The structure of the COPI coat determined within the cell
Figures
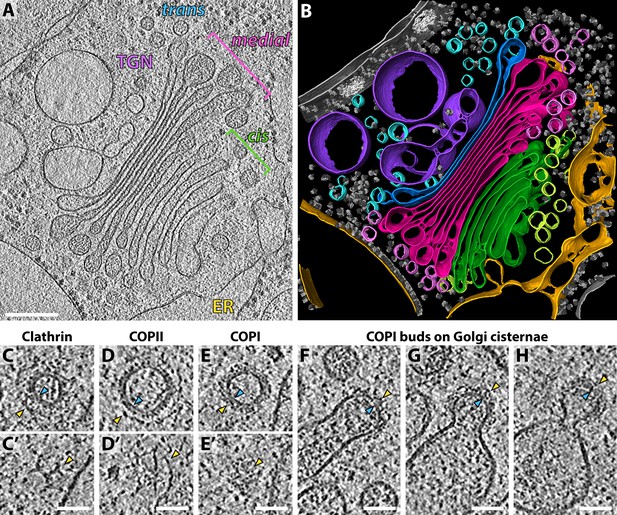
Molecular architecture of the Chlamydomonas Golgi apparatus and transport vesicles revealed by in situ cryo-ET.
(A) A slice through a cellular tomogram containing a representative Golgi stack and (B) corresponding 3D segmentation, showing the native morphology of the ER (yellow), four cis cisternae (green), cis vesicles (light green), four medial cisternae (magenta), medial vesicles (light pink), the trans cisterna (blue), trans vesicles (light blue) and the TGN (purple). Other membranes, the nuclear envelope, nuclear pore complexes and ribosomes are shown in grey. (C–E) Slices through (C, C’) clathrin, (D, D’) COPII and (E, E’) COPI coated transport vesicles found within the tomograms. Panels C, D and E are slices through the centers of the vesicles. Panels C’, D’ and E’ are top slices through the coats of the same vesicles, showing characteristic structures of (C’) clathrin triskelions, (D’) the triangular Sec13/31 COPII lattice, and (E’) the dense COPI coat. (F–H) Slices through three COPI buds in different stages of maturation. Yellow arrowheads: cytoplasmic boundary of the coat, blue arrowheads: vesicle or bud membrane. Scale bars: 200 nm in A-B, 50 nm in C-H.
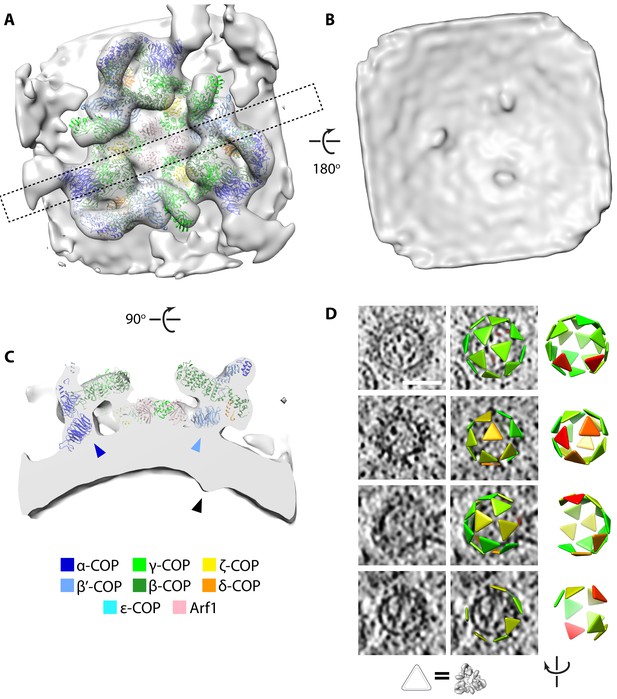
Determination of the native COPI coat structure and arrangement.
(A–C) Isosurface view of the triad density map determined within the cell. (A) View from the cytoplasmic side, dashed rectangle indicates the position of the cross-section shown in panel C. (B) View from the vesicle lumen side, displaying three cargo density protrusions. (C) Cross-section view, showing the position of the cargo density (black arrowhead) relative to the positions of the N-terminal β–propellers of α–COP (dark blue arrowhead) and β’-COP (light blue arrowhead). (D) Examples of coat arrangement on vesicles and buds. In each row, left: tomographic slice; middle: tomographic slice overlaid with triangles at the positions and orientations of each triad (correspondence between triangle and structure is illustrated below), colored by similarity to the structure in panels A-C (green to red = high to low correlation); right: view of the triad arrangement rotated to orient the budding scar towards the viewer. The vesicle in the bottom row has an incomplete coat and is likely undergoing uncoating. Scale bar: 50 nm.
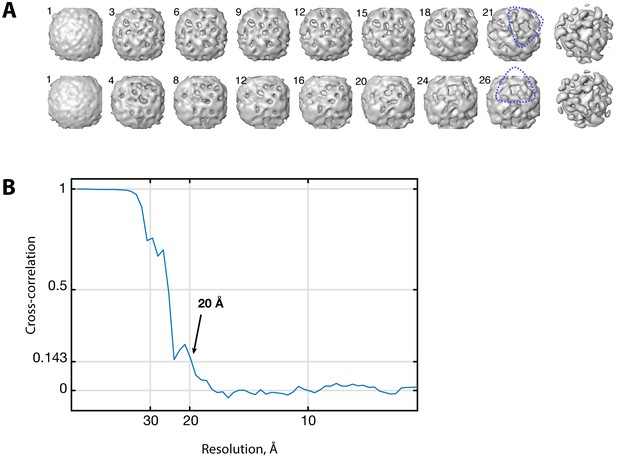
In situ COPI structure determination.
(A) Convergence of reference-free COPI reconstructions from two independent subsets of in situ data. Iteration number of the alignment is marked for each reference. After appearance of the triad structures (blue dashed lines), the triads were centered and aligned further applying three-fold symmetry to produce the structures on the right. (B) Fourier shell correlation curve for the final in situ COPI structure. The FSC between the two independent half-datasets is shown, indicating a resolution of 20 Å at the 0.143 criterion.
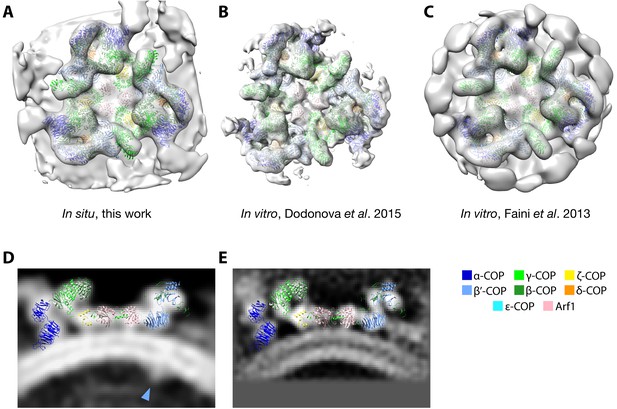
Comparison of in situ and in vitro structures.
(A–C) Isosurface views of COPI structures obtained in vitro and in situ. (A) The map from this work and EMDB maps (B) EMD-2985 (Dodonova et al., 2015) and (C) EMD-2084 (Faini et al., 2012). All structures are fit with PDB model 5A1U (Dodonova et al., 2015) as a rigid body (which at this resolution is essentially identical to PDB model 5NZR, derived from EMD-3720 (Dodonova et al., 2017)). (D, E) Comparison of orthographic slices through (D) the in situ map and (E) EMD-2985, both overlaid with PDB 5A1U. Luminal cargo density is highlighted in the in situ map (blue arrowhead).
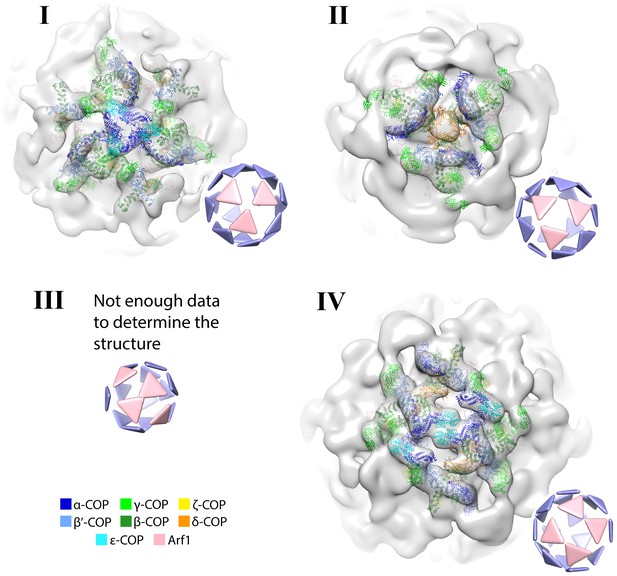
Structures of the in situ triad linkages.
Densities are displayed as isosurfaces. Insets show example vesicles from the dataset with the pattern of triads contributing to each linkage highlighted in pink. The linkage I structure is fit with PDB 5NZT, the linkage II structure is fit with PDB 5NZU, linkage III is rare and no structure was determined, the linkage IV structure is fit with PDB 5NZV (Dodonova et al., 2017). Note that linkage I is an asymmetric structure, and the three copies of the complex between ε-COP and the C-terminal domain of α–COP may not all be in equivalent positions (Dodonova et al., 2015). The asymmetry could not be resolved at this resolution. The central density of linkage II, corresponding to the µ-homology domain of δ-COP, is rather weak, which could indicate flexibility and/or asymmetry in the position of this domain in vivo. ε-COP and the µ-homology domain of δ-COP are both dispensable for COPI function (Arakel et al., 2016). We observed no additional density in the linkages that might correspond to bound cargo apart from the one found below the β’-COP subunit.
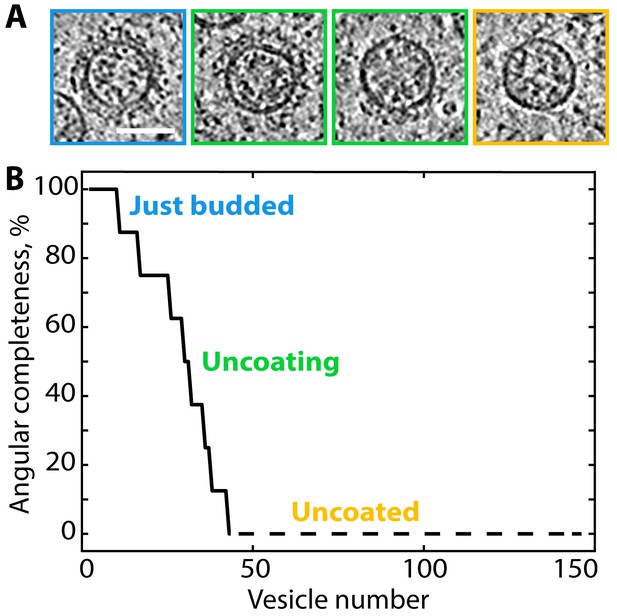
Ranked plot of COPI vesicle completeness indicates a rapid and processive uncoating process.
(A) Central slices through COPI vesicles at different stages of uncoating (from left to right). (B) Vesicles were sorted according to the angular completeness of the coat, as measured from central slices (see Materials and methods). Scale bar: 50 nm.
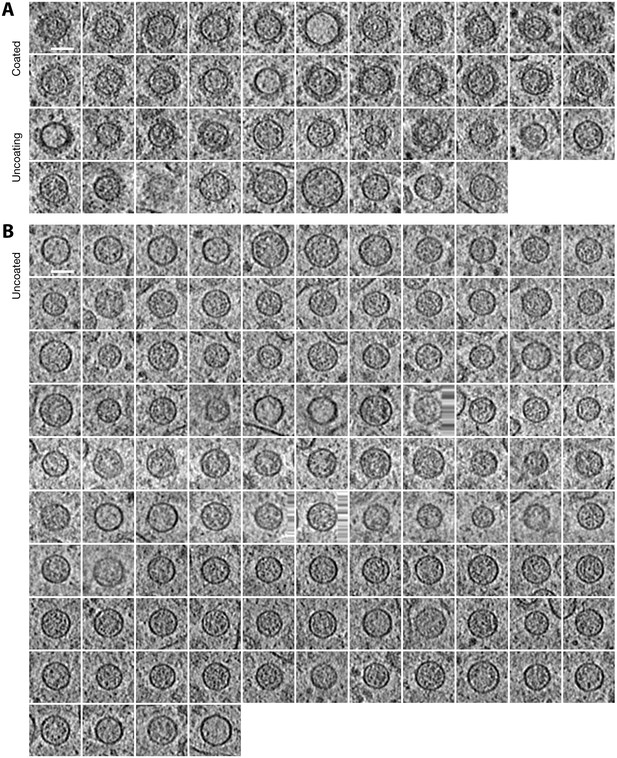

Examples of coated and uncoated COPI vesicles from five Golgi stacks.
(A) 42 vesicles with COPI coats of varying degrees of completeness. (B) 103 uncoated vesicles found within the ribosome exclusion zone surrounding the Golgi stack. Each image is an average of 13 consecutive tomographic slices (spacing 1.37 nm) at the vesicle center. Scale bars: 50 nm.
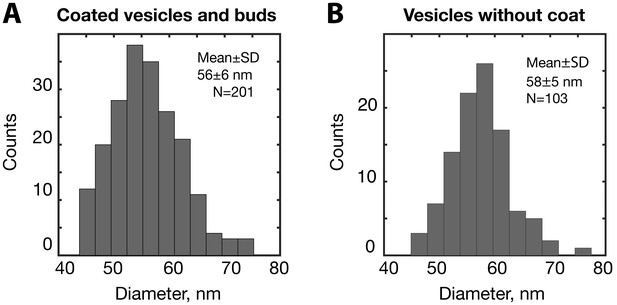
Diameter distributions of coated and uncoated COPI vesicles.
(A) Diameter distribution for COPI-coated late buds and vesicles, measured from membrane to membrane (excluding coats), (B) Diameter distribution of naked vesicles from Figure 3—figure supplement 1B. The size distribution is similar to that of the coated vesicles.
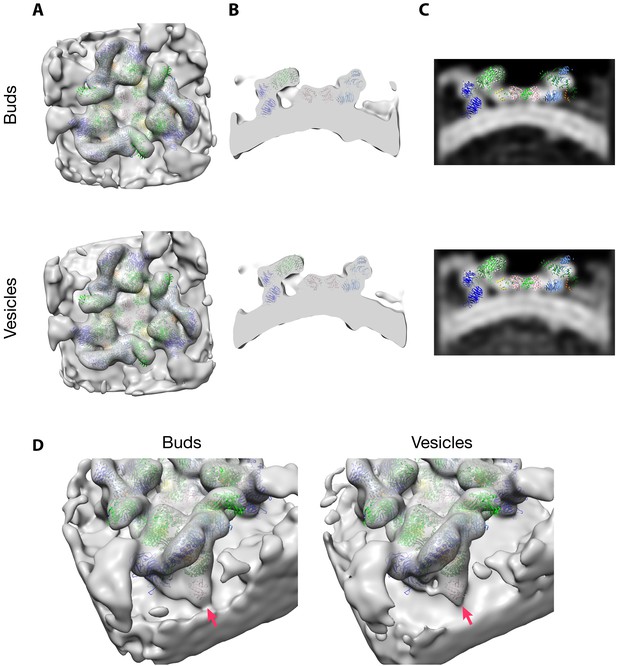
Comparison of the coat structures derived from buds and vesicles.
(A–C) First row: buds, second row: vesicles. (A) Isosurface overviews of the triads viewed from the cytoplasmic side. (B) Isosurface views sliced perpendicular to the membrane. (C) Orthoslice views as in panel B. (D) Isosurface views indicating the ‘outer’ Arf1 (red arrow). It could be hypothesized that this copy of Arf1 dissociates before other coat components, but we find that the Arf1 density is present in both buds (left) and released vesicles (right). Model color code is indicated in Figure 2 and Figure 2—figure supplement 2. Maps are shown at the same isosurface threshold. PDB 5A1U is fit as a rigid body. We note a slight difference in density between the Arf1 molecule bound to β–COP and the neighboring N-terminal β–propeller of α–COP. This is likely the result of the vesicle dataset containing a larger fraction of triads that are at the edge of the lattice. This is the position of a flexible loop region in α–COP and shows similar variability in previously described structures of different triad linkages (Dodonova et al., 2017).
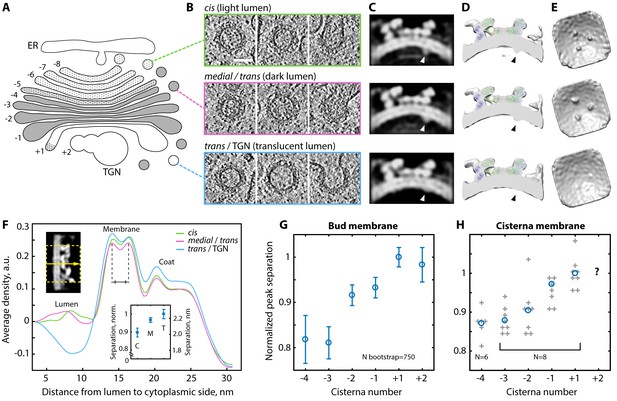
Variability of the COPI structure and membrane thickness throughout the Golgi.
(A) Schematic of the C. reinhardtii Golgi showing cisternae with cis (−8 to −6), intermediate (−5), medial (−4 to −1), and trans morphology (+1) and the TGN (+2). Cisternae are numbered relative to the transition from dark to translucent lumen (between −1 and +1). (B) Examples of buds and vesicles from three groups defined by position and luminal density (see Materials and methods). Top row: ‘cis’, middle row: ‘medial/trans’, bottom row: ‘trans/TGN’. (C–E) ~24 Å structures of the coat determined from the three groups. (C) Orthographic slices and (D) cross-section isosurface views through the averages. Arrowheads: cargo. (E) Isosurface views from inside the vesicle. While the coatomer structure remains constant, the ‘trans/TGN’ structure lacks bound cargo. (F) Density profiles through the three COPI structures. Top left inset: example orthographic slice through a straightened structure, dashed yellow rectangle and arrow: averaged region and direction of the profile. Bottom inset: bootstrapped mean bilayer leaflet separation measured for the ‘cis’ {C}, ‘medial/trans’ {M} and ‘trans/TGN’ {T} averages, error bars: bootstrapped SD. (G–H) Membrane bilayer thickness in (G) COPI buds and (H) non-coated membrane regions from individual medial and trans Golgi cisternae. In G, mean and error bars (SD) were bootstrapped (see Materials and methods). In H, grey crosses: individual cisternae, blue circles: means, TGN measurements were not obtained. Scale bar: 50 nm.
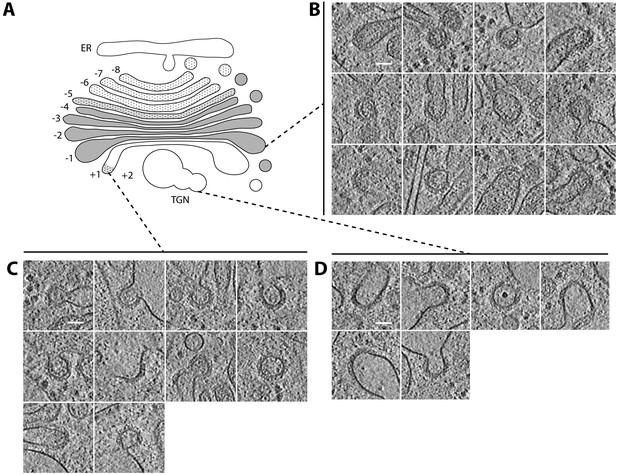
COPI bud and vesicle morphology varies across the Golgi stack.
(A) Schematic of the Golgi complex, see Figure 4A for explanation. (B–D) Examples of buds from the medial and trans Golgi, and the TGN. (B) On the last medial cisterna, the buds have dark luminal density similar to the donor cisternae. (C) On the trans cisterna, the buds typically have a darker lumen than the translucent donor cisternae. (D) On the swollen TGN, the buds have a translucent lumen similar to the donor compartment. Scale bar: 50 nm.
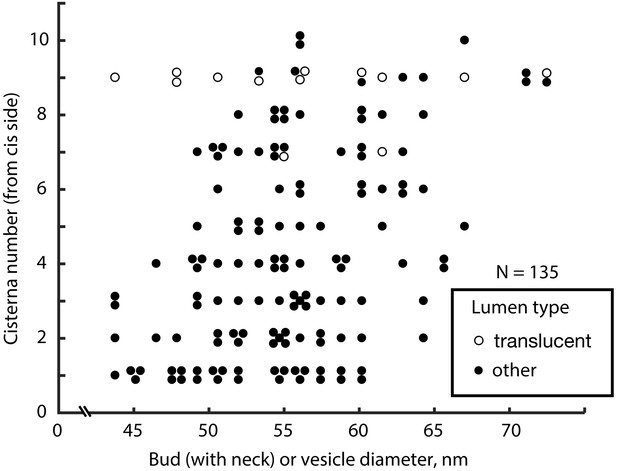
COPI vesicle and late bud diameter variation across the Golgi stack.
The diameter of vesicles and late buds, measured membrane to membrane, is plotted against position in the stack based on cisternae number counting from the cis face.
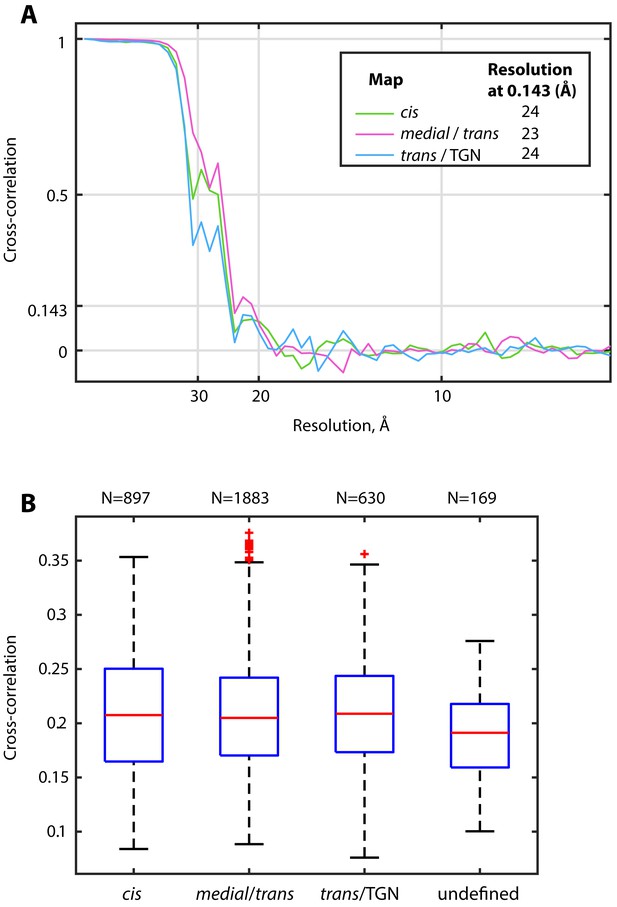
Structure determination of COPI obtained from different Golgi regions.
(A) Fourier shell correlation curves for the averages from the different Golgi regions depicted in Figure 4C–E indicate comparable resolutions of the structures. Resolutions are given in the legend. (B) Cross correlation coefficients of individual subtomograms from different Golgi regions to the common reference. The bottom and top of the boxes show 25 and 75 percentiles, the central red line shows 50 percentile. Whiskers show full data ranges, excluding outliers (red crosses). Outliers (red crosses) were calculated with default settings of the Matlab boxplot function. The number of subtomograms (N) for each group is displayed above the figure.
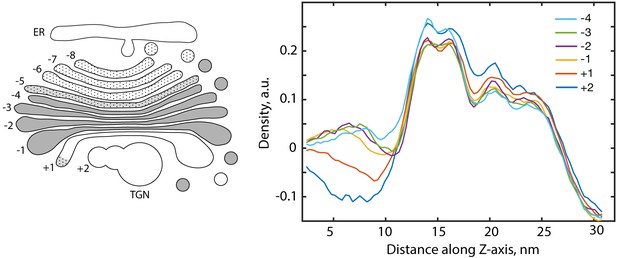
Density profiles through the averages derived from buds in individual trans and medial Golgi cisternae.
Donor cisternae morphology was defined according to the schematic on the left (numbered relative to the first translucent compartment). These profiles were used to measure the bilayer separations depicted in Figure 4G.
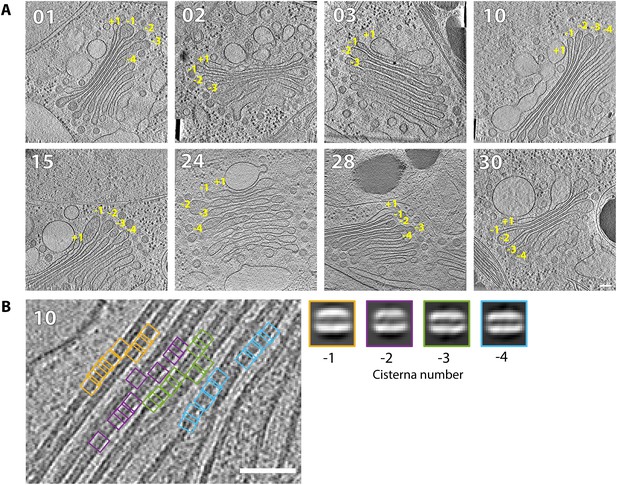

Membrane bilayer thickness measurements in individual cisternae of medial and trans Golgi.
(A) The eight Golgi stacks used for measurements. Cisternae were assigned numbers relative to the first translucent compartment, as diagrammed in Figure 4A; these numbers are indicated for each cisternae on each stack. (B) Close up view of the non-coated membranes from the central cisterna regions of Golgi 10, showing how the particles were extracted to obtain the bilayer 2D averages displayed on the right. Scale bars are 100 nm in panel (A) and 50 nm in panel (B).
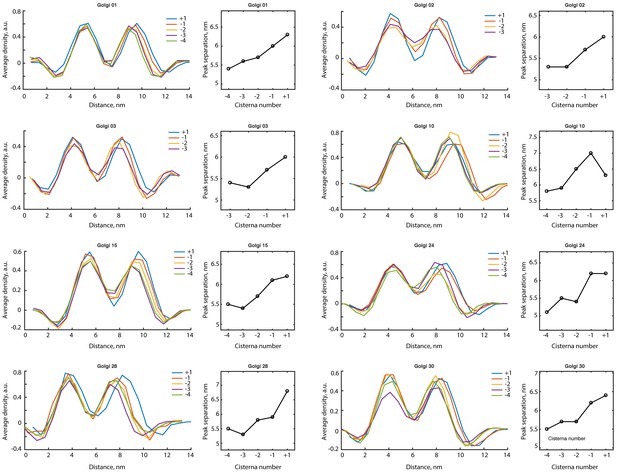
Membrane bilayer profiles measured in individual cisternae of medial and trans Golgi.
All measured density profiles and membrane bilayer separation distances for individual Golgi stacks. Each plot to the right of the profile summarizes the distance between the density peaks of each cisterna. An aggregate plot summarizing the dependence of bilayer thickness on cisterna number is shown in Figure 4H.
Videos
In situ cryo-electron tomogram of the native Chlamydomonas Golgi.
The movie slices back and forth in Z through the tomographic volume, then reveals the 3D segmentation, colored as in Figure 1B. The ER is yellow, four cis cisternae are green, cis vesicles are light green, four medial cisternae are magenta, medial vesicles are light pink, the trans cisterna is blue, trans vesicles are light blue and the TGN is purple. Other membranes, the nuclear envelope, nuclear pore complexes and ribosomes are shown in grey.
Overview of the in situ COPI structure and bound cargo density.
The movie shows an isosurface view of the native COPI triad structure. The PBD model 5A1U, derived from a structure of COPI determined in vitro (Dodonova et al., 2015), is fit into the structure as a rigid body. The isosurface is initially shown from the cytoplasmic side, before it is rotated 90° and sliced perpendicular to the membrane to show cargo density residing beneath the N-terminal β-propeller of β’-COPI but not beneath the analogous domain of α-COP. The isosurface is then rotated a further 90° and viewed from the vesicle lumen side to show three symmetric cargo densities.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| strain, strain background (Chlamydomonas reinhardtii) | mat3-4 mt+ | PMID: 11445540, DOI: 10.1101/gad.892101 | CC-3994, Chlamydomonas Resource Center | small cells improve vitrification |
| software, algorithm | SerialEM | PMID: 16182563, DOI: 10.1016/j.jsb.2005.07.007 | cryo-ET tilt-series acquisition | |
| software, algorithm | MotionCor2 | PMID: 28250466, DOI: 10.1038/nmeth.4193 | frame alignment | |
| software, algorithm | IMOD | PMID: 8742726, DOI: 10.1006/jsbi.1996.0013 | tomogram alignment, reconstruction | |
| software, algorithm | Amira | FEI/Thermo Fisher Sceintific | 3D segmentation of tomogram volumes | |
| software, algorithm | Matlab | Mathworks | subtomogram averaging, data analysis | |
| software, algorithm | TOM package for Matlab | PMID: 15721576, DOI:10.1016/j.jsb.2004.10.006 | subtomogram averaging | |
| software, algorithm | Av3 package for Matlab | PMID: 15774580, DOI: 10.1073/pnas.0409178102 | subtomogram averaging | |
| software, algorithm | RELION with helical processing | PMID: 28193500, DOI: 10.1101/095034 | averaging of 2D membrane profiles | |
| software, algorithm | USCF Chimera | PMID: 15264254, DOI: 10.1002/jcc.20084 | visualization |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.32493.020





