Autocrine regulation of stomatal differentiation potential by EPF1 and ERECTA-LIKE1 ligand-receptor signaling
Figures
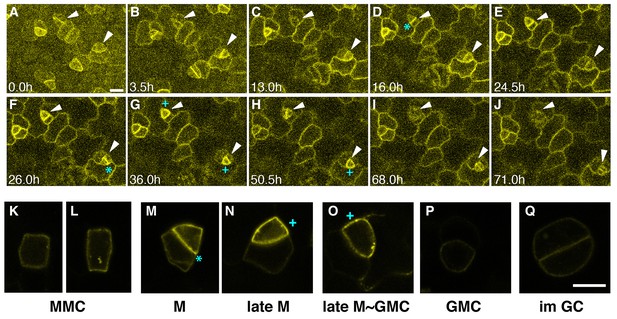
Expression patterns of ERL1 during stomatal development.
Expression pattern of ERL1pro::ERL1-YFP. (A–J) Time-lapse live imaging of developing abaxial cotyledon epidermis of the 1-day-old T4 seedling of ERL1pro::ERL1-YFP erl1-2. Time points after image collection are indicated in hours:(A) 0.0 hr; (B) 3.5 hr; (C) 13.0 hr; (D) 16.0 hr; (E) 24.5 hr; (F) 26.0 hr; (G) 36.0 hr; (H) 50.5 hr; (I) 68.0 hr; and (J) 71.0 hr. Arrowheads point to two representative cells. Images for (A–J) are taken at the same magnification. Scale bar, 10 µm. (K–L) High resolution live images of stomatal precursors expressing ERL1-YFP. (K, L) meristemoid mother cells (MMC); (M) meristemoid (M); (N) late meristemoid (late M); (O) late meristemoid to guard mother cell transition (late M~GMC); (P) GMC; (Q) immature guard cells (im GC). Images for (K–L) are taken at the same magnification. Scale bar, 7.5 µm. ERL1-YFP signals are detected at the plasma membrane from early meristemoids (A, K, L; arrowheads) and accumulate high at the asymmetric division site (e.g. D, F, M; cyan asterisks). ERL1-YFP signals boost during the late meristemoid-to-GMC transition (G, H, N, O; cyan plus), and diminishes after GMC symmetric division (I, J, P, Q; arrowheads). See accompanying Video 1. Experiments were repeated three times. Total seedlings analyzed; n = 9.
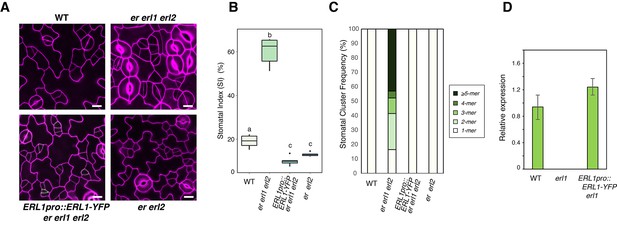
ERL1pro::ERL1-YFP is functional.
(A) Shown are confocal microscopy images of abaxial cotyledon epidermis from 4-day-old seedlings of: wild type (WT; top left); er erl1 erl2 (top right); ERL1pro::ERL1-YFP in er erl1 erl2 (bottom left); er erl2 (bottom right). Introduction of ERL1pro::ERL1-YFP rescues stomatal clustering phenotype of er erl1 erl2. T2 seedlings of ERL1pro::ERL1-YFP er erl1 erl2 were used for the analysis. Scale bars, 10 µm. Experiments were repeated three times. Total seedlings analyzed; n = 15. (B) Stomatal index (SI) of the cotyledon abaxial epidermis from 7-day-old seedlings of respective genotypes. One-way ANOVA followed by Tukey's HSD test was performed for comparing all other genotypes and classify their phenotypes into three categories (a, b, and c). n = 6. (C) Stomatal cluster frequency histogram. Total seedlings analyzed; n = 6 for each genotype. (D) Expression level of ERL1-YFP transgene is comparable to that of the endogenous ERL1. RT-PCR analysis was performed three times and normalized against eIF4A. p=0.074 (Student's t-test, unpaired).
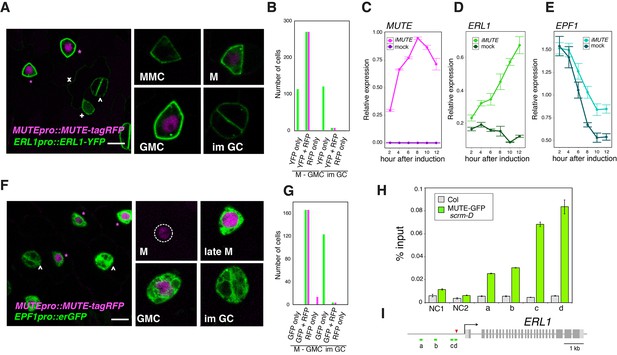
Overlapping expression of EPF1, ERL1, MUTE and direct upregulation of ERL1 by MUTE.
(A) Expression patterns of ERL1pro::ERL1-YFP and MUTEpro::MUTE-tagRFP from the abaxial true leaf epidermis of 7-day-old F1 seedlings. Co-expression of MUTE-tagRFP in nucleus and ERL1-YFP at the plasma membrane are detected in late meristemoids and GMCs (top; magenta asterisks) while only ERL1-YFP signals are visible in MMC, early meristemoid (+) and immature GCs (∧). Faint YFP signals are detected in the stomatal lineage ground cell (SLGC) differentiating into a pavement cell (x). (Insets) representative images of stomatal precursor cells. Scale bars, 7.5 µm. (B) Quantitative analysis of the number of stomatal precursor cells (meristemoids-to-GMC, and immature GCs) expressing ERL1pro::ERL1-YFP (green) and/or MUTEpro::MUTE-tagRFP (magenta). Total of 510 cells were analyzed from 12 seedlings. (C) Quantitative RT-PCR analysis of the total MUTE transcripts from 4-day-old seedlings that were either mock- or estradiol-treated for induced MUTE overexpression (iMUTE) at the time indicated. Relative expression represents qRT-PCR expression normalized over that of ACTIN2. Values are Mean ± standard deviation of three technical replicates from one representative experiment. (D) Quantitative RT-PCR analysis of the endogenous ERL1 transcripts from 4-day-old seedlings that were either mock- or iMUTE at the time indicated. Relative expression represents qRT-PCR expression normalized over that of ACTIN2. Values are Mean ± standard deviation of three technical replicates from one representative experiment. See Figure 2—figure supplement 2 for the two additional biological replicates. (E) Quantitative RT-PCR analysis of the endogenous EPF1 transcripts from 4-day-old seedlings that were either mock- or iMUTE at the time indicated. Relative expression represents qRT-PCR expression normalized over that of ACTIN2. Values are Mean ± standard deviation of three technical replicates from one representative experiment. See Figure 2—figure supplement 2 for the two additional biological replicates. (F) Expression patterns of EPF1pro::erGFP and MUTEpro::MUTE-tagRFP in the abaxial true leaf epidermis from 7-day-old F1 seedlings. MUTE-tagRFP expression precedes that of EPF1. Strong co-expression of MUTE-tagRFP and EPF1 can be detected in the late meristemoids and GMCs (top; magenta asterisks), while only EPF1 signals are visible in immature GCs (∧). (Insets) representative images of stomatal precursor cells. Scale bars, 7.5 µm. (G) Quantitative analysis of the number of stomatal precursor cells (meristemoids-to-GMC, and immature GCs) expressing EPF1pro::erGFP (green) and/or MUTEpro::MUTE-tagRFP (magenta). Total of 307 cells were analyzed from five seedlings. (H) Quantitative PCR on ERL1 promoter region after chromatin Immunoprecipitation against anti-GFP antibody in Col and transgenic plants expressing MUTE-GFP in scrm-D. Values are Mean ± S.E.M. of percent-input DNA of three technical replicates from one representative experiment. See Figure 2—figure supplement 1 for the other two biological replicates. NC, Negative Control; NC1, 5’ intergenic region of ACTIN2; NC2, promoter region of AGAMOUS; a, b, c and d, the ERL1 loci indicated below in (D). For raw data, see Supplementary file 3. (I) Diagram of the ERL1 loci tested. Green lines (a,b,c and d) below the ERL1 schematic, regions amplified by qPCR; Red arrowhead, SPCH binding peak; gray line, intergenic region or intron; light gray box, 5’ or 3’ untranslated region; dark gray box, exon; black arrow, transcription start site. See Figure 2—figure supplement 2 for the two additional biological replicates.
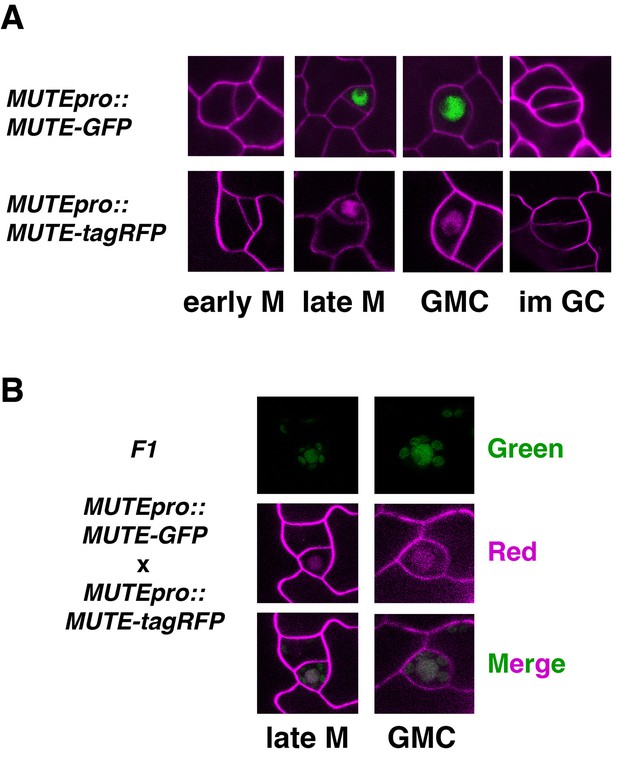
Expression patterns of MUTEpro::MUTE-GFP and MUTEpro::MUTE-tagRFP.
(A) Shown are representative stomatal precursor cells expressing MUTEpro::MUTE-GFP (top) and MUTEpro::MUTE-tagRFP from T5 transgenic seedlings. In early meristemoid (early M; left), no signals are visible. MUTE protein accumulates in the nucleus of the late meristemoids (late M; left middle) and guard mother cells (GMC; right middle). MUTE signals disappear in immature guard cells (im GC) following the final symmetric division (right). Cell outline was counter-stained with FM4-64. Experiments were repeated three times. Total seedlings analyzed; n = 6. (B) Co-expression of MUTE-GFP and MUTE-tagRFP signals. Shown are representative stomatal precursors from abaxial cotyledon epidermis of 5-day-old F1 seedlings co-expressing MUTEpro::MUTE-GFP and MUTEpro::MUTE-tagRFP. Signals were separately collected in green channel (Green) and red channel (red; false colored in magenta), and subsequently merged (Merged). Cell outline was counter-stained with FM4-64. Experiments were repeated three times. Total seedlings analyzed; n = 6.
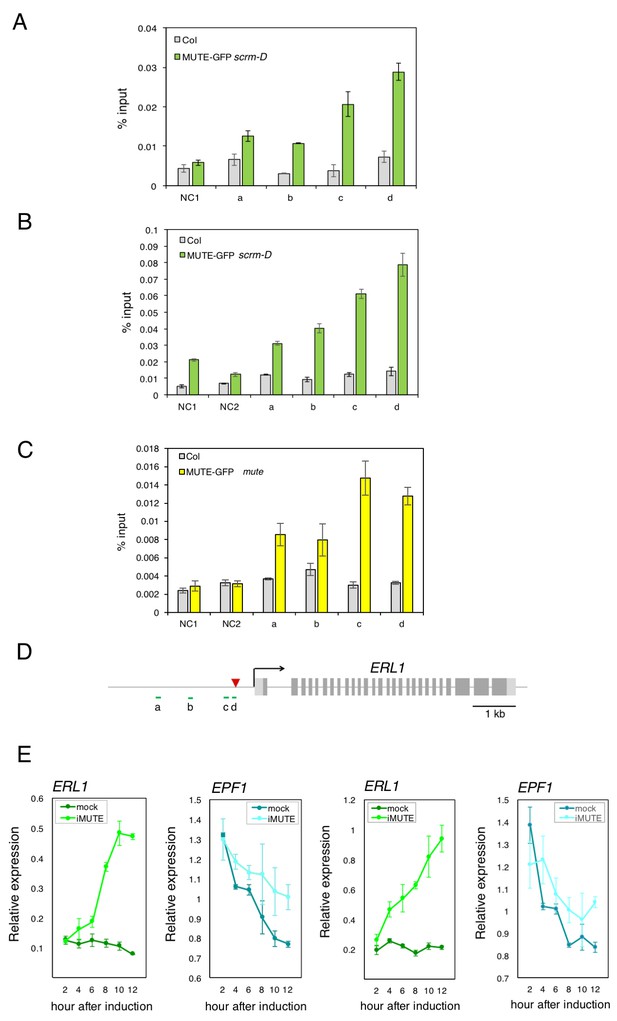
Additional two biological replicates of ChIP assays and qRT-PCR expression analysis of ERL1 and EPF1 upon induced MUTE overexpression.
(A and B) Quantitative PCR on ERL1 promoter region after chromatin Immunoprecipitation (CHIP) against anti-GFP antibody in Col and transgenic plants expressing MUTE-GFP in scrm-D. Values are Mean ± S.E.M. of percent-input DNA of three technical replicates from two independent experiments. NC, Negative Control; NC1, 5’ intergenic region of ACTIN2; NC2, promoter region of AGAMOUS; a, b, c and d, the ERL1 loci indicated below in (D). (C) Quantitative PCR on ERL1 promoter region after ChIP against anti-GFP antibody in Col and transgenic plants expressing MUTE-GFP in mute, which exhibit normal (wild-type) phenotype. Values are Mean ± S.E.M. of percent-input DNA of three technical replicates. NC, Negative Control; NC1, 5’ intergenic region of ACTIN2; NC2, promoter region of AGAMOUS; a, b, c and d, the ERL1 loci indicated below in (C). (D) Diagram of the ERL1 loci tested. (E) Quantitative RT-PCR analysis of the endogenous ERL1 and EPF1 transcripts from 4-day-old seedlings that were either mock- or estradiol-treated for induced MUTE overexpression (iMUTE) at the time indicated. Relative expression represents qRT-PCR expression normalized over that of ACTIN. Values are Mean ± standard deviation of three technical replicates from two independent experiments.
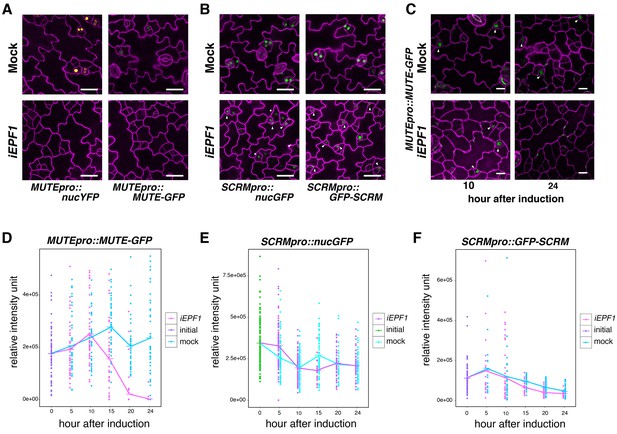
EPF1 signaling downregulates both MUTE promoter activity and MUTE protein accumulation.
(A) Representative confocal images of abaxial cotyledon epidermis from 4-day-old estradiol-inducible EPF1 seedlings carrying MUTEpro::nucYFP and MUTEpro::MUTE-GFP four days after mock-treated or estradiol-treated for induced EPF1 overexpression (iEPF1). F1 seedlings were used for the analysis. Scale bars, 25 µm. Experiments were repeated three times. Three seedlings were analyzed each time. (B) Representative confocal images of abaxial cotyledon epidermis from 4-day-old estradiol-inducible EPF1 seedlings carrying SCRMpro::nucGFP and SCRMpro::GFP-SCRM four days after mock-treated or estradiol-treated for induced EPF1 overexpression (iEPF1). F1 seedlings were used for the analysis. Scale bars, 25 µm. Experiments were repeated three times. Three seedlings were analyzed each time. (C) Representative Z-stack confocal image projection showing the MUTE-GFP levels after iEPF1 induction used for the quantitative analysis. Meristemoids accumulating MUTE-GFP are indicated by arrowheads. F1 seedlings were used for the analysis. Scale bars, 10 µm. Experiments were repeated three times. Three seedlings were analyzed each time. (D) Quantitative analysis of MUTE-GFP levels after iEPF1 induction at the time indicated on 3-day-old seedlings. Each dot represents the total intensity value of GFP signals from each nucleus expressing MUTE-GFP. To fully cover the entire nuclei expressing MUTE-GFP, serial Z-stack projection images were used for quantitative analysis (see Materials and methods). Mean value at each time point is connected by the line to visualize the average intensity change over time. Purple, initial values at time point 0; Cyan, mock treatment; Magenta, iEPF1 induction. Experiments were repeated three times; n = 6 for time point 0; n = 3 per subsequent time point. Total of 495 nuclei were analyzed. (E) Quantitative analysis of SCRMpro::nucGFP levels after iEPF1 induction at the time indicated on 3-day-old seedlings. Each dot represents the total intensity value of GFP signals from each nucleus expressing SCRMpro::nucGFP. To fully cover the entire nuclei expressing GFP, serial Z-stack projection images were used for quantitative analysis (see Materials and methods). Mean value at each time point is connected by the line to visualize the average intensity change over time. Green, initial values at time point 0; Cyan, mock treatment; Purple, iEPF1 induction. Experiments were repeated three times; n = 6 for time point 0; n = 3 per subsequent time point. Total of 817 nuclei were analyzed. (F) Quantitative analysis of GFP-SCRM levels after iEPF1 induction at the time indicated on 3-day-old seedlings. Each dot represents the total intensity value of GFP signals from each nucleus expressing SCRMpro:: GFP-SCRM. To fully cover the entire nuclei expressing GFP-SCRM, serial Z-stack projection images were used for quantitative analysis (see Materials and methods). Mean value at each time point is connected by the line to visualize the average intensity change over time. Purple, initial values at time point 0; Cyan, mock treatment; Magenta, iEPF1 induction. Experiments were repeated three times; n = 6 for time point 0; n = 3 per subsequent time point. Total of 368 nuclei were analyzed.
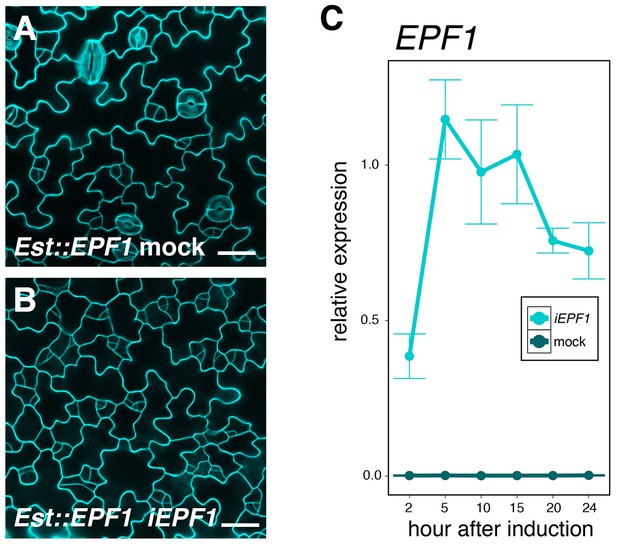
Quantitative and phenotypic analyses of induced overexpression of EPF1.
(A, B) Shown are confocal microscopy of cotyledon abaxial epidermis from 4-day-old T4 seedlings treated with control DMSO solution (A; mock) or 10 μM estradiol to induce EPF1 overexpression (B; iEPF1). Stomatal differentiation is inhibited by iEPF1, and arrested meristemoids are visible. Images are taken under the same magnification. Scale bar, 25 μm. Experiments were repeated three times. Three seedlings were analyzed each time. (C) Quantitative RT-PCR (qRT-PCR) analysis of EPF1 transcripts after inducing EPF1 overexpression (iEPF1) the identical time-course for the quantitative analysis of SPCH, MUTE, and SCRM transcriptional and translational reporter fluorescent protein accumulation in the nuclei. Relative expression represents qRT-PCR expression normalized over that of ACTIN2. Values are Mean ± standard deviation of three technical replicates from one representative experiment. The average fold increase in normalized EPF1 transcripts are: 137.1, 407.7, 347.6, 367.0, 269.1, and 257.5-fold increase in 2, 5, 10, 15, 20 and 24 hr after induction, respectively.
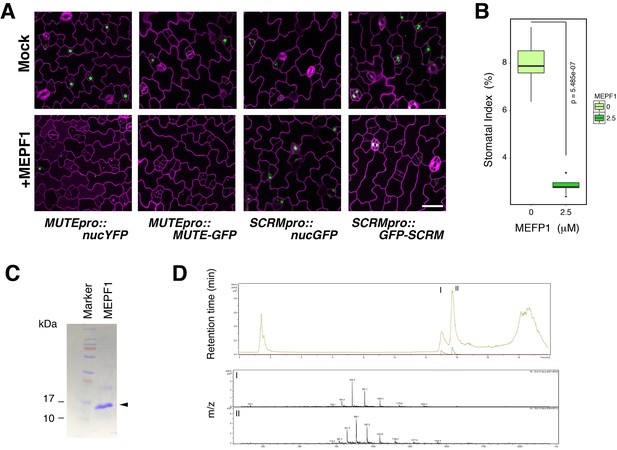
Predicted mature EPF1 peptide application reduces MUTE promoter activity and MUTE protein accumulation.
(A) Shown are confocal microscopy images of abaxial cotyledon epidermis from 4-day-old T4 seedlings expressing MUTEpro::nucYFP, MUTEpro::MUTE-GFP, SCRMpro::nucGFP, or SCRMpro::GFP-SCRM that were either mock treated or treated with predicted mature EPF1 peptide (MEPF1) (10 µM) for 3 days. MEPF1 treatment results in reduced MUTE promoter activity and MUTE-GFP protein amounts. Experiments were repeated three times. Three seedlings were analyzed each time. (B) Quantitative analysis of MEPF1 bioactivity. Shown is the stomatal index (SI) of abaxial cotyledon epidermis from 5-day-old seedlings that were mock treated or treated with MEPF1 (2.5 µM) for 4 days. Cotyledon images from six seedlings were subjected to analysis. Welch's Two Sample T-test was performed. (C) Purified and refolded MEPF1 peptide used for the assays. Shown is an image of SDS-PAGE gel stained with CBB. Arrowhead, MEPF1 peptide. (D) HPLC chromatograph and mass-spec analysis of the two major peaks (I and II), both corresponds to MEPF1, most likely different in refolding.
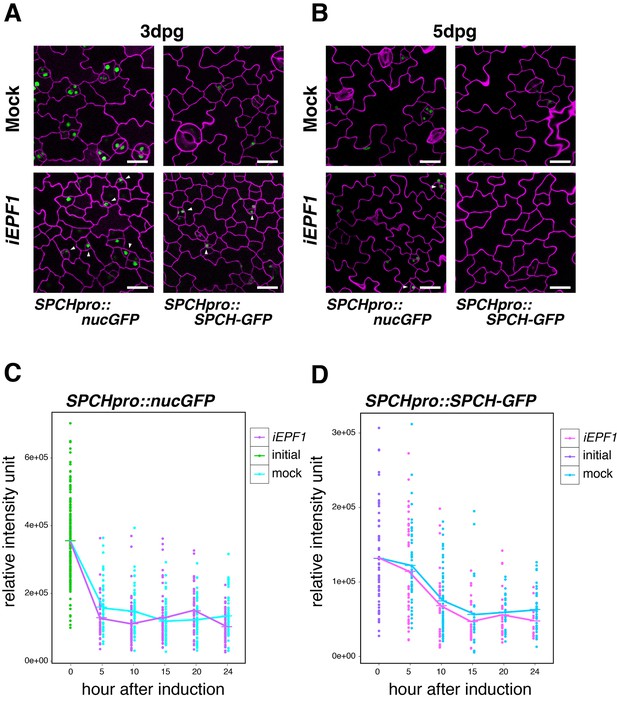
Induced EPF1 overexpression has no effect in SPCH promoter activity and SPCH protein levels.
(A) Confocal microscopy images of abaxial cotyledon epidermis from 3-day-old estradiol-inducible EPF1 seedlings expressing SPCHpro::nucGFP (left) and SPCHpro::SPCH-GFP (right) that were either mock treated (top) or treated with estradiol (bottom) from germination for induced EPF1 overexpression (iEPF1). Both SPCH promoter activity and SPCH-GFP protein are robustly detected in arrested meristemoids (arrowheads) by iEPF1 at 3-days post germination (dpg). F1 seedlings were used for the analysis. Scale bars, 20 µm. Experiments were repeated three times. Three seedlings were analyzed each time. (B) Confocal microscopy images of abaxial cotyledon epidermis from 5-day-old estradiol-inducible EPF1 seedlings expressing SPCHpro::nucGFP (left) and SPCHpro::SPCH-GFP (right) that were either mock treated (top) or treated with estradiol (bottom) from germination for induced EPF1 overexpression (iEPF1). At 5-days post germination (dpg), SPCH-GFP proteins have diminished, reflecting the stomatal cell-lineages have past the early stage. F1 seedlings were used for the analysis. Scale bars, 20 µm. Experiments were repeated three times. Three seedlings were analyzed each time. (C) Quantitative analysis of SPCHpro::nucGFP levels after iEPF1 induction at the time indicated on 3-day-old seedlings. Each dot represents the total intensity value of GFP signals from each nucleus expressing SPCHpro::nucGFP. To fully cover the entire nuclei expressing GFP, serial Z-stack projection images were used for quantitative analysis (see Materials and methods). Mean value at each time point is connected by the line to visualize the average intensity change over time. Green, initial values at time point 0; Cyan, mock treatment; Purple, iEPF1 induction. Experiments were repeated three times; n = 6 for time point 0; n = 3 per subsequent time point. Total of 790 nuclei were analyzed. (D) Quantitative analysis of SPCH-GFP levels after iEPF1 induction at the time indicated on 3-day-old seedlings. Each dot represents the total intensity value of GFP signals from each nucleus expressing SPCHpro::SPCH-GFP. To fully cover the entire nuclei expressing SPCH-GFP, serial Z-stack projection images were used for quantitative analysis (see Materials and methods). Mean value at each time point is connected by the line to visualize the average intensity change over time. Purple, initial values at time point 0; Cyan, mock treatment; Magenta, iEPF1 induction. Experiments were repeated three times; n = 6 for time point 0; n = 3 per subsequent time point. Total of 403 nuclei were analyzed.
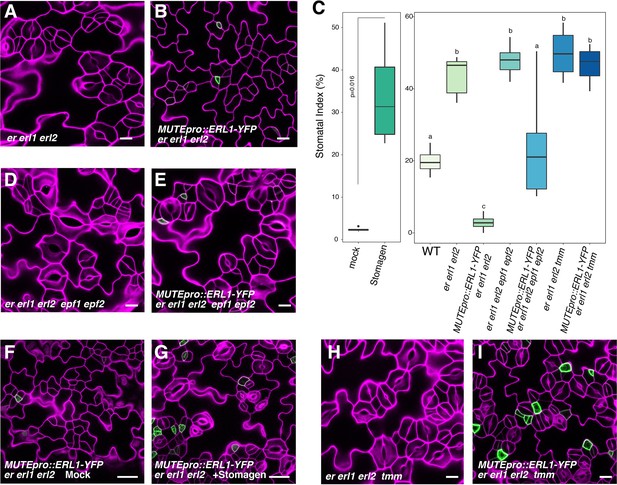
Absolute co-expression of ERL1 and MUTE confers meristemoid arrests.
Shown are confocal microscopy images of cotyledon abaxial epidermis from 7-day-old seedlings of the following genotypes: (A) er erl1 erl2; (B) MUTEpro::ERL1-YFP in er erl1 erl2; (D) er erl1 erl2 epf1 epf2; (E) MUTEpro::ERL1-YFP in er erl1 erl2 epf1 epf2; (F) MUTEpro::ERL1-YFP in er erl1 erl2 mock treated; (G) MUTEpro::ERL1-YFP in er erl1 erl2 treated with 5 µM Stomagen peptide; (H) er erl1 erl2 tmm; (I) MUTEpro::ERL1-YFP in er erl1 erl2 tmm. T1 transgenic seedlings of MUTEpro::ERL1-YFP er erl1 erl2; MUTEpro::ERL1-YFP er erl1 erl2 epf1 epf2; and MUTEpro::ERL1-YFP er erl1 erl2 tmm were used for the analysis. T2 seedlings of MUTEpro::ERL1-YFP er erl1 erl2 were used for the mock or Stomagen treatment. Scale bars, 10 µm (A, B, D, E, H, I), 25 µm (F, G). (C) Quantitative analysis. Stomatal index (SI) of the cotyledon abaxial epidermis from 7-day-old seedlings of respective genotypes. For each genotype, images from six seedlings were analyzed. Welch's Two Sample T-test was performed for mock vs. Stomagen application (Left). One-way ANOVA followed by Tukey's HSD test was performed for comparing all other genotypes and classify their phenotypes into three categories (a, b, and c).
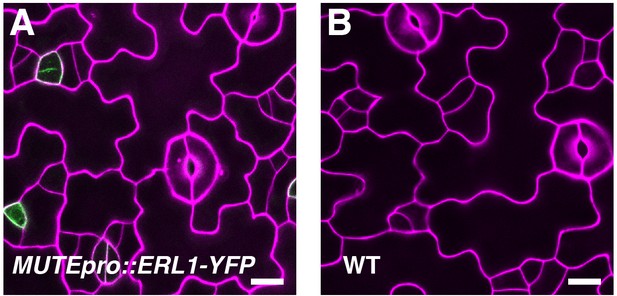
MUTEpro::ERL1-YFP does not cause meristemoid arrest in the presence of functional ERECTA-family genes.
Shown are confocal microscopy images of abaxial cotyledon epidermis from 4-day-old wild type seedlings with or without MUTEpro::ERL1-YFP. Introducing MUTEpro::ERL1-YFP into wild type does not cause arrested meristemoids. F1 seedlings were used for the analysis. Scale bars, 10 µm. Experiments were repeated three times. Total seedlings analyzed; n = 17.
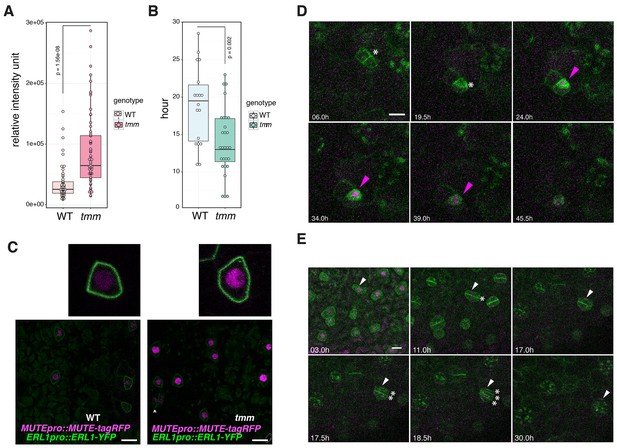
Perturbed EPF1 signaling results in higher amount of MUTE protein, accelerated stomatal differentiation, and symmetric division of GMC.
(A) Quantitative analysis of RFP signal intensity from the nuclei expressing MUTEpro::MUTE-tagRFP. WT and tmm refers to MUTEpro::MUTE-tagRFP ERL1pro::ERL1-YFP in erl1 and MUTEpro::MUTE-tagRFP ERL1pro::ERL1-YFP in erl1 tmm, respectively. F3 seedlings were used for the analysis. Experiments were repeated three times. Total numbers of nuclei analyzed; n = 120 (WT); n = 135 (tmm). Welch's Two Sample T-test was performed. (B) Quantitative analysis of a duration of proliferation-to-differentiation transition during stomatal development (hours from last asymmetric division of a meristemoid to onset of GMC symmetric division). MUTEpro::MUTE-tagRFP ERL1pro::ERL1-YFP in erl1 was referred as WT and MUTEpro::MUTE-tagRFP ERL1pro::ERL1-YFP in erl1 tmm was referred as tmm. F3 seedlings were used for the analysis. n = 18 (WT); n = 28 (tmm). Welch's Two Sample T-test was performed. (C) Z-stack confocal microscopy image projections of true leaf abaxial epidermis from 7-day-old F3 seedlings expressing MUTEpro::MUTE-tagRFP ERL1pro::ERL1-YFP in erl1 ('WT'); and MUTEpro::MUTE-tagRFP ERL1pro::ERL1-YFP in erl1 tmm ('tmm'). For quantitative comparison, image acquisition and processing were performed using the identical settings. Scale bars, 10 μm. Top insets, a representative late meristemoid from 'WT' (left) and 'tmm' (right). (D) Time-lapse imaging of stomatal differentiation in F3 seedlings of ‘wild type (WT)’; ERL1pro::ERL1-YFP and MUTEpro::MUTE-tagRFP in erl1. The site of asymmetric entry and amplifying divisions are indicated in white asterisks. The MUTE protein accumulates in the nucleus of the late meristeoid and disappears as the GMC differentiates (magenta arrowheads). Time points after image collection are indicated in hours. See accompanying Video 4. Scale bar, 10 μm. Experiments were repeated three times. Total seedlings analyzed; n = 9. (E) Time-lapse imaging of stomatal differentiation in F3 seedlings of ‘tmm’; ERL1pro::ERL1-YFP and MUTEpro::MUTE-tagRFP in erl1 tmm. Occasionally, a late meristemoid accumulating MUTE-tagRFP (white arrowhead) maintain MUTE accumulation after initial GMC symmetric division (*; in 11.0 hr), leading to an extra round of symmetric division (asterisks in 17.5 hr and 18.5 hr). Time points after image collection are indicated in hours. See accompanying Video 5. Scale bar, 10 μm. Experiments were repeated three times. Total seedlings analyzed; n = 9.
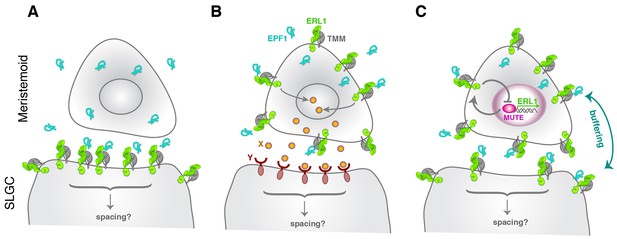
Model diagram of how EPF1-ERL1 peptide-receptor signaling regulates stomatal differentiation.
(A) Paracrine signaling model: Here, EPF1 peptide (cyan) emanating from a meristemoid is perceived by ERL1 (light green) and TMM (gray) localized at the plasma membrane of a neighboring stomatal lineage ground cell (SLGC). The activated signaling will in turn regulate asymmetric spacing division. (B) Autocrine signaling model: Facette and Smith (2012) proposed an alternate, autocrine signaling model based on the observations that EPF1, ERL1, and TMM all exhibit the highest expression in the meristemoid (Nadeau and Sack, 2002; Shpak et al., 2005; Hara et al., 2007). Here, EPF1 secreted from a meristemoid is perceived by ERL1-TMM on a surface of the same meristemoid. The downstream signaling leads to expression of a yet unidentified peptide (X: orange), which will be perceived by a yet unidentified receptor (Y: brown) on a surface of the neighboring stomatal lineage ground cell (SLGC). This triggers signal transduction regulating asymmetric spacing division. (C) Model based on the current study: Both ERL1 and TMM are expressed in a meristemoid and a stomatal lineage ground cell (SLGC), since they are both upregulated by SPCH at an earlier stage (Lau et al., 2014; Horst et al., 2015). During the meristemoid-to-GMC transition, MUTE upregulates ERL1 expression and this causes high accumulation of ERL1 in the meristemoid over the stomatal lineage ground cell (SLGC). Meanwhile, EPF1 is expressed and secreted from the late meristemoid by an independent mechanism. A perception of EPF1 by the meristemoid-expressed ERL1 and TMM triggers signal transduction leading to downregulation of MUTE, thereby restricting MUTE activity via an autocrine mechanism. The same pool of EPF1 is also perceived by ERL1 and TMM expressed in the neighboring stomatal lineage ground cell (SLGC), which trigger downstream signal transduction to ensure asymmetric spacing division via a paracrine mechanism. Consequently, receptors in the meristemoids and stomatal lineage ground cells (SLGCs) buffer the pool of EPF1 ligands. Note that our model does not exclude the possibility that yet another unknown signals (like X and Y) operate simultaneously during the stomatal differentiation process.
Videos
Time-lapse movie of ERL1-YFP dynamics during stomatal development.
Projection of z-stack images of ERL1pro::ERL1-YFP from the abaxial cotyledon epidermis of the 1-day-old T4 seedling were shown in 30 min time interval. Time points after image collection are indicated in hours. Arrowheads point to two representative cells.
Three-dimensional projection of the er erl1 erl2 phenotype.
Shown is a projection of z-stack images of er erl1 erl2 from the abaxial cotyledon epidermis of the 7-day-old seedling. Scale bar, 50 µm.
Three-dimensional projection of the MUTEpro::ERL1-YFP er erl1 erl2 phenotype.
Shown is a projection of z-stack images of MUTEpro::ERL1-YFP in er erl1 erl2 from the abaxial cotyledon epidermis of the 7-day-old T1 seedling. Numerous arrested meristemoids are visible with no stomatal differentiation. Scale bar, 50 µm.
Time-lapse movie of normal stomatal differentiation from the line co-expressing ERL1pro::ERL1-YFP and MUTEpro::MUTE-tagRFP.
Projection of z-stack images of ERL1pro::ERL1-YFP and MUTEpro::MUTE-TagRFP from the abaxial cotyledon epidermis of an 1-day-old erl1 seedling were shown in 30 min time interval. Time points after image collection are indicated in hours. F3 seedlings were used for the analysis.
Time-lapse movie of abnormal stomatal differentiation from tmm mutant line co-expressing ERL1pro::ERL1-YFP and MUTEpro::MUTE-tagRFP.
Projection of z-stack images of ERL1pro::ERL1-YFP and MUTEpro::MUTE-TagRFP from the abaxial cotyledon epidermis of an 1-day-old erl1 tmm seedling were shown in 30 min time interval. Time points after image collection are indicated in hours. The arrowhead points to one cell, which develops into two parallel-aligned stomata due to an extra round of symmetric division. F3 seedlings were used for the analysis.
Additional files
-
Supplementary file 1
Plasmid constructs generated in this study
List of plasmid constructs and selection markers.
- https://doi.org/10.7554/eLife.24102.020
-
Supplementary file 2
Oligo DNA primers and their sequences used in this study
List of oligo DNA names, sequence, and their use.
- https://doi.org/10.7554/eLife.24102.021
-
Supplementary file 3
Raw data of ChIP experiments
Numerical values of the ChIP experiments.
- https://doi.org/10.7554/eLife.24102.022
-
Supplementary file 4
Raw data and R-based analyses
List of original data for stomatal index, qRT-PCR, quantitative data of reporter fluorescent protein accumulation and cell-state transition. Also includes R-scripts written for the analyses and graph presentations.
- https://doi.org/10.7554/eLife.24102.023





