Primate TRIM5 proteins form hexagonal nets on HIV-1 capsids
Figures
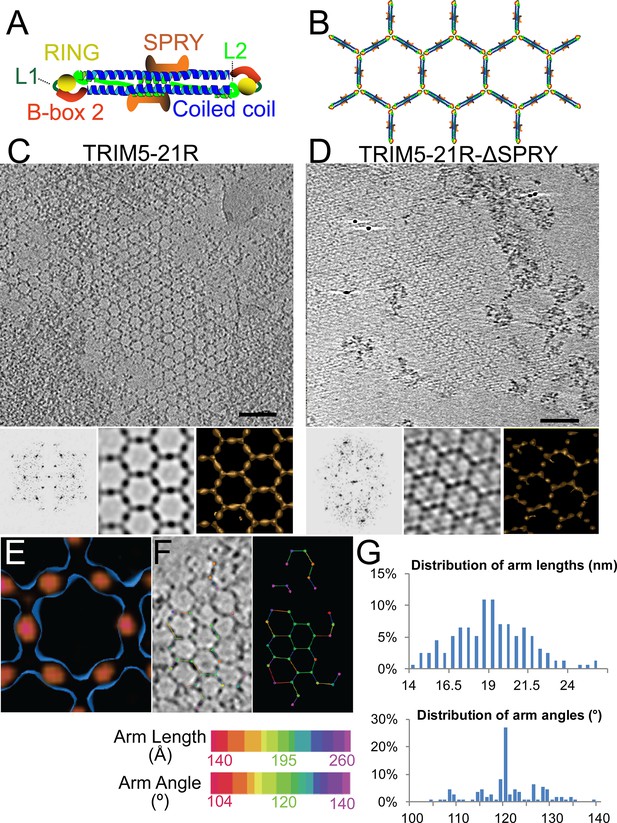
ECT analysis of TRIM5-21R 2D crystals.
(A) Schematic of the TRIM5 dimer. The two RING (yellow) and B-box 2 (red) domains are separated by a ~17 nm, antiparallel dimeric coiled-coil (blue). The two L2 linkers (green) fold back towards the 2-fold axis of the coiled-coil to orient two capsid-binding SPRY domains (orange). (B) Schematic of the TRIM5 hexagonal lattice model. (C and D) Tomographic slice (top) of (C) full-length TRIM5-21R, and (D) TRIM5-21RΔSPRY lattices. Scale bars are 100 nm. In both cases, the computed Fourier transform (bottom, left) and subtomogram average without imposed rotational symmetry (bottom, middle) exhibit six-fold symmetry. Iso-surface representations of the densities are also shown (bottom, right). (E) A density difference map of the subtomogram averages of full-length TRIM5-21R and TRIM5-21RΔSPRY lattices reveals positive density (red) at the center of each hexagon edge, corresponding to the SPRY domain position, supporting the TRIM5α dimer and hexamer models shown in (A and B). (F) Heat maps (bottom) of lattice arm lengths and angles measured from refined lattice points (top) selected from the TRIM5-21R tomogram in (C). (G) Histograms showing the distributions of measured arm lengths (n = 155) and angles (n = 111). The most abundant arm length (18.5–19 nm) and arm angle (120°) are consistent with the structure models in (A and B) and the p6 plane group symmetry of TRIM5-21R 2D crystals (Ganser-Pornillos et al., 2011). Note that this analysis probably underestimates the extent of hexamer variability owing to the initial selection of well-ordered lattice points.
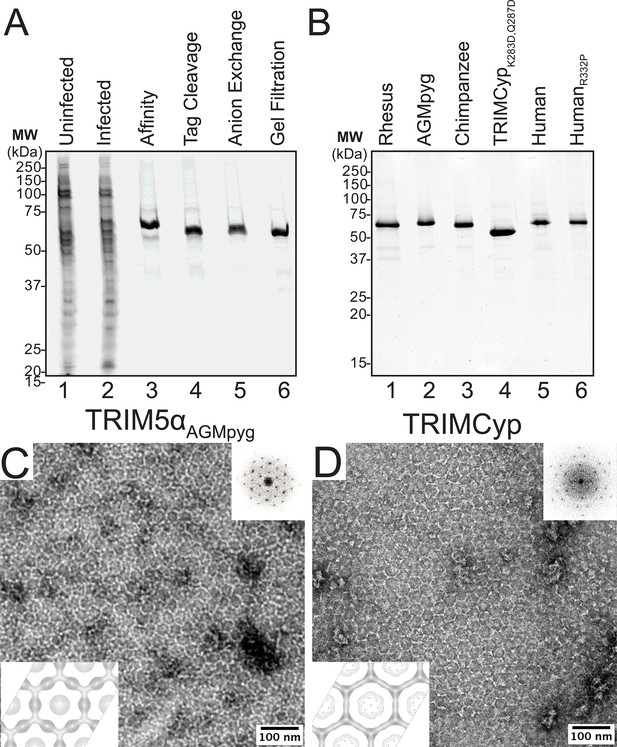
Purification and characterization of recombinant TRIM5 proteins.
(A) Coomassie-stained SDS-PAGE showing the stepwise purification of rhesus TRIM5α (TRIM5αrh). Samples correspond to: soluble lysate from control SF9 cells (Uninfected, lane 1); soluble lysate from SF9 cells expressing OSF-TRIM5αrh (Infected, lane 2); Strep-Tactin affinity-purified OSF-TRIM5αrh (Affinity, lane 3); TRIM5αrh after removal of the OSF tag by HRV14-3C protease treatment (Tag Cleavage, lane 4); dimeric TRIM5αrh purified by Q anion exchange chromatography (Anion Exchange, lane 5); dimeric TRIM5αrh purified by Superdex 200 gel filtration chromatography (Gel Filtration, lane 6). (B) Coomassie-stained SDS-PAGE showing 1.5 µg of purified rhesus, African green monkey pygerythrus (AGMpyg), chimpanzee TRIM5α, proteolysis-resistant owl monkey TRIMCypK283D,Q287D, human TRIM5α, and HIV-1-restricting human TRIM5αR332P. (C,D) TRIM5 hexagonal assembly is a conserved property of primate TRIM5 proteins. Negatively stained EM image of hexagonal arrays formed by (C) TRIM5αAGMpyg and (D) TRIMCyp. Computed Fourier transforms (top right insets) show clear hexagonal order and filtered projection density maps of 2-dimensional crystals (bottom left insets) also reveal hexagonal rings and density distributions reminiscent of TRIM5-21R lattices (Ganser-Pornillos et al., 2011). The unit cell parameters are a = 345 Å, b = 345 Å, γ = 120° (TRIM5αAGMpyg); and a = 345 Å, b = 344 Å, γ = 119° (TRIMCyp). Note that the TRIMCyp samples contained a mixture of full-length TRIMCyp and fragments that were proteolyzed to the C-terminus of residues K283 or Q287 (see Results and Materials and methods for details). The relatively thinner two fold density in the TRIMCyp projection map could either reflect low crystal occupancy of the CypA domain (due to proteolysis) or inherently flexible CypA domains in TRIMCyp as has been proposed by (Goldstone et al., 2014).
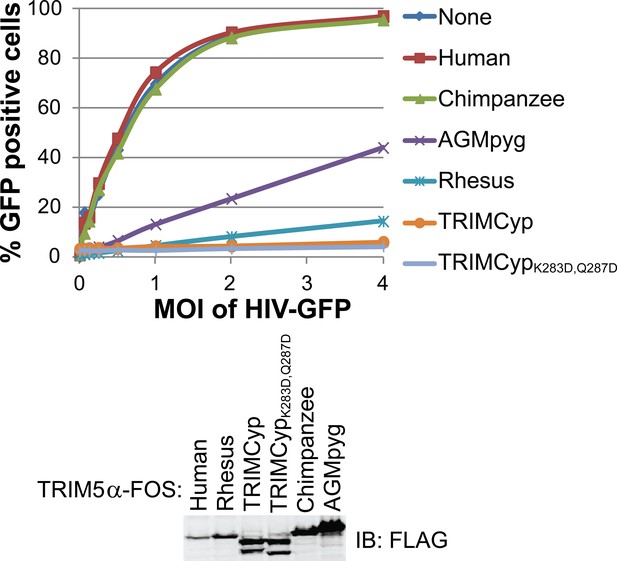
HIV-1 CA restriction activity of different TRIM5 alleles.
HeLa cells were transduced with vectors expressing TRIM5αhu, TRIM5αcpz, TRIM5αAGMpyg, TRIM5αrh, TRIMCyp, or TRIMCypK283D,Q287D. Cells express TRIM5 proteins at approximate similar levels as verified by western blotting with α-TRIM5α antibody (bottom). Integrated band intensities for this blot were calculated and normalized to the value of TRIM5αrh: 0.5 (TRIM5αhu), 1.0 (TRIM5αrh), 1.3 (TRIMCyp, upper band), 1.3 (TRIMCypK283D,Q287D, upper band), 1.8 (TRIM5αcpz) and 3.2 (TRIM5αAGMpyg). Cells were transduced with VSV-G pseudotyped HIV-GFP reporter virions at the designated MOI levels (top). Percentages of infected cells (GFP positive cells) were determined by FACS. As expected, TRIM5 proteins from AGMpyg, rhesus and owl monkey restricted HIV-1 whereas human and chimpanzee TRIM5 did not. Note also that introduction of proteolysis-resistant mutations (K283D, Q287D) into TRIMCyp did not affect restriction. Each experiment was repeated twice with similar results.
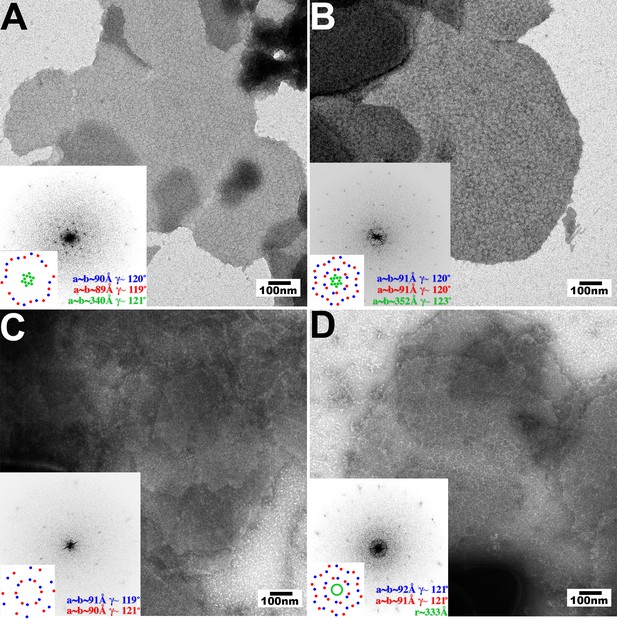
Assembly of restricting TRIM5 proteins on 2D crystals of HIV-1 CA.
Negative stain EM images of CA 2D crystals decorated with (A) TRIM5αrh, (B) TRIMCypK283D,Q287D, (C) TRIM5αhu (non-restricting allele), (D) TRIM5αhu,R332P (restricting mutant). Scale bars are 100 nm. Computed Fourier transforms (insets) and indexing (second insets) show the first and second order reflections of two CA lattices and their unit cell parameters (red and blue) as well as diffraction spots (A, B) or rings (D) corresponding to the first order reflections of the TRIM5 lattices (green).
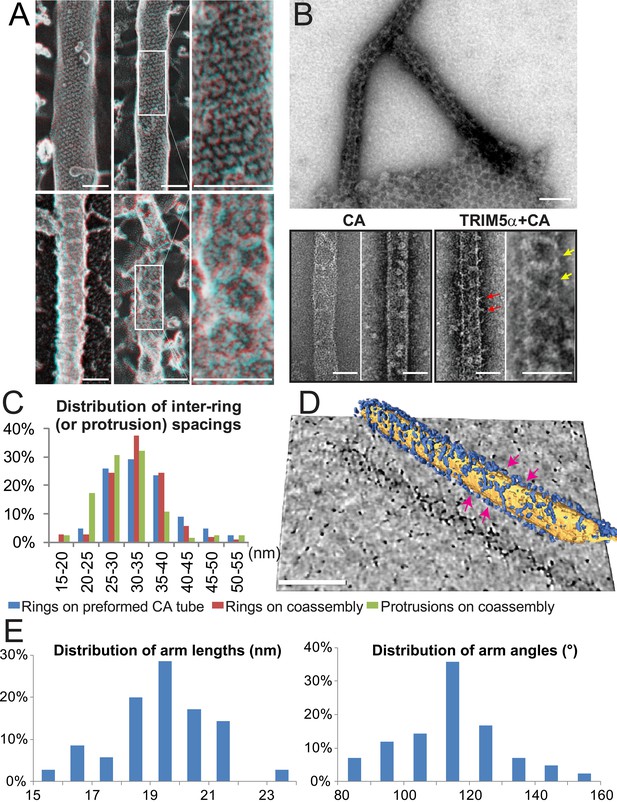
Assembly of TRIM5α proteins on HIV-1 CA tubes.
(A) Deep-etch electron micrographs of control hyperstable CA tubes (top) and TRIM5α decorated CA tubes (bottom) with blow-up views of boxed regions to the right. Scale bar is 50 nm. (B) Negative stain electron micrographs of co-assembled TRIM5αAGMpyg-coated CA tubes (top). Scale bar is 100 nm. Expanded views of negatively stained CA assembly in the absence (bottom left) or presence of TRIM5αAGMpyg proteins (bottom right). TRIM5α formed ring-like decorations (yellow arrows) on the tube surface and displayed protrusions along the edge of the tube (red arrows). Similar decorations were not observed in the control case. The protrusions were regularly spaced and we speculate that they are either well-ordered coiled-coil arms from adjacent hexagons wrapping around the CA tubes or possibly ordered RING domains projecting outward from the lattice. Scale bars are 50 nm. (C) Histograms showing the distributions of measured inter-ring spacings in the TRIM5αAGMpyg–decorated tubes (blue bars, n = 170) and in co-assemblies (red bars, n = 164), and of inter-protrusion spacings in co-assemblies (green bars, n = 166). The most abundant inter-ring spacing (30–35 nm) is consistent with the spacing of TRIM5-21R 2D crystals, indicating similar structures. (D) Electron cryotomography (ECT) reveals that TRIM5α forms hexagonal nets on the surface of CA tube. Hexagon-like rings are marked by magenta arrows. Scale bar is 80 nm. (E) Histograms showing the distributions of arm lengths (n = 35) and angles (n = 42) measured from three-fold vertices in ECT.
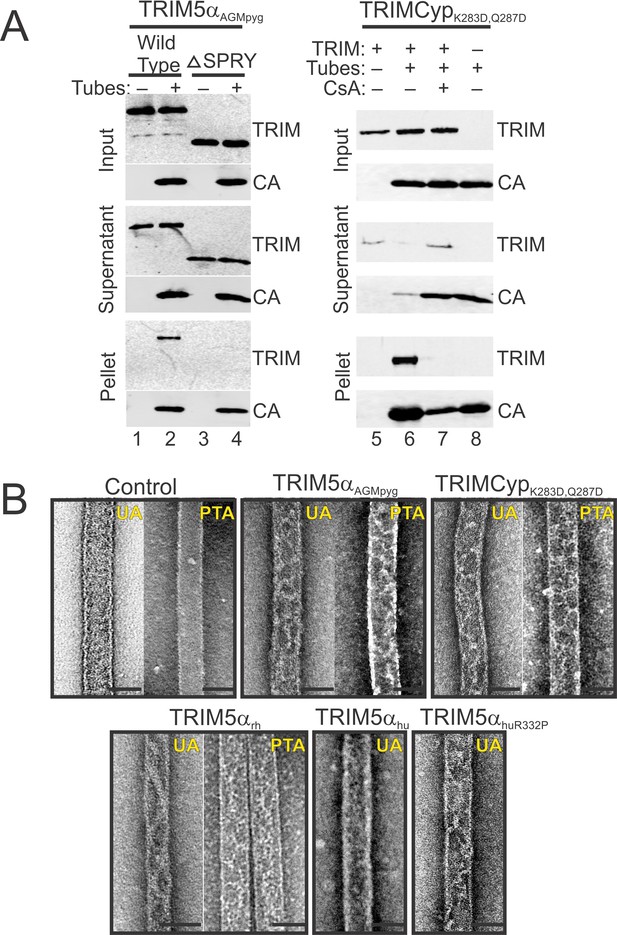
TRIM5 protein binding to hyperstable HIV-1 CA tubes.
(A) (left) Sucrose co-sedimentation binding experiments were performed using either wild type, full-length TRIM5αAGMpyg (Wild Type, lanes 1, 2), or a truncated TRIM5αAGMpyg that lacked the SPRY domain (ΔSPRY, lanes 3, 4), in the absence (lanes 1, 3) or presence of CA tubes (lanes 2, 4). Pelletable CA and associated TRIM5αAGMpyg (Pellet, 30% in total), were separated from unassembled and soluble CA proteins and unbound TRIM5αAGMpyg (Supernatant, 3% of total) by centrifugation through a sucrose cushion, and analyzed by western blotting, with the input levels of both proteins shown for reference (Input, 3% of total). (right) Analogous experiments performed with proteolysis-resistant TRIMCypK283D,Q287D in the absence (lane 5) or presence (lanes 6, 7) of helical CA tubes, without (lanes 5, 6) or with added cyclosporine A (CsA) (lane 7), a competitive inhibitor of the CypA-CA interaction. Lane 8 is a CA tube control, with no added TRIMCypK283D,Q287D or CsA. Representative results from one of three independent experiments are shown. (B) Representative electron micrographs of control HIV-1 CA tubes, and tubes decorated with different TRIM5α or TRIMCypK283D,Q287D and negatively stained with Uranyl Acetate (UA) or Phosphotungstate (PTA). Scale bars are 50 nm.
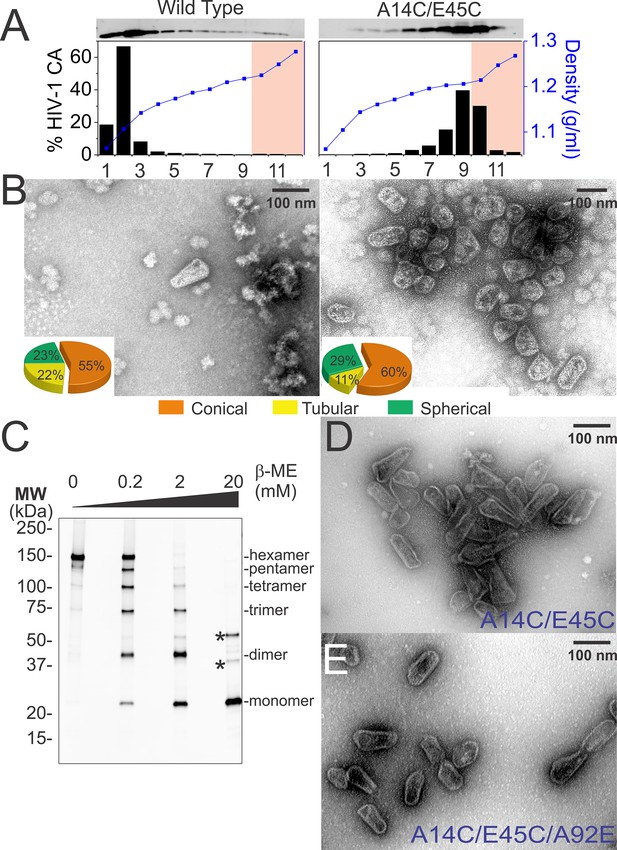
Purification of wild type and hyperstable HIV-1 cores.
(A) Sucrose-gradient purification profiles of wild type (left) and hyperstable A14C/E45C (right) HIV-1 cores (Kotov et al., 1999). (top) α-CA western blots of sucrose gradient fractions, (bottom) graph showing quantified CA levels (histogram) and solution density (blue line; g/ml) in each gradient fraction. The higher stability of crosslinked HIV-1 cores can be seen by comparing the amounts of wild type and CAA14C/E45C in fractions 10–12 (pink regions). Core-containing fractions were pooled, washed, and concentrated for the analyses in B and C. The experiment was repeated at least three times with similar results. (B) Negative stain electron micrographs of wild type (left) and CAA14C/E45C (right) cores. Pie charts (inset) of observed morphologies of wild type (left; n = 143 cores) and CAA14C/E45C cores (right; n = 353 cores) reveal that the introduced Cys crosslinks do not alter HIV-1 core morphologies significantly. (C) Non-reducing α-CA western blots showing that CAA14C/E45C cores are crosslinked (compare % hexamer as a function of [β-ME]). Asterisks indicate unprocessed Gag fragments (p55 and p41) that co-purifed with mature cores during purification. (D, E) Negative stain electron micrographs showing the relative abundance and purity of cores purified by the affinity method. (D) CAA14C/E45C crosslinked cores and (E) CAA14C/E45C/A92E crosslinked cores. Note that the additional A92E mutation reduced core clustering. Scale bar is 100 nm.
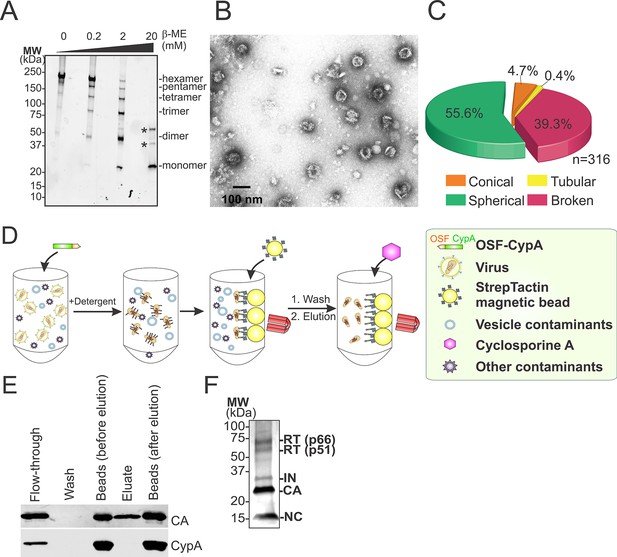
Hyperstable core particle characterization and purification.
(A-–C) CAA14C/E45C assemblies in fractions 7–9 are also composed of crosslinked hexamers. (A) Non-reducing α-CA western blot of CAA14C/E45C assemblies in fractions 7–9. (B) Negative stain electron micrograph of fractions 7–9. (C) Pie chart showing the atypical morphological distribution of particles in fractions 7–9. (D–E) HIV-1 core affinity purification and characterization. (D) Purification scheme (key at right). Pure, recombinant OneSTrEP-FLAG-cyclophilin A (OSF-CypA) was added to HIV-1 virions and their limiting membrane was stripped by detergent treatment. OSF-CypA-core complexes were captured using Strep-Tactin sepharose magnetic beads. The beads were restrained magnetically and vesicles and other bound contaminants were removed by rigorous washing. Cores (but not CypA) were then eluted by addition of cyclosporine A. (E) α-CA (top) and α-FLAG (CypA; bottom) western blots of samples from different steps in HIV-1 core affinity purification (see Materials and methods for details). Note that the eluate, which contains purified cores, is free from bound CypA. (F) SDS-PAGE gel with silver staining reveals that the expected protein components of HIV-1 cores are all present in purified hyperstable cores.
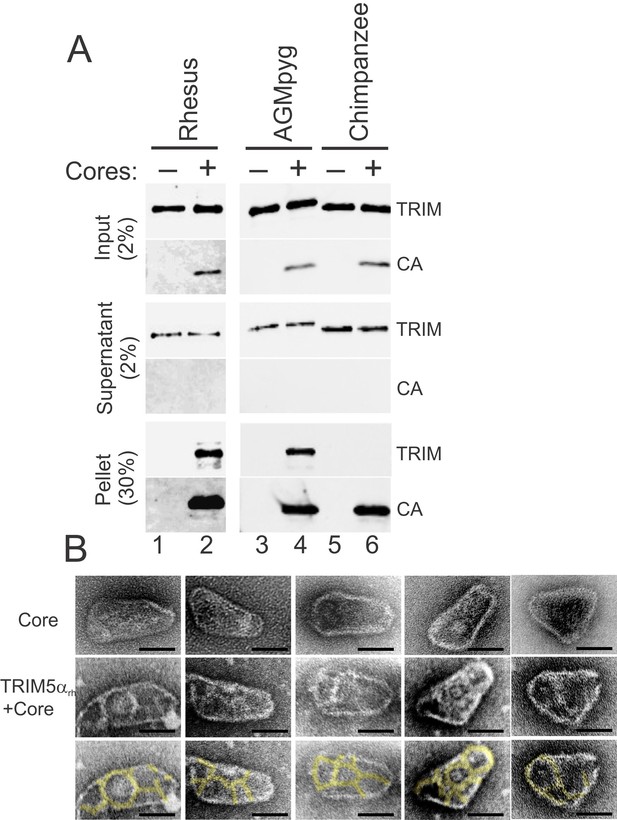
TRIM5α proteins bind directly to HIV-1 cores.
(A) Sucrose cushion co-sedimentation binding assay for TRIM5α-HIV-1 core interactions. TRIM5α proteins were incubated in the absence of cores (lanes 1, 3, 5) or in the presence of hyperstable HIV-1 cores (lanes 2, 4, 6), and the mixtures were subjected to centrifugation through the sucrose cushion. Pelletable cores and bound TRIM5α (Pellet, 30% of total) and unbound TRIM5α (Supernatant, 2% of total) were analyzed by western blotting for TRIM5α and CA proteins. The input levels of both proteins are also shown for reference (Input, 2% of total). Representative results from one of three independent experiments are shown. (B) Representative electron micrographs of control HIV-1 cores (first row), and cores decorated with TRIM5αrh (second row) and negatively stained with uranyl acetate. TRIM5α decorations were highlighted in yellow (third row). Scale bars are 50 nm.
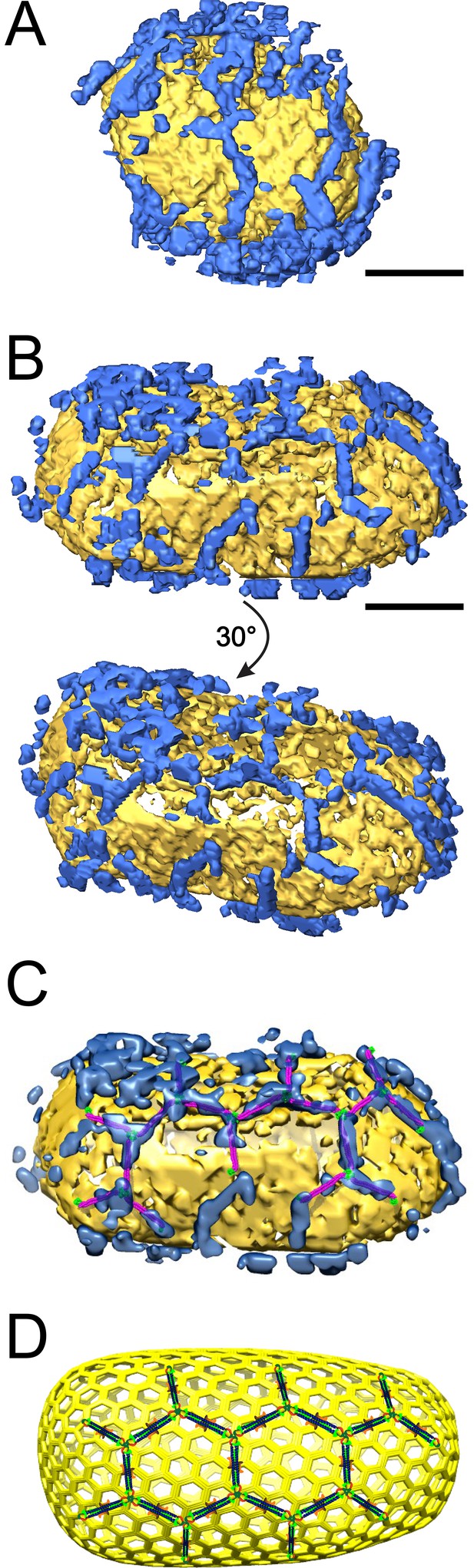
ECT reveals that TRIM5α forms flexible hexagonal nets on hyperstable HIV-1 cores.
(A) Segmented HIV-1 core (yellow) decorated with TRIM5α (blue). (B) Segmented HIV-1 core decorated with TRIM5α and subjected to mild sulfo-EGS crosslinking prior to vitrification. (C) TRIM5α structural model docked into the cryoEM volume of the tomogram shown in (B). (D) Idealized schematic model of an HIV-1 fullerene cone bound by a TRIM5α hexagonal net. Domains and linkers of TRIM5α are colored as described in Figure 1A. Scale bars are 35 nm.
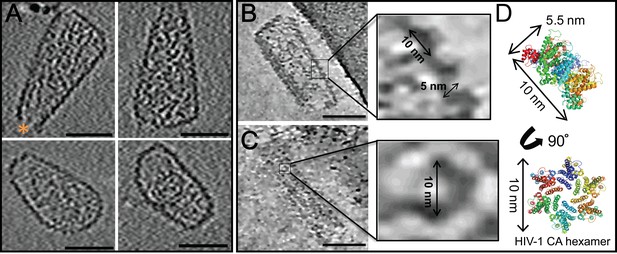
ECT of hyperstable HIV-1 cores.
(A) Tomographic slices of four representative HIV-1 cores, displaying no densities above background noise around their exterior. A hole at the tip is marked by an orange asterisk (Yu et al., 2013). Scale bar is 50 nm. (B, C) Tomographic slices of non-linear anisotropic diffusion (NAD)-filtered cores display side- and front-facing views of individual CA hexamers. Blow-up views show densities with dimensions similar to those of the crystal structure of the HIV-1 CA hexamer. (D) Crystal structure of the HIV-1 CA hexamer whose dimensions match those of the densities highlighted in (B) and (C).
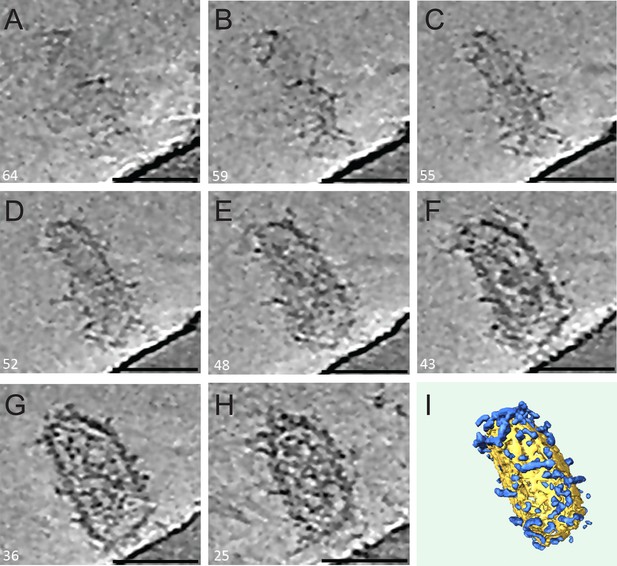
ECT of a HIV-1 hyperstable core in complex with TRIM5α.
(A–H) Series of slices through the Z axis of a tomogram of one HIV-1 core, displaying densities above background noise around their exterior (sulfo-EGS crosslinked). The tomograms were SIRT reconstructed and NAD filtered. Slice numbers are indicated in the bottom left corner of each frame. Scale bars are 50 nm. (I) Segmentation of the tomogram shown in (A–H) reveals the distribution of TRIM5α (blue) on the HIV-1 core surface (yellow).
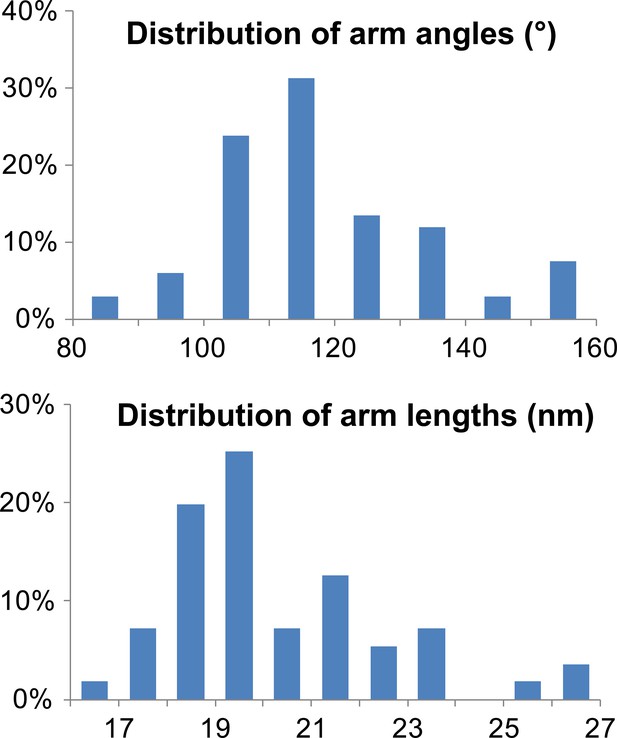
Characterization of TRIM5 hexagonal nets on HIV-1 cores.
Histograms showing the distribution of arm angles (n = 68, top) and lengths (n = 51, bottom) of TRIM5 hexamers in ECT.
Videos
HIV-1 CA tube co-assembled with TRIM5α and crosslinked.
ECT of HIV-1 CA tube (yellow) decorated with TRIM5α (blue). Video corresponds to Figure 4D.
HIV-1 core decorated with TRIM5α (without crosslinking). ECT of HIV-1 core (yellow) decorated with TRIM5α (blue).
Video corresponds to Figure 7A.
TRIM5α-HIV-1 cores associated with the air/water interface.
Representative ECT showing the interaction of TRIM5α-HIV-1 cores with the air/water interface. Video corresponds to Figure 7.
HIV-1 core decorated with TRIM5α and crosslinked.
HIV-1 core (yellow) decorated with TRIM5α (blue) were subjected to crosslinking and imaged by ECT. Video corresponds to Figure 7B and C.
Additional files
-
Supplementary file 1
Plasmids and pure recombinant TRIM5 proteins used in this study.
(A) List of plasmids used in this study. (B) TRIM5 constructs and their molecular weights.
- https://doi.org/10.7554/eLife.16269.020





