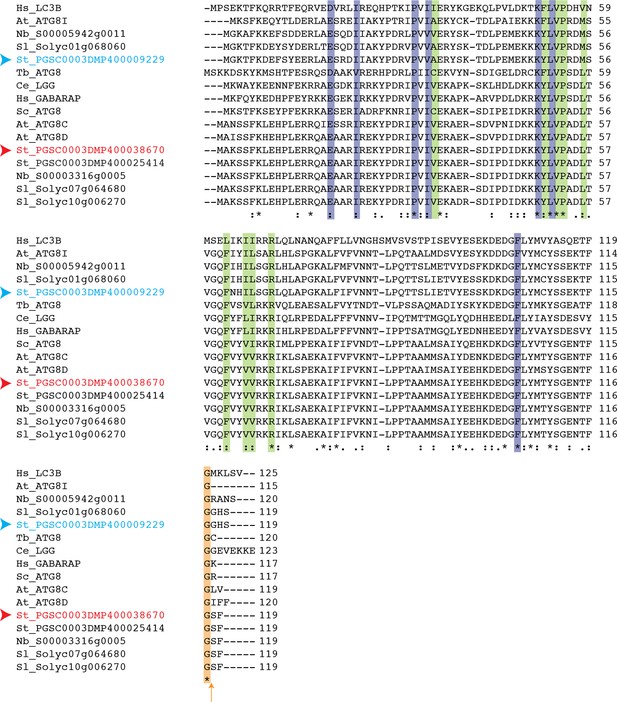

Conservation of terminal glycine residue in ATG8 proteins from various species.
Amino acid sequence alignment of ATG8 variants from different species that belong to different kingdoms shows conservation of terminal glycine residue (highlighted with an orange arrow). Residues that are responsible for interaction with core ATG8 interacting motif (AIM) residues are highlighted in purple (W pocket) and green (L pocket). Red and blue arrowheads point to ATG8CL and ATG8IL variants, respectively. Abbreviations for species: Hs: Homo sapiens; Tb: Trypanosoma brucei; Sc:Saccharomyces cerevisiae; Ce: Caenorhabditis elegans; At: Arabidopsis thaliana; Sl: Solanum lycopersium, Nb: Nicotiana benthamiana.
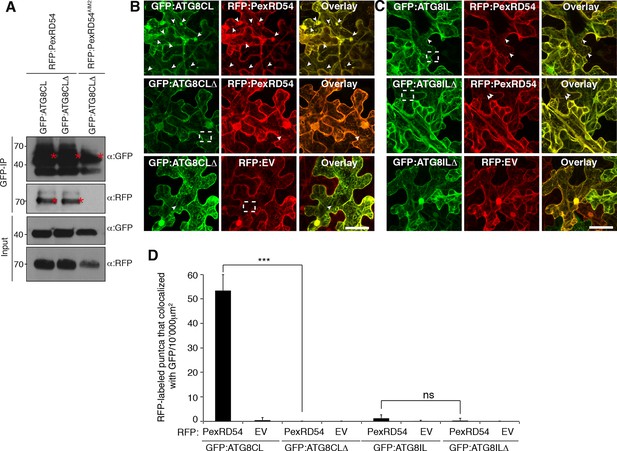
ATG8CL terminal glycine deletion mutant prevents formation of PexRD54 labelled puncta in a dominant negative manner.
(A) A glycine to alanine mutation in the conserved terminal glycine residue of GFP:ATG8CL does not prevent binding to RFP:PexRD54. GFP:ATG8CL or GFP:ATG8CLΔ were transiently co-expressed with RFP:PexRD54 or RFP:PexRD54AIM2 in N. benthamiana leaves and total proteins were extracted two days after infiltration and used in immunoprecipitation experiments (IPs). IPs obtained with anti-GFP and total protein extracts were immunoblotted with appropriate antisera. Stars indicate the expected band sizes. (B) GFP:ATG8CL or GFP:ATG8CLΔ were transiently co-expressed with RFP:EV or RFP:PexRD54 in N. benthamiana leaves and examined by confocal laser scanning microscopy 3 days post infiltration. GFP:ATG8CL showed distinct punctate structures, whereas GFP:ATG8CLΔ was mainly cytosolic with few visible puncta. When RFP:PexRD54 was co-expressed with GFP:ATG8CLΔ, the punctate localization was lost consistent with the recruitment of PexRD54 to ATG8CL labelled autophagosomes. (C) Co-expression of GFP:ATG8ILΔ and RFP:PexRD54, did not alter punctate localization of RFP:PexRD54. Arrowheads point to puncta labelled with GFP or RFP fusion constructs. Regions where GFP or RFP labelled puncta do not colocalize are indicated with dotted squares. Scale bar=50 μm (D) Bar charts confirming punctate distribution of PexRD54 depends on localization at ATG8CL labelled autophagosomes. The data are representative of 500 individual images from two biological replicates. Each replicate consists of five independent Z stacks with 50 images each, acquired from five independent leaf areas. (***p<0.001, ns=statistically not significant). Error bars represent ± SD.
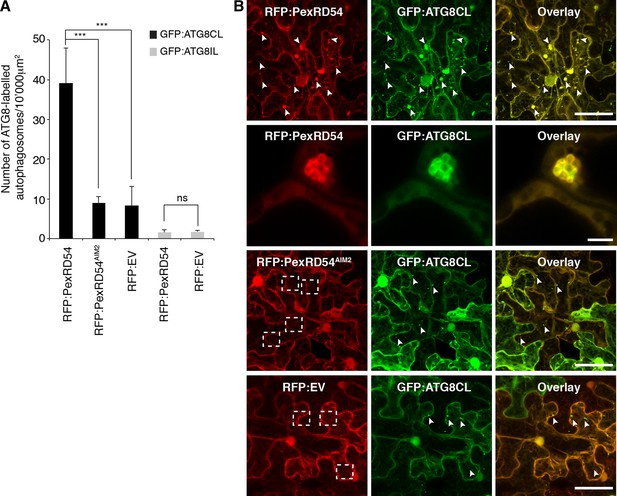
PexRD54 increases the number of ATG8CL autophagosomes.
(A) Co-expression of RFP:PexRD54, but not RFP:PexRD54AIM2 or RFP:EV significantly enhanced the number of GFP:ATG8CL-labelled autophagosomes in N. benthamiana. Bar charts display the number of GFP:ATG8CL or GFP:ATG8IL-labelled autophagosomes in the presence of RFP:PexRD54, RFP:PexRD54AIM2 or RFP:EV. GFP:ATG8CL autophagosomes were significantly enhanced by the expression of RFP:PexRD54 (***p<0.001). GFP:ATG8IL autophagosomes were not significantly enhanced by expression of RFP:PexRD54 (ns=statistically not significant). The data are representative of 500 individual images from two biological replicates. Each replicate consists of five independent Z stacks with 50 images each, acquired from five independent leaf areas. (B) GFP:ATG8CL was transiently co-expressed with RFP:PexRD54, RFP:PexRD54AIM2 or RFP:EV in N. benthamiana leaves and examined by confocal laser scanning microscopy 3 days post infiltration. Maximum projections of images show that RFP:PexRD54 increases the number of GFP:ATG8CL- labelled autophagosomes. RFP:PexRD54AIM2 and RFP:EV were used as controls. Arrowheads point to punctate structures. Regions where GFP or RFP-labelled puncta do not overlap are indicated with dotted squares. Scale bar=50 μm. Zoomed single plane images shown in the second panel indicate that larger puncta co-labelled by RFP:PexRD54 and GFP:ATG8CL are ring-shaped autophagosome clusters. Scale bar=10 μm.
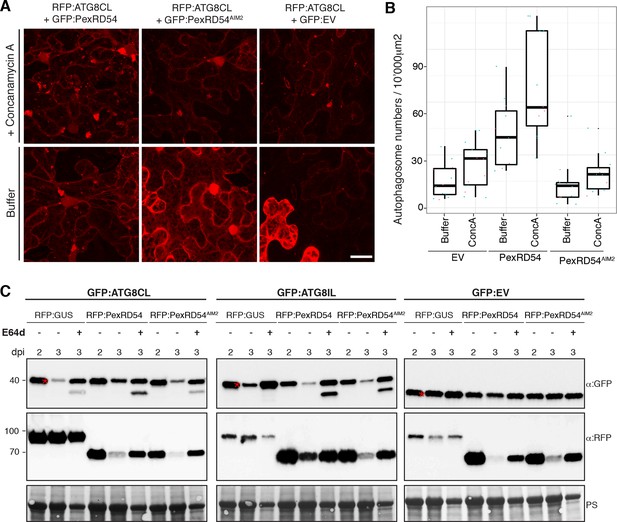
PexRD54 does not block autophagic flux.
(A–B) ConcanamycinA treatment increases the number of autophagosomes in PexRD54 expressing samples.RFP:ATG8CL was transienly coexpressed with PexRD54, PexRD54AIM2 and empty vector controls in N. benthamiana. Two days after infiltration, leaves were treated with concanamycinA (conA) or infiltration buffer and number of autophagosomes was counted 24 hr after treatment. ConA treatment significantly increased the number of autophagosomes in PexRD54 expressing cells (p<0.05), confirming PexRD54 does not block autophagic flux. Scale bar=10 μm. (C) E64D treatment increases ATG8CL protein levels in PexRD54 expressing samples. GFP:ATG8CL, GFP:ATG8IL and GFP:EV were transiently coexpressed with RFP:GUS, RFP:PexRD54 or RFP:PexRD54AIM2 in N. benthamiana leaves and protein levels in total extracts were determined two and 3 days post infiltration (dpi). RFP:PexRD54 increased protein levels of GFP:ATG8CL but not GFP:ATG8IL consistent with stronger binding affinity of PexRD54 to ATG8CL. RFP:PexRD54 did not increase protein levels of GFP:EV, suggesting that protein level increase depends on ATG8CL binding and that PexRD54 does not increase protein levels in general. The samples were also treated with E64d to measure autophagic flux. In RFP:PexRD54 coexpressed 3 dpi samples, E64d treatment increased ATG8CL protein levels even more suggesting PexRD54 does not block autophagic flux. Hence, protein level increase is a result of stimulation of autophagy. The blots were stained with Ponceau stain (PS) to show equal loading.
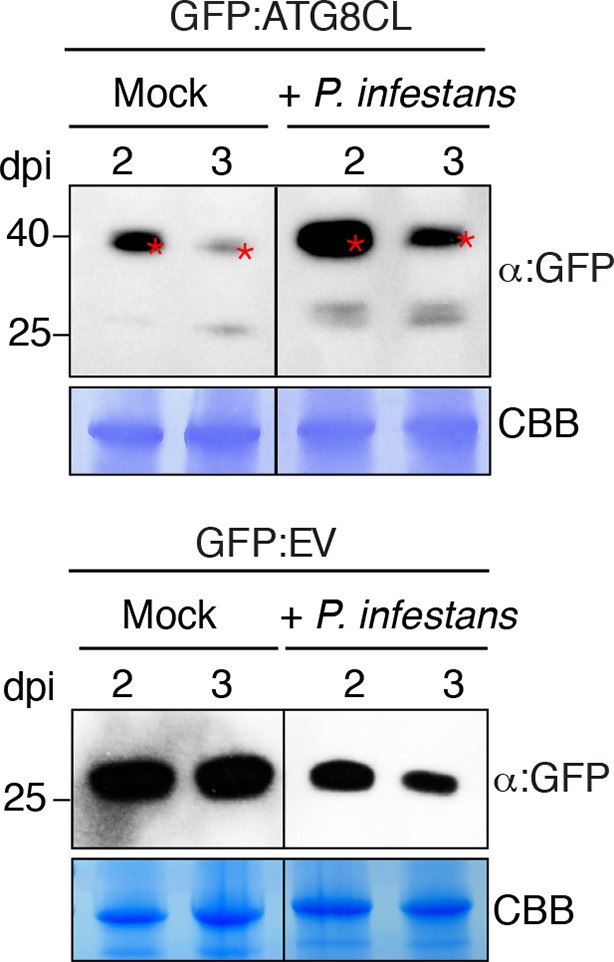
GFP:ATG8CL protein levels increase in N. benthamiana during P. infestans infection.
N. benthamiana leaves transiently expressing GFP:ATG8CL or GFP:EV were sprayed with P. infestans zoospore solution or water control (mock treatment). Protein levels of GFP:ATG8CL or GFP were determined 2 and 3 days after infection. The samples were run on the same gel and blots were developed with the same conditions. Gels were stained with coomassie brilliant blue (CBB) to show equal loading. Stars indicate expected band sizes.
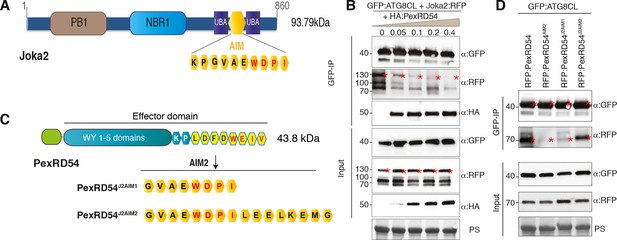
PexRD54 is a competitive antagonist of the plant selective autophagy cargo receptor Joka2.
(A) Domain organization of Joka2. (B) PexRD54 reduces binding of Joka2 to ATG8CL in a dose-dependent manner. GFP:ATG8CL (OD=0.2) and Joka2:RFP (OD=0.2) were transiently co-expressed with varying Agrobacterium concentrations (from OD=0 to OD=0.4) carrying HA:PexRD54 construct in N. benthamiana (OD=optical density of Agrobacterium cells). Joka2:RFP is depleted in GFP:ATG8CL pulldowns as the expression of HA:PexRD54 increased. Immunoprecipitates (IPs) obtained with anti-GFP antiserum and total protein extracts were immunoblotted with appropriate antisera. Stars indicate expected band sizes. (C) Schematic illustrations of PexRD54 variants (PexRD54J2AIM1 and PexRD54J2AIM2) with Joka2 AIM peptides. (D) Replacement of PexRD54 AIM with Joka2 AIM fragments decreases ATG8CL binding affinity. Immunoblots showing binding affinity of PexRD54, PexRD54AIM2, PexRD54J2AIM1 or PexRD54J2AIM2 to GFP:ATG8CL. IPs obtained with anti-GFP antiserum and total protein extracts were immunoblotted with appropriate antisera. Stars indicate expected band sizes.
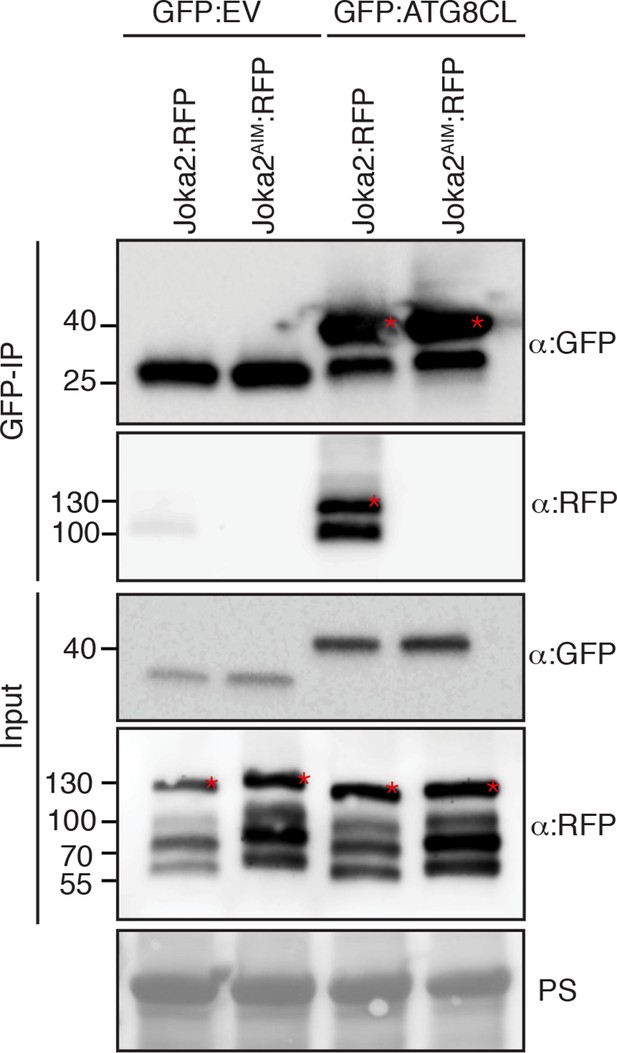
Joka2 has a functional ATG8 interacting motif (AIM).
Mutation of the predicted AIM in Joka2 (819-WDPI-822) to ADPA resulted in loss of ATG8CL binding. GFP:EV and GFP:ATG8CL were co-expressed with Joka2:RFP and Joka2AIM:RFP in N. benthamiana and proteins were extracted two days after infiltration and used in immunoprecipitation experiments (IPs). IPs were obtained with anti-GFP antiserum and total protein extracts were immunoblotted with appropriate antisera. Stars indicate expected band sizes.
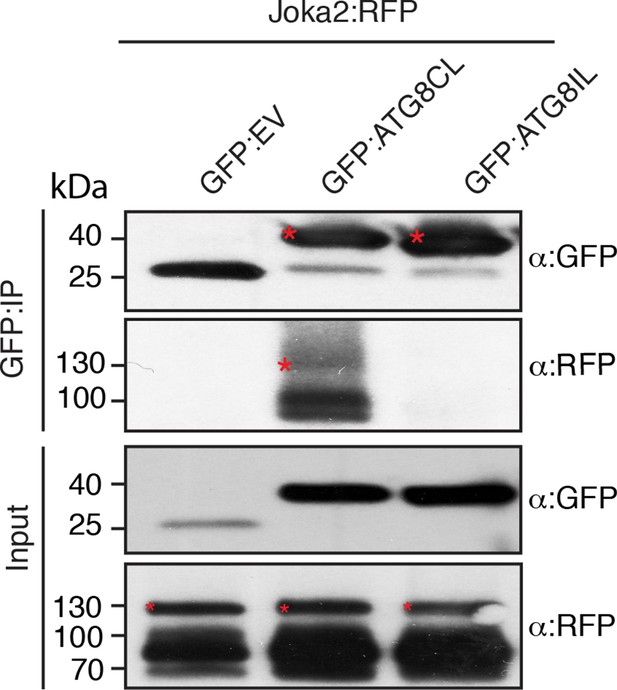
Joka2 has higher binding affinity to ATG8CL than ATG8IL.
Joka2:RFP was transiently co-expressed with GFP:ATG8CL, GFP:ATG8IL or GFP:EV in N. benthamiana and proteins were extracted two days after infiltration and used in immunoprecipitation experiments (IPs). IPs obtained with anti-GFP and total protein extracts were immunoblotted with appropriate antisera. IP experiments confirmed higher binding affinity of Joka2 to ATG8CL than ATG8IL. Stars indicate expected band sizes.
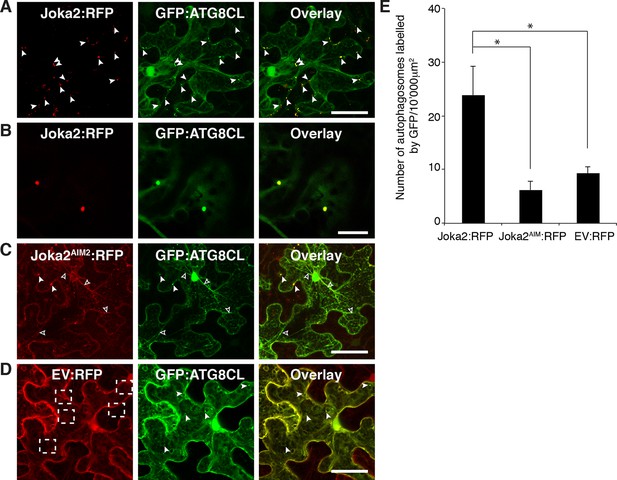
Joka2 increases the number of ATG8CL labelled autophagasomes.
(A) Joka2:RFP partially colocalizes with GFP:ATG8CL. Joka2:RFP and GFP:ATG8CL were transiently co-expressed in N. benthamiana leaves and examined by confocal laser scanning microscopy 3 days post infiltration. Maximum projections of stacks of images indicate that Joka2:RFP colocalizes with GFP:ATG8CL-labelled autophagosomes. Scale bar= 50 μm (B) Zoomed single plane images of the puncta co-labelled by Joka2:RFP and GFP:ATG8CL. Scale bar= 10 μm (C) Joka2AIM:RFP and GFP:ATG8CL were transiently co-expressed in N. benthamiana leaves and examined by confocal laser scanning microscopy 3 days post infiltration. Arrowheads point to puncta co-labelled by Joka2AIM:RFP and GFP:ATG8CL. Scale bar= 50 μm (D) EV:RFP and GFP:ATG8CL were transiently co-expressed in N. benthamiana leaves and examined by confocal laser scanning microscopy 3 days post infiltration. Arrowheads point to puncta labelled by GFP:ATG8CL. Dotted squares highlight the absence of control RFP signal at the GFP:ATG8CL puncta. Scale bar= 50 μm (E) Bar charts showing colocalization of GFP:ATG8CL with Joka2:RFP, Joka2AIM:RFP or EV:RFP. Joka2AIM:RFP or EV:RFP display significantly low co-localization with GFP:ATG8CL-labelled puncta in contrast to Joka2:RFP. The data are representative of 500 individual images from two biological replicates. Each replicate consists of five independent Z stacks with 50 images each, acquired from five independent leaf areas. (*p<0.05). Error bars represent ± SD.
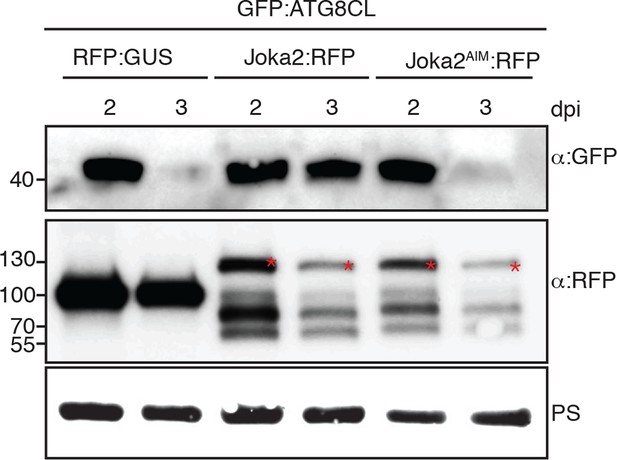
Joka2 increases ATG8CL protein levels.
GFP:ATG8CL was transiently co-expressed with RFP:GUS, Joka2:RFP or Joka2AIM:RFP in N. benthamiana leaves and protein levels in total extracts were determined 2 and 3 days after infiltration, using appropriate antisera. At 3 dpi GFP:ATG8CL protein levels were higher in Joka2:RFP expressing samples compared to Joka2AIM:RFP or RFP:GUS expressing samples. Blots were stained with Ponceau stain (PS) to show equal loading.
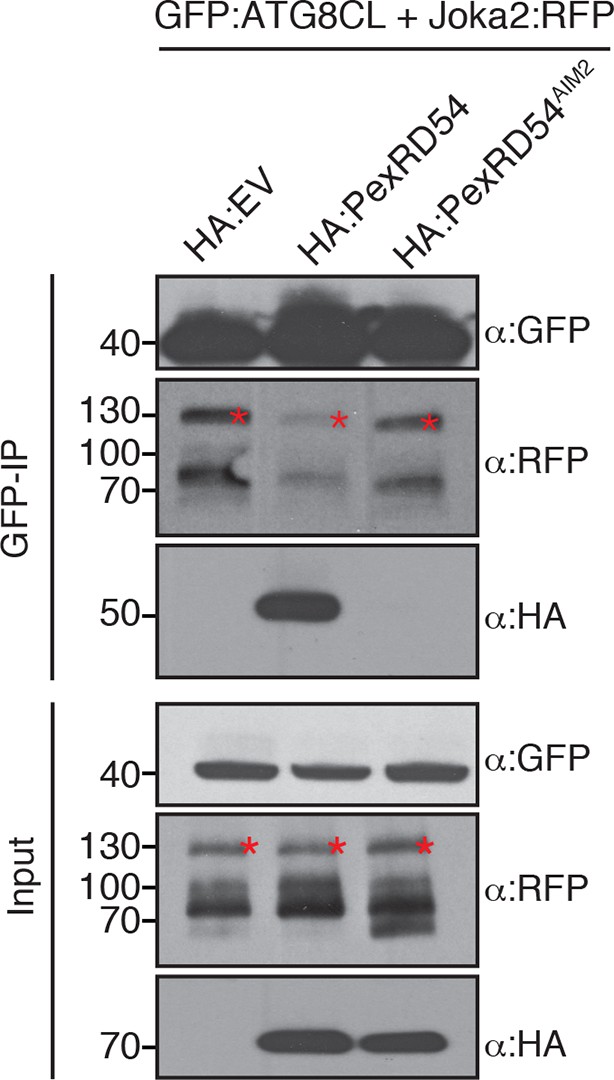
PexRD54 outcompetes Joka2 for ATG8CL binding.
Joka2:RFP and GFP:ATG8CL were transiently co-expressed with HA:EV, HA:PexRD54 and HA:PexRD54AIM2 in N. benthamiana leaves and proteins were extracted two days after infiltration and used in immunoprecipitation experiments (IPs). IPs obtained with anti-GFP and total protein extracts were immunoblotted with appropriate antisera. In the presence of HA:PexRD54, Joka2:RFP was depleted from GFP:ATG8CL complexes.
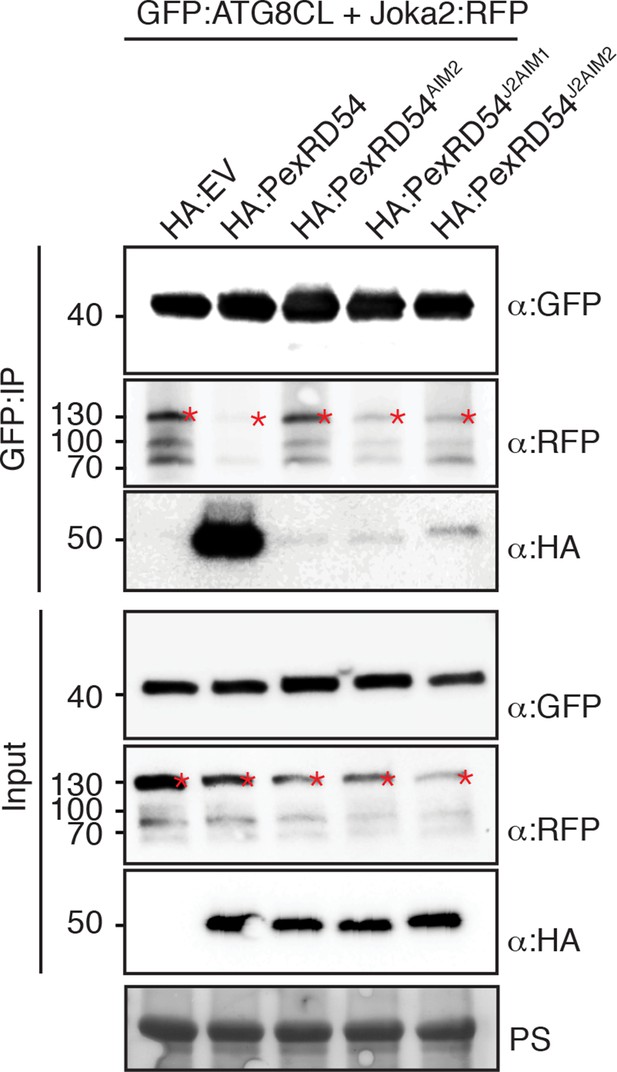
PexRD54 AIM has a higher affinity to ATG8CL than Joka2 AIM.
GFP:ATG8CL and Joka2:RFP were transiently co-expressed with HA:PexRD54, HA:PexRD54AIM2, HA:PexRD54J2AIM1, HA:PexRD54 J2AIM2 in N. benthamiana and proteins were extracted two days after infiltration and used in immunoprecipitation experiments (IPs). IPs obtained with anti-GFP and total protein extracts were immunoblotted with appropriate antisera. Joka2:RFP levels in GFP:ATG8CL complexes were higher in the presence of HA:PexRD54AIM2, HA:PexRD54J2AIM1 and HA:PexRD54J2AIM2 compared to HA:PexRD54 expressing samples suggesting PexRD54 variants carrying Joka2 AIM peptides (HA:PexRD54J2AIM1, HA:PexRD54J2AIM2) were less effective in depleting Joka2:RFP from GFP:ATG8CL complexes.
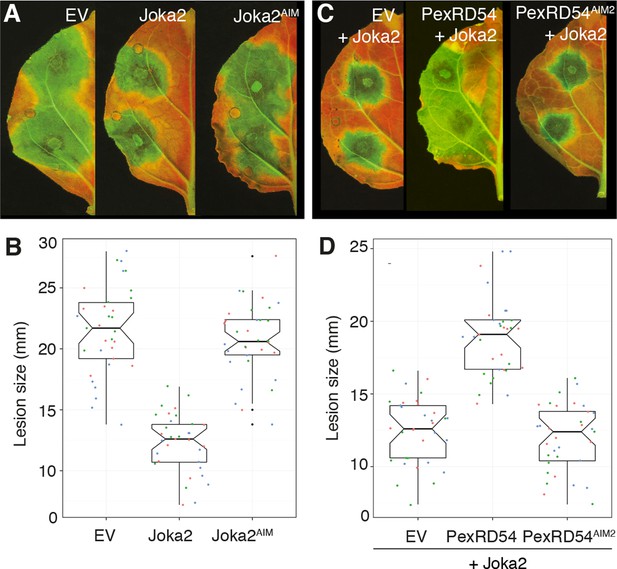
PexRD54 counteracts the enhanced resistance conferred by Joka2.
(A) Overexpression of Joka2:RFP limits P. infestans colonization. Halves of N. benthamiana leaves expressing RFP:EV, Joka2:RFP and Joka2AIM:RFP were infected with P. infestans and pathogen growth was determined by lesion sizes measured 6 days post-inoculation. (B) Categorical scatter plots show lesion diameters of 11 infections sites from three independent biological replicates pointed out by three different colors. Similar p values (p<0.001) were obtained in three independent biological repeats. (C) PexRD54 counteracts the effect of Joka2 on P. infestans colonization. Joka2:RFP was co-expressed with HA:EV, HA:PexRD54 or HA:PexRD54AIM2 in N. benthamiana leaves which are then inoculated with P. infestans. Joka2:RFP failed to limit pathogen growth in the presence of PexRD54, whereas it could still restrict pathogen growth in the presence of PexRD54AIM2 or vector control (EV). (D) Categorical scatter plots show lesion diameters of 11 infections sites from three independent biological replicates pointed out by three different colors. Similar p values (p<0.001) were obtained in three independent biological repeats.
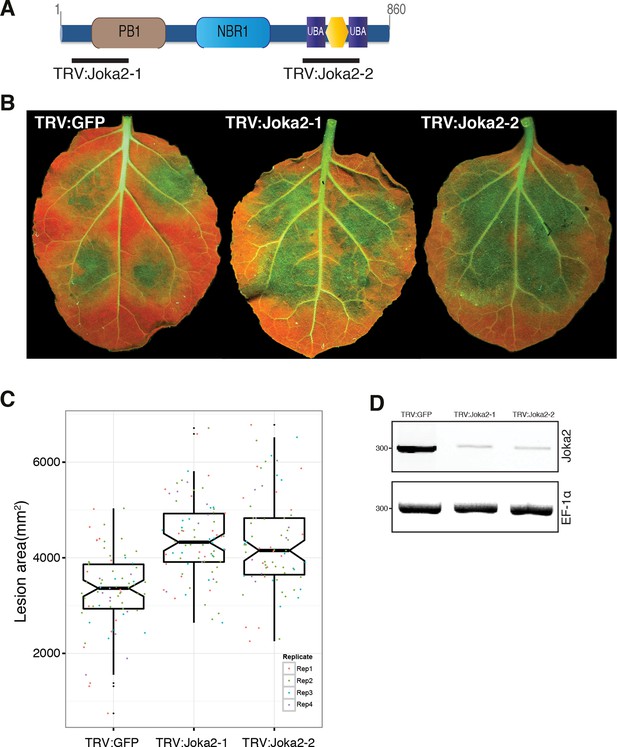
Silencing of Joka2 enhances susceptibility to P. infestans.
(A) Cartoon showing Joka2 sites targeted for VIGs. (B) TRV:Joka2-1, TRV:Joka2-2 and TRV:GFP control plants were inoculated with P. infestans 88,069 and pathogen growth was determined by lesion areas measured 5 days post-inoculation. (C) Categorical scatter plots show lesion areas of 10 infections sites from four independent biological replicates pointed out by four different colors. Similar p values (p<0.001) were obtained in four independent biological repeats. (D) RT-PCR confirmed Joka2 silencing. cDNA generated from silenced and control plants were used for the RT-PCR experiments. Transcript abundance was normalized using NbEF1α.
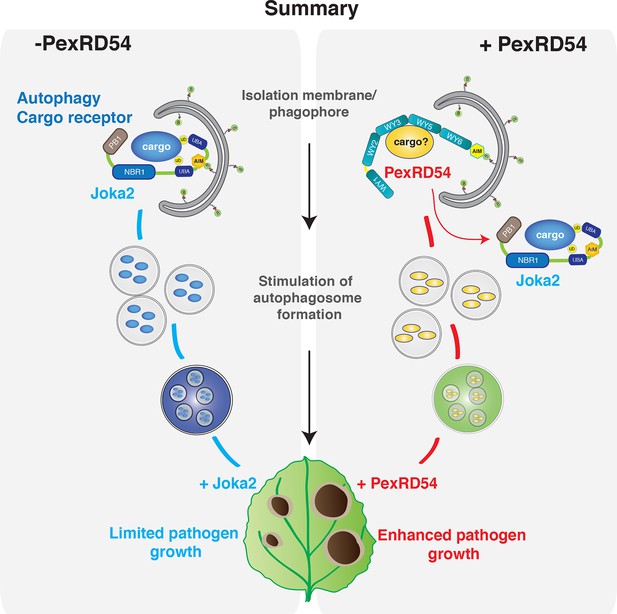
PexRD54 outcompetes the autophagy cargo receptor Joka2 and enhances virulence.
https://doi.org/10.7554/eLife.10856.032Videos
PexRD54 localizes to mobile endomembrane compartments.
GFP:PexRD54 was transiently expressed in N. benthamiana leaves and examined by confocal laser scanning microscopy, 3 days post infiltration. (Related to Figure 3)
PexRD54 and ATG8CL colocalize at mobile endomembrane compartments.
RFP:PexRD54 was transiently co-expressed with GFP:ATG8CL in N. benthamiana leaves and examined by confocal laser scanning microscopy, 3 days post infiltration. (Related to Figure 3)
RFP:PexRD54 and GFP:ATG8CL colocalize at mobile ring shaped clusters.
RFP:PexRD54 was transiently co-expressed with GFP:ATG8CL in N. benthamiana leaves and examined by confocal laser scanning microscopy, 3 days post infiltration. (Related to Figure 4)
RFP:ATG8CL and Joka2:GFP colocalize at mobile endomembrane compartments.
RFP:ATG8CL was transiently coexpressed with Joka2:GFP in N. benthamiana leaves and examined by confocal laser scanning microscopy, 3 days post infiltration. (Related to Figure 6)
Additional files
-
Supplementary file 1
Plant proteins that associate with PexRD54 as identified by mass spectrometry after immunoprecipitation (IP).
FLAG:PexRD54 was transiently expressed in N. benthamiana leaves and proteins were extracted two days after infiltration and used in IP experiments. Unique spectral counts are given for each control and PexRD54 sample. Proteins related to vesicle trafficking and autophagy are highlighted in yellow. ATG8CL is highlighted in blue and found in several replicates including the stringent IP experiment.
- https://doi.org/10.7554/eLife.10856.033
-
Supplementary file 2
Unique peptides obtained from liquid chromatography tandem mass spectrometry (LC-MS/MS) analysis of PexRD54 immunoprecipitation experiments suggest specific association with ATG8CL.
Although ATG8 protein family is highly conserved, and there are eight ATG8 variants in N. benthamiana, unique peptides obtained from LC-MS/MS analysis enabled specific identification of ATG8CL variant (Nb_S00003316g0005) from N. benthamiana. ATG8CL variant of N. benthamiana has identical amino acid sequence with the potato ATG8CL variant that is used for other experiments.
- https://doi.org/10.7554/eLife.10856.034
-
Supplementary file 3
Primers used in this study.
- https://doi.org/10.7554/eLife.10856.035





