Structure of the active form of human origin recognition complex and its ATPase motor module
Figures
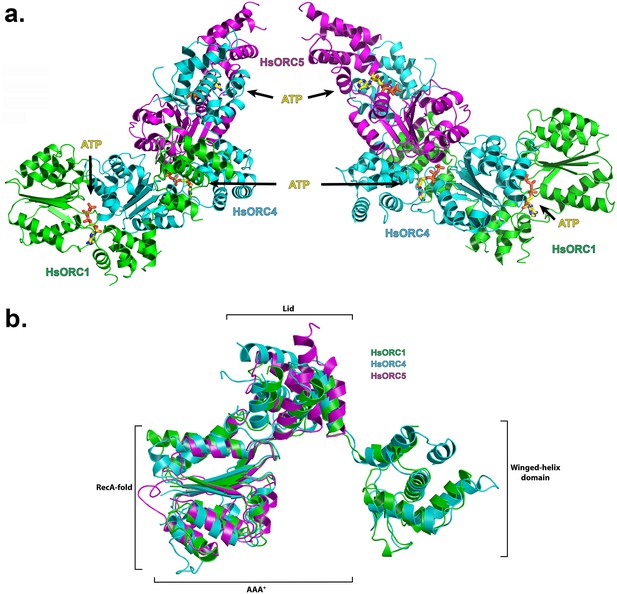
Structure of the motor module of HsORC.
(a) Ribbon diagram of the motor module resembling a cashew nut, with ORC1 in green, ORC4 in cyan and ORC5 in purple. The three ATP molecules nestled between the domains are shown in stick representation. The RecA-fold domains or ORC4 and ORC5 are cupped between the RecA-fold and lid domains of ORC1 and ORC4, respectively. The WHDs of ORC1 and ORC4 are positioned above the RecA-fold domains of ORC4 and ORC5, respectively. (b) Superposition of ORC1, ORC4 and ORC5, showing an identical organization between the domains of each ORC subunit.
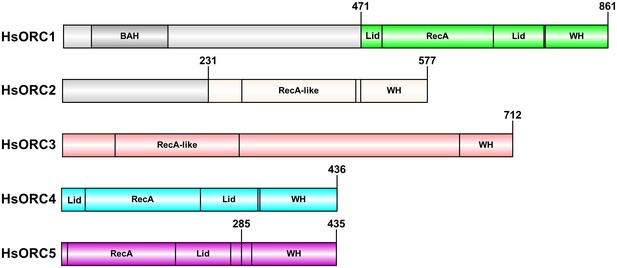
HsORC constructs used in this study.
Bar diagram illustrating the constructs and domain structure of HsORC subunits. The colored regions were used in this study. For ORC5, the construct used for the motor module extends till residue 285, whereas full-length ORC5 was used in the EM reconstruction.
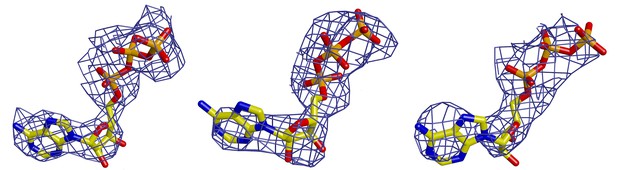
ATP nucleotides at the motor module subunit interfaces.
SigmaA-weighted omit maps of three of the six ATP nucleotides at the three different interfaces from left to right: at the ORC1-4 interface, at the ORC4-5 interface, at ORC5 ATP-binding site.
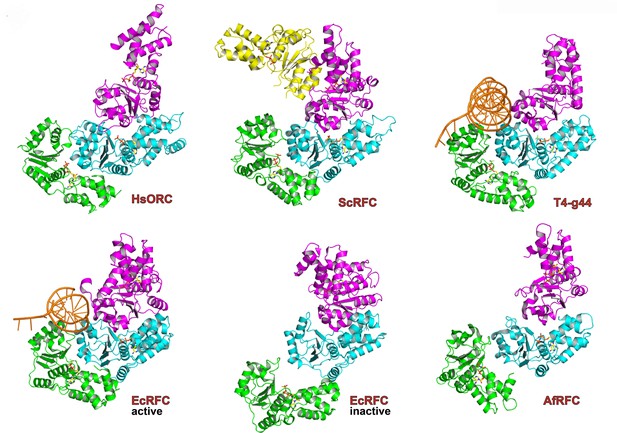
Similarity of the HsORC motor module with DNA polymerase clamp loader complexes.
Top row: HsORC1/4/5 (this work); yeast RFC-B, -C, -D, -E subunits with nucleotide from the RFC clamp loader bound to the PCNA clamp (Bowman et al., 2004) (1SXJ); gp44 subunits with nucleotide from the T4 bacteriophage clamp loader bound to an open clamp and DNA (Kelch et al., 2011) (1U60). Bottom row: γ subunits with nucleotide from the E. coli clamp loader bound to DNA (Simonetta et al., 2009) (3GLF); γ subunits from the inactive E. coli clamp loader (Kazmirski et al., 2004) (1XXH); small subunits bound to ADPNP from the Archaeoglobus fulgidus RFC complex (Seybert et al., 2006) (2CHQ). All proteins were aligned based on the subunit depicted in cyan (ORC4 in the case of HsORC).
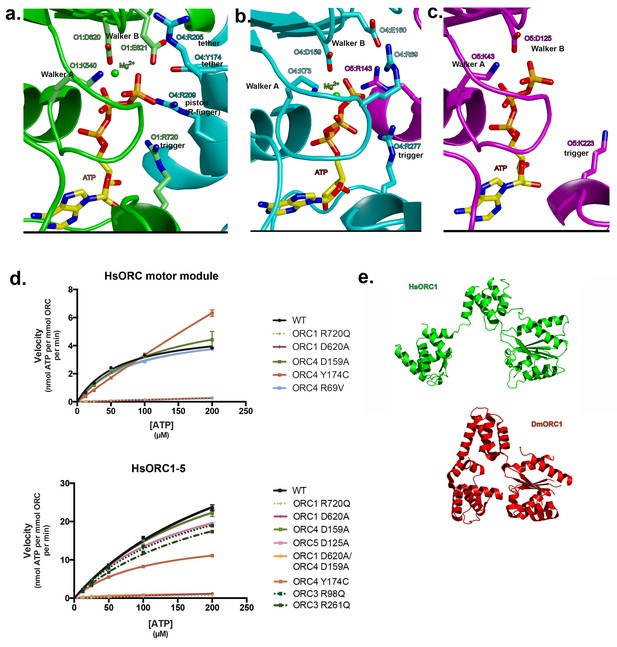
Motor module ATPase.
(a) ATP-binding site at the ORC1/4 interface. (b) ATP-binding site at the ORC4/5 interface. (c) ATP-binding site of ORC5. ORC1 is shown in green, ORC4 in cyan and ORC5 in purple. ATP and key residues discussed in the text are shown in stick representation. (d) Michaelis-Menten curves for ATPase activity of the motor module (top), HsORC1-5 (bottom), for the wild type and several mutants from experiments performed in triplicate. (e) Different domain organization of HsORC1 (green) compared with the more compact DmORC1 (red).
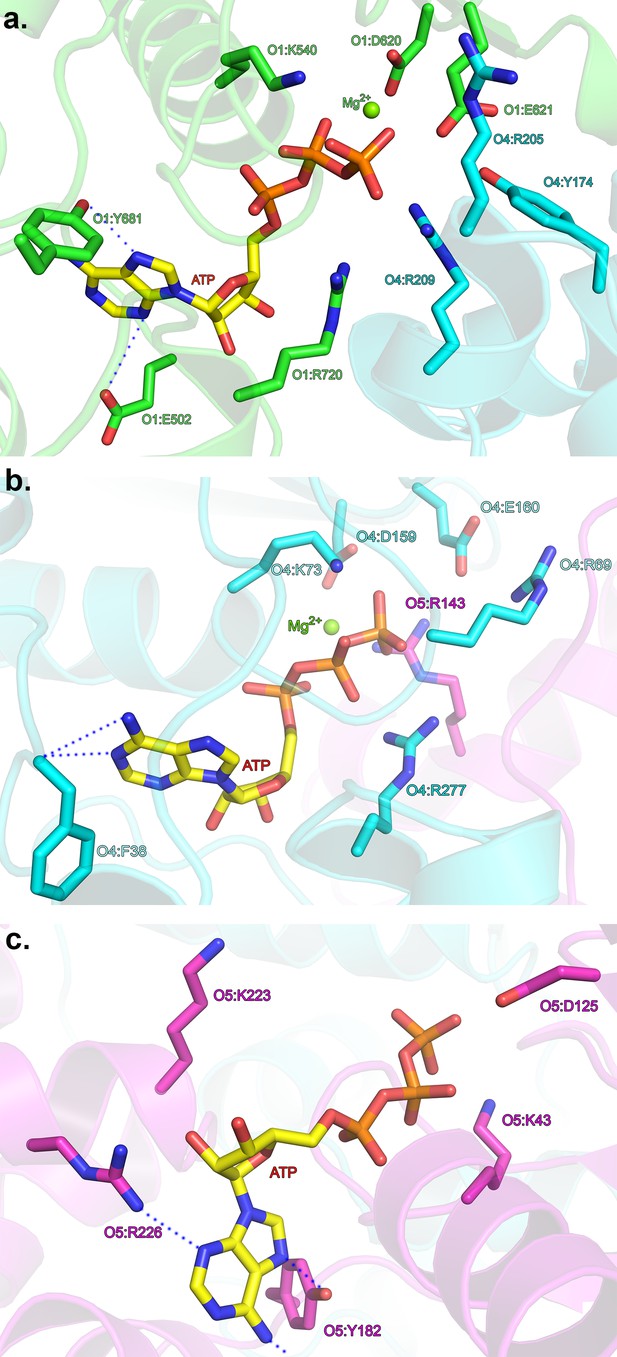
Interactions with the adenine base at the ATP-binding sites.
(a) ATP-binding site at the ORC1/4 interface. (b) ATP-binding site at the ORC4/5 interface. (c) ATP-binding site of ORC5. ORC1 is shown in green, ORC4 in cyan and ORC5 in purple. ATP and key residues discussed in the text are shown in stick representation.
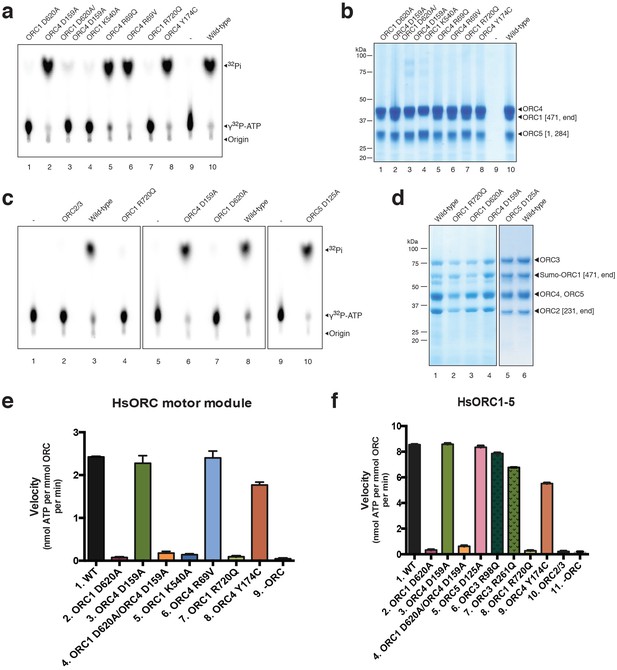
HsORC ATPase activities.
Representative samples from ATPase assays. (a) Thin layer chromatography (TLC) of ATPase activity assays of the ORC motor module and mutants as indicated. (b) Coomassie-stained SDS-PAGE gel showing purity and expression levels of the various wild type and mutant motor module complexes used in the reactions. (c) TLC of ATPase activity assays of the ORC1-5 complex and ORC2/3 subcomplex. (d) As in b for the ORC1-5 complex. (e) Histogram showing the rates of ATPase from wild type or mutant ORC motor module at 50 μM ATP. (f) Same as e for the ORC1-5 complex. Experiments were performed in triplicate.
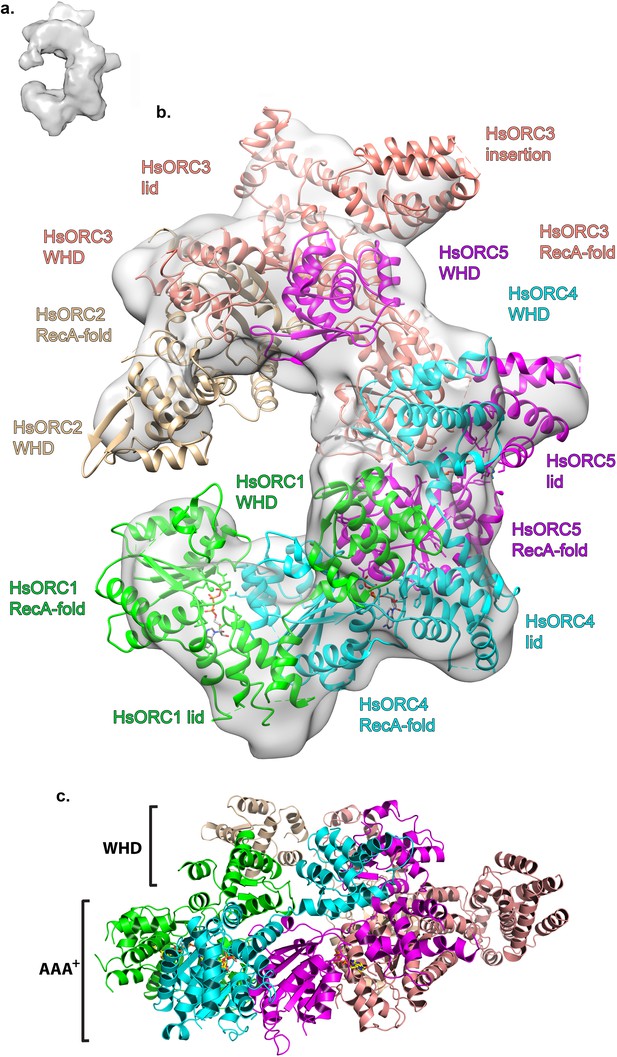
Structure of HsORC.
(a) Cryo-EM density of HsORC1-5, back view. Note that the top and bottom are flipped compared with the maps shown in the supplementary Figures. (b) Ribbon diagram of HsORC modeled into cryo-EM density. The ORC motor module is colored as in Figure 1. ORC2 is shown in wheat and ORC3 is shown in salmon. ATP is shown in stick. (c) Side view with the WHD layer on top and the AAA+ layer on the bottom. The ORC3 insertion can be seen extending from the particle on the right.
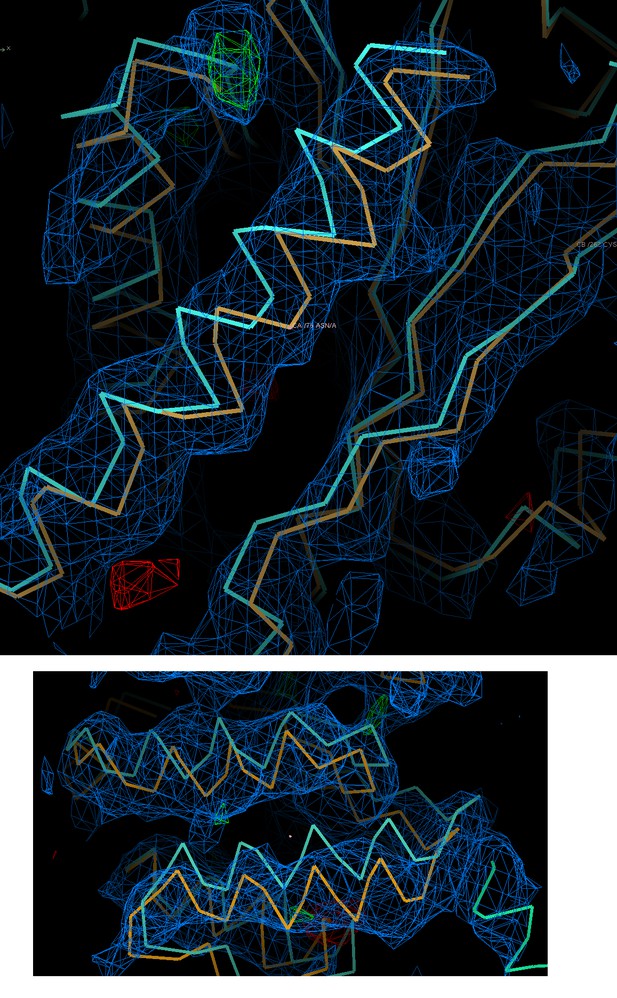
Electron-density maps for the HsORC2/3 complex.
Electron-density map directly following molecular replacement using DmORC2/3 as a search model contoured at 1σ. This illustrates that even at this modest resolution the crystallographic data has reasonable phasing ability. Shown are two different regions with the search model in cyan and the fitted model in orange.
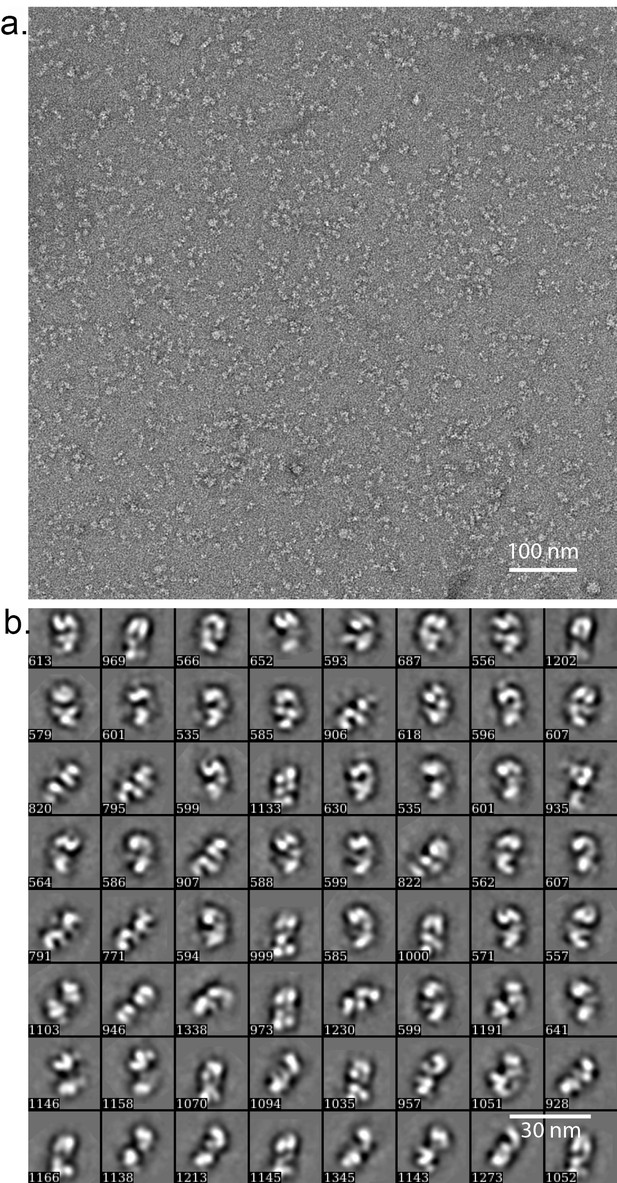

Negative stain EM of the truncated human ORC1-5.
(a) A typical raw micrograph of the HsORC particles stained in uranyl acetate. (b) 2D class averages. Numbers at the lower left corner in each panel refers to the number of particles used to compute the average images.
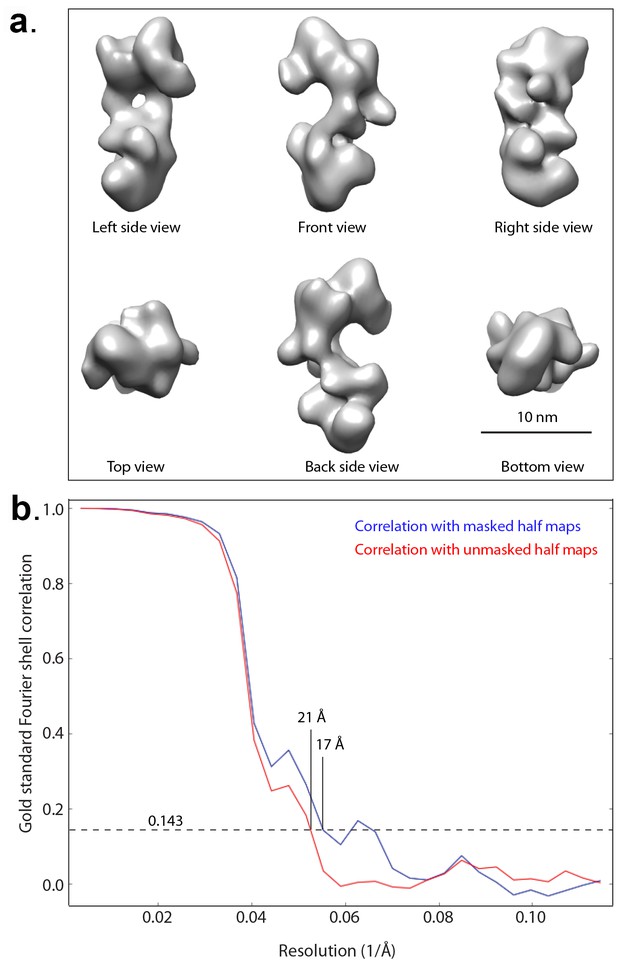
Negative stain 3D EM map of the truncated HsORC.
(a) The 3D map in six different views. Note that in these views, the top and bottom are flipped (upside down) compared with Figure 4. (b) Gold standard Fourier Shell correlation of the reconstructed 3D EM map.
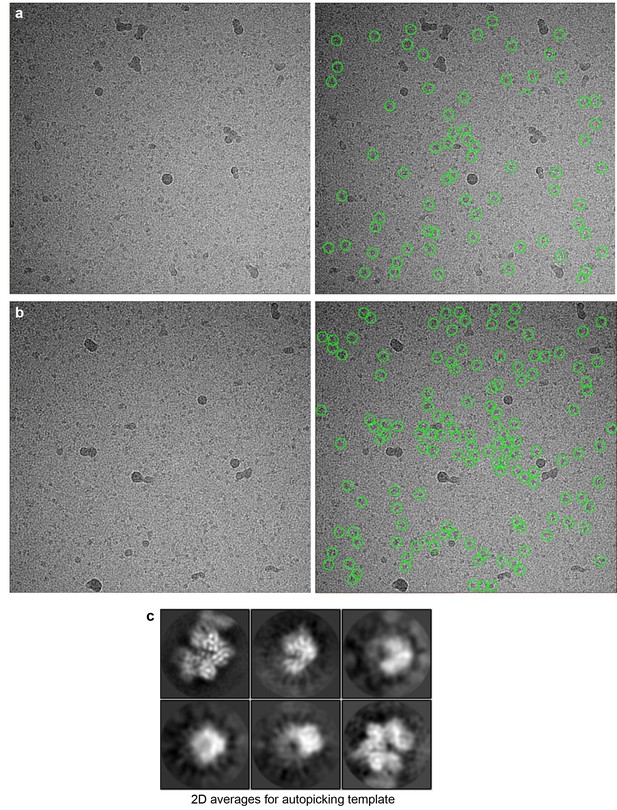
Generation of templates for automatic particle picking.
(a) Left panel, a typical raw image for initial manual particle picking. Right panel, around 100 particles marked by green circles were picked manually. (b) Left panel, a second raw image used for manual particle picking. Right panel, Selected particles were marked by green circles. (c) Following 2D classification, six class averages were used as templates for autopicking in Relion.
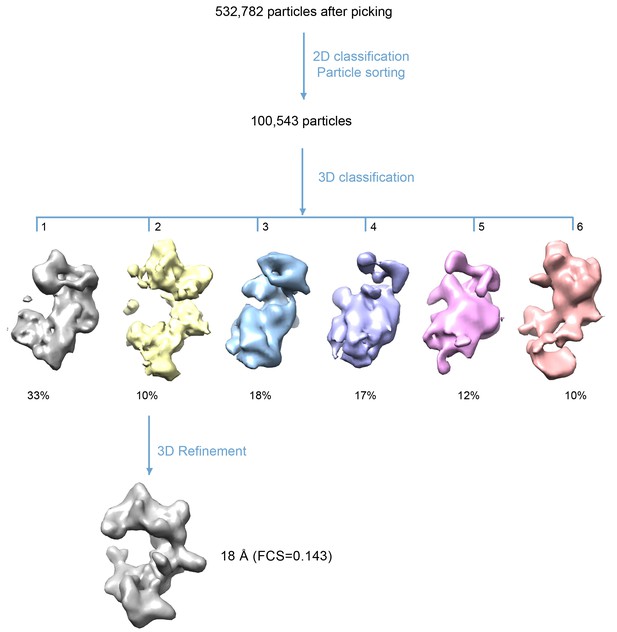
3D classification procedure used to derive the final 3D-density map of the HsORC.
532,782 raw particles were selected from drift corrected electron micrographs. 2D classification and sorting rejected ~80% of particles that did not yield good averages or appeared to be broken particles. 3D classification into six groups resulted in one map of the expected shape that was similar to the previously published EM structures of the yeast or Drosophila ORC. The other five maps were either partial structures or distorted. Note that in these views, the top and bottom are flipped (upside down) compared with Figure 4. Refinement using the selected ~10,500 particles led to the final reported 3D map.
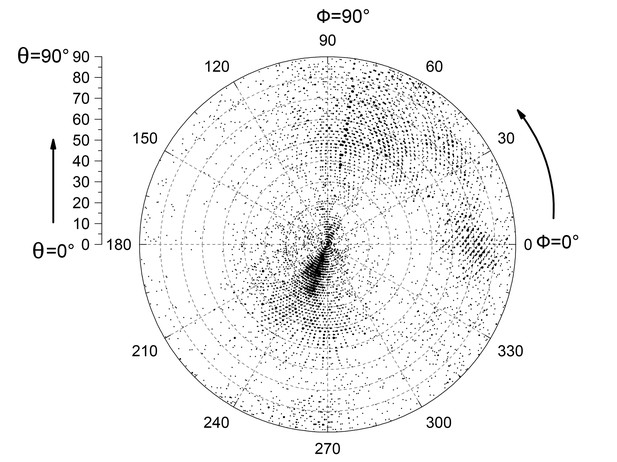
Eulerian angle distribution of the cryo-EM particles used in 3D refinement.
Euler angle distribution of 10,357 particles used in refinement of the 20 Å 3D map. The particle orientation covers nearly all angular space.
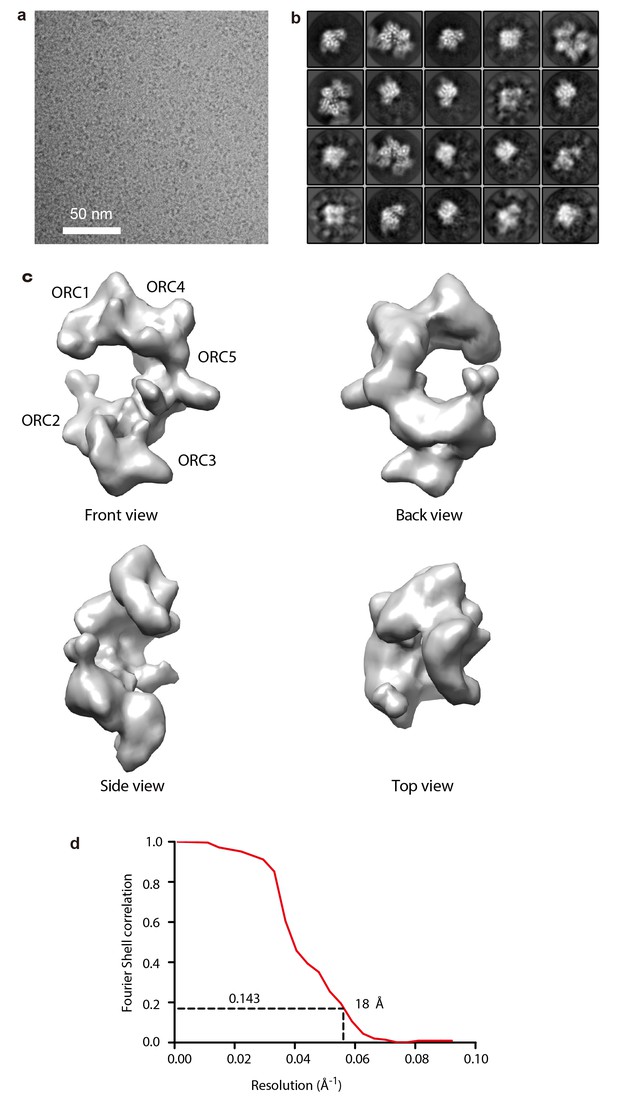
Cryo-EM of the truncated human ORC1-5 complex.
(A) A typical motion-corrected raw image of HsORC particles frozen in vitreous ice recorded on a K2 direct detector. (B) Selected 20 2D class averages. (C) 3D cryo-EM map of HsORC, front, back, side and top views are showed separately. Note that in these views, the top and bottom are flipped (upside down) compared with Figure 4. (D) The gold standard Fourier shell correlation suggests a resolution of 18 Å at the 0.143 threshold. Due to drop off of the FSC curve starting as early as at ~30 Å, the map may only contain information up to ~20 Å resolution.
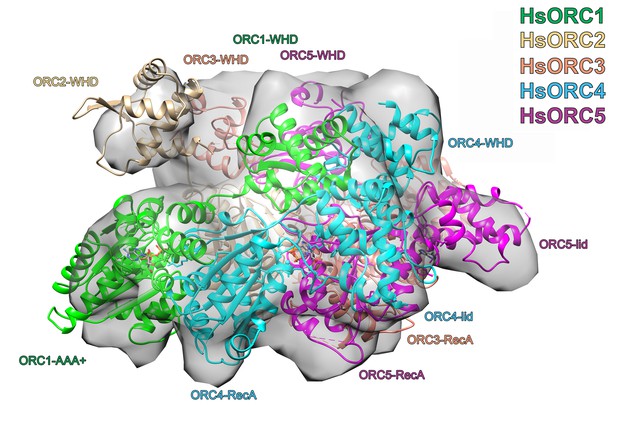
Structure of HsORC.
Ribbon diagram of HsORC modeled into cryo-EM density in an orthogonal view to that shown in Figure 3b. The ORC motor module is colored as in Figure 1. ORC2 is shown in wheat and ORC3 is shown in salmon. ATP is shown in stick. Many of the domains are labeled.
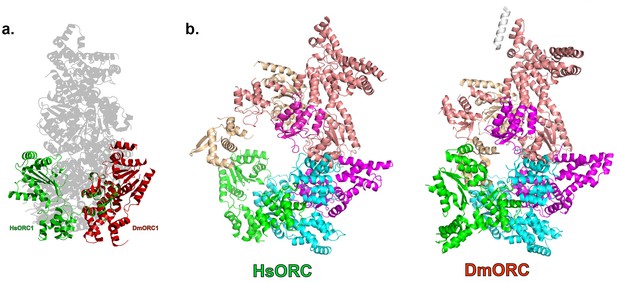
HsORC is in an active conformation.
(a) Different positions of HsORC1 and DmORC1 in the context of the bigger particle. HsORC1 is shown in green, DmORC1 is shown in red and the rest of the HsORC complex is shown in grey. (b) The subunit organization in HsORC results in a more open architecture compared with DmORC.
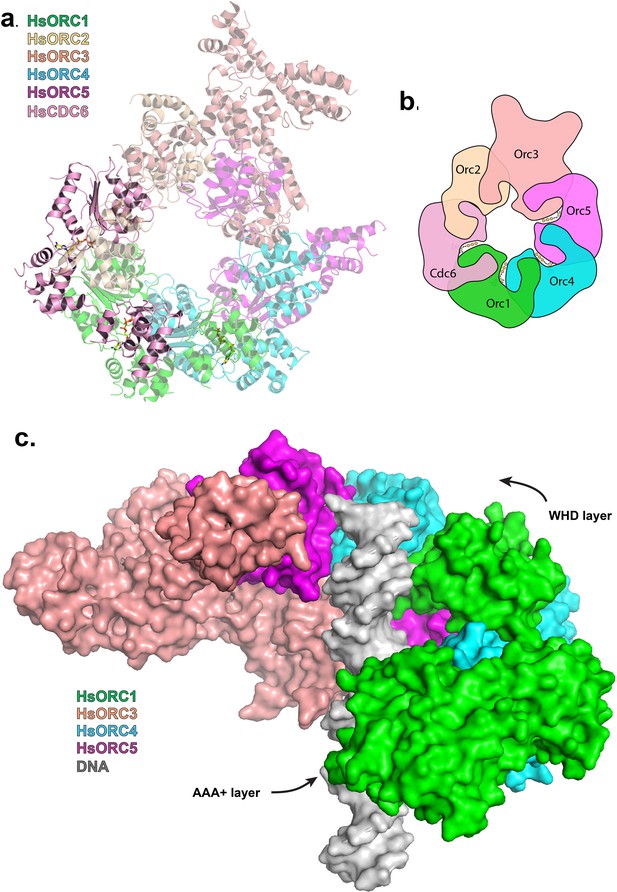
Modeled interaction of HsORC with CDC6 and DNA.
(a) Model of HsORC-CDC6 with CDC6 in pink. HsORC1-5 subunits are shown in a transparent rendering. (b) Cartoon depiction of HsORC-CDC6 showing the organization of the particle with the smaller WHDs forming one layer on top of the bigger AAA+ layer on the bottom. It also clearly illustrates the WHD from one subunit sitting on top of the RecA-fold domain of the neighboring subunit. (c) Model of HsORC with DNA. ORC2 and CDC6 were removed for clarity.
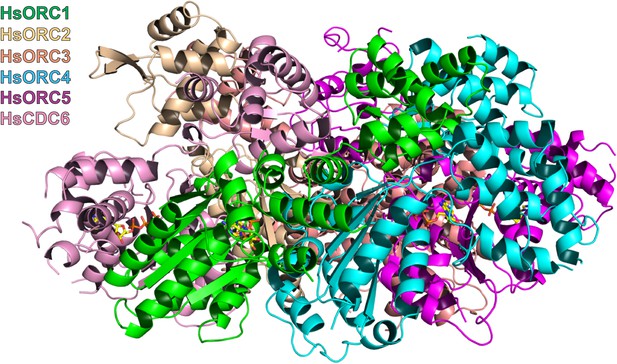
Model of interaction of HsORC with CDC6.
Side view of the model of HsORC-CDC6 with CDC6 in pink. CDC6 completes the ring without any additional movements.
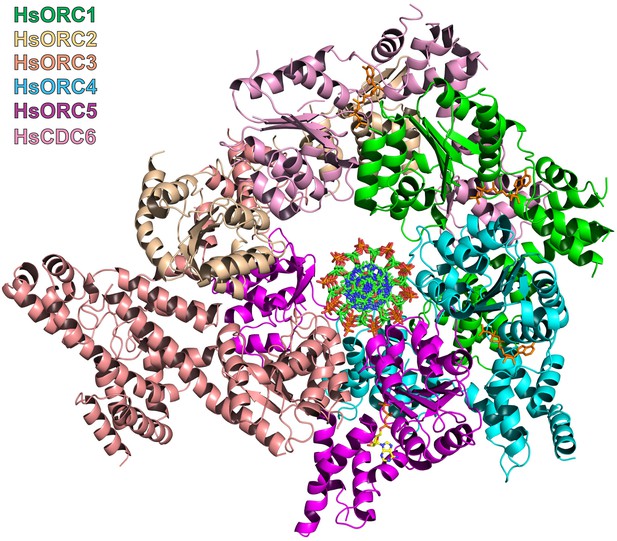
Model of interaction of HsORC with DNA.
Top view of the model of HsORC-CDC6 with DNA.
Videos
Ribbon diagram of HsORC modeled into cryo-EM density.
ORC1 is shown in green, ORC4 in cyan, ORC5 in purple, ORC2 in wheat and ORC3 in salmon. ATP is shown in stick.
A morphing of the structure of DmORC into HsORC.
HsORC1 moves to form an active ATP interface with HsORC4. HsORC2 moves out and up from the more ‘collapsed’ placement in DmORC.
Tables
Data collection and refinement statistics for HsORC motor module*.
| Data collection | |
|---|---|
| Wavelength (Å) | 0.9793 |
| Resolution range (Å) | 19.88–3.39 (3.52–3.39) |
| Space group | P21 |
| Unit cell (Å,°) | 120.89 81.14 151.95 90 97.25 90 |
| Total reflections | 148448 (14543) |
| Unique reflections | 39700 (3849 ) |
| Multiplicity | 3.7 (3.8) |
| Completeness (%) | 0.97 (0.96) |
| Mean I/s(I) | 8.75 (1.19) |
| Wilson B-factor (Å2) | 104.85 |
| R-merge | 0.1385 (1.406) |
| R-measure | 0.1619 (1.641) |
| CC1/2 | 0.997 (0.524) |
| CC* | 0.999 (0.829) |
| Refinement | |
|---|---|
| Resolution range (Å) | 19.88–3.39 (3.52–3.39) |
| Reflections used in refinement | 39700 (3849) |
| Reflections used for R-free | 1963 (198) |
| R-work | 0.2421 (0.3745) |
| R-free | 0.2811 (0.4015) |
| CC(work) | 0.959 (0.662) |
| CC(free) | 0.923 (0.591) |
| Number of non-hydrogen atoms | 15870 |
| Macromolecules | 15678 |
| Ligands | 192 |
| Protein residues | 1947 |
| RMSD bonds (Å) | 0.004 |
| RMSD angles (°) | 0.72 |
| Ramachandran favored (%) | 95 |
| Ramachandran allowed (%) | 4.6 |
| Ramachandran outliers (%) | 0 |
| Rotamer outliers (%) | 0.57 |
| Clashscore | 3.59 |
| Average B-factor (Å2) | 127.05 |
| Macromolecules | 127.23 |
| Ligands | 112.92 |
| Number of TLS groups | 31 |
-
*Values in parentheses are for the highest resolution shell.
Data collection and refinement statistics for HsORC2/3*.
| Data collection | |
|---|---|
| Wavelength (Å) | 0.9793 |
| Resolution range (Å) | 20.07–6.00 (6.21–6.00) |
| Space group | P21 |
| Unit cell (Å,°) | 87.26 114.96 316.46 90 90.72 90 |
| Total reflections | 52200 (8302) |
| Unique reflections | 15430 (2286) |
| Multiplicity | 3.4 (3.6) |
| Completeness (%) | 95.0 (99.1) |
| Mean I/s(I) | 5.7 (1.0) |
| Wilson B-factor (Å2) | 316 |
| R-merge | 0.183 (>1) |
| R-measure | 0.24 (>1) |
| CC1/2 | 0.991 (0.422) |
| Refinement | |
|---|---|
| Resolution range (Å) | 20.07–6.00 (6.45–6.00) |
| Reflections used in refinement | 15179 (1498) |
| Reflections used for R-free | 753 (1498) |
| R-work | 0.3180 (0.3804) |
| R-free | 0.3685 (0.4019) |
| Number of non-hydrogen atoms | 24148 |
| Protein residues | 2944 |
| RMSD bonds (Å) | 0.013 |
| RMSD angles (°) | 1.30 |
| Ramachandran favored (%) | 87 |
| Ramachandran allowed (%) | 12 |
| Ramachandran outliers (%) | 0.8 |
-
*Values in parentheses are for the highest resolution shell.
Cryo-EM data collection and refinement statistics of HsORC1-5.
Data Collection | |
|---|---|
EM equipment | FEI Titan Krios |
Voltage (kV) | 300 |
Detector | Gatan K2 |
Pixel size (Å) | 1.01 |
Electron dose (e-/Å2) | 50 |
Defocus range (µm) | −1.5 ~ −3.5 |
Reconstruction | |
|---|---|
Software | RELION 1.4 |
Number of final particles Resolution (Å) | 10,357 20 |
Map sharpening B-factor (Å2) | −479 |





