Increasing Notch signaling antagonizes PRC2-mediated silencing to promote reprograming of germ cells into neurons
Figures
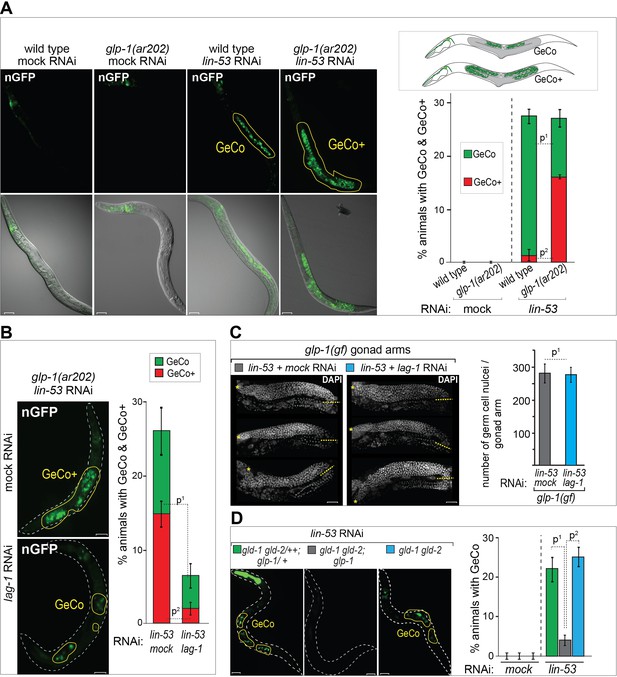
GLP-1Notch signaling promotes reprograming of germ cells.
(A) GLP-1Notch enhances germ cell conversion (GeCo) into neuronal-like cells. Left: Fluorescent (top) and combined fluorescent/differential interference contrast (DIC) micrographs (bottom) of adult animals. All animals ectopically expressed the pro-neuronal transcription factor CHE-1 from a heat-shock promoter. glp-1(ar202) is a temperature-sensitive gain-of-function allele of the Notch receptor. Animals were subjected to either mock (control) or lin-53 RNAi. Reprogrammed cells expressed a GFP reporter driven from the neuronal gcy-5 promoter (here an in other figures nGFP) and are outlined here and elsewhere in yellow. Any signal outside the outlined region comes from somatic tissues. GeCo+ indicates animals that displayed a strongly enhanced GeCo phenotype. Scale bars = 10 μm. The cartoons depicting the GeCo and GeCo+ phenotypes are on the top right. The gonads are shaded in grey and GFP-positive converted germ cells are green. Fractions of animals displaying GeCo and GeCo+ are indicated below. At least 250 animals were quantified per condition. P-values were calculated using Student's t-test: p1<0,0001; p2=0,0006. Error bars represent SEM. (B) The transcriptional effector of the GLP-1Notch signaling pathway, LAG-1, is required for the GLP-1Notch–mediated enhancement of GeCo. Left: Fluorescent micrographs of adults expressing CHE-1–induced nGFP as explained above. GeCo is diminished upon the depletion of LAG-1. White dashed lines outline the animal body. Scale bars = 10 μm. Right: The corresponding quantifications. At least 400 animals were quantified per condition. P-values were calculated using Student's t-test: p1<0,0001; p2=0,0018. Error bars represent SEM. (C) GLP-1Notch signaling enhances GeCo independently from germ cell proliferation. Shown are DAPI-stained gonads of glp-1(ar202) animals, expressing CHE-1–induced nGFP, treated with either mock or lin-53 RNAi. Germ cells were counted from the DTC (yellow asterisk) to the turn of the gonad arm (dashed yellow line). 15 gonad arms per condition were counted. Scale bars = 10 μm. Quantifications are on the right. While greatly inhibiting GeCo, lag-1 RNAi did not change the number of germ cells. P-values were calculated using Student's t-test: p1=0,89. Error bars represent SEM. (D) GLP-1Notch enhances GeCo independently from proliferation. Left: Fluorescent micrographs of adults (with indicated genotypes), expressing CHE-1–induced nGFP. The first panel on the left shows a control, heterozygous (wild-type) gld-1 gld-2/++; glp-1/+ animal. The other panels show the homozygous gld-1(q497) gld-2(q485) mutants, carrying either a loss-of-function (q175, center) or a wild-type (right) allele of glp-1. Despite proliferating, germ cells in the gld-1 gld-2; glp-1 gonads have lost the ability to undergo GeCo. Scale bars = 10 μm. Right: the corresponding quantifications. At least 250 animals were quantified per condition. P-values were calculated using Student's t-test: p1=0,0478; p2=0,0201. Error bars represent SEM.
-
Figure 1—source data 1
Quantification of GeCo in glp-1(gf) and lag-1 RNAi animals.
(A) Quantification of GeCo+ phenotype upon RNAi against lin-53 in glp-1(ar202) mutants. (B) Quantification of GeCo dependency on LAG-1. (C) Quantification of germ cells in glp-1 (ar202) gonads with our without lag-1 RNAi treatment. (D) Quantification of GeCo in different genetic backgrounds with highly proliferative germlines upon RNAi against lin-53. Figure 1—figure supplement 1A source data: Quantification of gfp-positive germ cells. Figure 1—figure supplement 2 source data: Quantification of germ cells in different genetic backgrounds with highly proliferative germlines. Figure 1—figure supplement 3A source data: Quantification of GeCo after cell cycle block with HU treatment. More details can be found in the corresponding figure legends.
- https://doi.org/10.7554/eLife.15477.004
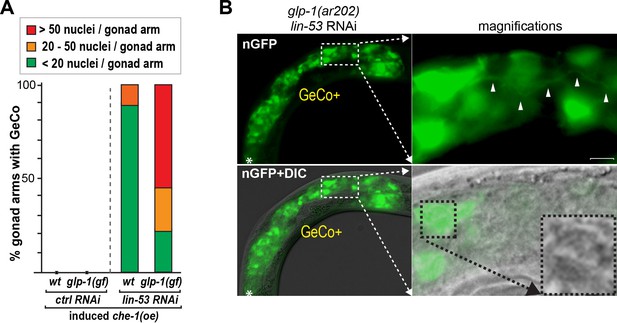
glp-1(gf) gonads contain more than twice the number of converted cells which display neuronal characteristics.
(A) For the quantification of gcy-5::gfp-positive cells per gonadal arm only the GeCo category of wt vs. glp-1(gf) was used because GeCo+ animals already show an extensive area of the gonad filled with gcy-5::gfp-positive cells with usually >100 cells/gonad making reliable counting impossible. Notably, animals with a seemingly similar extend of GeCo in wt vs. glp-1(gf) show a clear increase of gcy-5::gfp-positive cells per gonadal arm from approx. 10 in wt to > 30 in glp-1(gf). n(wt) = 75 gonaldal arms, n (glp-1(ar202)) = 221 gonadal arms. The background of the loss of function allele glp-1(q175) leads to a significant decrease in GeCo as shown previously (Tursun et al., 2011). (B) A magnified view of gcy-5::gfp-positive (nGFP) cells, in a GeCo+ gonad from a glp-1(gf) animal. The converted cells show axo-dendritic projections (white arrow heads). The inset in the corresponding DIC image, magnified in the right-bottom corner, shows the nuclear morphology of a converted germ cell, which has lost the germ cell-specific ‘fried-egg’-like shape and instead shows nuclear speckles characteristic of a neuronal cell. Scale bar = 1 μm.
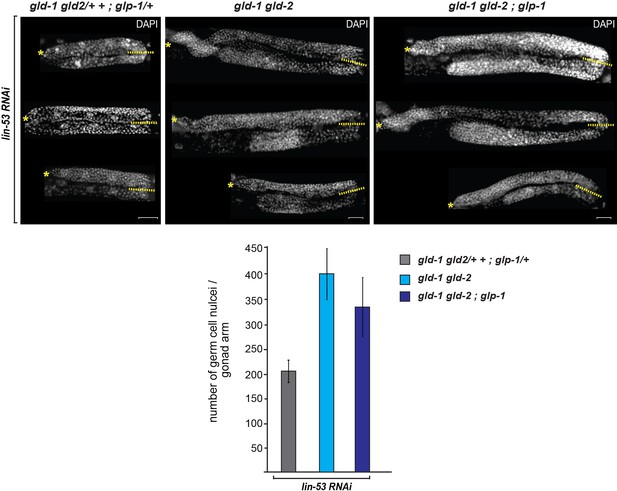
Germ cell numbers are similar between gld-1 gld-2 double and gld-1 gld-2; glp-1 triple mutants.
DAPI-stained gonads of gld-1(q497) gld-2(q485) or gld-1(q497) gld-2(q485); glp-1(q175) mutants, carrying the hsp::che-1 and gcy-5::gfp transgenes. The gonads were imaged by fluorescent microscopy using Z-stack acquisitions. Germ cells from the DTCs (yellow asterisks) to the turn of the gonad arm (dashed lines) were counted. Below:15 gonad arms per condition of L4 animals were counted. The numbers of germ cells differ only slightly (15%) in the double mutant vs. triple mutants background. Scale bars = 10 μm. Error bars represent SEM.
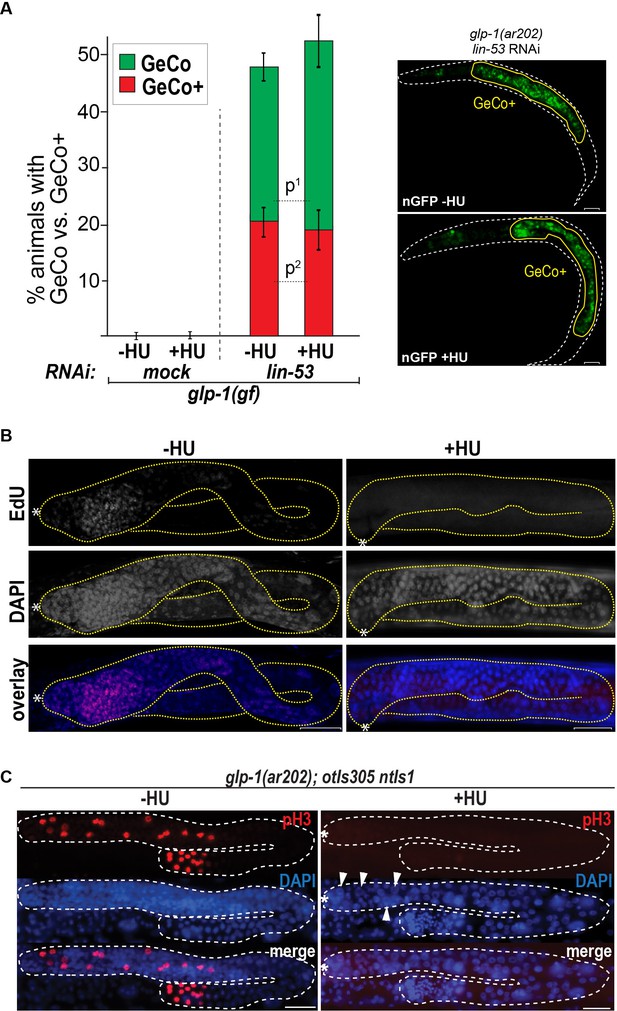
Blocking the cell cycle with hydroxyurea does not inhibit GeCo+.
(A) We used hydroxyurea (HU) treatment for 5 hr to chemically block the cell cycle, which makes germ cells arrest in the S phase of the cell cycle. This arrest does not diminish the GeCo+ phenotype in glp-1(gf) gonads upon lin-53 RNAi and che-1(oe). At least 150 animals were quantified per condition. P-values were calculated using Student's t-test: p1=0,1409; p2=0,4583. Error bars represent SEM. The right panel shows examples of GeCo+ displaying animals based on gcy-5::gfp (nGFP) for HU-untreated (-HU) and HU-treated (+HU) animals. (B) The gonads were stained for EdU incorporation. Dashed lines outline gonads. Asterisks indicate distal tips of gonads. Scale bars = 10 μm. (C) The gonads of glp-1(ar202) gf animals, which were treated with HU for 12 hr and stained with DAPI and H3Ser10ph (pH3) antibody. The pH3-positive cells indicate condensed chromosomes of dividing cells. After 12 hr of HU treatment, the gonads contained, as expected, enlargement nuclei (arrowheads) (Gartner et al., 2004; Fox et al., 2011). The loss of pH3-positive cells indicates a cell cycle arrest. Asterisks indicate distal tips of gonads, dashed lines outline gonad. Scale bars = 10 μm.
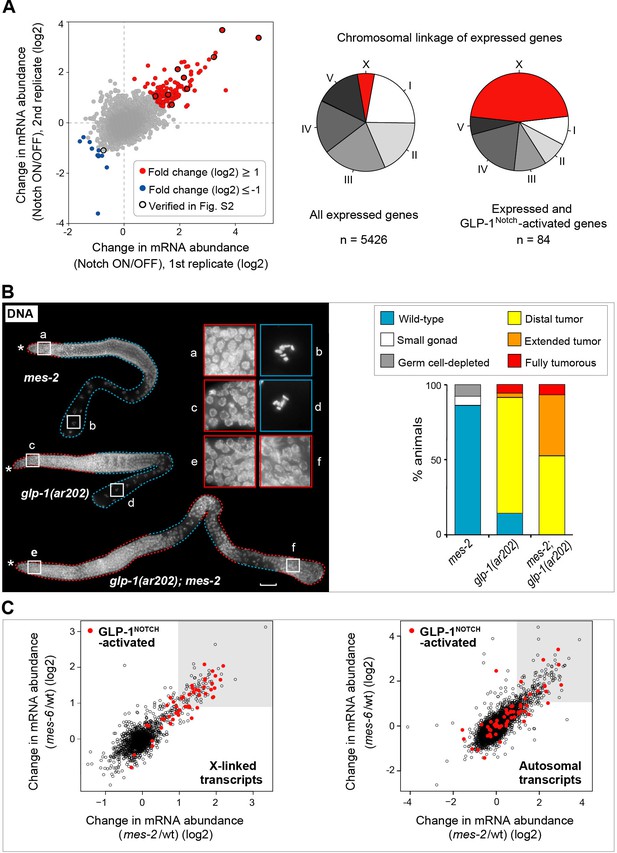
GLP-1Notch and PRC2 regulate common targets and are functionally connected.
(A) Notch-activated genes are biased for the sex chromosome linkage. Left: Changes in transcript abundance in the ‘Notch ON’ versus ‘OFF’ dissected gonads (genotypes explained in Figure 2—figure supplement 1A–B) were analyzed by microarrays. Transcripts upregulated at least 2-fold in the ‘Notch ON’ gonads are marked in red, those downregulated at least 2-fold in blue. Selected transcripts verified by RT-qPCR in Figure 2—figure supplement 2A are additionally circled in black. Right: 5426 genes can be considered expressed in the gonad, based on the bimodal distribution of expression values. Only 3% of those expressed genes are X-linked. In contrast, nearly half (46%) of the expressed and Notch-activated transcripts are X-linked (see Figure 2—figure supplement 2B for numbers). (B) GLP-1Notch and PRC2 interact genetically. Left: DAPI-stained gonads from animals of the indicated genotypes. The mes-2(bn11) M+Z- single mutant gonads have wild-type appearance at 20°C. The glp-1(ar202) gain-of-function mutants have an almost wild-type appearance at this temperature, except for an extended proliferative zone in the gonad, referred to as 'distal tumor'. At the same temperature, mes-2(bn11) M+Z-; glp-1(ar202) double mutants developed germline tumors in 32/32 of the examined gonads. The insets show close-ups from the indicated gonadal regions: the distal-most regions contain undifferentiated, proliferative germ cells in all mutants (a, c, e). However, while the single mutants contain oocytes with characteristically condensed chromosomes in the proximal gonads (b, d), the proximal gonads of the double mutants harbor proliferative germ cells (f). Scale bar = 30 μm. Right: quantification of the phenotypes. 'Distal tumor' indicates the presence of an elongated distal proliferative zone (approximately ½ of the distal gonad arm). 'Extended' tumor indicates an extended distal tumor, few oocytes, and frequently also a proximal tumor. 'Fully tumorous' indicates the absence of all differentiated cell types except for sperm produced during larval development. (C) GLP-1Notch and PRC2 target the same genes on the X chromosomes. The plots correlate changes in gene expression in M+Z- mes-2 mutants with changes in gene expression changes in M+Z- mes-6 mutants. Results are shown separately for X-linked (left) and autosomal (right) transcripts. Notch-activated genes (red in Figure 2A) are marked in red. Lightly shaded areas indicate transcripts that are at least 2-fold upregulated. The overlap between transcripts upregulated by GLP-1Notch and transcripts upregulated by the loss of CePRC2 is highly significant, particularly for the X-linked genes. The significance of the correlation was measured by hypergeometric distribution; X-linked Notch-activated vs. mes-2 derepressed: p=1.31e-31; X-linked Notch-activated vs. mes-6 de-repressed: p=7.41e-25; autosomal Notch-activated vs. mes-2 derepressed: p=1.47e-22; autosomal Notch-activated vs. mes-6 de-repressed: p=1.8e-12.
-
Figure 2—source data 1
Microarray results.
(A) Microarray results of differentially expressed genes in Notch ON/OFF gonads. Figure 2—figure supplement 3 source data: Quantification of germlines with wild-type morphology versus germlines with tumors upon germline-autonomous RNAi against genetic interactors of PRC2. More details can be found in the corresponding figure legends.
- https://doi.org/10.7554/eLife.15477.009
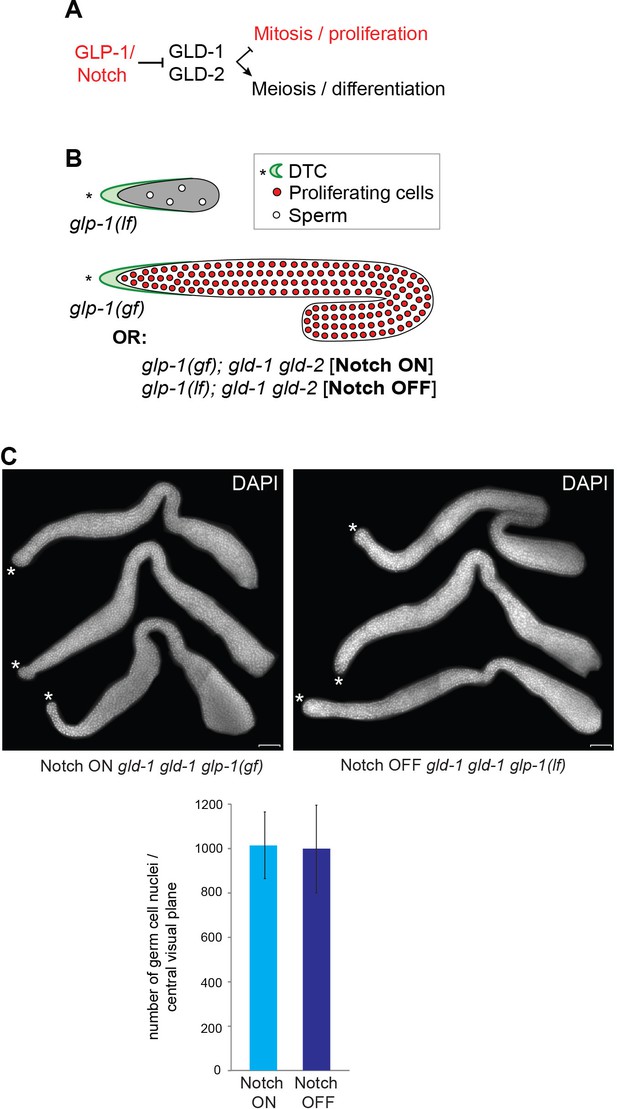
Examining transcriptional effects of GLP-1Notch signaling.
(A) GLP-1Notch signaling counteracts the meiosis-promoting activity of several RNA binding proteins, of which GLD-1 and -2 are indicated. (B) Mutants that were used in this study: while the loss-of-function (lf) alleles of glp-1 cause the loss of germ cells, gain-of-function (gf) alleles result in a constitutive proliferation of germ cells. However, in the absence of the meiosis-promoting GLD-1 and GLD-2 proteins, germ cells continue to proliferate in the absence of GLP-1Notch activity. (C) Three representative gonads of Notch ON: gld-2(q497) gld-1(q485); glp-1(ar202) and Notch OFF: gld-2(a497) gld-1(q485); glp-1(e2144) animals are shown after dissection and DAPI staining. The central planes of the gonads were imaged. Nuclei were counted from those images using the CellCounter plugin with ImageJ. For each genetic background, germ cells in the entire gonad of 10 dissected gonads were counted. The quantification below revealed that the numbers of germ cells in both backgrounds are not changed. Error bars represent SD. Scale bar = 10 μm.
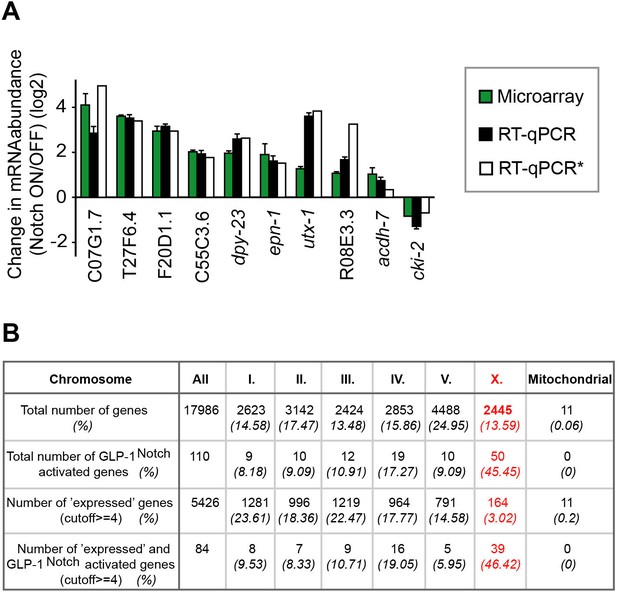
Analysis of Notch-activated genes.
(A) Changes in the abundance of several transcripts (marked black in Figure 2A) were verified independently from the genomic quantification by RT-qPCR. Colors correspond to fold-changes detected by microarrays (green), by RT-qPCR on genetically identical gonads (black), or by RT-qPCR on gonads with a different loss-of-function glp-1 allele, q175 (white). *This experiment was performed only once. The error bars represent SEM. (B) Although the number of genes on the arrays is almost equally distributed between the different chromosomes, the expression of X-linked genes is largely silenced in the germline. The table displays the numbers and fractions of genes on the different chromosomes, and the distributions of the Notch-activated genes. The cutoff between 'not expressed' and 'expressed' genes was set according to the bimodal distribution of expression values in the Notch ON and OFF arrays.
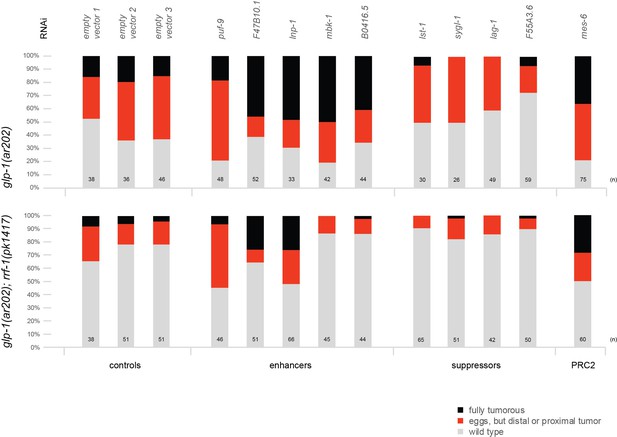
The PRC2 component MES-6 and most enhancers/suppressors of glp-1(ar202) induced tumors appear to interact genetically with GLP-1Notch signaling in a germline-autonomous manner.
Each bar indicates the relative proportion of germlines with wild-type morphology (grey), germlines that contain eggs but also a proximal or distal tumor (red), and germlines that are fully tumorous (black). 'n' indicates the number of DAPI-stained gonads scored for each column. Empty vector 1–3 represent three independent replicates of the empty vector control and demonstrate the robustness of the experiment. We observed that, for an unknown reason, the rrf-1(pk1417); glp-1(ar202) double mutants were less likely to produce tumors at the semi-permissive temperature of 20°C. Nonetheless, the double mutant strain reacts to enhancers, suppressors, and depletion of PRC2 components in a similar manner as the glp-1(ar202) single mutant strain, with the exception of mbk-1 and B0416.5 RNAi. Depletion of mes-2 and mes-3 by RNAi was ineffective, since it did not enhance the tumourous phenotype in either of the two strains.
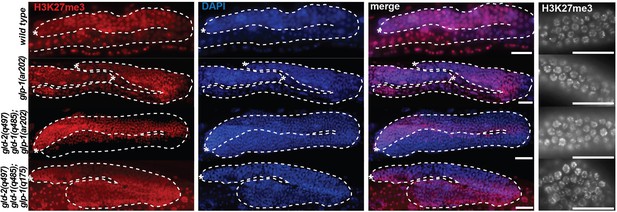
Global levels of H3K27me3 are unaffected by neither loss-of-function nor gain-of-function mutations in glp-1.
Stainings of wild-type, gain-of-function glp-1(ar202), Notch ON (gld-2(q497) gld-1(q485); glp-1(ar202)) and Notch OFF (gld-2(a497) gld-1(q485); glp-1(q175)) gonads with antibodies against H3K27me3 performed on whole worms (left panel, scale bars = 10 μm) or dissected gonads (right panel, scale bars = 1 μm). The H3K27me3 levels do not differ globally between the different mutant backgrounds. Asterisks indicate distal tips of gonads, dashed lines outline gonad.
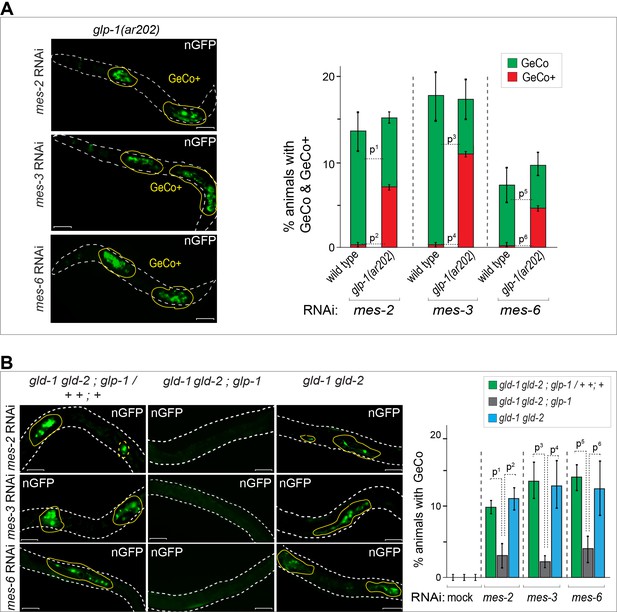
GLP-1Notch enhances reprograming upon the depletion of PRC2.
(A) Notch and PRC2 genetically interact in GeCo. Left: Fluorescent micrographs of glp-1(ar202) gain-of-function mutants expressing CHE-1–induced neuronal GFP. The animals were subjected to control RNAi or RNAi against PRC2 components (MES-2, 3, and 6), as indicated. Increased GLP-1Notch signaling enhanced the GeCo+ phenotype upon PRC2 depletion. Control RNAi (mock) for each genetic background did not result in any GeCo (images not shown – see quantification). Right: The corresponding quantifications. P-values were calculated using Student's t-test: p1=0,0006; p2<0,0001; p3=0,0536; p4=0,0001; p5=0,4035; p6=0,0003. At least 200 animals were scored per condition. Error bars represent SEM. (B) GLP-1Notch is required for GeCo in PRC2-depleted animals independently from proliferation. Left: Fluorescent micrographs of adults expressing CHE-1–induced nGFP, with the genotypes indicated above the panels. The animals were subjected to RNAi as indicated on the left. The first column shows heterozygous, the other two homozygous animals carrying the loss of function alleles gld-1(q497), gld-2(q485) and, in the central panels, glp-1(q175). The animals were subjected to control RNAi or RNAi against PRC2 components (MES-2, 3, and 6). In the absence of GLP-1Notch, depletion of PRC2 components did not induce GeCo. Scale bars = 10 μm. Right: The corresponding quantifications. P-values were calculated using Student's t-test: p1<0,0456; p2=0,0337; p3=0,0070; p4=0,0637; p5=0,0080; p6=0,1259. At least 70 animals were scored per condition. Error bars represent SEM.
-
Figure 3—source data 1
Quantification of GeCo upon PRC2 depletion.
(A) Quantification of GeCo+ phenotype upon RNAi against PRC2 subunits in glp-1(ar202) mutants. (B) Quantification of GeCo in different genetic backgrounds with highly proliferative germlines upon RNAi against PRC2 subunits. More details can be found in the corresponding figure legends.
- https://doi.org/10.7554/eLife.15477.015
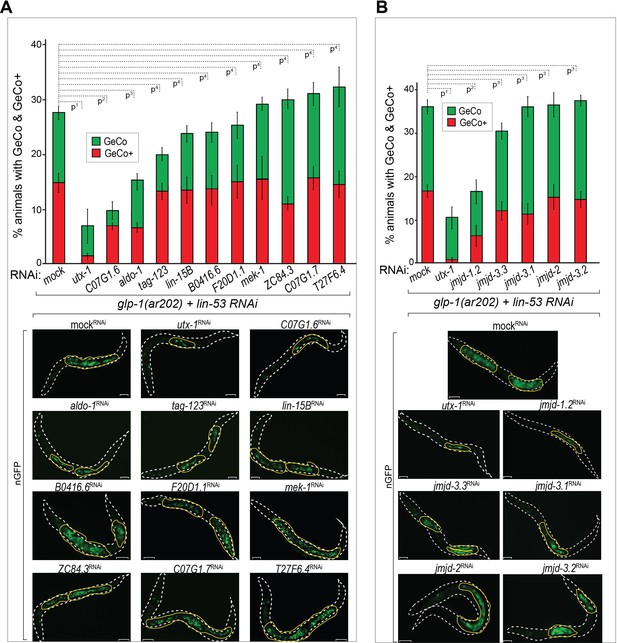
The H3K27 demethylase UTX-1 is required for GeCo enhancement.
(A) UTX-1 is critical for GeCo enhancement. Candidate Notch-activated genes, selected from Supplementary file 2 with available RNAi clones, were assayed for a role in GeCo in glp-1(ar202) animals, expressing CHE-1–induced nGFP and treated with lin-53 RNAi. While the additional depletion of utx-1 had the strongest impact on GeCo+ and GeCo, the depletion of C07G1.6 and aldo-1 had a weaker effect. Representative fluorescence micrographs are below the quantification chart. White dashed line outline the animal body, yellow lines outline gonadal areas with GeCo. P-values for GeCo+ were calculated using Student's t-test: p1=0,000013; p2=0,026; p3=0,021; p4>0,1. At least 250 animals were scored per condition. Error bars represent SEM. nGFP = gcy-5::gfp. Scale bars = 10 μm. (B) As in A, but RNAi was performed against jmjd-1.2 (H3K9/27me2 demethylase); jmjd-3.1, jmjd-3.2, and jmjd-3.3, (H3K27me2/3 demethylases); and jmjd-2 (H3K9/36 demethylase). Only RNAi against jmjd-1.2 suppresses GeCo+, though to a lesser degree compared to utx-1 RNAi. Representative fluorescence micrographs are below the quantification chart. P-values for GeCo+ were calculated using Student's t-test: p1=0,0042; p2=0,035; p3>0,2. At least 190 animals were scored per condition. Error bars represent SEM. Scale bars = 10 μm.
-
Figure 4—source data 1
Quantification of GeCo upon double RNAi against lin-53 and Notch-activated genes.
(A) Quantification of GeCo+ upon RNAi against lin-53 and Notch-activated genes. (B) Quantification of GeCo+ upon RNAi against lin-53 and Histone demethylases. Figure 4—figure supplement 1A source data: Quantification of GeCo+ upon RNAi against lin-53 and utx-1 with and without rrf-1 (pk1417) background. More details can be found in the corresponding figure legends.
- https://doi.org/10.7554/eLife.15477.017
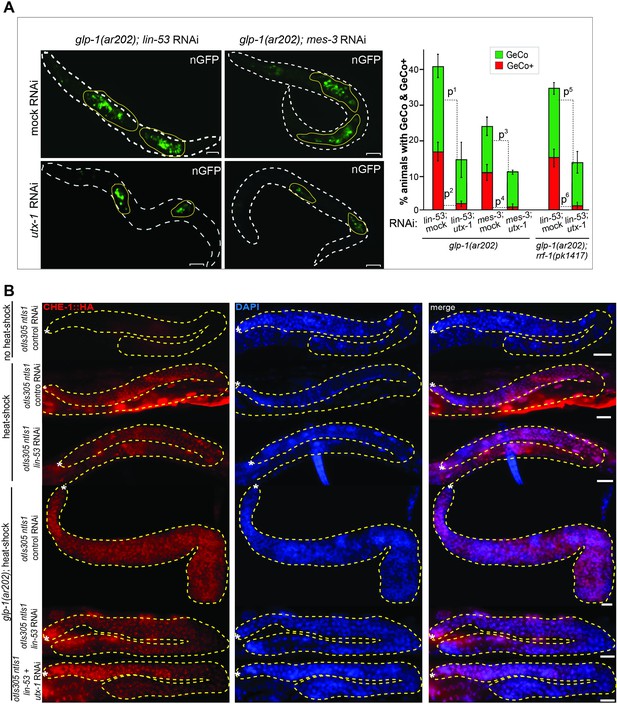
UTX-1 is required for the GeCo+ enhancement upon the depletion of PRC2.
(A) Adult glp-1(ar202) animals treated with lin-53 or mes-3 RNAi were additionally subjected to either control or utx-1 RNAi. Depletion of utx-1 strongly suppressed the GeCo+ phenotype. Suppression upon utx-1 co-depletion with lin-53 is also detectable in the rrf-1(pk1417) background which is permissive for RNAi in the germline but not in the somatic gonad and the DTC. Right: quantification: n = 715 were scored for lin-53; utx-1 RNAi; n = 500 for mes-3; utx-1 RNAi and n = 270 were scored for lin-53; utx-1 RNAi in rrf-1(pk1417). P-values were calculated using Student's t-test: p1=0,0588; p2=0,0042; p3=0,2454; p4=0,01713; p5=0,40479; p6=0,00271. Error bars represent SEM. (B) anti-HA antibody staining for the 3xHA-tagged CHE-1 protein, which is being induced after heat-shock treatment in the different genetic backgrounds: glp-1(ar202); otIs305 (hspprom::che-1::3xHA) ntIs1 (gcy-5prom::gfp) treated with our without RNAi against lin-53 and utx-1. As additional controls the strain otIs305 (hspprom::che-1::3xHA) ntIs1 (gcy-5prom::gfp) with or without lin-53 RNAi and heat shock treatment was used. No obvious changes in the induction of CHE-1::3xHA in the germlines of the different genetic backgrounds can be detected. Scale bars = 1 μm.
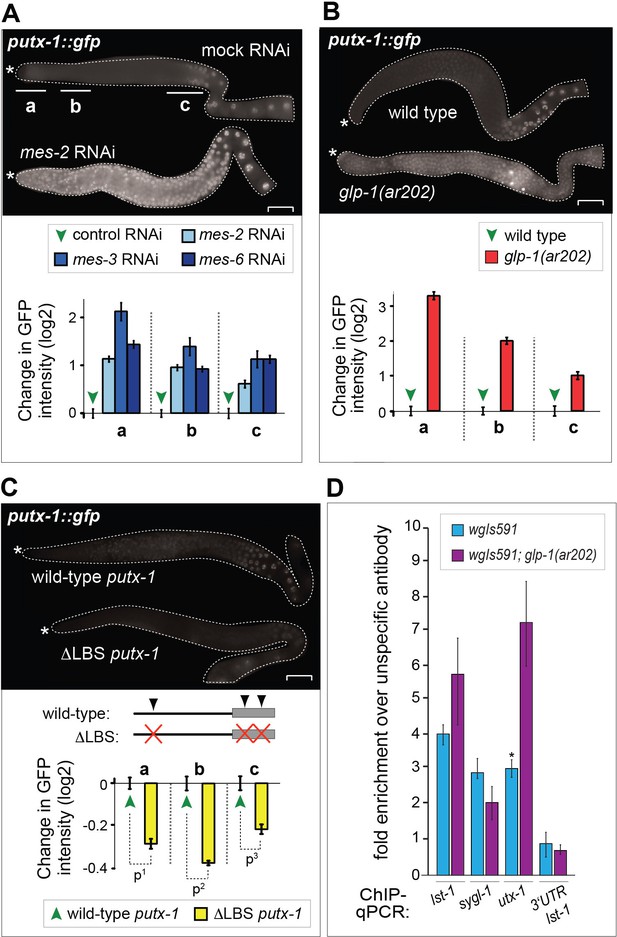
UTX-1 is regulated by GLP-1Notch and PRC2.
(A–C) Expression of utx-1 is regulated by PRC2 and GLP-1Notch. Top: dissected gonads expressing a GFP reporter, driven from the utx-1 promoter (putx-1::GFP, fused to histone 2B for nuclear localization to facilitate quantification), subjected to the indicated RNAi (A), crossed into the indicated genetic background (B) or carrying the indicated mutations in the reporter gene (C). a, b, and c indicate gonadal regions containing germ cells in mitosis (a), and leptotene/zygotene (b) or pachytene (c) stages of meiosis. Below: the corresponding GFP quantifications. The diagrams show GFP intensities relative to the control (indicated by green arrows) in regions a-c. Results are represented as average changes in the GFP intensity (relative to mock RNAi-ed or untreated animals). The error bars represent SEM. The numbers of analyzed gonads were as follows: n = 44 for wild-type reporter; n = 36 for glp-1(ar202); n = 55 for wild-type reporter on control RNAi; n = 48 for mes-2 (RNAi); n = 15 for mes-3 (RNAi); n = 29 for mes-6 (RNAi), and n = 20 for the LAG-1 binding sites deleted reporter. (A) The putx-1::GFP reporter is repressed by PRC2. In all mes-depleted gonads, the reporter was de-repressed in proliferating cells (a) as well as in more proximal gonadal regions (b-c). (B) The putx-1::GFP reporter is upregulated by GLP-1Notch. In the gain-of-function glp-1(ar202) mutant, the reporter was strongly derepressed in the proliferating cells in the distal-most gonad (a). Its expression was also increased in the more proximal regions (b-c), which, in this mutant background, contain proliferating cells instead of meiotic cells. (C) putx-1::GFP expression depends on the predicted LAG-1/CSL binding sites in the promoter. Upon deletion of putative LAG-1/CSL binding sites, the reporter expression was abolished. The changes in GFP intensities were highly significant (p-values were measured by independent t-tests) p1=4.85–13, p2=1.38–20, p3=1.18–7. (D) LAG-1 binds the utx-1 promoter. Lysates of animals expressing FLAG-tagged LAG-1 (strain wgIs591; lag-1::TY1::EGFP::3xFLAG), either in wild-type or glp-1(ar202) background, were subjected to ChIP-qPCR analysis of the indicated genes. Negative controls and additional tested genes are shown in Figure 5—figure supplement 3. The qPCR amplicons were tested in at least three independent experiments. The results are shown as fold enrichment in anti-FLAG IP compared to IP with unspecific antibody. The 3’UTR of lst-1 serves as a negative control. Interestingly, LAG-1 binding in the glp-1(ar202) gain-of-function background is stronger to the utx-1 promoter than to the reported GLP-1Notch targets lst-1 and sygl-1. The asterisk indicates a p-value < 0.05 (Students t-test). Error bars represent SEM.
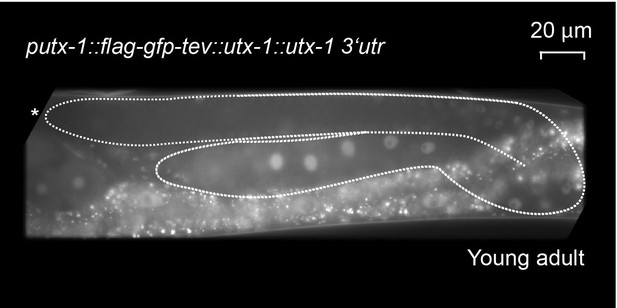

The functional utx-1 transgene is expressed in the same pattern as a GFP reporter coupled to the utx-1 promoter.
Shown is an adult with an outlined gonad. The full-length GFP-tagged functional UTX-1 (rrrSi189) is repressed in the distal, proliferative part. Nuclei entering meiosis and developing oocytes express the fusion protein progressively stronger. This expression pattern is identical with the pattern seen in the utx-1 promoter reporter strains (rrrSi185, rrrSi281).
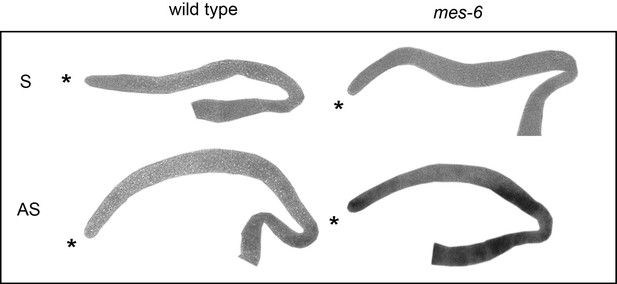

The endogenous utx-1 mRNA is upregulated in the absence of the PRC2-component MES-6.
Shown are representative in-situ hybridisations against endogenous utx-1 mRNA in dissected gonads. In M+Z- mes-6(bn66) mutants, utx-1 is upregulated throughout the gonads compared to the control wild-type gonads. 'AS' and 'S' indicate antisense and sense probes.
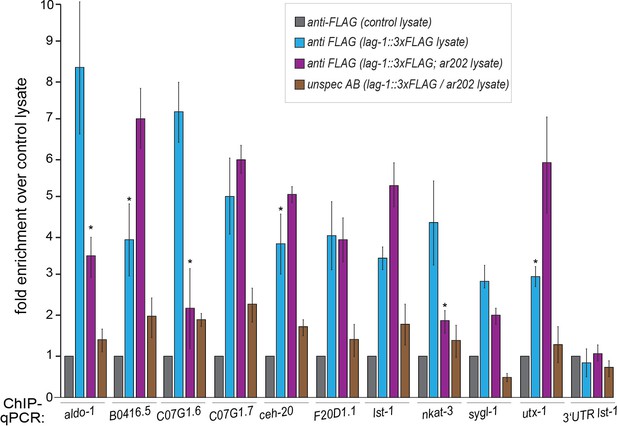
Testing LAG-1 binding to additional genes by ChIP.
Worm lysates (corresponding to 4 mg protein) of animals, with or without lag-1::TY1::EGFP::3xFLAG (wgIs591) transgene, were used for ChIP. Samples were incubated with 50 µl of FLAG (‘specific’ FLAG antibody) or HA antibodies (‘unspecific’ HA antibody) coupled to µMACS microbeads (Milteny). As negative control, lysate N2 or glp-1(ar202) worm lysates, which do not express the recombinant target protein, were used. Both negative control lysates did not show any differences during the ChIP experiment when tested with either specific antibody (anti-FLAG coupled to µMACS beads) or unspecific antibody (anti-HA coupled to µMACS beads). Lysates of worms expressing the recombinant target protein in N2 or glp-1(ar202) background were incubated with specific (anti-FLAG) and unspecific (anti-HA) antibodies coupled to µMACS beads. The qPCR amplicons were tested in a minimum of three independent ChIP-qPCR experiments. Quantification results are shown as fold enrichment of anti-FLAG µMACS™ beads using wgIs591 lysate over anti-FLAG µMACS beads using lysate without wgIs591 (no lag-1::TY1::EGFP::3xFLAG). Primer for qPCRs (sequence details above) were designed using Primer3Plus (Untergasser et al., 2007). The FLAG-beads using lysates with lag-1::TY1::EGFP::3xFLAG show specific enrichment for tested target genes thereby validating the specificity of the ChIP. The asterisks indicates p-values < 0.05 (Students t-test). Error bars represent SEM.
Additional files
-
Supplementary file 1
Enhancement or suppression of tumorous phenotypes.
- https://doi.org/10.7554/eLife.15477.023
-
Supplementary file 2
In situ hybridization patterns of Notch-activated genes.
- https://doi.org/10.7554/eLife.15477.024
-
Supplementary file 3
Information on used C. elegans strains, RNAi clones and primers.
(A) Information on C. elegans strains used in the study. (B) Information on RNAi clones used in this study. (C) Information on primer design and sequences.
- https://doi.org/10.7554/eLife.15477.025





