Radially patterned cell behaviours during tube budding from an epithelium
Figures
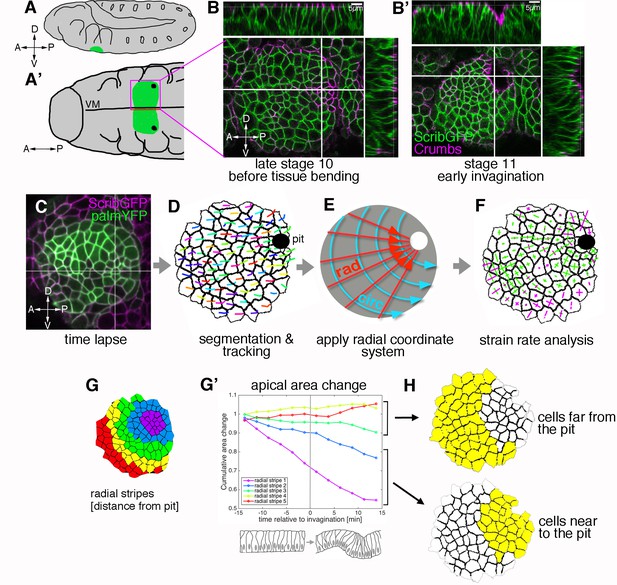
Morphogenetic events in the salivary gland placode show a radial organisation.
(A, A’). Schematic of a stage 11 Drosophila embryo highlighting the position of the salivary gland placode (green) in lateral (A) and ventral (A’) views; A: anterior, P: posterior; D: dorsal; V: ventral. (B, B’). Surface and cross-section views of the salivary gland placode just prior to the first tissue bending (B) and once the initial pit of invagination has formed (B’). Lateral membranes are labelled by ScribbleGFP (green) and apical cell outlines by Crumbs (magenta). Scale bars are 5 µm. (C-F) Workflow of the morphometric analysis. Early salivary gland placode morphogenesis is recorded by time-lapse analysis using markers of cell outlines (C), cells are segmented and tracked over time (D; exemplary tracks of individual cells are indicated by coloured lines). Data are recorded and expressed in a radial coordinate system with the invaginating pit as the origin (E; ‘rad’ is the vectorial contribution radially towards the pit, ‘circ’ is the vectorial circumferential contribution). Various derived measures are projected onto the radial coordinate system. Here, example projected small domain deformation (strain) rates are shown (F; contraction in green, expansion in magenta; see Figure 2B). See also Videos 1 and 2. (G-H) Cumulative time-resolved analysis of apical area change for nine embryos (G’), the cumulative area strain is plotted, grouping the placodal cells into radial stripes (G) for the analysis. This reveals a clear split in behaviour between cells far from the pit (H, top; negligible area change) and cells near the pit (H, bottom; constriction, consistently negative area change). See also Figure 1—figure supplement 1.
-
Figure 1—source data 1
Apical area change in radial stripes.
Related to Figure 1G.
- https://doi.org/10.7554/eLife.35717.005
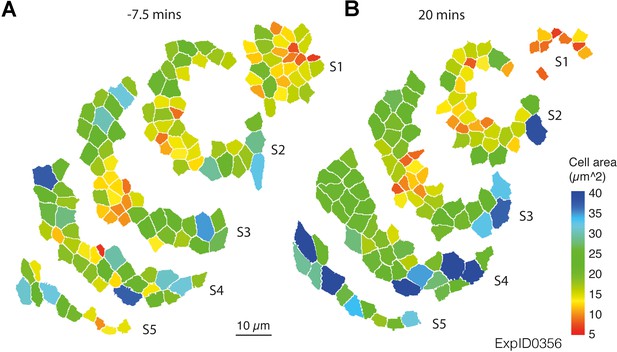
Morphogenetic events in the salivary gland placode show a radial organisation.
For an example movie with perfect tracking (ExpID0356) the change in apical area is shown for even-width radial stripes (defined at t = −7.5 min, A). Stripes S1-S5 correspond to the radial stripes 1–5 in Figure 1G,G’. (A) Start of movie at t = −7.5 min; (B) end of movie at t = 20.0 min. Apical area is colour-coded according to the legend.
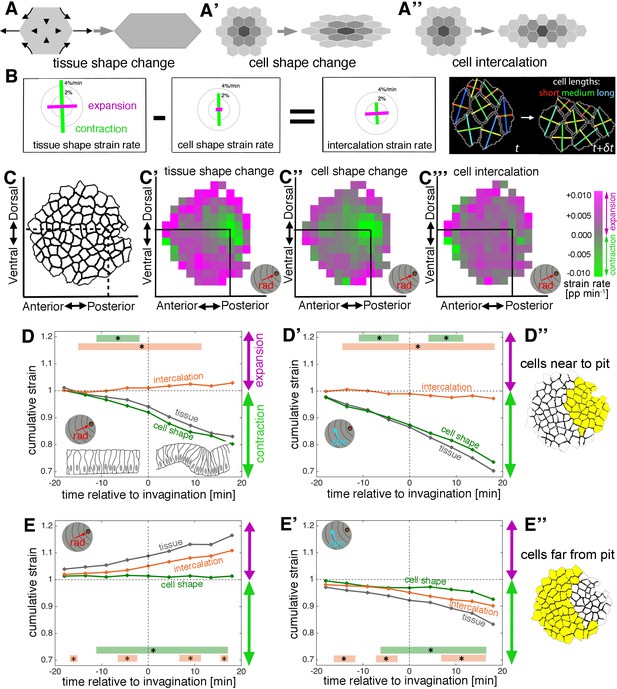
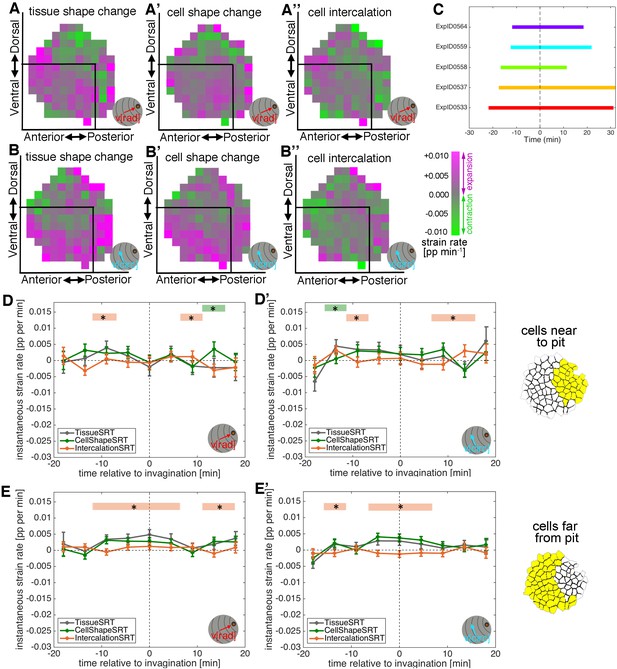
Apical strain rate analysis of early events during salivary gland placode invagination.
(A–A’’) During early salivary gland placode development the change in tissue shape (A) can be accounted for by cell shape changes (A’), cell intercalation (A’’) or any combination of the two. During this phase, cell division and gain or loss of cells from the epithelium (other than cells invaginating into the pit) do not occur. (B) For small domains of a focal cell and its immediate neighbours, tissue shape change is the sum of two additive contributions, cell shape change and cell intercalation (Blanchard et al., 2009). Both tissue shape change and cell shape change can be measured directly from the segmented and tracked time-lapse movies, so the amount of cell intercalation can be deduced. Strain rates are depicted as orthogonal lines, with contraction in green and expansion in magenta, and the length of the line proportional to the rate of strain (grey circles mark 2 and 4% change per minute). The last panel shows the group of cells (central cell and first corona of neighbours) for which the example strain rates in (B) were calculated, with orthogonal lines indicating the lengths of the major and minor axes of the cells. The vertical contraction in the group of cells can be clearly seen as a combination of vertical cell shortening and continuous rearrangement/intercalation (evidenced by one topological change). (C–C’’’) Spatial maps summarising the strain rate analysis covering 18 min prior to 18 min post commencement of tissue bending. Mapped onto the shape of the placode (C) the strain rate contribution towards the pit (‘rad’) is shown, quantified from data from nine embryos (see Figure 2—figure supplement 1). (C’) Tissue contraction (green) dominates near the pit, with expansion (magenta) towards the periphery. The tissue contraction near the pit is mostly due to cell constriction near the pit (C’’), whereas the tissue expansion is contributed to by both cell expansion and cell intercalation far from the pit (magenta in C’’ and C’’’). Strain rates are given in proportion (pp) of change per minute. (D–E’’) Regional breakdown of time-resolved cumulative strain. In the area near the pit (D’’) tissue constriction dominates (grey curves in D and D’) and is due to isotropic constriction at the cell level (green curves in D and D’), whilst intercalation only plays a minor role in this region (orange curves in D, D’). Far from the pit (E’’), the tissue elongates towards the pit (E, grey curve), with a corresponding contraction circumferentially (E’, grey curve), and this is predominantly due to cell intercalation (orange curves in E and E’). Statistical significance based on a mixed-effects model (see Materials and methods) and a p<0.05 threshold (calculated for instantaneous strain rates [see Figure 2—figure supplement 1]), is indicated by shaded boxes at the top or bottom of each panel (tissue vs cell shape is green and tissue vs intercalation is orange). See also Video 3. Cumulative strains here and in all subsequent plots are calculated as the exponent of the cumulative instantaneous strain rates, which are shown in Figure supplements.
-
Figure 2—source data 1
Cumulative strain and statistics for cells near to the pit and far from the pit.
Related to Figure 2D–E.
- https://doi.org/10.7554/eLife.35717.012
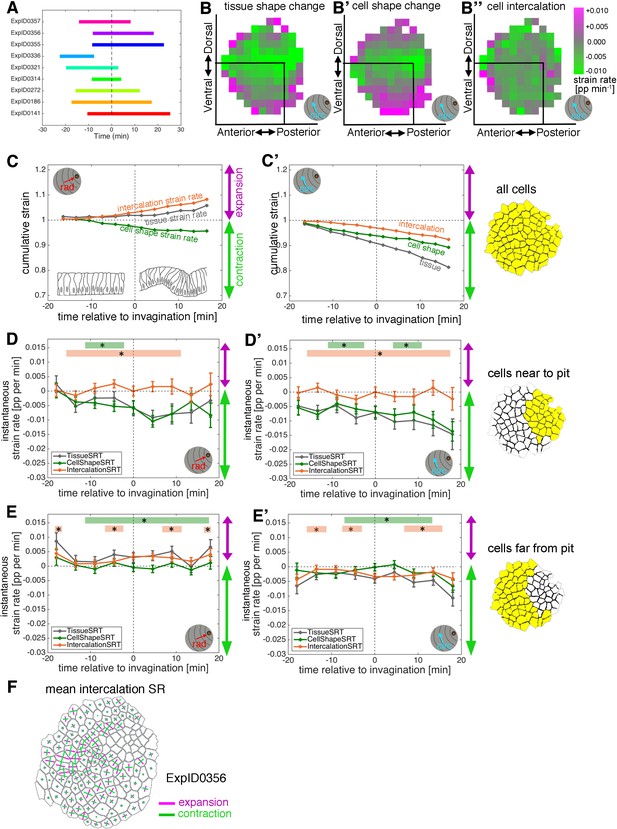
Apical strain rate analysis of early events during salivary gland placode invagination.
(A) Schematic of time windows covered by the time-lapse movies analysed for the apical strain rate calculations. (B–B’’) Spatial maps summarising the strain rates projected onto the circumferential orientation, covering 18 min prior to 18 min post-commencement of tissue bending. (C–C’) Cumulative strain of all pooled placodal cells. Overall, the tissue elongates a little radially towards the pit whilst contracting circumferentially (grey curves). Cell shapes constrict nearly isotropically (green curves), with intercalation extending the tissue towards the pit and contracting it circumferentially (orange curves). (D–E’) Instantaneous strain rates corresponding to the cumulative strain plots in Figure 2D–E’’. Error bars show an indicative confidence interval of the mean, calculated as the mean of within-embryo variances. Statistical significance based on a mixed-effects models and a p<0.05 threshold is indicated by shaded boxes at the top of each panel (see Materials and methods for details of statistics): tissue vs cell shape (green) and tissue vs intercalation (orange). The same conventions for displaying confidence intervals and statistical significance are used in all subsequent instantaneous strain rate plots. (F) Example placode showing the radial organisation of intercalation strain rates. Intercalation strain rates calculated between the first (−7.5 min) and last (+20 min) frames of placode movie ExplD0356 are shown, drawn on a segmented image of the central time point of the time-lapse movie. Strain rates are calculated for small domains of a focal cell (on which the strain rate is drawn) and its immediate neighbours and visualised here projected onto the central cell. Magenta indicates expansion and green contraction, the length of the line is proportional to the rate of strain in a given direction. The overall circumferential tissue convergence (green) and radial expansion (magenta) due to intercalation is clearly visible. Strain rates were not calculated for cells that disappeared into the pit before the end of the movie.
-
Figure 2—figure supplement 1—source data 1
Cumulative strain for all cells and instantaneous strain rates for cells near to the pit and far from the pit.
Related to Figure 2—figure supplement 1C,D and E.
- https://doi.org/10.7554/eLife.35717.011
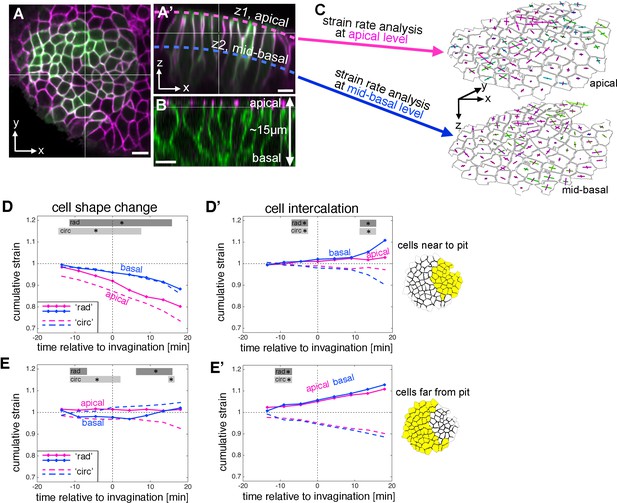
Strain rate analysis of early tubulogenesis using a 3D proxy.
(A–B) Epithelial cells of the placode are about 2–5 µm in apical diameter but extend about 15 µm into the embryo along their apical-basal axis. To assess behaviour of the tissue and cells at depth, we analysed a mid-basal level, about 8 µm basal to the apical surface. Scale bars are 5 µm. (C) Illustration of the method used: we segmented and tracked cells at a depth of ~8 µm, and repeated the strain rate analysis at this mid-basal level (blue) to compare with apical (pink). (D–D’) Comparison of apical (pink, as shown in Figure 2) and basal (blue) cumulative strains, for cell shape changes (D, E) and cell intercalation (D’, E’), near the pit (D,D’) and far from the pit (E,E’). Note how cell shape changes apical versus basal near the pit suggest progressive cell wedging as apices contract isotropically more rapidly than basally (D), and how the rate of cell intercalation across the tissue is highly coordinated between apical and basal, especially in the cells far from the pit (D’,E’). Statistical significance based on a mixed-effects model and a p<0.05 threshold (calculated for instantaneous strain rates [see Figure 2—figure supplement 1 and Figure 3—figure supplement 2]), is indicated by shaded boxes at the top of each panel: apical ‘rad’ vs basal ‘rad’ (dark grey) and apical ‘circ’ vs basal ‘circ’ (light grey).
-
Figure 3—source data 1
Cumulative strain and statistics for cells near to the pit and far from the pit at apical and mid-basal depth.
Related to Figure 3D–E.
- https://doi.org/10.7554/eLife.35717.019
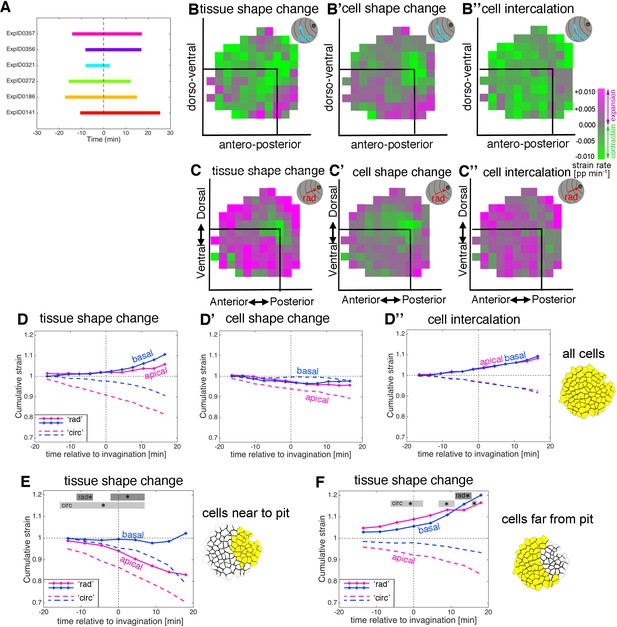
Strain rate analysis of early tubulogenesis using a 3D proxy.
(A) Schematic of time windows covered by the time-lapse movies analysed for the basal strain rate calculations. (B–C’’) Spatial summary of the basal strain rates (calculated from −15 to + 18 min), showing circumferential contribution (B–B’’, circ) and radial contribution, (C–C’’, ‘rad’). Constriction (green) still dominates the pit region at the tissue level (B, C), due to cell constriction (B’, C’), with strong expansion (magenta) towards the pit radially, predominantly due to cell intercalation (C’’). (D–D’) Comparison of the average basal and apical cumulative strain of all pooled placodal cells. Overall, basally (blue curves) the tissue elongates towards the pit whilst contracting circumferentially, in a more isotropic fashion than apically (pink curves). Basally, cell shape change is weakly contractile radially, like apical, while it is near zero circumferentially, unlike apical (D’), with intercalation extending the tissue towards the pit and contracting it circumferentially almost identically in apical and basal (D’’). (E, F). Comparison of apical (pink, as shown in Figure 2) and basal (blue) cumulative strain, for tissue shape change (E, F). Statistical significance based on a mixed-effects model and a p<0.05 threshold (calculated for instantaneous strain rates [Figure 2—figure supplement 1 and Figure 3—figure supplement 2]), is indicated by shaded boxes at the top of each panel: apical ‘rad’ vs basal ‘rad’ (dark grey) and apical ‘circ’ vs basal ‘circ’ (light grey).
-
Figure 3—figure supplement 1—source data 1
Cumulative strain for all cells at apical and mid-basal depth and tissue strain rate for cells near to the pit and far from the pit from an apical and basal depth.
Related to Figure 3—figure supplement 1D,E and F.
- https://doi.org/10.7554/eLife.35717.016
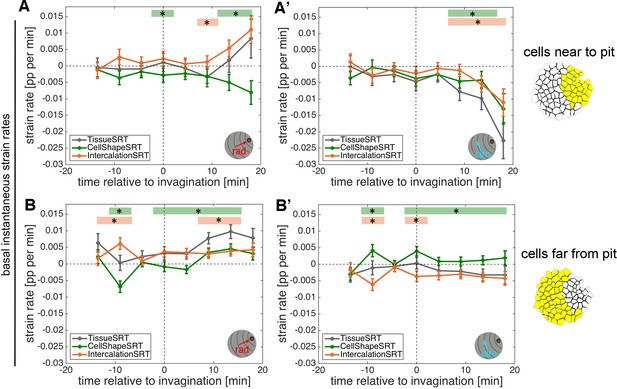
Strain rate analysis of early tubulogenesis using a 3D proxy.
(A–B’) Instantaneous mid-basal strain rates corresponding to the cumulative strain plots in Figure 3D–E’ and Figure 3—figure supplement 1E,F. Error bars depict the mean of within-embryo variance of six movies. Significance using a mixed-effects model of p<0.05 is indicated by shaded boxes at the top of each panel: tissue vs cell shape (green) and tissue vs intercalation (orange). See Materials and methods for details of statistics.
-
Figure 3—figure supplement 2—source data 1
Instantaneous strain rate for mid-basal depth.
Related to Figure 3—figure supplement 2A–D.
- https://doi.org/10.7554/eLife.35717.018
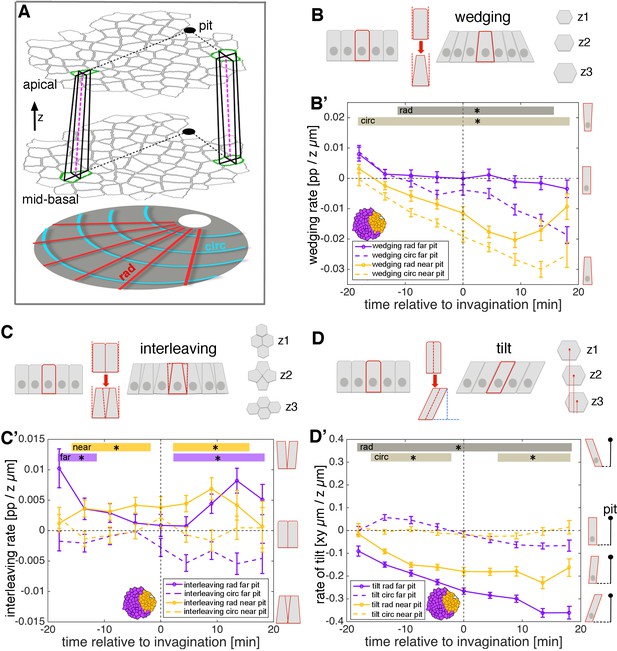
3D cell geometries during early tube budding.
(A) Illustration of the method used for calculating 3D domain geometries using tracked cells at apical and mid-basal levels throughout the salivary placode. Although for simplicity only two matched cells are shown here (green outlines in both z-levels), all cells were matched accurately between levels (see text and Video 4). Small domains of a focal cell and its immediate neighbours were used to calculate local rates of wedging, interleaving and tilt (from cell ‘in-lines’, pink dashed lines) using z-strain rate methods (see text and Figure 4—figure supplement 1A–D). Measures were projected onto the pit-centred radial (dotted black lines) and circumferential axes. In this tilted side-view cartoon, the z distance between apical and mid-basal levels has been exaggerated. (B) Schematic of cell wedging in a cross-section and as individual z-levels (z1–z3) in the marked red cell. (B’) Cell wedging is patterned across the placode, increasing most strongly in cells near to the pit, in accordance with the isotropic apical constriction observed here (orange lines). Away from the pit, wedging contributes mainly in the circumferential direction (purple dashed line). (C) Schematic of cell interleaving in a cross-section and as individual z-levels (z1–z3) in the group of marked red cells. (C’) Radial cell interleaving (solid line) is always more positive than circumferential interleaving (dashed lines), often significantly so. Interleaving contributes to a radial expansion apically and a concomitant circumferential contraction. (D) Schematic of cell tilt in a cross-section and as individual z-levels (z1–z3) in the marked red cell. (D’) Cell tilt increased continuously in the radial direction (solid lines) apically towards the pit over the period of our analysis. A stronger rate of tilt was observed for the cells further from the pit (purple solid line), which is expected from the radial wedging seen near the pit. (B’, C’, D’) Error bars show the mean of the within-embryo variances for five movies. Significance at p<0.05 using a mixed-effect model (see Materials and methods) is depicted as shaded boxes at the top of the panel: B’) wedging in radial orientation, near to vs far from pit (dark grey, ‘rad’) and wedging in circumferential orientation, near to vs far from pit (light grey, ‘circ). (C’) Interleaving near to the pit, radial vs circumferential (orange, ‘near’) and interleaving far from the pit, radial vs circumferential (purple, ‘far’). (D’) Tilt in radial orientation, near to vs far from pit (dark grey, ‘rad’) and tilt in circumferential orientation, near to vs far from pit (light grey, ‘circ).
-
Figure 4—source data 1
Wedging, interleaving and tilt mean ± confidence intervall (CI) and statistics.
Related to Figure 4B’, C’ and D’.
- https://doi.org/10.7554/eLife.35717.023
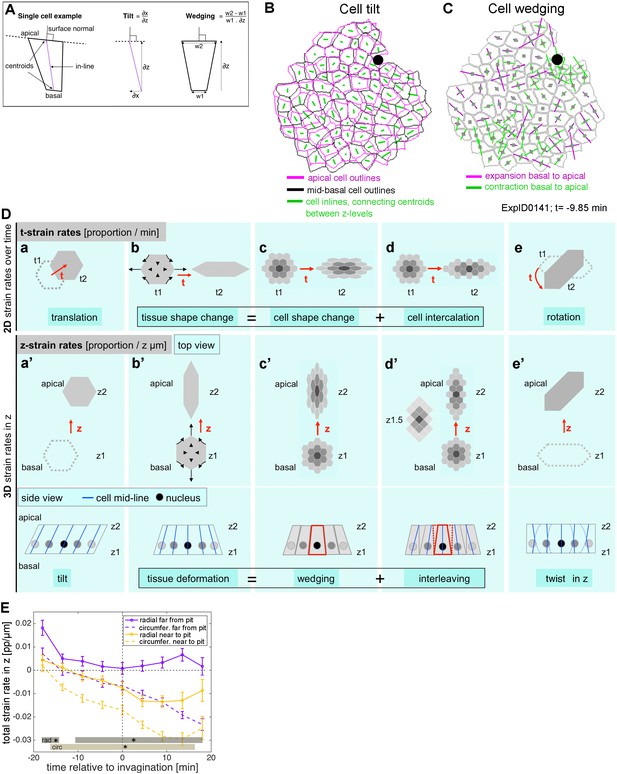
3D cell geometries during early tube budding.
(A) Schematic illustrating wedging and tilt definitions in the context of a single cell. (B) Illustration of cell arrangement in an example movie (ExpID0141) at time point t = −9.85 min showing segmented apical cell outlines in magenta and segmented basal cell outlines in black. Green lines are cell inlines, connecting centroid positions between z-levels, thereby giving an indication of the cell tilt, whereas mis-match between apical and mid-basal outlines give an indication of wedging interleaving. (C) Illustration of cell wedging in an example movie (ExpID0141) at t = −9.85 min, with amounts of individual cell wedging shown for each cell (mapped onto segmented apical cell outlines). An expansion of an individual cell from basal to apical is shown in magenta, and a contraction of an individual cell from basal to apical is shown in green. The extent of change is proportional to the length of the lines shown. (D) For domains of small numbers of cells, z-strain rates are calculated analogously to t-strain rates. Overall tissue deformation in depth (b’) is addition of wedging (c’) and interleaving (d’), just as tissue shape change (b) over time is the addition of cell shape change (c) and cell intercalation (d). Translation in 2D (a) equates to cell tilt in z, and cell rotation in 2D (e) is represented by tissue twist in z (e’). Tissue and cell schematics are shown in top views in (a–e) and in both top and side views in (a’–e’). (E) Total strain rate in z, representing the sum of the wedging and interleaving strain rates shown in Figure 4B’, C’. Confidence intervals and representation of statistics are as in Figure 4.
-
Figure 4—figure supplement 1—source data 1
Total strain rate in Z and statistics.
Related to Figure 4—figure supplement 1E.
- https://doi.org/10.7554/eLife.35717.022
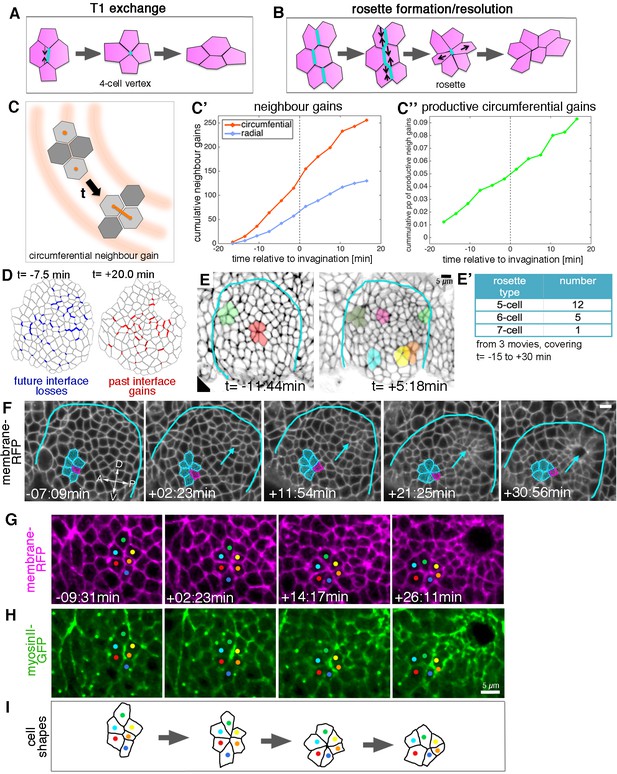
Cell intercalation during tube formation combines T1 exchanges and rosette-formation/resolution.
(A,B) Depending on the number of cells involved, intercalation can proceed through formation and resolution of a four-cell vertex, a so-called T1-exchange (A), or through formation and resolution of a rosette-like structure than can involve 5–10 or more cells (B). (C–C’’) Quantification of neighbour gains as a measure of T1 and intercalation events, with an example of a circumferential neighbour gain (leading to radial tissue expansion) shown in (C). (C’) Circumferential neighbour gains dominate over radial neighbour gains. (C’’) Rate of productive gains, defined as the amount of circumferential neighbour gains leading to radial tissue elongation and expressed as a proportion (pp) of cell-cell interfaces tracked at each time point. Data are pooled from seven embryo movies. (D) Represented for a single exemplary movie analysed (ExpID0356), interfaces that will be lost (blue) or have been gained (red) are shown, mapped onto a segmented version of the placode at the start of the movie for future losses (t = −7.5 min) and at the end of the movie for past gains (t = +20.0 min). Note that lost interfaces are mostly oriented circumferentially, whereas gained interfaces are mostly radial (see alternative visualisation in Figure 5—figure supplement 1). (E) Stills of two time-lapse movies used for the strain rate analysis, illustrating the appearance of rosette structures prior to (t= −11:44 min) and after (t = +5:18 min) tissue bending. (E’) The majority of rosettes observed in the salivary gland placode consist of five to six cells. Data are pooled from three embryo movies. (F–H) Stills of a time-lapse movie of an example of rosette formation-resolution. (F) The cluster of cells contracts along the circumferential direction of the placode, and resolution from the six-cell-vertex is oriented towards the pit (arrow). Label is Ubi-RFP-CAAX. (G,H) The same rosette as in (F) in close-up, showing Ubi-RFP-CAAX to label membranes in magenta (G) and myosin II-GFP (sqhGFP) in green (H). Note the prominent but transient myosin accumulation at the centre of the rosette-forming group of cells. (I) is a schematic of the rosette formation-resolution analysed in (F–H). See also Video 5. Scale bars in (E, F, H) are 5 µm.
-
Figure 5—source data 1
Cumulative neighbour gains and cumulative proportions of circumferential productive neighbour gains.
Related to Figure 5C.
- https://doi.org/10.7554/eLife.35717.027
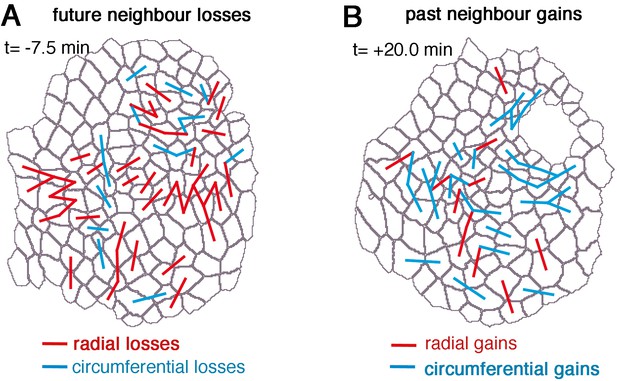
Cell intercalation during tube formation combines T1 exchanges and rosette-formation/resolution.
For an exemplary movie (ExpID0356) neighbour losses and gains are shown. (A) Future neighbour losses are shown on the segmented cell outlines of the initial time point (t = −7.5 min). (B) Past neighbour gains are shown on the segmented cell outlines of the last time point (t = +20.0 min). Red lines connecting centroids indicate radial losses or gains of the shared interface, blue lines connecting centroids indicate circumferential losses or gains of the shared interface.
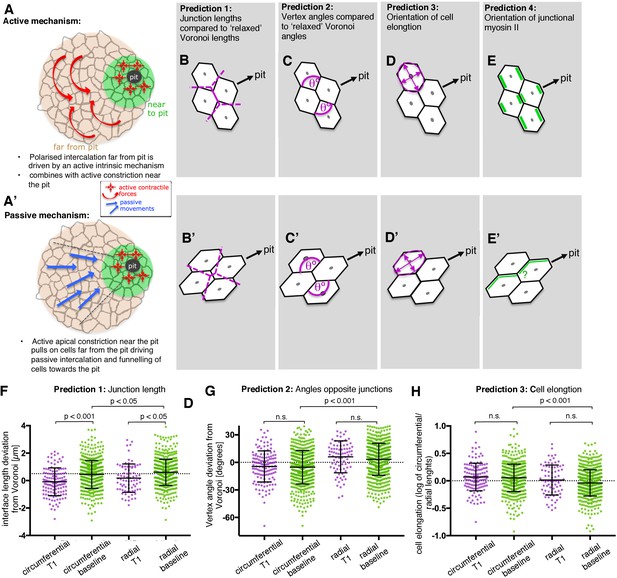
Analysis of signatures of active versus passive cell intercalation.
(A, A’). Cell intercalation in cells far from the pit could either be an active process, helping to draw cells together circumferentially and to extend tissue towards the pit (A), or the active pulling from the contractile pit could lead to a passive funnelling of cells towards the pit with associated passive intercalations (A’). We considered four measures should allow to distinguish between the above active and passive mechanisms. (B, B’) The lengths of actively shrinking circumferential junctions we predicted would be shorter than equivalent junction lengths of a Voronoi tessellation seeded with real cell centroid locations. We used the Voronoi tessellation as a proxy for ‘relaxed’ (not actively or passively altered) junction lengths and angles. Junctions pulled radially by the pit would be longer than predicted by Voronoi tessellation. (C,C’) Angles at vertices opposite actively shrinking circumferential junctions we predicted to be more acute than angles derived from a relaxed Voronoi tessellation, with the opposite predicted for pulled radial junctions. (D, D’) Active intercalation should lead to circumferential elongation of cells connected at vertices to shrinking junctions, whereas pulling from the pit would lead to radial cell elongation. (E, E’) Junctional actomyosin is predicted to be enriched in circumferential junctions that are actively shrinking. A response to pulling from the pit could lead to no enrichment or radially enriched junctional myosin (see text). (F) The junction lengths of both circumferentially and radially oriented junctions undergoing a T1 exchange showed an active signature (shorter than expected) compared to similarly oriented non-intercalating junctions. N = 135, 523, 78 and 618 junctions for the four distributions, respectively, from seven embryos. (G) Circumferentially oriented junctions far from the pit had an active signature of their vertex angles being more acute compared to radially oriented junctions, though junctions undergoing a T1 did not have more acute angles than junctions not directly involved in a T1. N = 139, 571, 84 and 675 junctions for the four distributions, respectively, from seven embryos. (F) Cells far from the pit had an active signature of being more elongated in the circumferential orientation. Cells at the ends of junctions involved in a T1 did not behave differently to junctions not involved in a T1. N = 135, 523, 78 and 618 junctions for the four distributions, respectively, from seven embryos. Error bars in (F–H) are ±SD.
-
Figure 6—source data 1
Predictions (junction length, angles opposite shrinking junctions and cell elongation) for circumferential T1, circumferential baseline, radial T1 and radial baseline and their statistics.
Related to Figure 6F–H.
- https://doi.org/10.7554/eLife.35717.033
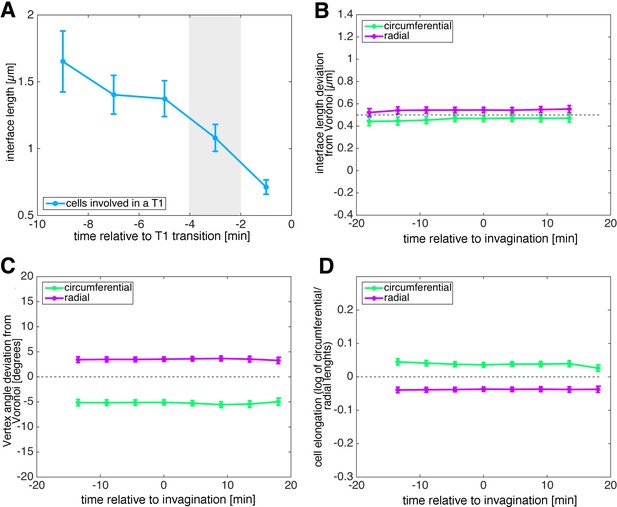
Analysis of signatures of active versus passive cell intercalation.
(A) Shortening of interfaces in the run up to a neighbour exchange event (t = 0 min). We used data in the window −4 to −2 min as T1 data to compare to non-intercalating baseline junctions. (B–D) Signatures of non-intercalating junctions (baseline) over developmental time. Non-intercalating junctions were defined as those that were more than 10 min away from being involved in a T1, or were never involved in a T1. All interface length deviation from Voronoi (B), vertex angle deviation from Voronoi (C) and cell elongation of cells opposite junctions (D) showed no temporal trend to their baseline (see Materials and methods).
-
Figure 6—figure supplement 1—source data 1
Interface length during a T1 transition and baselines for predictions indicated above.
Related to Figure 6—figure supplement 1.
- https://doi.org/10.7554/eLife.35717.032
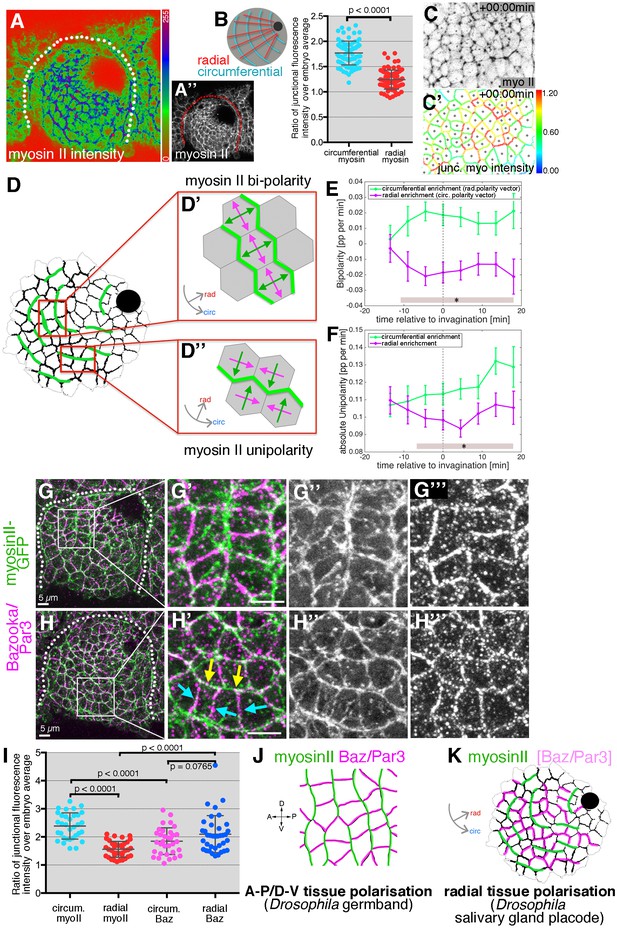
Junctional myosin II shows a strong circumferential polarisation during tube formation.
(A–B) Example of junctional myosin II intensity, visualised using sqh[AX3]; sqhGFP in fixed embryos; (A) shows a heat map of intensity. (B) Quantification of apical junctional myosin polarisation, depicted as the ratio of junctional intensity (circumferential or radial) over embryo average (5 placodes from 5 embryos, 83 circumferential junctions versus 83 radial junctions; significance calculated using unpaired t-test; embryo average is average apical myosin intensity in the epidermis outside the placode). (C, C’) Myosin II enrichment was quantified from segmented and tracked time-lapse movies. (C) depicts the sqhGFP signal from a single time point of a movie, (C’) shows the calculated junctional myosin II intensity. See also Videos 6 and 7. (D–F) Myosin enrichment at junctions can occur in two flavours: (D’, E) Myosin II bi-polarity is defined as myosin II enrichment at two parallel oriented junctions of a single cell, calculated as the magnitude of a vector pointing at the enrichment (D’). Circumferential myosin II bi-polar enrichment (i.e. the radial bi-polarity vector, green in (D’) pointing at myosin II enrichment), increases between −15 min and + 18 min (E), green curve). (D’’, F) Myosin II unipolarity is defined as myosin II enrichment selectively on side of a cell (D’’). Circumferential myosin II uni-polar enrichment (i.e. the radial uni-polarity vector, green, in (D’’) pointing at myosin II enrichment), increases between −15 min and +18 min (F), green curve), whereas radial uni-polar enrichment (i.e. the circumferential uni-polarity vector in magenta in D’’), does not increase (F), magenta curve). (E, F) Error bars represent the mean of within-embryo variances of four movies, and significance between radial and circumferential bi- and uni-polarity at p<0.05 using a mixed-effects model (see Materials and methods) is depicted as shaded boxes at the bottom of panels E and F. See also Video 7. (G–H’’’) Examples of myosin II and Bazooka/Par3 polarisation in fixed embryos, two different regions of two placodes are shown. Note the complementary localisation of myosin II (green) and Bazooka (magenta). Scale bars are 5 µm. (I) Quantification of myosin II and Bazooka/Par3 polarisation in areas of strong junctional myosin II enrichment at circumferential junctions as shown in (G’) and (H’). Both circumferential (yellow arrows in H’) and radial (turquoise arrows in H’) fluorescence intensity are shown as ratios over embryo average, both for myosin II and Bazooka in the same junctions (from five embryos; mean and SEM are shown; paired t-test for comparison in the same junctions, unpaired t-tests for comparison of circumferential myo vs radial myo and circumferential Baz vs radial Baz; N = 33 circumferential junctions and N = 38 radial junctions across five embryos; embryo average is average apical myosin II or bazooka intensity in the epidermis outside the placode). (J, K) In contrast to the well-documented A-P/D-V polarity of cells in the Drosophila germband during elongation in gastrulation (J), the salivary gland placode appears to show a radial tissue polarisation, with a radial-circumferential molecular pattern imprinted onto it that instructs the morphogenesis (K).
-
Figure 7—source data 1
Myosin and Bazooka fluoresce intensity in radial and circumferential junctions and Myosin unipolary and bipolarity over time.
Related to Figure 7B,E,F and I.
- https://doi.org/10.7554/eLife.35717.035
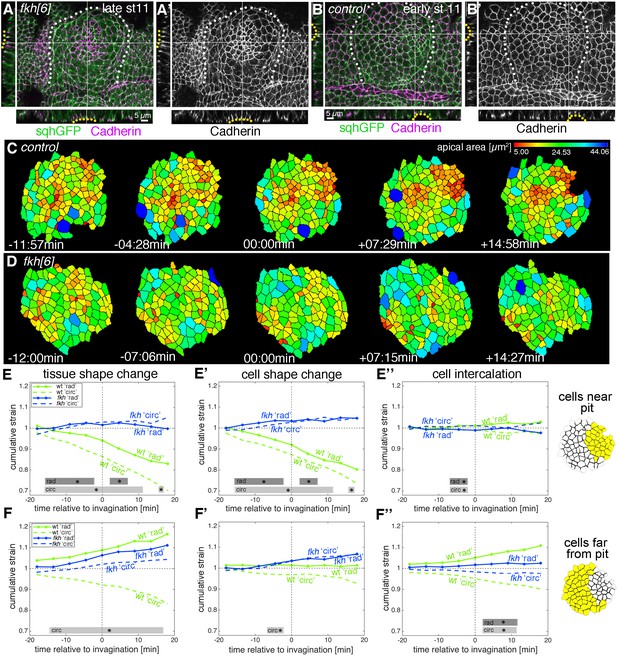
Loss of radial patterning of cell behaviours in salivary gland placodes lacking Fkh.
(A–B’) Examples of fkh mutant and wild-type placodes illustrating the lack of pit formation. A control placode at early stage 11 shows clear constriction and pit formation in the dorsal-posterior corner (yellow dotted line in cross-sections in B, B’), whereas even at late stage 11 a fkh mutant placode (beyond the time frame of our quantitative analysis) does not show a pit in the dorsal-posterior corner, instead a shallow central depression forms with some constricted central apices (yellow dotted line in cross-sections in A), (A’). Myosin II (sqhGFP) is in green and DE-Cadherin in magenta, white dotted lines in main panels outline the placode boundary. (C, D) Stills of segmented and tracked time lapse movies for control (C) and fkh[6] mutant (D) placodes, apical cell outlines are shown and colour-coded by apical cell area. Note the lack of contraction at the tissue and cell level in the fkh[6] mutant. (E–F’’) Analysis of cumulative apical strains in fkh mutant placodes (from five movies) in comparison to the analysis of wild-type embryos (as shown in Figure 2). Over the first 36 min of tube budding centred around the first appearance of tissue-bending in the wild-type and an equivalent time point in the fkh mutants, the fkh mutant placodes show only a slight expansion at the tissue level in the cells far from the predicted pit position (F) due to cell shape changes (F’) with very little intercalation contributing to the change (E’’, F’’). Statistical significance based on a mixed-effects model and a p<0.05 threshold (calculated for instantaneous strain rates [see Figure 2—figure supplement 1 and Figure 8—figure supplement 2]), is indicated by shaded boxes at the top of each panel: wt ‘rad’ vs fkh ‘rad’ (dark grey) and wt ‘circ’ vs fkh ‘circ’ (light grey).
-
Figure 8—source data 1
Cumulative strain and statistics for cells near to the pit and far from the pit for wild type and fkh[6] mutant.
Related to Figure 8E and F.
- https://doi.org/10.7554/eLife.35717.041
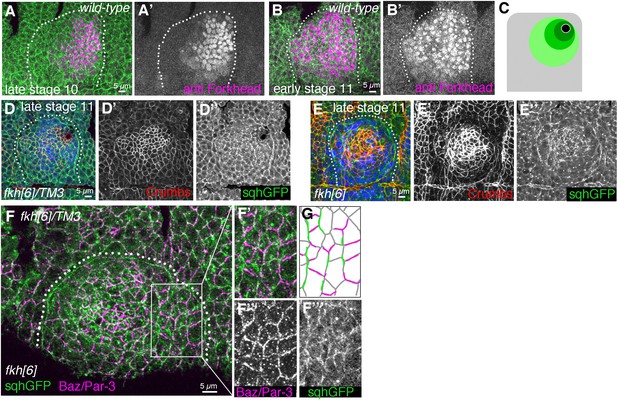
Loss of radial patterning of cell behaviours in salivary gland placodes lacking Fkh.
(A–C) Fkh expression (magenta in A, B) initiates around the future point of invagination in the dorsal-posterior corner at late stage 10 (A’) and then spreads radially across the secretory part of the placode in early stage 11 (B’). (C) Schematic of radially expanding expression pattern of Fkh. (D–E’’) Comparison of Crumbs (red) and myosin II (sqhGFP; green) localisation in control (D–D’’) and fkh mutant (E–E’’) placodes. In the control, Crumbs is strongest near the invagination point and levels drop with radial distance from the pit, whereas in the fkh[6] mutant beyond late stage 11 Crumbs is stronger in the central shallow depression, with less towards the periphery. Areas of remaining circumferential myosin II polarisation are visible in the fkh[6] mutant (E’’). (F–G) Example fkh[6] mutant placode showing remaining myosin II (green) and Bazooka (magenta) polarisation, (G) shows a schematic representation of the area magnified in (F’–F’’’). Dotted lines in panels indicate the placode boundary. Scale bars in (A, B, D, E and F) are 5 µm.
Loss of radial patterning of cell behaviours in salivary gland placodes lacking Fkh.
(A–B’’) Spatial summary of average apical strain rates in fkh[6] mutants covering 15 min prior to 18 min post-commencement of tissue bending, showing the radial (A–A’’) as well as circumferential (B–B’’) contributions, expressed as proportion (pp) change per minute. Expansion is magenta, contraction is green. (C) Schematic of time windows covered by the time-lapse movies analysed for the apical strain rate calculations in fkh[6] mutant embryos. (D–E’) Instantaneous apical strain rates in fkh[6] mutants corresponding to the cumulative plots in Figure 8E–F’’. Error bars depict the mean of within-embryo variances of five movies. Significance using a mixed-effects model of p<0.05 is indicated by shaded boxes at the top of each panel: tissue vs cell shape (green) and tissue vs intercalation (orange), see Materials and methods for details of statistics.
-
Figure 8—figure supplement 2—source data 1
Instantaneous strain rates for fkh[6] mutant.
Related to Figure 8—figure supplement 2D–E.
- https://doi.org/10.7554/eLife.35717.040
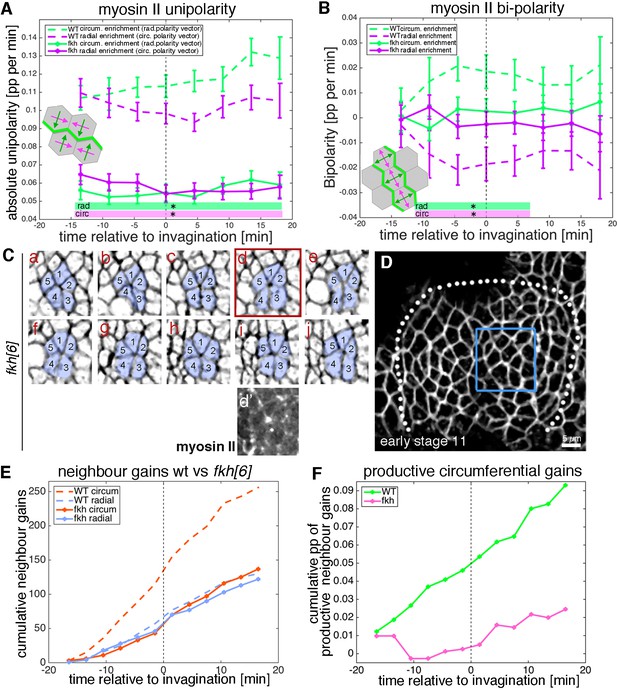
Analysis of myosin patterns and intercalation behaviour in fkh mutants.
(A,B) Analysis of myosin II unipolarity (A) and bi-polarity (B) in fkh[6] mutant placodes compared to wild-type (as shown in Figure 7, see also Videos 7 and 8). Overall myosin II levels in fkh[6] mutant placodes are lower (not shown) and both unipolarity and bi-polarity are decreased, with bi-polarity near zero (B, solid curves). Error bars show intra-embryo variation of five embryo movies for fkh[6] and four embryo movies for wt. Statistical significance at p<0.05 using a mixed-effect model is indicated as shaded boxed at the bottom of the panels: wt vs fkh[6] for the radial vector/circumferential enrichment (green, ‘rad’) and wt vs fkh for the circumferential vector/radial enrichment (purple, ‘circ’). (C–F) Neighbour exchanges still occur in fkh[6] mutant placodes. (C) Sill pictures from a tracked and segmented movie of a fkh[6] mutant placode (labelled with membrane-RFP), frames (labelled a–j) are 1:55 min apart; the full placode view in (D) is from time point (a, d’) shows the myosin accumulation at the centre of the rosette structure observed in (d). (E) In comparison to wt where circumferential neighbour gains dominate over radial ones (dashed lines), in the fkh[6] mutant placodes both occur with equal frequency (solid lines). This leads to nearly no productive circumferential neighbour gains in the mutant, significantly fewer than in the wt (F); Kolmogorov-Smirnov two sample test D = 0.5833, p=0.0191).
-
Figure 9—source data 1
Unipolarity and bipolarity and dynamics of neighbour gains in fkh[6] mutant.
Related to Figure 9A–B and E–F.
- https://doi.org/10.7554/eLife.35717.044
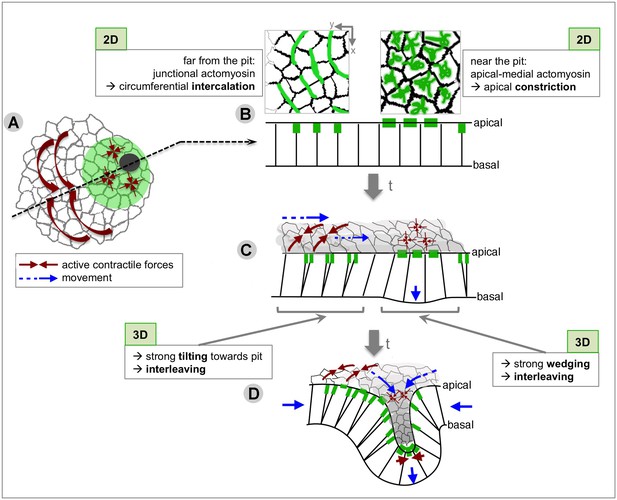
Summary of the radial patterning of 2D and 3D cell behaviours and actomyosin pools across the salivary gland placode during early tube formation.
(A) Circumferential tissue convergence through intercalation and apical constriction at the pit combine to result in radial tissue expansion towards the invagination point. (B) These 2D behaviours are associated with different actomyosin pools: far from the pit, circumferential junctional actomyosin underlies active intercalation through junction shrinkage, and near to the pit a pulsatile apical-medial actomyosin underlies apical constriction (Booth et al., 2014). (C) Quasi-3D analyses revealed that active apical intercalation (thick brown arrows) and isotropic constriction (thin brown arrows) lead in 3D to strong wedging of cells near the pit, strong tilting of cells far from the pit always towards the pit, as well as interleaving (i.e. change of neighbour connectivity along the apical-basal axis) across the tissue, aiding circumferential convergence and radial extension (cell and tissue movement indicated by blue arrows). (D) Once tissue bending has commenced at the pit, active apical constriction in and around the pit (thick and thin brown arrows) cooperates with active circumferential intercalations (medium brown arrows) to feed the elongation of the pit tube (blue arrows).
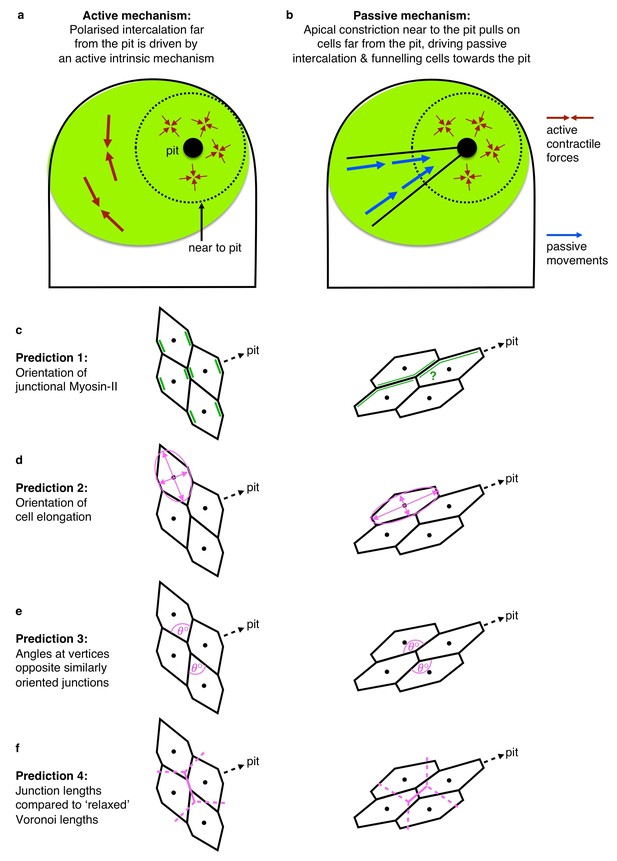
Active versus Passive mechanisms driving intercalation.
https://doi.org/10.7554/eLife.35717.049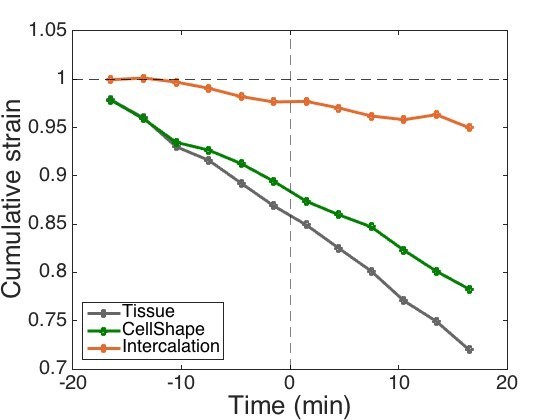
New cumulative strain graph to be used.
https://doi.org/10.7554/eLife.35717.050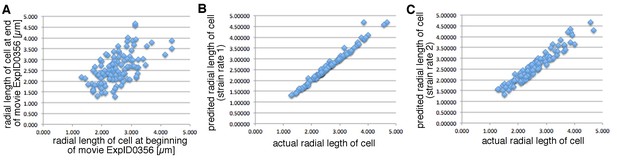
Videos
Example movie of early salivary gland placode morphogenesis in 3D.
Embryo of the genotype Scribble-GFP/fkhGal4::UAS-palmYFP as shown if Figure 1C and Figure 3A. Time stamp indicates time before and after initiation of tissue bending at t = 0. Scale bar 20 µm.
Example movie showing evolution of apical cell identities and apical area.
Movie ExpID0356 with 100% tracking, is shown. Cell identities are randomly colour-coded on the left, apical area on the right. Area colour codes are identical to Figure 8C, please refer to this scale. Time stamp indicates time before and after initiation of tissue bending at t = 0. Scale bar 20 µm.
Example movies of Tissue strain rate tensor (SRT), Cell Shape SRT and Intercalation SRT.
Local strain rates are extracted from segmented and tracked movies. Rates of tissue shape change (left), cell shape change (centre) and intercalation (right) are shown. Green vectors represent contraction and magenta vectors represent expansion. Time stamp indicates time before and after initiation of tissue bending at t = 0. See also Figure 2.
Example of matching cells through depth.
Apical cell identities matched to basal cell identities are shown within the placode. Stamp refers to the depth in the tissue. Scale bar 20 µm. See also Figure 4.
Example movie of apical rosette formation/resolution.
Embryo of the genotype sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX, only the UbiRFP label is shown. A group of cells going through rosette formation/resolution is highlighted, still of the movie are shown in Figure 5F. Time stamp indicates time before and after initiation of tissue bending at t = 0. Scale bar 20 µm.
Example movie of cell shape and myosin II localisation in a control embryo.
Embryo of the genotype sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX used for the myosin II uni- and bi-polarity quantifications as shown in Figure 7. Time stamp indicates time before and after initiation of tissue bending at t = 0. Scale bar 20 µm.
Example movie of automatic junctional myosin II quantification.
Myosin II uni-polarity vectors (left) and bi-polarity vectors (right) are shown. Junctions are colour coded according to their myosin II intensity levels as shown in Figure 7C’ (middle). Time stamp indicates time before and after initiation of tissue bending at t = 0. Scale bar 20 µm. See also Figure 7.
Example movie of cell shape and myosin II localisation in a fkh[6] mutant embryo.
Embryo of the genotype sqh::sqhGFP42, UbiRFP-CAAX; fkh[6] used for the myosin II uni- and bi-polarity quantifications as shown in Figure 9. Time stamp indicates time before and after initiation of tissue bending at t = 0. Scale bar 20 µm.
Tables
Number of cells in time-lapse experiments.
https://doi.org/10.7554/eLife.35717.006| Figure | Cells | Number of embryos | number of cells per time point at [−18, −13.5, −9, −4.5, 0, 4.5, 9, 13.5, 18] min |
|---|---|---|---|
| Figure 2 | Apical cells near to pit | 9 | [123, 599, 724, 1132, 980, 737, 468, 219, 80] |
| Figure 2 | Apical cells far from pit | 9 | [156, 837, 1183, 1880, 1731, 1484, 1151, 744, 319] |
| Figure 3 | Basal cells near to pit | 6 | [260, 453, 726, 569, 546, 506, 360, 89] |
| Figure 3 | Basal cells far from pit | 6 | [382, 762, 1309, 1200, 1145, 1133, 770, 247] |
| Figure 4 | 3D proxy cells near to pit | 5 | [36, 307, 420, 479, 336, 410 260 148 58] |
| Figure 4 | 3D proxy cells far from pit | 5 | [37, 350, 515, 652, 479, 564, 394, 243, 131] |
| Figure 8 | Apical fkh6 cells near to pit | 5 | [27, 161, 248, 269, 215, 238, 233, 186, 85] |
| Figure 8 | Apical fkh6 cells far from pit | 5 | [52, 343, 626, 735, 615, 706, 655, 550, 239] |
| Figure | Cells | Number of embryos | number of cells per time point at [−13.5, −9, −4.5, 0, 4.5, 9, 13.5, 18] min |
| Figure 7 | Uni and Bi-polarity WT | 4 | [207, 522, 1115, 918, 1091, 1125, 1047, 484] |
| Figure 9 | Uni- and Bi-polarity fkh6 | 5 | [528, 783, 815, 712, 809, 786, 708, 307] |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (D. melanogaster) | Fork Head | NA | FLYB:FBgn0000659 | |
| Gene (D. melanogaster) | non-muscle myosin II/sqh | NA | FLYB:FBgn0003514 | |
| Gene (D. melanogaster) | Bazooka/Par3 | NA | FLYB:FBgn0000163 | |
| Genetic reagent (D. melanogaster) | Scribble-GFP | Kyoto Drosophila Genomic Research Centre | ||
| Genetic reagent (D. melanogaster) | fkh-Gal4 | PMID:10625560 | ||
| Genetic reagent (D. melanogaster) | Fkh-Gal4::UAS-palmYFP | PMID:10625560, PMID:21297621 and this study | stock generated upon recombination of Brainbow Cassette stock | |
| Genetic reagent (D. melanogaster) | sqh[AX3]; sqh::sqhGFP42 | PMID:14657248 | ||
| Genetic reagent (D. melanogaster) | sqh::sqhGFP42, UbiRFP-CAAX | Kyoto Drosophila Genomic Research Centre Number 109822 | ||
| Genetic reagent (D. melanogaster) | fkh[6] | Bloomington Drosophila Stock Center, PMID:2566386 | FLYB:FBal0004012 | |
| Antibody | anti-DE-Cadherin (rat monoclonal) | Developmental Studies Hybridoma Bank at the University of Iowa | DSHB:DCAD2 | (1:10) |
| Antibody | anti-Crumbs (mouse monoclonal) | Developmental Studies Hybridoma Bank at the University of Iowa | DSHB:Cq4 | (1:10) |
| Antibody | anti-Bazooka (rabbit polyclonal) | PMID:10591216 | (1:500) | |
| Antibody | anti-Forkhead (guinea pig plyclonal) | PMID:2566386 | (1:2000) | |
| Antibody | Alexa Fluor 488/549/649- coupled secondary antibodies | Molecular Probes | ||
| Antibody | Cy3-, Cy5- coupled secondary antibodies | Jackson Immuno Research | ||
| Software | otracks | PMID:19412170, PMID:24914560 | Software file (custom software written in IDL) | |
| Software | nd-safir | PMID:19900849 | Denoising algorithm. Available at http://serpico.rennes.inria.fr/doku.php?id=software:nd-safir:index |
Genotypes used for Figure Panels.
https://doi.org/10.7554/eLife.35717.046| Figure | Panel | Embryo genotypes | Number of embryos used for quantitative analysis | |
|---|---|---|---|---|
| Figure 1 | B, B' | Scribble-GFP | panel representative of genotype (>30 embryos inspected) | fixed |
| Figure 1 | C | fkhGal4,UASpalm-YFP/Scribble-GFP | panel representative of genotype (>30 embryos inspected) | live |
| Figure 1 | G,G’ | fkhGal4,UASpalm-YFP/Scribble-GFP and sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 1 and 8 (i.e. 9 in total) | live |
| Figure 2 | C,D,E | fkhGal4,UASpalm-YFP/Scribble-GFP and sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 1 and 8 (i.e. 9 in total) | live |
| Figure 3 | A,B | fkhGal4,UASpalm-YFP/Scribble-GFP | panels representative of genotype (>30 embryos inspected) | live |
| Figure 3 | D,E,F | sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 5 | live |
| Figure 4 | C,E,G | fkhGal4,UASpalm-YFP/Scribble-GFP and sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 1 and 4 (i.e. 5 in total) | live |
| Figure 5 | C',C'', C''' | fkhGal4,UASpalm-YFP/Scribble-GFP and sqh[AX3]; sqh::sqhGFP42, UbiRFP | 1 and 6 (i.e. 7 in total) | live |
| Figure 5 | E,E' | sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 3 | live |
| Figure 5 | F,G,H | sqh[AX3];sqh::sqhGFP42, UbiRFP-CAAX | panels representative of genotype (>30 embryos inspected) | live |
| Figure 6 | F,G,H | fkhGal4,UASpalm-YFP/Scribble-GFP and sqh[AX3]; sqh::sqhGFP42, UbiRFP | 1 and 6 (i.e. 7 in total) | live |
| Figure 7 | A | sqh[AX3]; sqh::sqhGFP42 | panel representative of genotype (>30 embryos inspected) | fixed |
| Figure 7 | B | sqh[AX3]; sqh::sqhGFP42 | 5 | fixed |
| Figure 7 | C | sqh[AX3]; sqh::sqhGFP42 UbiRFP-CAAX | panels representative of genotype (>30 embryos inspected) | live |
| Figure 7 | E,F | sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 4 | live |
| Figure 7 | G | sqh[AX3]; sqh::sqhGFP42 | panels representative of genotype (>30 embryos inspected) | fixed |
| Figure 7 | H | sqh[AX3]; sqh::sqhGFP42 | panels representative of genotype (>30 embryos inspected) | fixed |
| Figure 7 | I | sqh[AX3]; sqh::sqhGFP42 | 5 | fixed |
| Figure 8 | A | sqh::sqhGFP42, UbiRFP-CAAX; fkh[6] | panels representative of genotype (>30 embryos inspected) | fixed |
| Figure 8 | B | sqh::sqhGFP42, UbiRFP-CAAX; fkh[6]/TM3 | panels representative of genotype (>30 embryos inspected) | fixed |
| Figure 8 | C | sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | panels representative of genotype (>30 embryos inspected) | live |
| Figure 8 | D | sqh::sqhGFP42, UbiRFP-CAAX; fkh[6] | panels representative of genotype (>30 embryos inspected) | live |
| Figure 8 | E,F (as in Figure 2C,D,E) | fkhGal4,UASpalm-YFP/Scribble-GFP and sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 1 and 8 (i.e. 9 in total) | live |
| Figure 8 | E,F | sqh::sqhGFP42, UbiRFP; fkh[6] | 5 | live |
| Figure 9 | A,B | sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 4 | live |
| Figure 9 | A,B | sqh::sqhGFP42, UbiRFP-CAAX; fkh[6] | 5 | live |
| Figure 9 | C,D | sqh::sqhGFP42, UbiRFP-CAAX; fkh[6] | 3 | live |
| Figure 9 | E | sqh[AX3];sqh::sqhGFP42, UbiRFP-CAAX | 6 | live |
| Figure 9 | E | sqh::sqhGFP42, UbiRFP-CAAX; fkh[6] | 5 | live |
| Figure 9 | F | fkhGal4,UASpalm-YFP/Scribble-GFP and sqh[AX3]; sqh::sqhGFP42, UbiRFP | 1 and 6 (i.e. 7 in total) | live |
| Figure 9 | F | sqh::sqhGFP42, UbiRFP-CAAX; fkh[6] | 5 | live |
| Figure Supplement | Panel | Embryo genotype | Number of embryos used for quantitative analysis | |
| Figure 2—figure supplement 1 | B-E' | fkhGal4,UASpalm-YFP/Scribble-GFP and sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 1 and 8 (i.e. 9 in total) | live |
| Figure 2—figure supplement 1 | F | sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 1 | live |
| Figure 3—figure supplement 1 | B-F | sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 5 | live |
| Figure 3—figure supplement 2 | A-B’ | sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 5 | live |
| Figure 4—figure supplement 1 | B,C | sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 1 | live |
| Figure 4—figure supplement 1 | E | fkhGal4,UASpalm-YFP/Scribble-GFP and sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 1 and 4 (i.e. 5 in total) | live |
| Figure 5—figure supplement 1 | A,B | sqh[AX3]; sqh::sqhGFP42, UbiRFP-CAAX | 1 | live |
| Figure 6—figure supplement 1 | fkhGal4,UASpalm-YFP/Scribble-GFP and sqh[AX3]; sqh::sqhGFP42, UbiRFP | 1 and 6 (i.e. 7 in total) | live | |
| Figure 8—figure supplement 1 | A-B' | sqh::sqhGFP42 | panels representative of genotype (>30 embryos inspected) | fixed |
| Figure 8—figure supplement 1 | D-F’’’ | sqh::sqhGFP42, UbiRFP-CAAX; fkh[6] | panels representative of genotype (>30 embryos inspected) | fixed |
| Figure 8—figure supplement 2 | A-E' | sqh::sqhGFP42, UbiRFP-CAAX fkh[6] | 5 | live |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.35717.047





