Helical jackknives control the gates of the double-pore K+ uptake system KtrAB
Figures
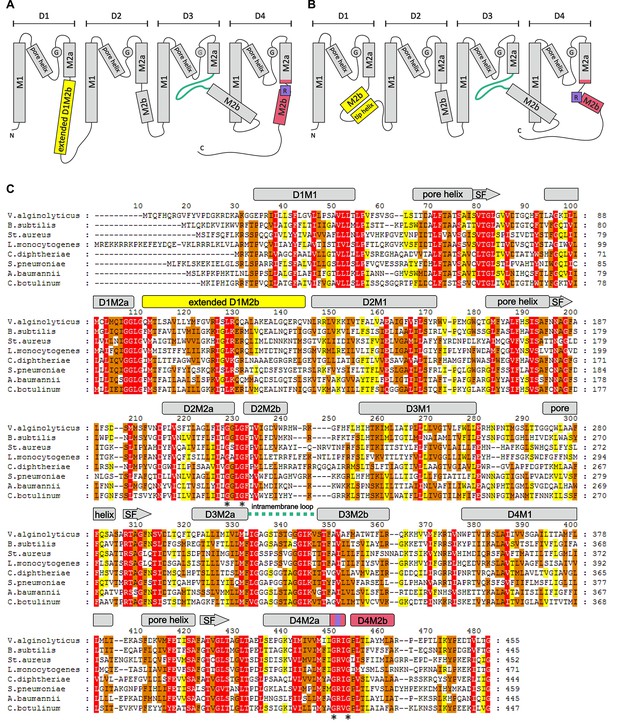
Overview of the KtrB subunit.
Topology diagram of KtrB subunit structure in the presence of ADP (A) and ATP (B). (C) Amino acid sequence alignment of different KtrBs using Clustal Omega and GeneDoc and secondary structure of VaKtrB in the presence of ADP. α-helices and β-strands are shown as rectangles and arrows, respectively. Sequence conservation color-coded: White on red, 100%; black on orange, ≥80%; black on yellow, ≥60%. SF = selectivity filter. Discussed motifs are highlighted: intramembrane loop is colored in green, D1M2b helix and tip helix in yellow, D4M2b helix in magenta and the conserved arginine in purple. Asterisks (*) mark conserved gating hinge glycine residues.
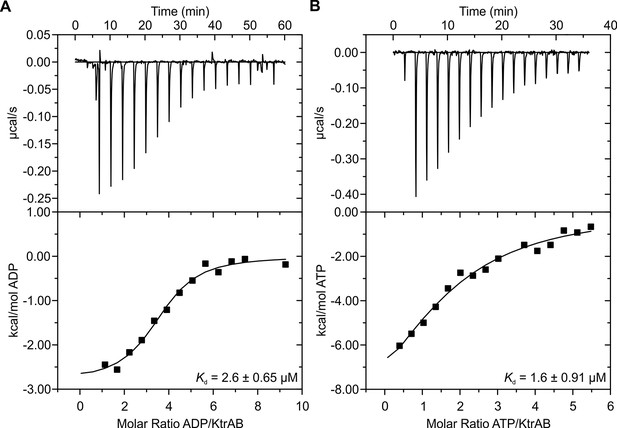
Binding of KtrAB to nucleotides examined by ITC measurements.
Upper panels: raw heat exchange data, associated with (A) ADP or (B) ATP binding to detergent-solubilized KtrAB. Lower panel: integrated injection heat pulses, normalized per mole of injection, reveal differential binding curves calculated by a one-site binding model with determined Kd values of 3 μM for ADP and 2 μM for ATP.
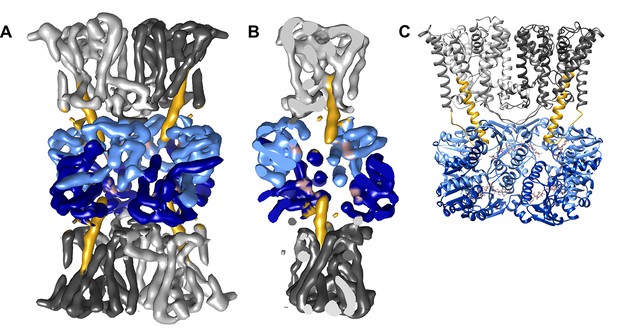
Three-dimensional reconstruction of ADP-bound KtrAB.
(A) Side view of the cryo-EM map colored by protein subunits. (B) 90° rotated side view of the density map cut open to highlight the extended helices. (C) Structural model of ADP-bound KtrAB in the same orientation as in A. KtrBs, dark and light grey with helices D1M2, yellow; octameric ring of KtrA, dark and light blue; ADP in rosy brown.
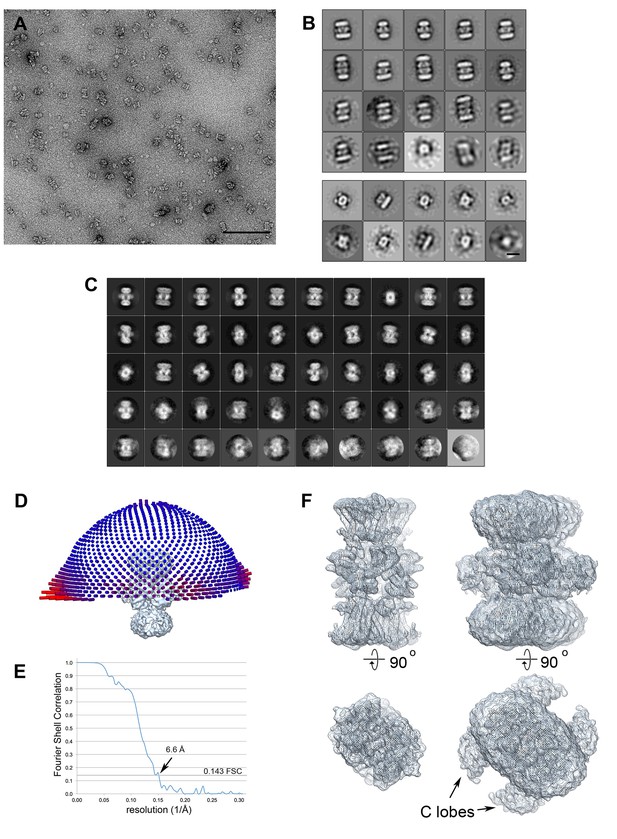
Electron microscopy data.
(A) KtrAB negatively stained with uranyl acetate. The scale bar is 100 nm. (B) Class averages from negatively stained particles. The top panel shows classes from a data set of ~1600 large particles. Most classes show a KtrB2A8B2 arrangement, a few are KtrB2A8A8B2 complexes. The lower panel shows a classification of ~500 small particles. Most classes show an elliptic top view with four KtrA C-terminal dimers; two classes show a side view of a KtrAB complex. Scale bar: 10 nm. (C) 2D class averages of the final cryo-EM data set of 20,500 particles. The classes show different views of the complex, many with high-resolution detail. (D) The final cryo-EM density map with the orientation distribution. The length of each cylinder represents the number of particles viewed from that direction. Although the particles have preferred orientations (long red cylinders), there are no missing orientations. (E) Fourier Shell Correlation curve indicating a resolution of 6.6 Å using the 0.143 criterion (dotted line). (F) Top view and side view of the density map with high (left) and low (right) contour level. At low contour level, the detergent micelle around KtrB and the characteristic peripheral C lobes around the octameric KtrA ring are visible.
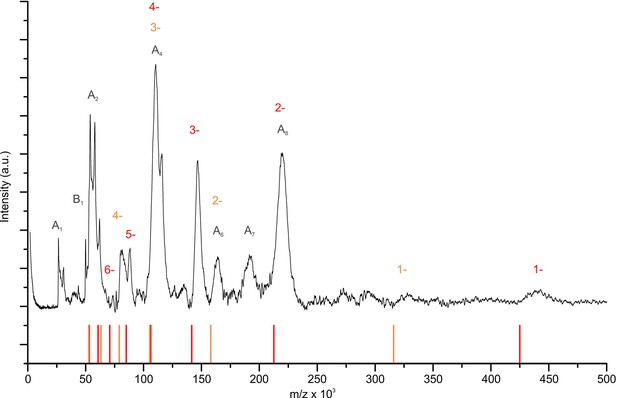
Mass spectrum of the detergent-solubilized KtrAB complex.
The LILBID-MS measurement reveals the presence of the following complexes: four KtrBs associated to eight KtrAs (red), and of two KtrBs bound to eight KtrAs (orange). Additionally, subunits KtrA and KtrB are shown in different complexations (grey).
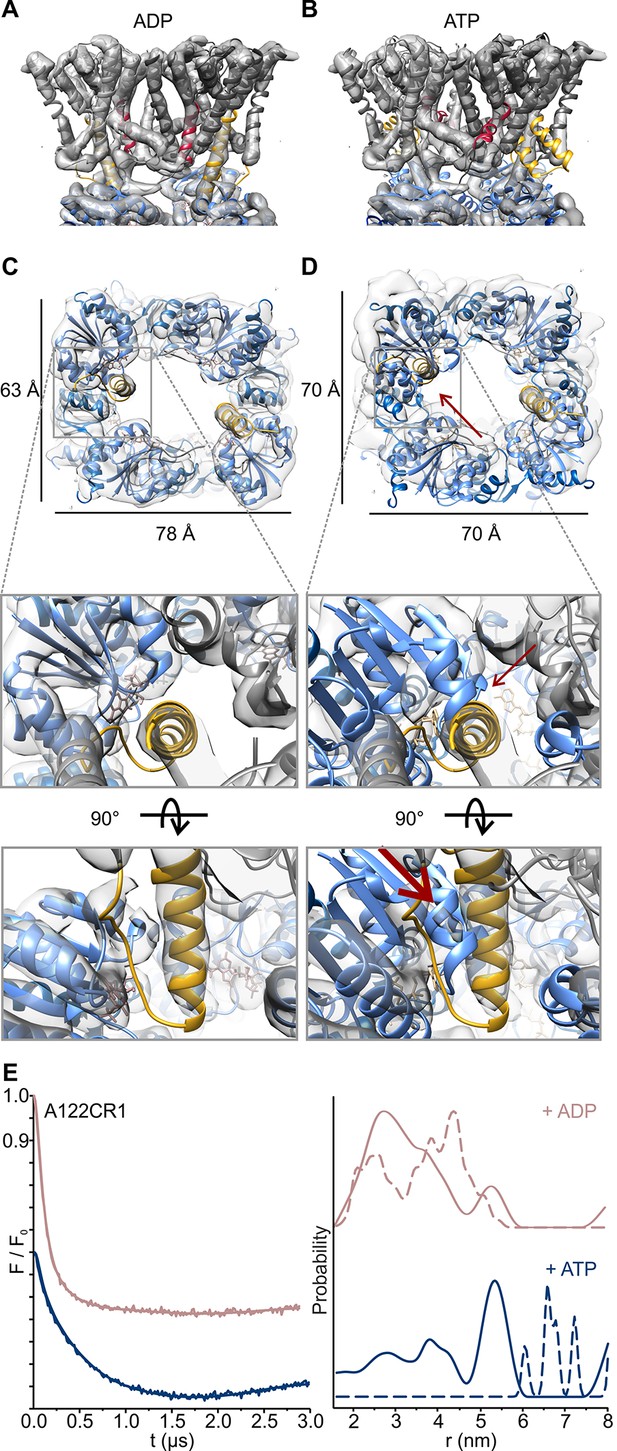
Nucleotide-induced conformational changes of the KtrAB complex.
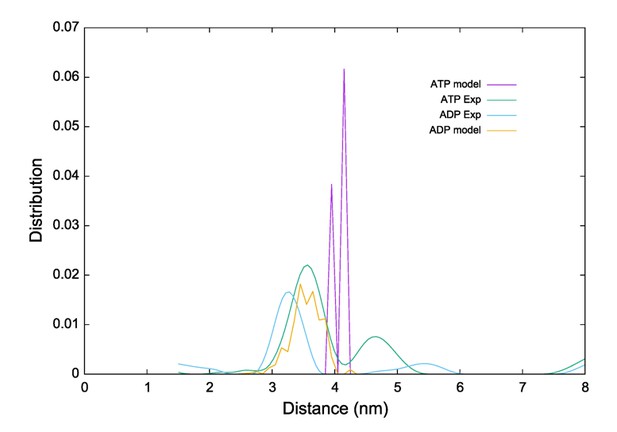
Overlay of the KtrAB density map in the presence of ADP with KtrAB model in the presence of (A) ADP and (B) ATP. KtrB subunits are depicted in light and dark grey, D1M2b helices in yellow, D4M2 helices in magenta, and RCK domains in blue. (C), (D) Top and side views of overlays of the density map with the ADP-bound KtrB model and the ADP- (C) or the ATP-bound (D) model of KtrA. KtrBs, grey and light grey with helices D1M2b, yellow; octameric ring of KtrA, blue and light blue; ADP, rosy brown; ATP, tan. Red arrows indicate steric hindrance of helix D1M2 with the ATP-bound KtrA. (E) Pulsed EPR data of spin-labeled variant KtrAB A122CR1 in detergent solution recorded at Q band (34 GHz). Dipolar evolution functions and the corresponding interprotomer distances obtained by Tikhonov regularization were determined in the presence of ADP (rosy brown) and ATP (blue), respectively. Dashed lines represent the predicted distance distributions of the ADP- and ATP- bound models, respectively, using the rotamer library analysis (Polyhach et al., 2011).
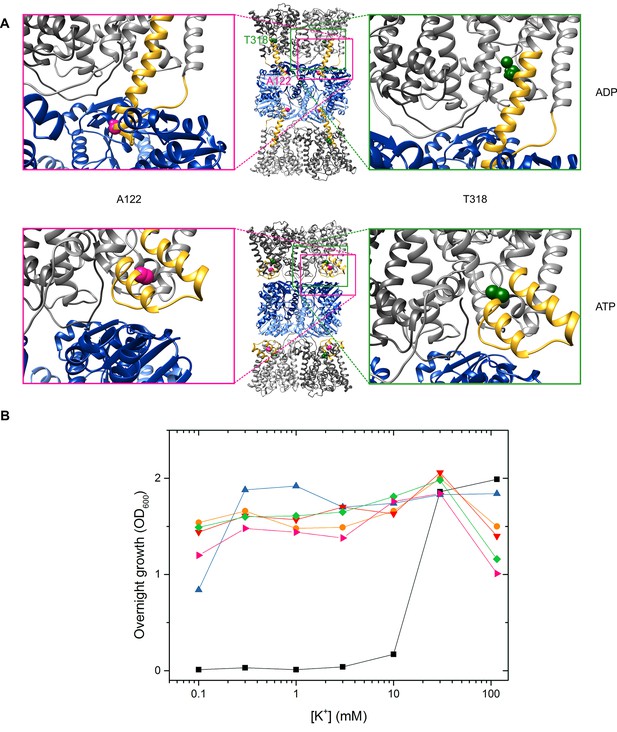
Functionality of protein variants.
(A) Location of spin-labeled residues A122CR1 (pink) and T318CR1 (green) in the ADP- and ATP-bound models. (B) Complementation assay for determination of K+ uptake activity of VaKtrAB variants. Different KtrAB variants-containing E. coli LB2003 cells were grown overnight in minimal media with glycerol as a carbon source at the K+ concentrations indicated.  , empty vector pBAD18;
, empty vector pBAD18;  , pEL903, encoding for VaKtrB-His6;
, pEL903, encoding for VaKtrB-His6;  , pIH301, encoding for His10-VaKtrAB;
, pIH301, encoding for His10-VaKtrAB;  , pEL903-100 T318C, encoding for Cys-less VaKtrB-His6 with T318C in KtrB;
, pEL903-100 T318C, encoding for Cys-less VaKtrB-His6 with T318C in KtrB;  , pIH301-100 T318C, encoding for Cys-less His10-VaKtrAB with T318C in KtrB;
, pIH301-100 T318C, encoding for Cys-less His10-VaKtrAB with T318C in KtrB;  pIH301-100 A122C, encoding for Cys-less His10-VaKtrAB with A122C in KtrB.
pIH301-100 A122C, encoding for Cys-less His10-VaKtrAB with A122C in KtrB.
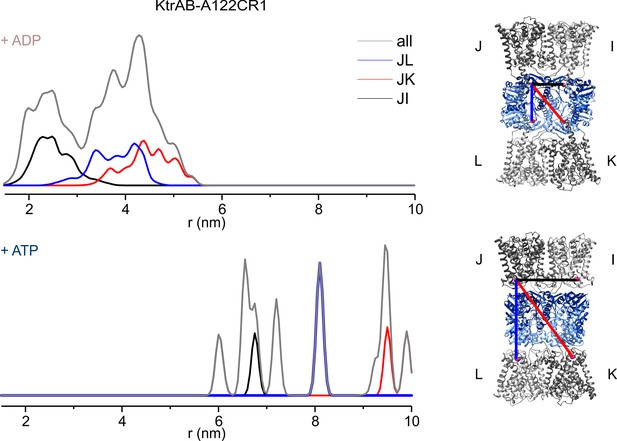
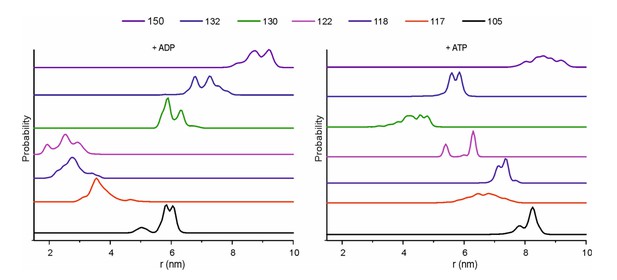
Inter-protomer distances within the detergent-solubilized KtrB2A8B2 complex predicted by rotamer library analysis.
Inter-protomer distance distributions P(d) obtained by the rotamer library analysis on the ADP- (upper panel) and ATP-bound model (lower panel) for the J–L (blue), J–K (red) and J–I (black) protomers. In grey, the sum of the contributions of the three protomers pairs (as in Figure 5).
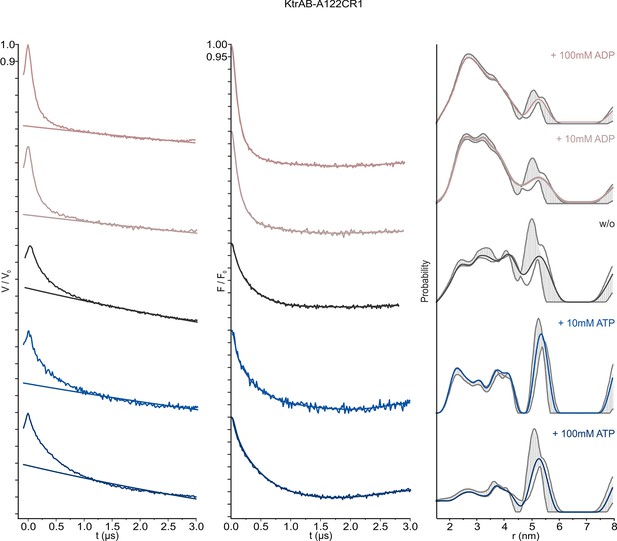
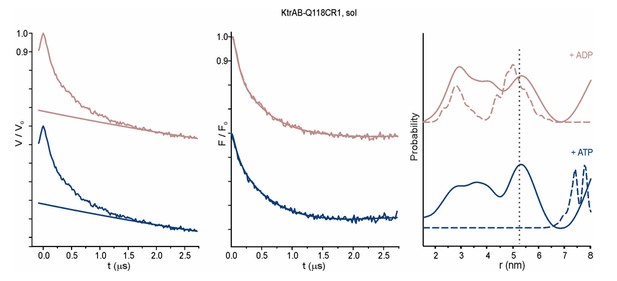
Pulsed EPR data of spin-labeled variant KtrAB A122CR1 in detergent solution recorded at Q band (34 GHz).
KtrAB in the presence of ADP, rosy brown and ATP, blue; KtrAB in the absence of nucleotides, dark grey. Left, experimental pulsed EPR data with fitted background function; middle, dipolar evolution function; right, interspin distance distribution obtained by Tikhonov regularization. Grey areas indicate the full variation of possible distance distributions. The lower and upper error estimates (grey lines) represent the respective mean value minus and plus two times its standard deviation, respectively.
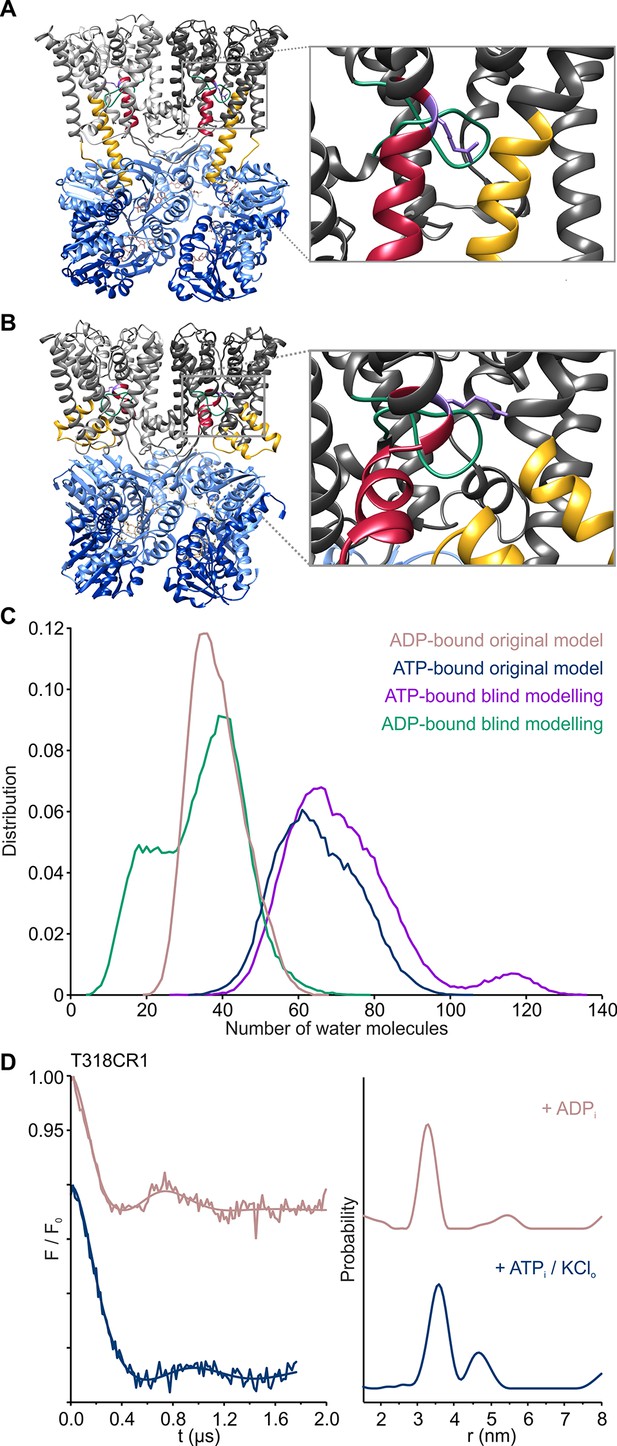
Gating mechanism of the KtrAB complex.
(A), (B) Side views of one gating region of the KtrAB complex in the ADP- (A) and ATP-bound (B) conformation with helices at the front removed. KtrBs, grey and light grey with helices D1M2b, yellow and D4M2b, magenta; octameric ring of KtrA, blue and light blue; intramembrane loop, green; R427, purple. (C) Number of water molecules in the cavity below the gating region during MD simulations in the ADP-bound (rosy brown; simulation code: ABDGVA+ABDNVA) and ATP-bound states (blue; simulation code: ABTGVA+ABTNVA). For reference, averaged hydration numbers are shown from 10 additional simulations each in the ADP-bound (green; simulation code: ABDGBVA) and ATP-bound states (purple; simulation code: ABTGBVA), starting from structures in which the intramembrane loop was modeled blindly. The lower end of the cavity volume was consistently defined by a plane passing through the lipid head groups. (D) Pulsed EPR data of spin-labeled variant KtrAB T318CR1 reconstituted in liposomes recorded at Q band (34 GHz). Dipolar evolution functions and the corresponding interprotomer distances obtained by Tikhonov regularization were determined in the presence of internal ADP (rosy brown) and in the presence of internal ATP and external K+ (blue), respectively.
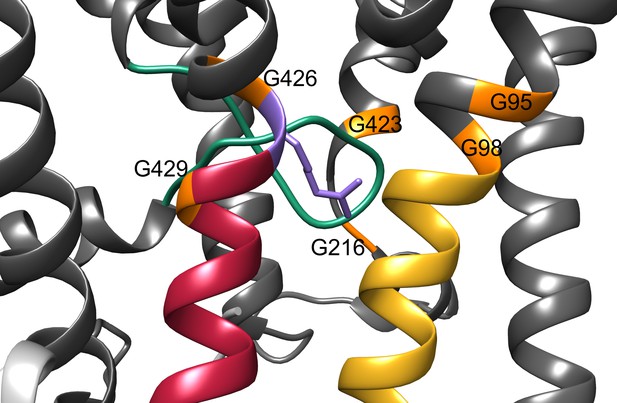
Structural gating features of the KtrB subunit.
Close up of the gating region of the KtrB (grey) subunit. The intramembrane loop is colored in green, the gating hinge glycines in orange, D1M2b helix in yellow, D4M2b helix in magenta and the conserved arginine in purple.
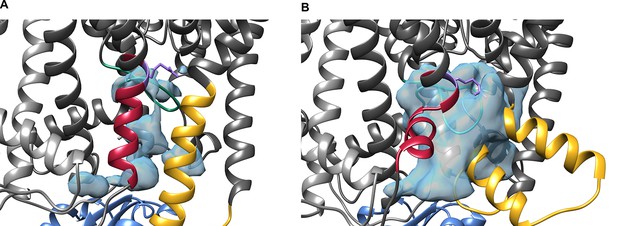
Comparison of water accessibility in the cavity below the gating region from MD simulations of ADP- (A) and ATP-bound (B) states of KtrAB.
The light-blue surfaces indicate the boundaries of the water-occupied cavity below the selectivity filter determined from 200-ns MD simulations (simulation codes of the used systems for the figure are ABDNVA and ABTNVA for A and B sections, respectively). The surfaces are cut off at the bottom by a plane defined by the lipid head groups. The intramembrane loop is colored in green, the D1M2b helix in yellow, the D4M2b helix in magenta, and the conserved arginine in purple.
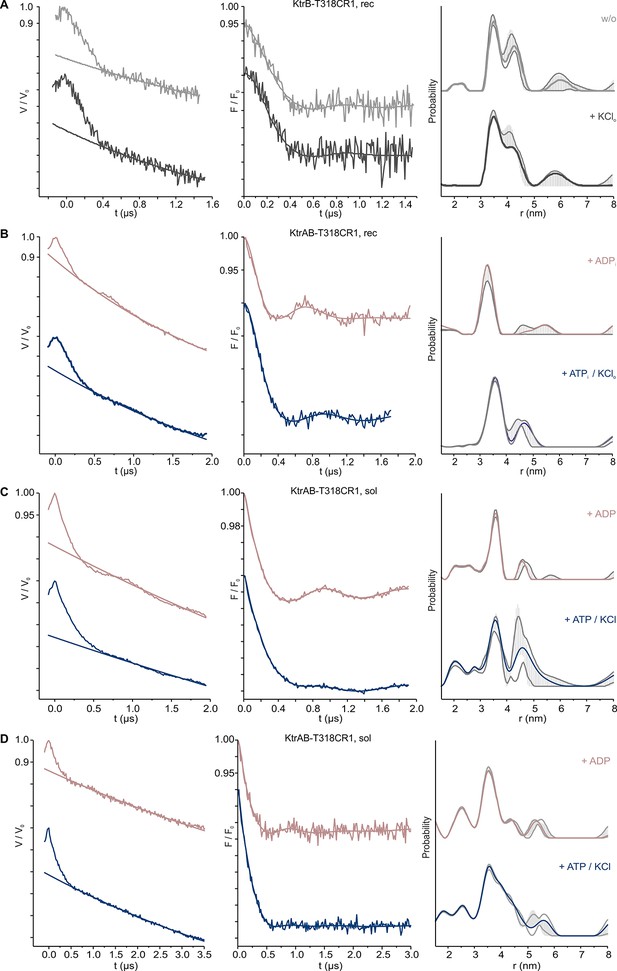
Pulsed EPR data of spin-labeled variants KtrAB/KtrB T318CR1.
(A) Pulsed EPR data of spin-labeled variant KtrB T318CR1 in liposomes recorded at X-band (9.4 GHz). (B) Pulsed EPR data of spin-labeled variant KtrAB T318CR1 in liposomes recorded at Q band (34 GHz). (C) Pulsed EPR data of spin-labeled variant KtrAB T318CR1 in detergent micelles recorded at Q band (34 GHz) for 2 µs. (D) Pulsed EPR data of spin-labeled variant KtrAB T318CR1 in detergent micelles recorded at Q band (34 GHz) for 3.5 µs. KtrAB in the presence of ADP, rosy brown and ATP, blue; KtrB in the presence of external K+, dark grey, and in its absence, light grey. Left, experimental pulsed EPR data with fitted background function; middle, dipolar evolution function; right, interspin distance distribution obtained by Tikhonov regularization. Grey areas indicate the full variation of possible distance distributions. The lower and upper error estimates (grey lines) represent the respective mean value minus and plus two times its standard deviation, respectively.
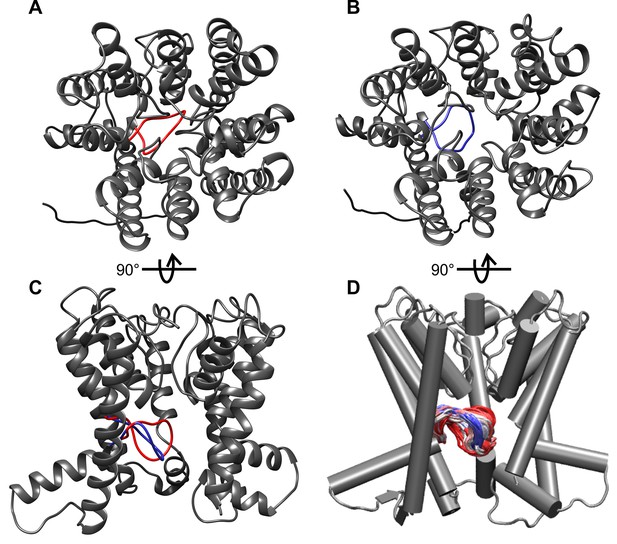
Fluctuations of the intramembrane loop in MD simulations of B. subtilis KtrB in its ATP-bound conformation.
(A) Top view of the most closed conformation. (B) Top view of the most open conformation. (C) Side view of both the most closed (red) and the most open conformation (blue) with helices at the front removed. (D) Superposition of the intramembrane loop conformations at 1-ns intervals from 400-ns MD simulations (simulation code: BTGBS) (beginning of simulation: red, end of simulation: blue; helices at the front removed).
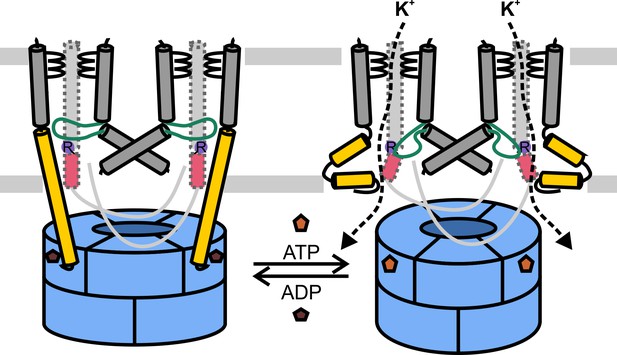
Model for KtrAB activation.
Cartoons showing the ADP- and ATP-bound conformations. With ADP bound, the extended helices D1M2b of KtrB dimers interact with residues in KtrAs close to the ADP binding sites, and with the gating regions of KtrBs. The gates are locked in the closed conformation. In the presence of ATP, the D1M2 helices break, forming helix hairpins at the membrane surface, and helices D4M2b move away from the pore. The disruption of interactions in the gating regions allows the intramembrane loops to move. In general, K+ flux is enabled. KtrAs, blue; KtrBs, grey with helices D1M2b, yellow; helices D4M2b, magenta; intramembrane loop, green, and conserved arginine, purple; ADP, dark brown; ATP, light brown.
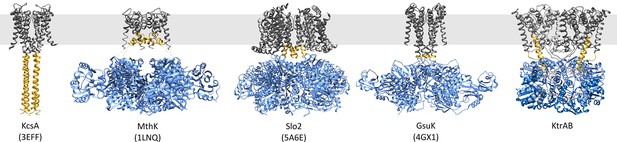
Comparison of proteins regulated by RCK domains.
Side views of KcsA (PDB code 3EFF [Uysal et al., 2009]), MthK (PDB code 1LNQ [Jiang et al., 2002b]), Slo2 (PDB code 5A6E [Hite et al., 2015]), GsuK (PDB code 4GX1 [Kong et al., 2012]) and ADP-bound KtrAB. Membrane domains/protomers, grey; flexible linker or extended helices, yellow; octameric RCK domains/protomers, blue. The grey bar represents the membrane.
Videos
Conformational differences between the ADP- and ATP-bound KtrAB complex.
Linear morph between the ADP- and ATP-bound KtrAB models providing a visual aid for the undergone conformational changes shown in Figure 6. Subunits and structural features are colored according to Figure 6.
Tables
Description of the different MD simulation systems and setups performed in this study.
| Simulation code | System | Conformation | Organism | MD package | Length |
|---|---|---|---|---|---|
| ABTGVA | KtrAB | ATP-bound | V. alginolyticus | GROMACS | 530 ns |
| ABTNVA | KtrAB | ATP-bound | V. alginolyticus | NAMD | 200 ns |
| ABDGVA | KtrAB | ADP-bound | V. alginolyticus | GROMACS | 450 ns |
| ABDNVA | KtrAB | ADP-bound | V. alginolyticus | NAMD | 200 ns |
| BTGBS | KtrB | ATP-bound | B. subtilis | GROMACS | 400 ns |
| BTNBS | KtrB | ATP-bound | B. subtilis | NAMD | 240 ns |
| ABTGBVA | KtrAB | ATP-bound | V. alginolyticus | GROMACS | 10 × 60 ns |
| ABDGBVA | KtrAB | ADP-bound | V. alginolyticus | GROMACS | 10 × 60 ns |





