Allosteric control of an asymmetric transduction in a G protein-coupled receptor heterodimer
Figures
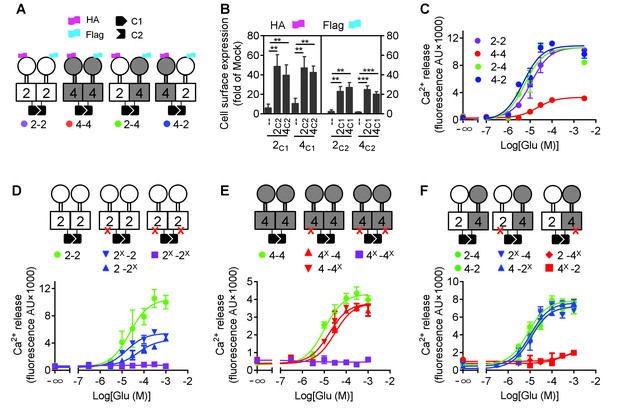
mGlu4 activates G protein in the mGlu2-4 heterodimer.
(A) Cartoons illustrating mGlu2 and mGlu4 homodimers, and mGlu2-4 (2–4 or 4–2) heterodimers with each subunit carrying the quality control C1 or C2 system as C terminal tails, and the indicated HA or Flag tag at their N terminus. (B) Quantification of cell surface expressed HA-tagged or Flag-tagged constructs by ELISA on intact cells transfected with the indicated subunits (2C1, 2C2, 4C1, 4C2) alone or together. Data are expressed as means ± SEM (n ≥ 3). **p<0.01, ***p<0.001 (unpaired t test). (C, D, E, F) Intracellular Ca2+ responses mediated by the indicated subunits upon stimulation with increasing concentrations of glutamate, in the presence of the chimeric Gqi9, with the control subunits (C), the mGlu2 homodimer with no, one or both subunits mutated (D), same with mGlu4 homodimer (E) or mGlu2-4 heterodimer (F). The red cross indicates the subunit carries the FS mutation that prevents G protein activation. Data are expressed as means ± SEM of triplicates from a typical experiment repeated at least three times.
-
Figure 1—source data 1
Glutamate potency at the indicated heterodimers.
Intracellular Ca2+ responses mediated by the indicated subunits upon stimulation with increasing concentrations of glutamate. 2X and 4X indicated subunits carrying the F756S (mGlu2) or F781S (mGlu4) mutation preventing G protein activation. Data represent the means ± SEM of (n) independent experiments. N.D., not determined.
- https://doi.org/10.7554/eLife.26985.006
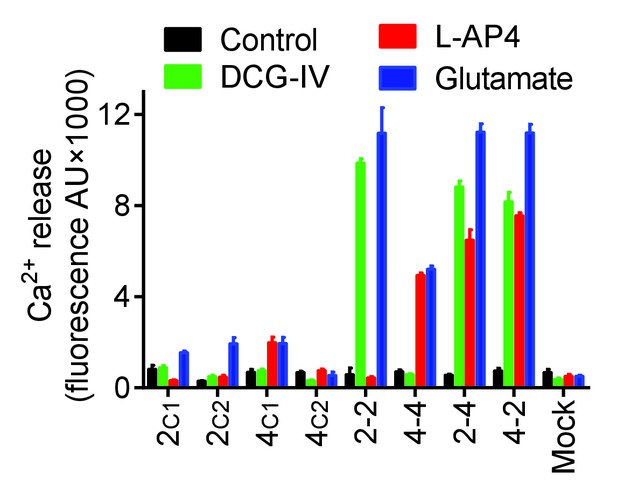
G protein coupling of the mGlu subunits with C1 or C2 tail.
Intracellular Ca2+ response mediated by indicated subunits co-expressed with the chimeric Gqi9 upon stimulation with DCG-IV (30 μM, green), L-AP4 (30 μM, red) or glutamate (1 mM, blue). Data are means ±SEM of triplicates from a typical experiment repeated at least three times.
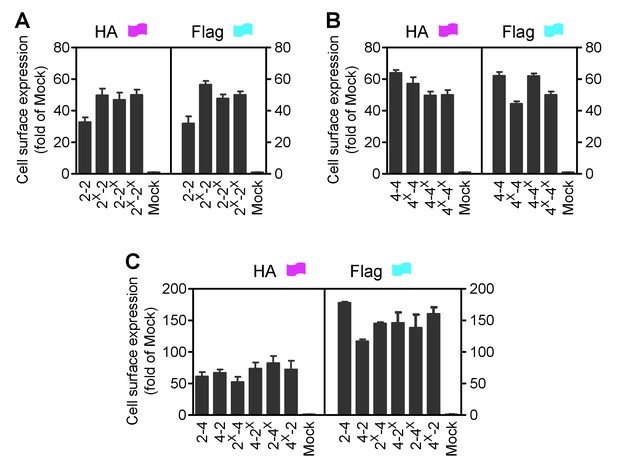
Cell surface and total expression of various mGlu dimer combination.
Quantification of cell surface-total expression of HA-tagged or Flag-tagged constructs (as indicated in Figure 1) by ELISA on intact cells transfected with indicated subunits. Data were obtained from a same cells used for with Figure 1D,E,F and expressed as means ± SEM of triplicates from a typical experiment repeated at least three times.
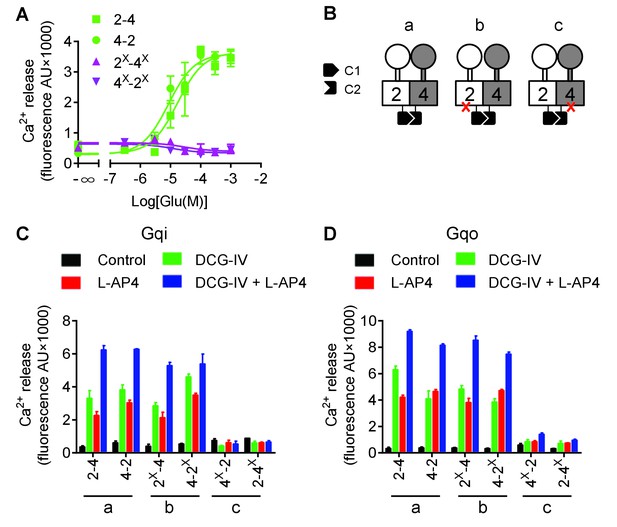
mGlu4 is responsible for G protein coupling in the mGlu2-4 heterodimer.
(A) Intracellular Ca2+ responses mediated by indicated subunits upon stimulation with increasing concentration of glutamate, in the presence of the chimeric Gqi9. (B) Cartoons illustrating the receptor combinations used in panels C and D. (C) Intracellular Ca2+ response mediated by indicated subunits co-expressed with Gqi9 upon stimulation with DCG-IV (30 μM), L-AP4 (30 μM), or DCG-IV +L-AP4. (D) Same as in C with Gqo. Data are means ±SEM of triplicates from a typical experiment repeated at least three times.
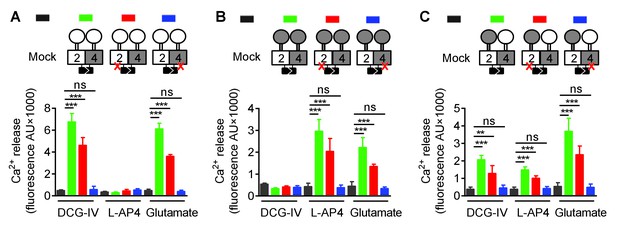
Asymmetric transduction results from the HDs in the mGlu2-4 heterodimer.
In (A), (B) and (C) cartoons illustrating the heterodimer combinations used with one or both subunits carrying the FS mutation (red cross) that prevents G protein activation are indicated on the top. For each subunit, white domains are from mGlu2, while the grey domains are from mGlu4. The chimeric protein made of mGlu2 ECD and mGlu4 HD is named 2ECD4HD and the reverse chimera named 4ECD2HD. The intracellular Ca2+ responses mediated by the indicated subunit compositions (color coded, as indicated on top of the cartoons) upon stimulation with DCG-IV (30 μM), L-AP4 (30 μM) or glutamate (1 mM) shown at the bottom. (A) Data obtained with heterodimers containing both ECDs (VFT and CRD) from mGlu2. (B) Data obtained with heterodimers containing both ECDs from mGlu4. (C) Data obtained with heterodimers in which the ECDs were swapped between the two subunits. Data are means ±SEM (n ≥ 3). **p<0.01, ***p<0.001 (unpaired t test).
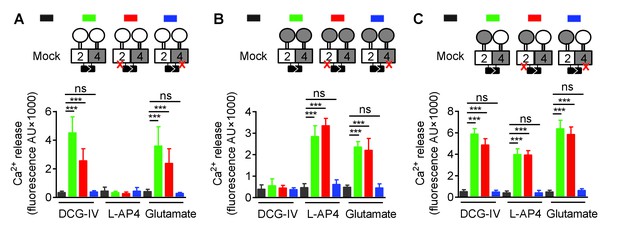
Asymmetric transduction results from the HDs in the mGlu2-4 heterodimer.
Same as Figure 2 excepted that only the VFT is swapped, and not the entire CRD. In (A), (B) and (C) cartoons illustrating the heterodimer combinations used with one or both subunits carrying the FS mutation (red cross) that prevents G protein activation are indicated on the top. For each subunit, white domains are from mGlu2, while the grey domains are from mGlu4. The chimeric protein made of mGlu2 VFT and mGlu4 CRD + HD is named 2VFT4HD and the reverse chimera named 4VFT2HD. The intracellular Ca2+ responses mediated by the indicated subunit compositions (color coded, as indicated on top of the cartoons) upon stimulation with DCG-IV (30 μM), L-AP4 (30 μM) or glutamate (1 mM) shown at the bottom. (A) Data obtained with heterodimers containing both VFT from mGlu2. (B) Data obtained with heterodimers containing both VFTs from mGlu4. (C) Data obtained with heterodimers in which the VFTs were swapped between the two subunits. Data are means ±SEM (n ≥ 3). **p<0.01, ***p<0.001 (unpaired t test).
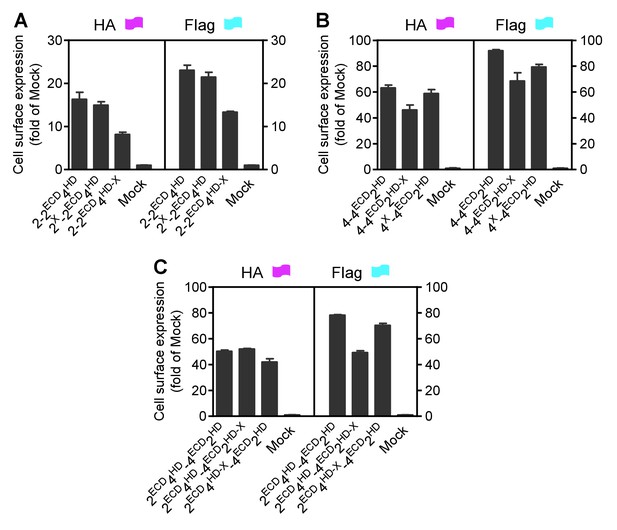
Cell surface and total expression of various mGlu dimer combination.
Quantification of cell surface-total expression of HA-tagged or Flag-tagged constructs by ELISA on intact cells transfected with the indicated subunits as in Figure 2. Data were generated from the same cells used in Figure 2 and are means ±SEM of triplicates from a typical experiment repeated at least three times.
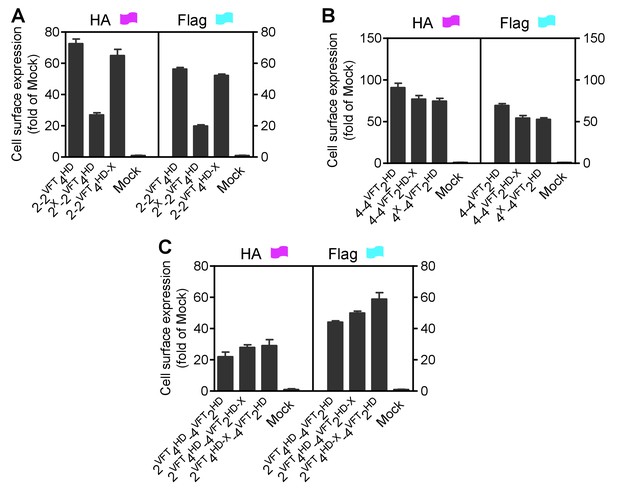
Cell surface and total expression of various mGlu dimer combination.
Quantification of cell surface-total expression of HA-tagged or Flag-tagged constructs by ELISA on intact cells transfected with the indicated subunits as in Sup Figure 4. Data were generated from the same cells used in Sup Figure 4 and are means ±SEM of triplicates from a typical experiment repeated at least three times.
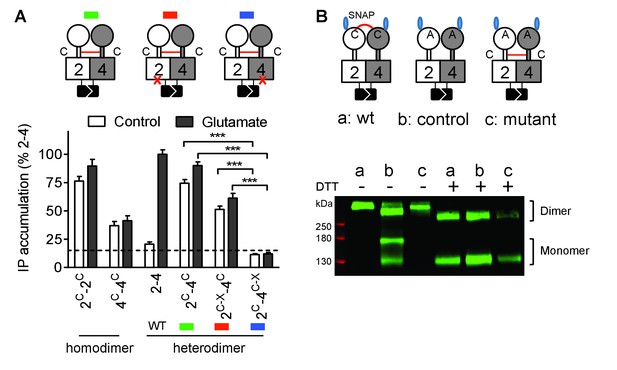
Constitutive activity of disulfide-tethered mGlu2-4 heterodimer is mediated by the mGlu4 subunit.
(A) Cartoons illustrating the heterodimer combinations used with mGlu2 or mGlu4 subunits carrying the FS mutation (red cross) that prevents G protein activation (top). The red line linking both CRDs indicates the disulfide bridge that constrains the dimer into an active state. Inositol phosphate (IP) accumulation in cells expressing the dimer combinations after incubation with or without glutamate (1 mM). Data are means ±SEM (n ≥ 3). ***p<0.001 (unpaired t test). (B) On the top, the cartoons indicate the heterodimeric combinations analyzed by western blots (bottom) with or without DTT treatment. The natural inter-subunit disulfide bridge in the control dimer (wt) is indicated in (a), leading to the lack of monomers in the non-reducing conditions. When mutating both Cys involved in this natural crosslink (C121A in mGlu2 and C136A in mGlu4), both subunits can dissociate into monomers even in the absence of DTT (b). Adding a new disulfide bridge in the CRD (L521C in mGlu2 and H523C in mGlu4) (c) restores the subunit cross-linking. By using SNAP-tag labeling with a cell-impermeant fluorescein substrate, only the cell surface subunits are labeled, and then detected on the blot. Data are from a typical experiment repeated three times.
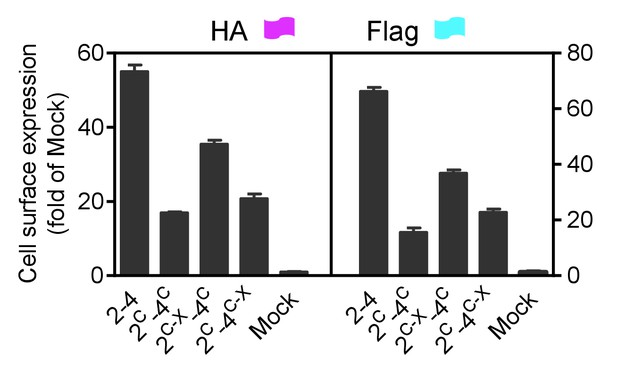
Cell surface and total expression of various mGlu dimer combination.
Quantification of cell surface-total expression of HA-tagged or Flag-tagged constructs by ELISA on intact cells transfected with the indicated subunits as in Figure 3A. Data were generated from the same cells used in Figure 3A and are means ±SEM of triplicates from a typical experiment repeated at least three times.
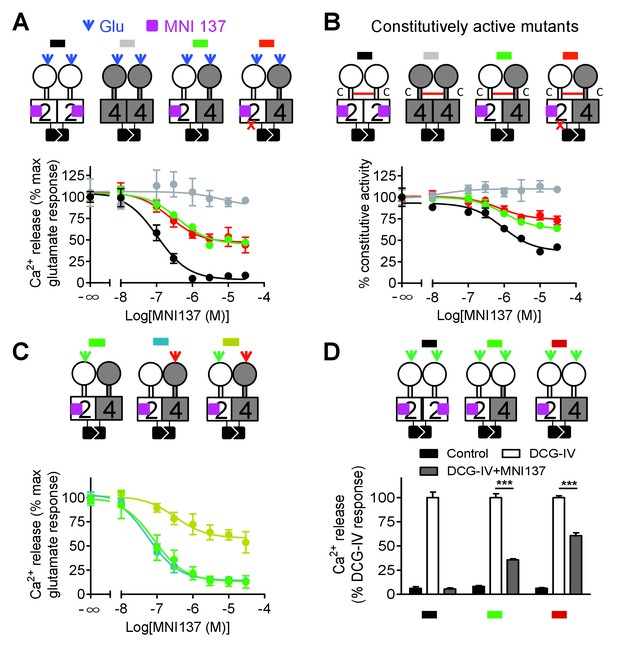
Allosteric regulation of mGlu4-induced signaling by mGlu2 HD.
In each panel, cartoons (color coded) illustrating the dimer compositions used are indicated on the top, and intracellular Ca2+ responses mediated by indicated dimer combinations upon stimulation with glutamate (1 mM) and increasing concentration of the mGlu2 NAM, MNI137 (purple square). The inactivating FS mutation is shown as a red cross. (A) Effect of MNI137 on homodimeric mGlu2 and mGlu4 receptors, and on the mGlu2-4 heterodimer carrying or not the FS mutation in the mGlu4 subunit activated by glutamate (blue arrow). (B) Effect of MNI137 on the constitutively active dimers resulting from the CRD disulfide cross-linking. (C) Effect of increasing concentrations of MNI137 on the mGlu2-4 heterodimer activated by the mGlu2 agonist DCG-IV (30 μM, green arrow), L-AP4 (30 μM, red arrow) or both. (D) Intracellular Ca2+ response under control condition, or after stimulation with DCG-IV (30 μM, green arrow) with or without MNI137 (10 μM) with the indicated dimer combinations. Data are means ±SEM of triplicates from a typical experiment repeated at least three times (A, B, C), or from three independent experiments (D). ***p<0.001 (unpaired t test).
-
Figure 4—source data 1
MNI137 potency at the indicated mGlu dimers.
Intracellular Ca2+ response mediated by indicated subunits upon stimulation with glutamate (1 mM) and inhibited by increasing concentration of MNI137. Data represent the means ± SEM of (n) independent experiments. N.D., not determined.
- https://doi.org/10.7554/eLife.26985.014
-
Figure 4—source data 2
MNI137 potency at the indicated mGlu dimers.
Inositol phosphate (IP) accumulation in cells expressed indicated subunits, which have constitutive activation, upon inhibition with increasing concentration of MN137. Data represent the means ± SEM of (n) independent experiments. N.D., not determined.
- https://doi.org/10.7554/eLife.26985.015
-
Figure 4—source data 3
MNI137 potency at the indicated heterodimers.
Intracellular Ca2+ response mediated by indicated heterodimer upon stimulation with DCG-IV (30 μM), L-AP4 (30 μM) or DCG-IV (30 μM)+L-AP4 (30 μM) and inhibited by increasing concentration of MNI137. Data represent the means ± SEM of (n) independent experiments. N.D., not determined.
- https://doi.org/10.7554/eLife.26985.016
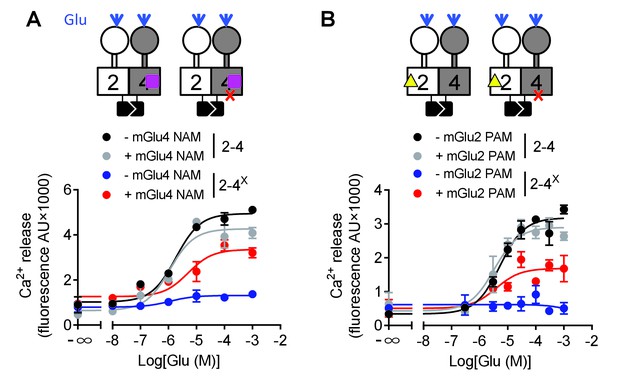
Switching of the G protein coupling subunit in the mGlu2-4 heterodimer by mGlu2 PAM and mGlu4 NAM.
Intracellular Ca2+ response mediated by the indicated subunits upon stimulation with increasing concentration of glutamate with/without a mGlu4 NAM (optoGluNAM4.1, purple square, 30 μM) or a mGlu2 PAM (LY487379, yellow triangle, 10 μM). (A) The mGlu4 NAM allows mGlu2 HD coupling to G proteins in the heterodimer. (B) The mGlu2 PAM allows mGlu2 HD coupling to G proteins. Data are means ±SEM of triplicates from a typical experiment repeated at least three times.
-
Figure 5—source data 1
Glutamate potency at the indicated heterodimers and in the presence or absence of the indicated allosteric modulators.
Intracellular Ca2+ response mediated by indicated subunits upon stimulation with increasing concentration of glutamate with/without optoGluNAM4.1 (30μM) or LY487379 (10 μM). Data represent the means ± SEM of (n) independent experiments. N.D., not determined.
- https://doi.org/10.7554/eLife.26985.018
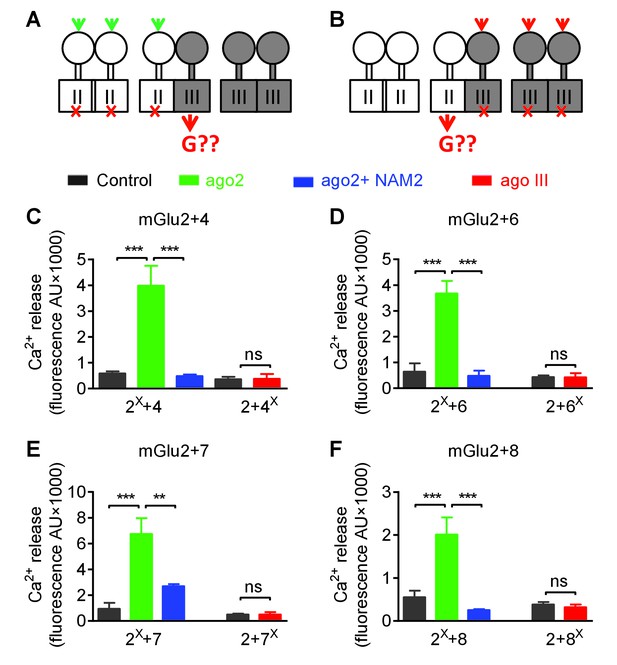
Asymmetric transduction by mGlu2-groupIII heterodimers.
(A–B) schemes illustrating the method used to study the coupling properties of mGlu heterodimers composed of mGlu2 (group-II) and a group-III subunit with the wild-type C-terminal tails. In (A), activating specifically the mGlu2 subunit unable to activate G protein (F756S, red cross) can generate a signal only if associated with a functional group-III subunit. (B) Same as in (A) with the inactive group-III subunit (F781S, F773S, F784S, F777S in mGlu4-6-7-8, respectively) and a specific group-III agonist. (C–F) functional coupling of the indicated subunits under the condition indicated on the top (black, control; green, group-II agonist (DCG-IV, 30 μM); blue, group-II agonist with mGlu2 NAM (DCG-IV, 30 μM and MNI137 10 μM); red, group-III agonist (L-AP4, 30 μM for mGlu4-6-8, LSP4-2022, 300 μM for mGlu7). (C) Data obtained with cells expressing both mGlu2 and mGlu4, with either the inactive mGlu2 (2X) or the inactive mGlu4 (4X). (D, E and F), same as in C using mGlu6, mGlu7 or mGlu8 constructs, respectively. Data are means ±SEM (n ≥ 3). **p<0.01, ***p<0.001 (unpaired t test).
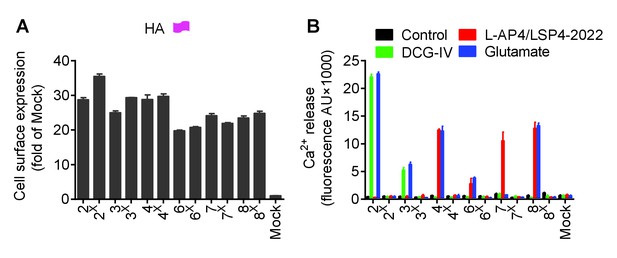
Expression and function of the indicated subunits.
(A) Quantification of cell expression of HA-tagged constructs by ELISA on cells transfected with the indicated subunits. (B) Intracellular Ca2+ response mediated by the indicated subunits upon stimulation with DCG-IV (30 μM) for mGluR2-3, L-AP4 (30 μM) for mGluR4-6-8, LSP4-2022 (300 μM) for mGluR7 or glutamate (1 mM). Data in A and B were generated from the same transfected cells, and are means ±SEM of triplicates from a typical experiment repeated at least three times.
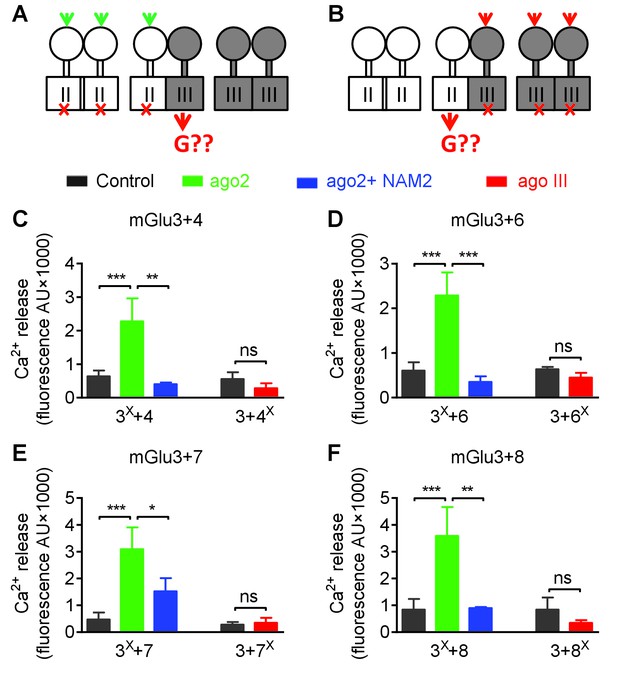
Asymmetric transduction by mGlu2-groupIII heterodimers.
Same as Figure 7, except mGlu3 was used instead of mGlu2. (A) and (B) schemes illustrating the method used to study the coupling properties of mGlu heterodimers composed of mGlu3 (group-II) and a group-III subunit. In (A), activating specifically the mGlu3 subunit unables to activate G protein (F765S, red cross) can generate a signal only if associated with a functional group-III subunit. (B) Same as in (A) with the inactive group-III subunit (F781S, F773S, F784S, F777S in mGlu4-6-7-8, respectively) and a specific group-III agonist. (C-D) functional coupling of the indicated subunits under the condition indicated on the top (black, control; green, group-II agonist (DCG-IV, 30 μM); blue, group-II agonist with mGlu2/3 NAM (DCG-IV, 30 μM and MNI137 10 μM); red, group-III agonist (L-AP4, 30 μM for mGlu4-6-8, LSP4-2022, 300 μM for mGlu7). (C) Data obtained with cells expressing both mGlu3 and mGlu4, with either the inactive mGlu3 (3X) or the inactive mGlu4 (4X). (D, E and F) same as in C using mGlu6, mGlu7 or mGlu8 constructs, respectively. Data are means ±SEM (n ≥ 3). **p<0.01, ***p<0.001 (unpaired t test).
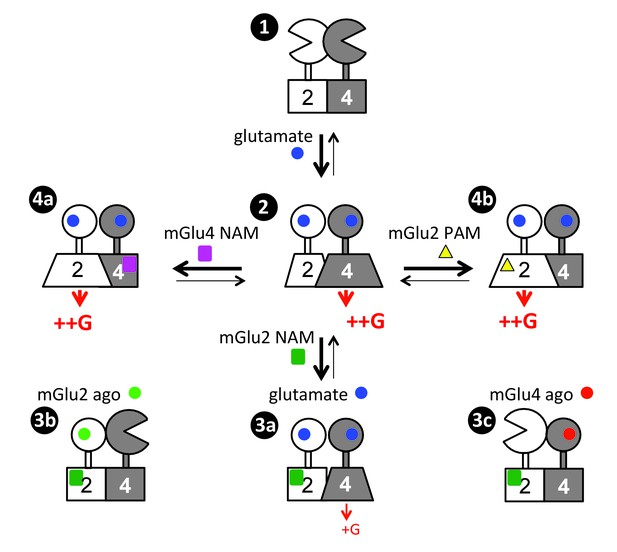
Scheme illustrating the activation mechanism and allosteric control of mGlu2-4 heterodimer.
State 1. the inactive heterodimer in its basal state. State 2: Glutamate (blue disk) activation of both subunits leads to G protein activation by mGlu4 HD, also involving a conformational change in the mGlu2 HD. State 3: the addition of mGlu2 NAM (green square) largely decreases coupling efficacy of the mGlu2-4 heterodimer activated by glutamate (3a), or suppress detectable coupling if either the mGlu2 (3b) or mGlu4 (3c) is specifically activated. State 4: heterodimeric mGlu2-4 coupling through the mGlu2 HD thanks to the addition of a mGlu4 NAM (purple square, 4a), or a mGlu2 PAM (yellow triangle, 4b).
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.26985.023





