Length-dependent flagellar growth of Vibrio alginolyticus revealed by real time fluorescent imaging
Figures
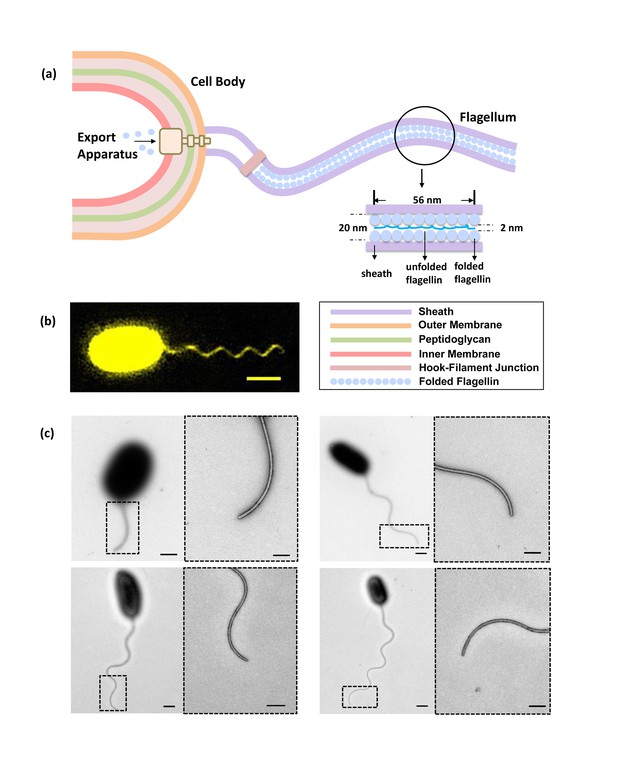
Bacterial flagellum of the Vibrio alginolyticus.
(a) Schematic showing major components of the bacterial flagellar system. (b) Representative high-resolution fluorescence image of Vibrio alginolyticus polar flagellum taken by the Structured Illumination Microscopy (SIM, Materials and methods). Scale bar, 1.5 μm. (c) TEM images of cells with different lengths of flagella (1.8 μm, 4.5 μm, 5.3 μm and 7.8 μm, respectively), which are all tightly covered by sheath. Scale bars, 500 nm (200 nm in magnified area).
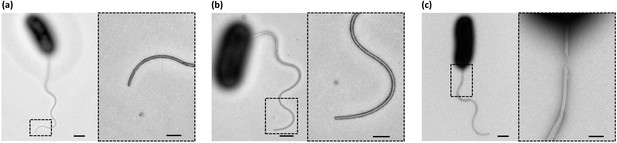
Negative-staining EM images of the sheathed polar flagella of Vibrio alginolyticus.
To confirm that flagellar growth and sheath growth are synchronized, we performed TEM analysis to observe the fine structure of the sheathed flagella of V. alginolyticus. We examined three types of samples: (a) the control group: cells cultured in normal growth medium; (b) the NanoOrange group: cells cultured in growth medium plus NanoOrange; and (c) the fixation group: the control group cells after fixation with formalin. Scale Bars, 500 nm (and 200 nm in the magnified area). For the control group, images showed that there was no visual difference along whole flagellum including the tip, suggesting the sheath always tightly wrapping the inner flagellum. For the NanoOrange group, flagella stained with NanoOrange were morphologically identical to that in the control group, indicating that NanoOrange labeling does not affect flagellar growth nor sheath growth. For the fixation group, we saw the sheath detached from the inner flagellum due to fixation, as previously reported (Glauert et al., 1963). Only after formalin fixation do flagella lose their sheath, which helps us to recognize the two-layered structure of sheath and the core flagellar filament. Consistent with the previous study (Glauert et al., 1963), the sheathed flagellum has an over-all diameter of about 300 Å and the inner flagellum is only ~120 Å in diameter. By these experiments, we confirmed the flagella of Vibrio alginolyticus are wrapped by sheath and the growth of flagella and sheath are tightly synchronized. Therefore, labeling flagellar sheath by NanoOrange is a robust method to quickly visualize the flagellar growth in V. alginolyticus..
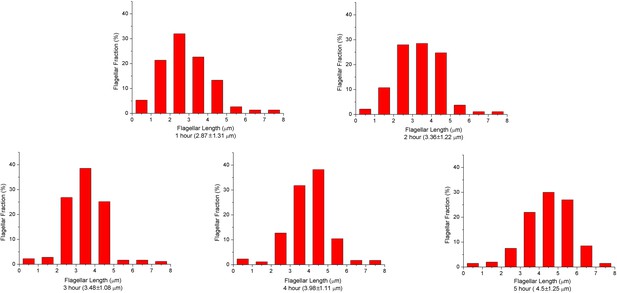
V.alginolyticus flagellar length depends on culturing time.
Distributions of flagellar lengths at different incubation times, binned at 1 μm intervals. The mean values and standard deviation of the lengths at different culturing times are also presented. From 1 to 5 hr, the number of flagella we analyzed were 75, 186, 179, 174 and 200, respectively. The experimental results demonstrate the average flagellar length of a bacterial population increases with the culturing time.
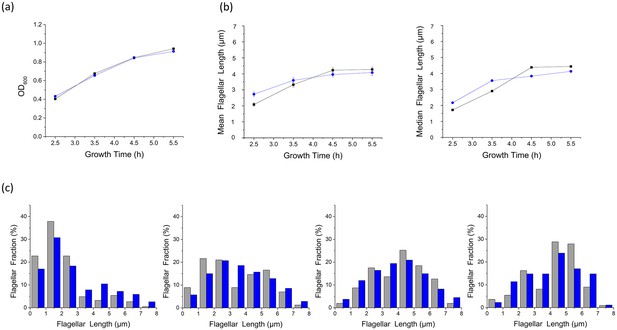
NanoOrange labeling does not affect cell viability or flagellar growth.
(a) Growth curves of the two culture groups. The cells' optical density at 600 nm is plotted according to the culture time at 2.5, 3.5, 4.5 and 5.5 hr (gray: cells in normal imaging medium, blue: cells cultured with NanoOrange). The result showed that cells grown in the medium with NanoOrange exhibited identical growth curve to cells in the normal medium, indicating that labeling by NanoOrange does not affect cell growth; (b) Plots showing the mean and median values of flagellar lengths of the two culture groups at different culturing times (gray: cells in normal imaging medium, blue: cells cultured with NanoOrange); (c) Distributions of flagellar lengths of the two culture groups at different incubation times, binned at 1 μm intervals. From 2.5 to 5.5 hr in sequence, the number of flagella we analyzed were 185, 157, 103 and 111 for the group with normal medium and 153, 140, 134 and 88 for the group with NanoOrange (gray column: cells in normal imaging medium, blue column: cells cultured with NanoOrange). The distributions of flagellar lengths in two culture groups at different incubation times were roughly identical, demonstrating that NanoOrange labeling does not affect flagellar growth, either.
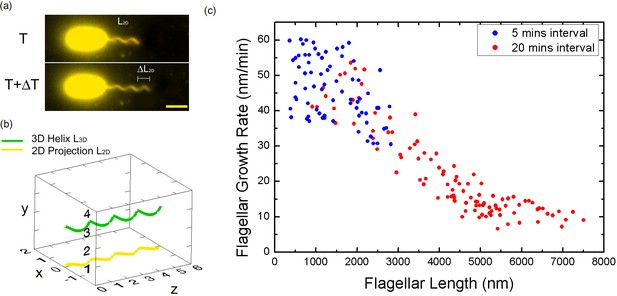
Bacterial flagellar growth rate measurements.
(a) Representative images of a growing bacterial flagellum, taken at different time points, ΔT = 30 mins. The flagellar length increases by length ΔL2D. The growth rate of the flagellum can be calculated as ΔL/ΔT. Scale bar: 2 μm. (b) Schematic of a helical flagellum in 3D space L3D (green) and its 2D projection L2D (yellow). (c) Length dependence of flagellar growth rate of Vibrio alginolyticus from measuring growth rate at different flagellar lengths at 5 min intervals for short flagella (blue points) and 20 min intervals for longer flagella (red points). Flagellar growth rate is constant for short flagella (<1500 nm), and it decreases dramatically with increasing flagellar length.
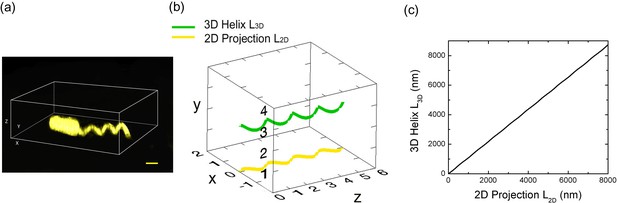
Measurement of flagellar length.
(a) 3D SIM image of a Vibrio alginolyticus cell with a helical flagellum. Scale bar = 1 μm; (b) Schematic plot of a 3D helix (green) and it's 2D projection (yellow); (c) the arc length of 3D helix (L3D) vs. 2D projection (L2D) (Materials and methods).
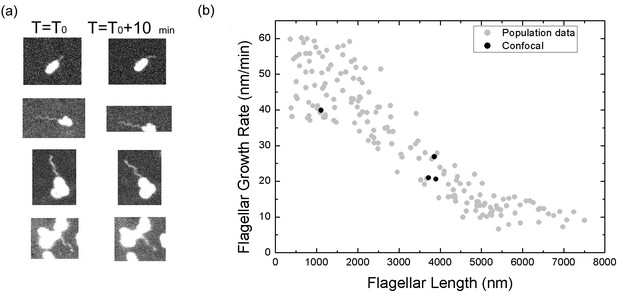
Flagellar growth rate measured from non-tethering flagella.
(a) Pairwise confocal images of bacterial flagella at time T0 and T0 +10 mins. (b) The growth rate of free moving flagella (block dots) versus flagellar length, compared to the population measurement data (gray dots) using tethered flagella.
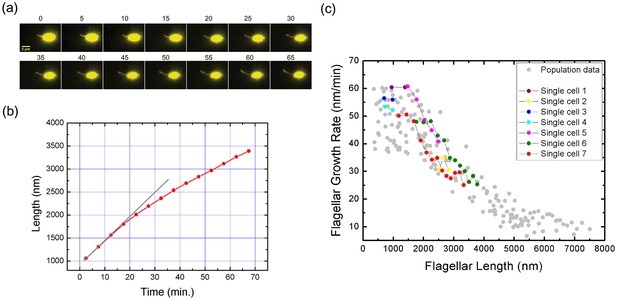
Real-time tracking of single flagellar growth by fluorescence microscopy.
(a) Continuous time-lapse recording of a single growing flagellum with images taken every 5 mins. (b) Flagellar length versus time for the cell presented in (a) (red dots). Gray line shows initial slope of flagellar length versus time. (c) Population measurement data for flagellar growth (grey dots) with measurements from cells tracked over more than two frames indicated in color. The cell in (a) and (b) is indicated in red.
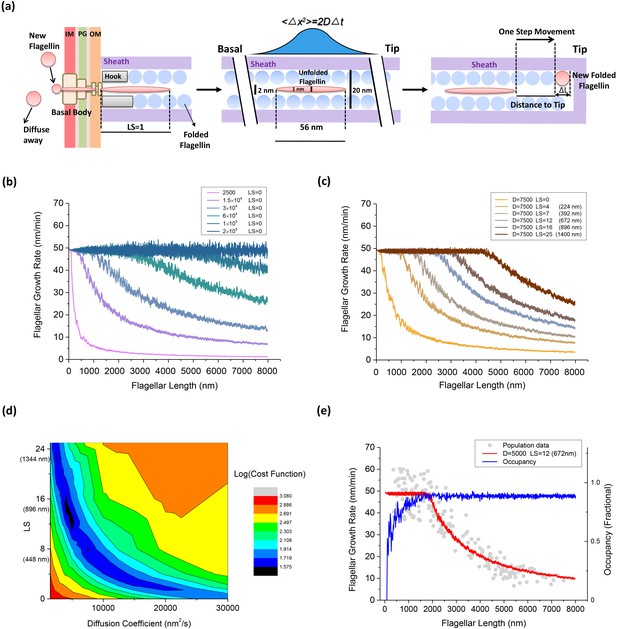
An injection-diffusion model explaining the length dependent flagellar growth.
(a) Schematic illustration of the key elements of our one-dimensional diffusion model. At the base of the BFM, flagellin monomers are inserted into the channel by the T3FSS at a constant speed Sload. Once pumped into the channel, the flagellin diffuses as an unfolded peptide chain of 56 nm long. When a flagellin monomer reaches the tip of the growing flagellum, it crystallizes immediately and extends the flagellum by 0.47 nm. (b) Simulated flagellar growth rate vs. length curves with loading strength equal to zero (LS = 0) and varying values of D. (c) The simulated flagellar growth rate vs. length curves with D = 7500 nm2/s and varying values of LS. (d) The contour plot representing the cost function of the model fitting to experimental dataset in the parameter space formed by D and LS. (e) The predicted flagellar growth rate vs. length curve of the best-fit parameter set (LS = 12 and D = 5000 nm2/s, red line) and the channel occupancy vs. length (blue line).





