The shape of the bacterial ribosome exit tunnel affects cotranslational protein folding
Figures
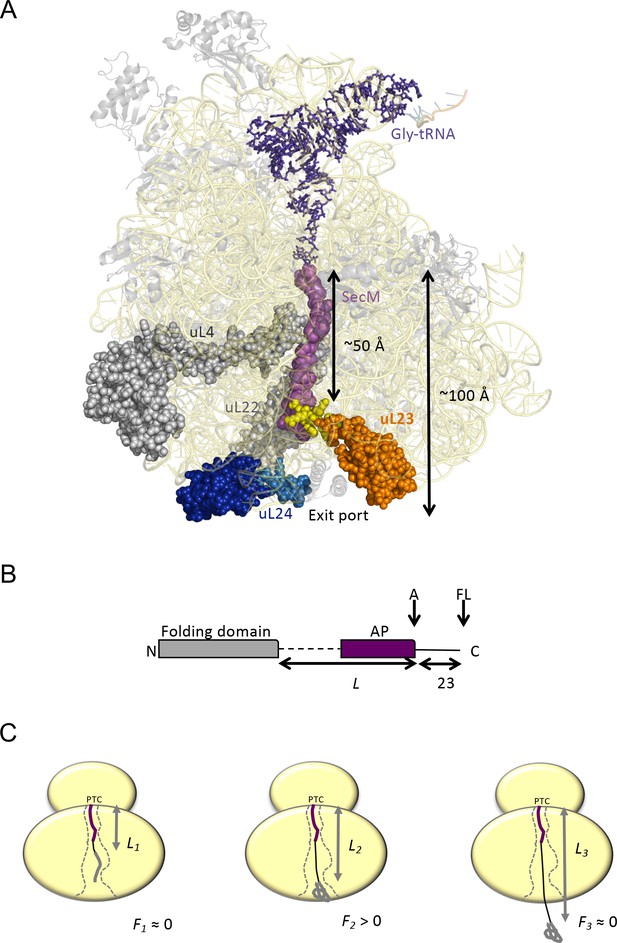
Cotranslational protein folding assay.
(A) Front view of the 50S subunit of the E.coli ribosome adapted from PDB 3JBU (Zhang et al., 2015), with tunnel proteins uL4 and uL22 indicated in gray. The globular domain of uL23 is indicated in orange with the β-hairpin loop depicted in yellow. uL24 is shown in dark blue, with the loop at the tunnel exit shown in light blue. The exit tunnel, outlined by a stalled SecM nascent chain (purple), is ~100 Å in length. (B) The arrest-peptide assay (Nilsson et al., 2015). The domain to be studied is placed L residues upstream of the critical proline at the C-terminal end of the 17-residue long arrest peptide (AP) from the E. coli SecM protein. A 23-residue long stretch of the E. coli LepB protein is attached downstream of the AP, allowing us to separate the arrested (A) and full-length (FL) products by SDS-PAGE after translation. Constructs are translated in the PURExpress in vitro translation system supplemented with WT, uL23 Δloop, or uL24 Δloop high-salt washed ribosomes for 20 min. The relative amounts of arrested and full-length protein were estimated by quantification of SDS-PAGE gels, and the fraction of full-length protein was calculated as fFL = IFL/(IA +IFL) where IA and IFL are the intensities of the bands corresponding to the A and FL products. (c) fFL is a proxy for the force F that cotranslational folding of a protein domain exerts on the AP. At short linker lengths, both F and fFL ≈ 0 because the domain is unable to fold due to lack of space in the exit tunnel. At intermediate linker lengths, F and fFL > 0 because the domain pulls on the nascent chain as it folds. At longer linker lengths, F and fFL ≈ 0 because the domain is already folded when the ribosome reaches the end of the AP.
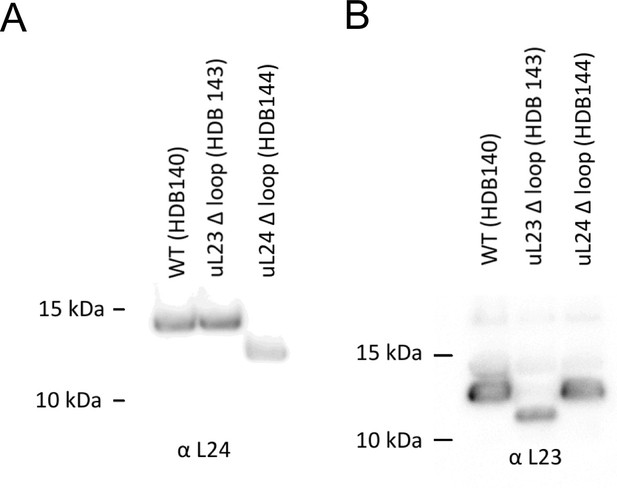
A260 = 300 units (6.9 μM) of high-salt washed ribosomes were separated on a 12% Bis-Tris gel and transferred by Western blotting onto a nitrocellulose membrane and detected with antibodies against uL24 (panel A) or uL23 (panel B).
https://doi.org/10.7554/eLife.36326.003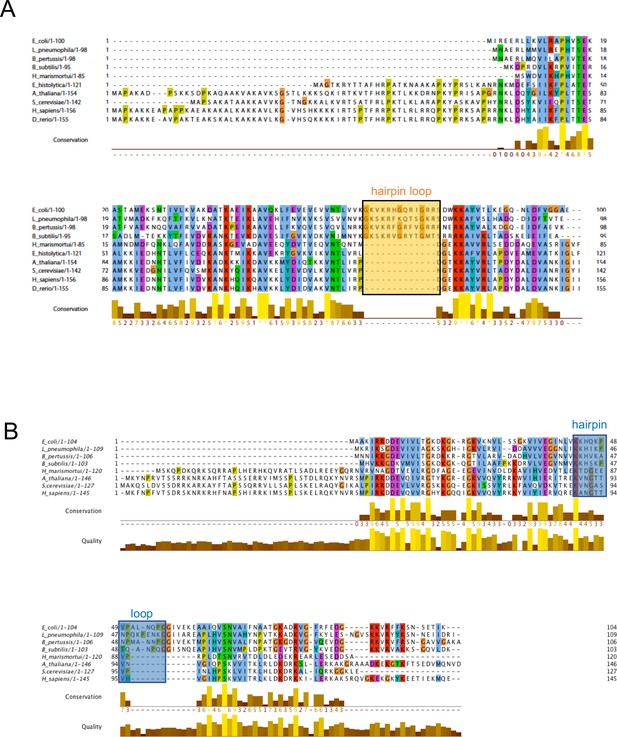
Multiple sequence alignment of uL23 and uL24.
(A) and uL24 (B). The uL23 and uL24 b-hairpin loops, boxed, respectively, in orange and blue, are conserved among Gram-negative bacteria, but are short or absent in archaea and eukaryotes. In eukaryotes and archaea, part of the function of uL23 is proposed to be fulfilled by ribosomal protein eL39.
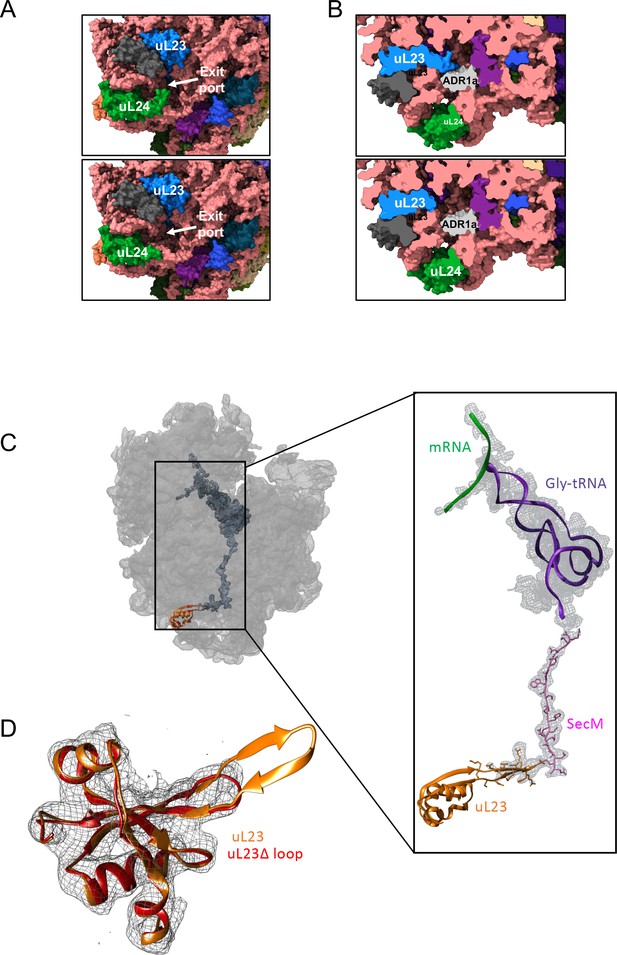
Structural consequences of removing the hairpin loops in uL24 and uL23 modeled after PDB 3JBU of the SecM stalled ribosome.
(A) In wildtype ribosomes, the loop in uL24 partially obstructs the exit tunnel (top panel). Its removal in uL24 Δloop ribosomes creates a wide opening into the tunnel (bottom panel). (B) In wildtype ribosomes, the loop in uL23 extends into the exit tunnel (top panel). Its removal in uL23 Δloop ribosomes creates an open space around the area where the ADR1a domain is known to fold (Nilsson et al., 2015). The ADR1a structure is from PDB 5A7U. (C) Cryo-EM structure of the uL23 Δloop 70 S ribosome (EMD-4319), fitted to PDB 3JBU (that includes a Gly-tRNA and a 26-residue long arrested SecM AP) to locate uL23 (orange) and the exit tunnel. The enlarged region shows a difference map (in mesh) obtained by subtracting the cryo-EM map of the uL23 Δloop 70 S ribosome from a map generated from 3JBU in Chimera. The difference map shows that the only difference in volume between the two maps is the tRNA (in magenta), the SecM AP (in pink), and the loop deleted from uL23. (D) Extracted cryo-EM density (in mesh) for uL23 in the uL23 Δloop ribosome EMD-4319. Wildtype uL23 (orange) and a de novo-built model for the mutant uL23 Δloop protein (PDB 6FU8; red) are shown in ribbon representation.
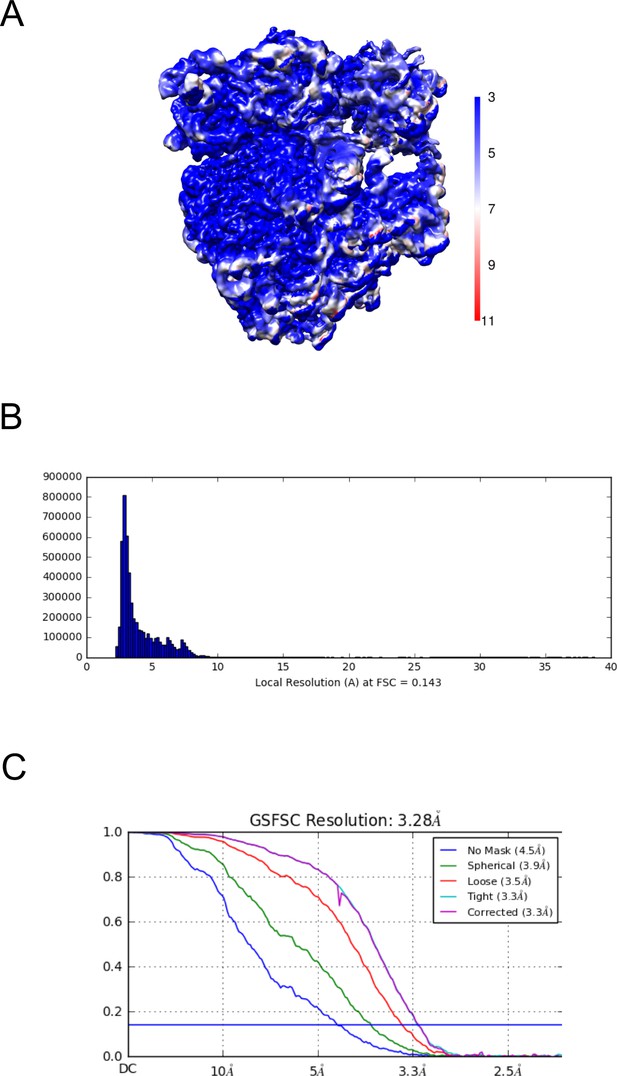
Resolution map of the uL23Δ loop ribosome.
(A) Calculation of the local resolution using cryoSPARC. (B) Resolution histogram at FSC 0.143. (C) Fourier-Shell correlation (FSC) curve of the refined map at 0.143 indicating an average resolution of 3.28 Å.
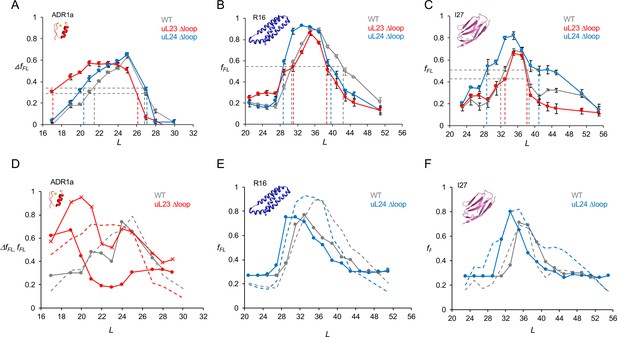
Cotranslational folding in WT, uL23 delta-loop, and uL24 delta-loop ribosomes.
(A) ΔfFL profiles (ΔfFL = fFL(50 μM Zn2+) – fFL(50 μM TPEN)) for ADR1a constructs translated in the PURE system supplemented with in-house purified WT (gray), uL23 Δloop (red), and uL24 Δloop (blue) E. coil ribosomes. (B) fFL profiles for spectrin R16 constructs translated in the PURE system supplemented with in-house purified WT (gray), uL23 Δloop (red), and uL24 Δloop (blue) E. coli ribosomes. (C) fFL profiles for titin I27 constructs translated in the PURE system supplemented with in-house purified WT (gray), uL23 Δloop (red), and uL24 Δloop (blue) E. coli ribosomes. Error bars in panels a-c show SEM values calculated from at least three independent experiments. Dashed lines indicate Lonset and Lend values, c.f., Table 1. fFL profiles for non-folding mutants of R16 and I27 are found in (Nilsson et al., 2017; Tian et al., 2018). (D) Simulated fFL profiles (full lines) for ADR1a, spectrin R16, and titin I27 obtained with WT (gray), uL23 Δloop (red), and uL24 Δloop (blue) ribosomes. The corresponding experimental fFL profiles from panels a-c are shown as dashed lines. The simulated ADR1a fFL profile marked by X’s was obtained with a uL23 Δloop(70-72) ribosome model. Simulated fFL profiles for ADR1a with uL24 Δloop ribosomes, and for R16 and I27 with uL23 Δloop ribosomes, are essentially identical to the corresponding profiles obtained with WT ribosomes, and are shown in Figure 3—figure supplement 10.
-
Figure 3—source data 1
Experimental fFL values.
- https://doi.org/10.7554/eLife.36326.019
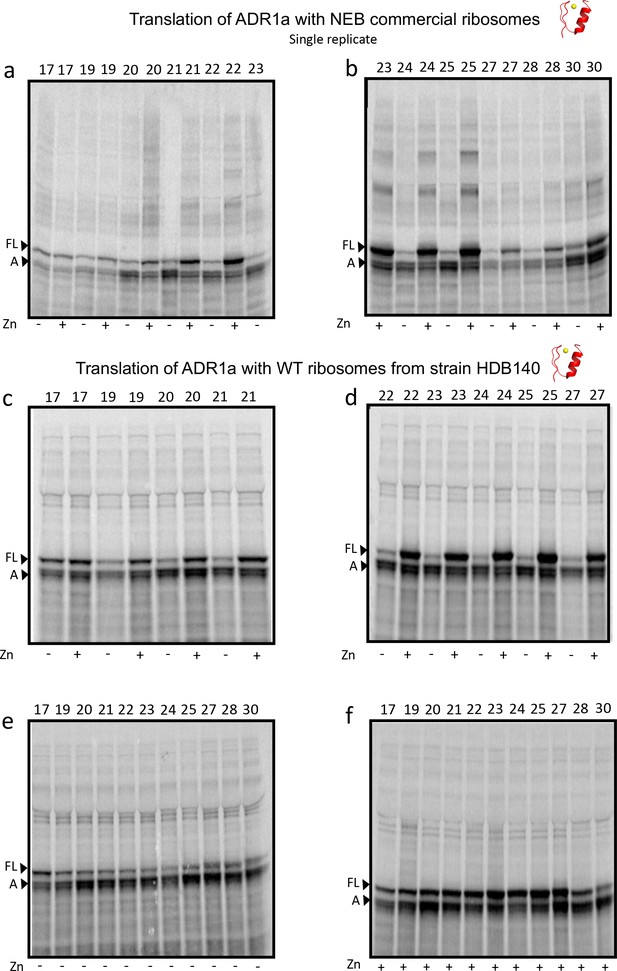
SDS PAGE showing ADR1 constructs translated in PURExpress Δ-Ribosome kit supplemented with high-salt-washed ribosomes isolated from HDB140, HDB143 (uL23 Δloop), or HDB144 (uL24 Δloop) as indicated.
Translations were run on 12% Bis-Tris gels with MOPS running buffer.
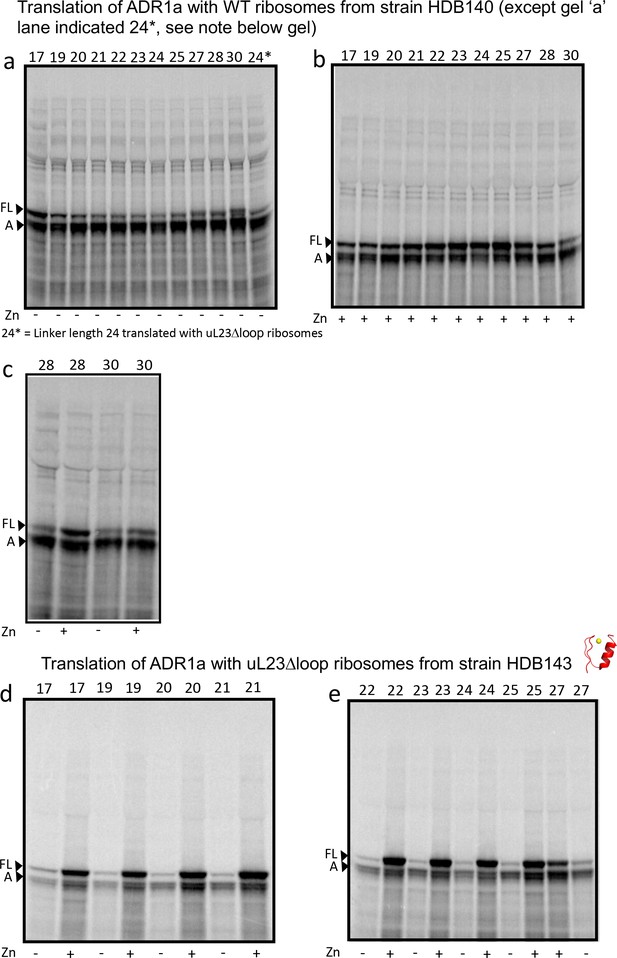
SDS PAGE showing ADR1 constructs translated in PURExpress Δ-Ribosome kit supplemented with high-salt-washed ribosomes isolated from HDB140, HDB143 (uL23 Δloop), or HDB144 (uL24 Δloop) as indicated.
https://doi.org/10.7554/eLife.36326.010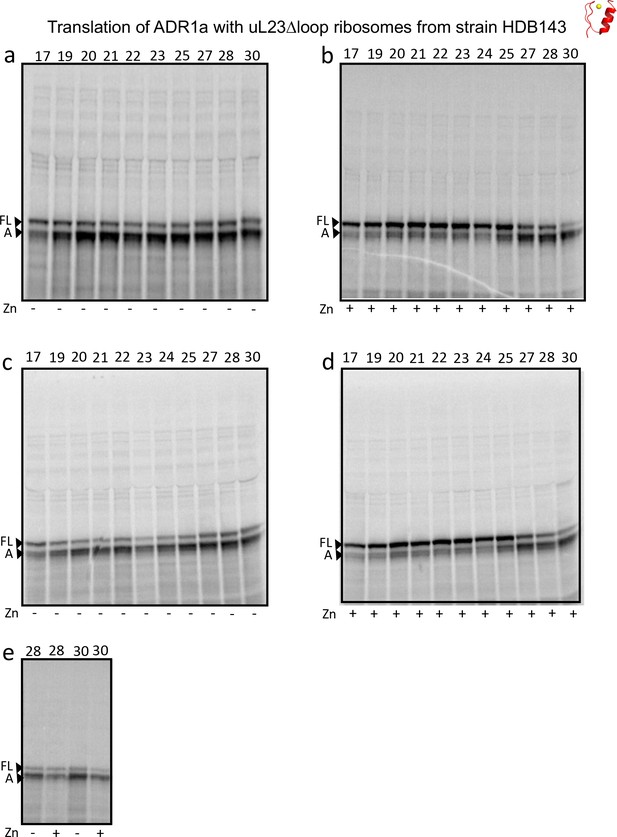
SDS PAGE showing ADR1 constructs translated in PURExpress Δ-Ribosome kit supplemented with high-salt-washed ribosomes isolated from HDB140, HDB143 (uL23 Δloop), or HDB144 (uL24 Δloop) as indicated.
https://doi.org/10.7554/eLife.36326.011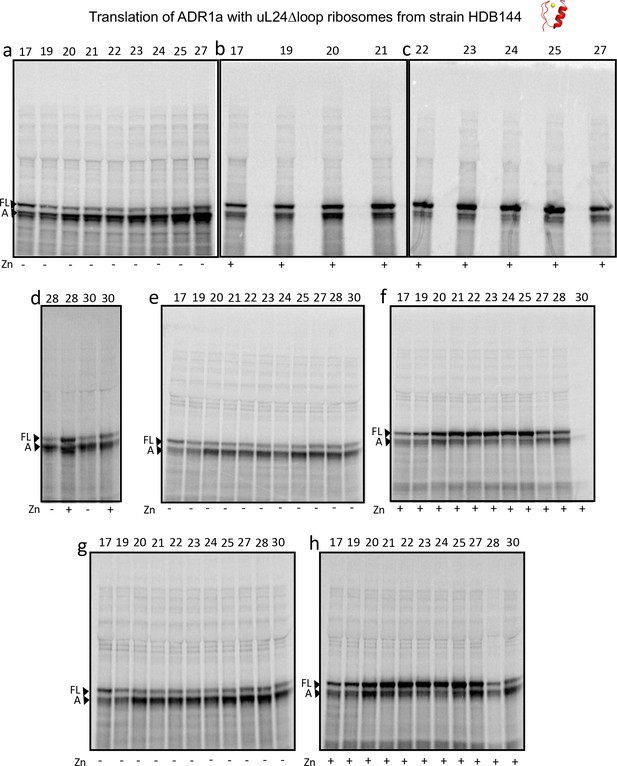
SDS PAGE showing ADR1 constructs translated in PURExpress Δ-Ribosome kit supplemented with high-salt-washed ribosomes isolated from HDB140, HDB143 (uL23 Δloop), or HDB144 (uL24 Δloop) as indicated.
https://doi.org/10.7554/eLife.36326.012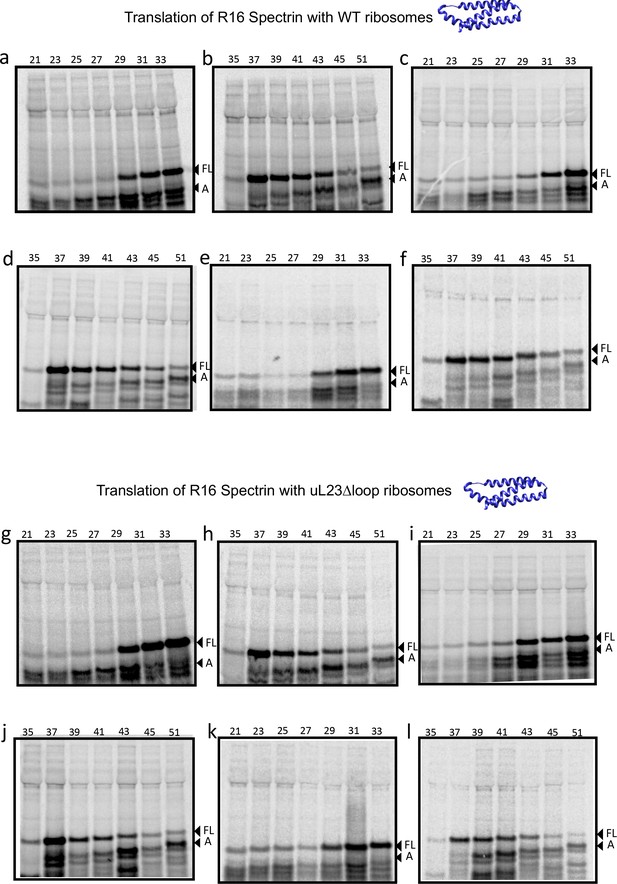
SDS PAGE showing Spectrin R16 constructs translated in PURExpress Δ-Ribosome kit supplemented with high-salt-washed ribosomes isolated from HDB140, HDB143 (uL23 Δloop), or HDB144 (uL24 Δloop) as indicated.
Translations were run on 12% Bis-Tris gels with MES running buffer.
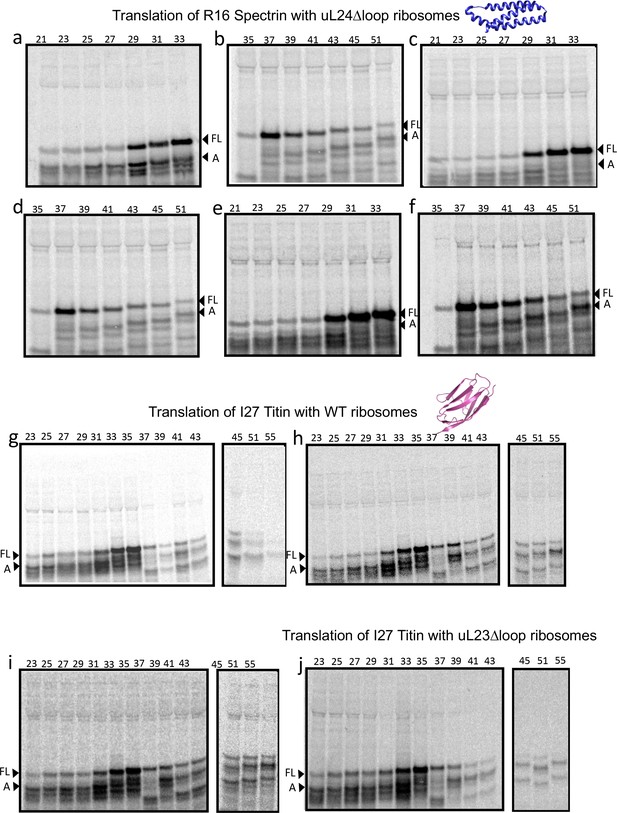
SDS PAGE showing Spectrin R16 and Titin I27 constructs translated in PURExpress Δ-Ribosome kit supplemented with high-salt-washed ribosomes isolated from HDB140, HDB143 (uL23 Δloop), or HDB144 (uL24 Δloop) as indicated.
Translations were run on 12% Bis-Tris gels with MES running buffer.
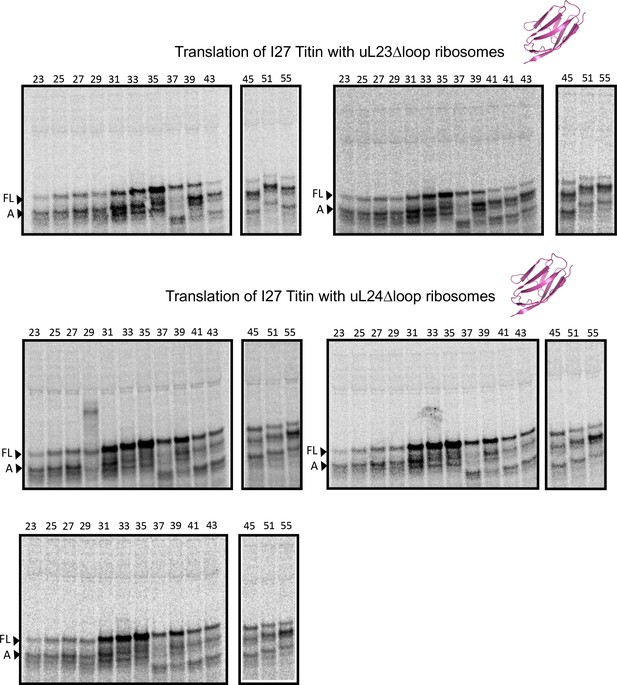
SDS PAGE showing titin I27 constructs translated in PURExpress Δ-Ribosome kit supplemented with high-salt-washed ribosomes isolated from HDB140, HDB143 (uL23 Δloop), or HDB144 (uL24 Δloop) as indicated
https://doi.org/10.7554/eLife.36326.015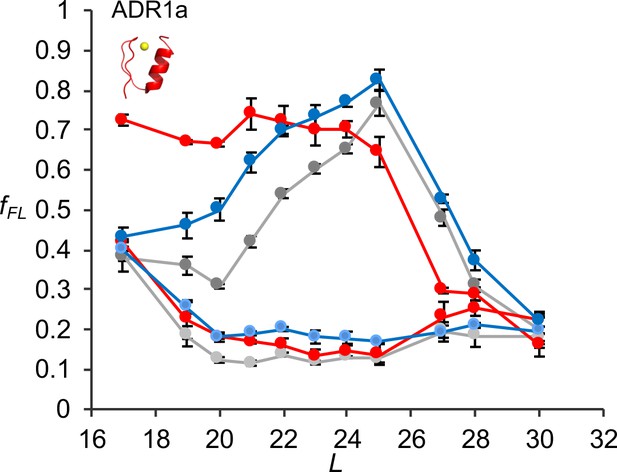
fFL profiles for ADR1a constructs translated in PURE by WT, uL23 Δloop, and uL24 Δloop ribosomes, either in the presence of Zn2+ or of the Zn2+ chelator TPEN.
Averages and standard errors calculated from three independent translation reactions are shown.

Sequences of the longest and the shortest constructs used for each protein and, a depiction of the location of the sequences the ribosome exit tunnel when the last residue of the AP is in the P-site (lower panel, yellow box).
ADR1 is indicated in red, spectrin R16 in blue, titin I27 in pink, and the SecM AP in magenta. The linker between each domain and the AP is in gray. The part of LepB added to the N-terminus of ADR1a is in black.
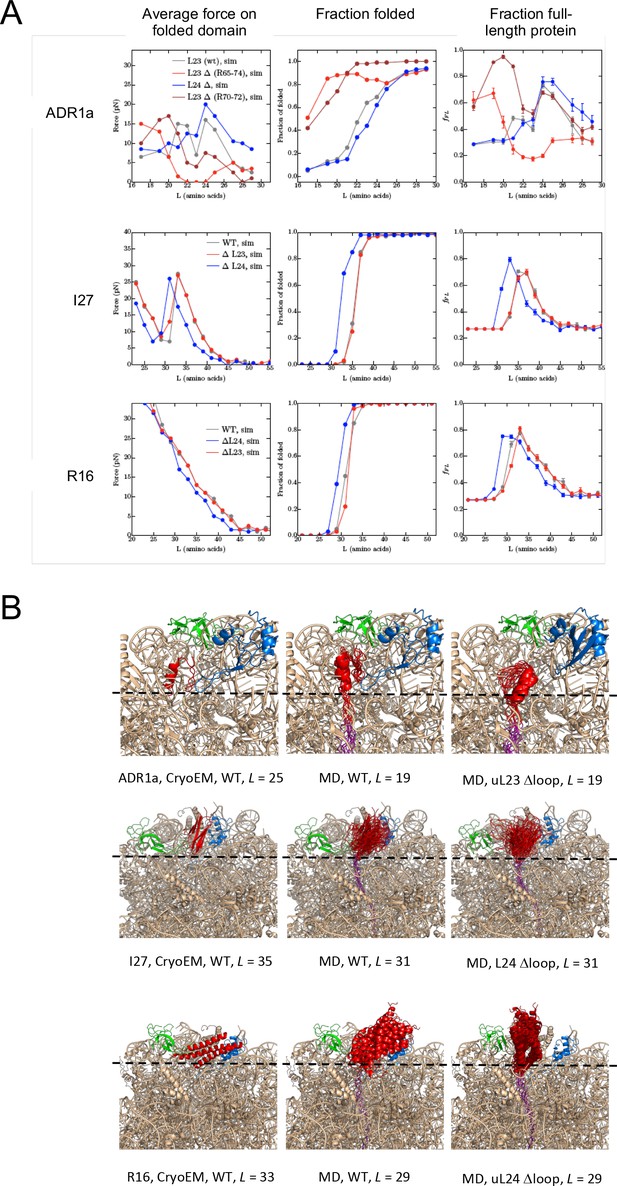
Summary of results from coarse-grained MD simulations.
(A) Average forces exerted on the AP by the folded state (first column), fraction folded protein (second column), and fFL values (last column) for ADR1a, I27, and R16 at different linker lengths L. (B) Snapshots of folded ADR1a, I27, and R16 domains in wildtype (WT), uL23 Δloop, and uL24 Δloop ribosomes at L values ≈ Lonset. Note that the folded proteins are located at similar depths in the exit tunnel in the cryo-EM structures and the simulations for WT ribosomes (this holds also for ADR1a in uL24 Δloop ribosomes as well as for I27 and R16 in uL23 Dloop ribosomes, c.f. panel A). The C terminus of folded ADR1a is located ~6 Å deeper in the exit tunnel in uL23 Δloop than in WT ribosomes, while folded I27 and R16 are located, respectively,~15 Å and ~13 Å deeper in the exit tunnel in uL24 Δloop ribosomes (dashed guide lines). Note that the linker is more stretched in the WT ribosome simulations compared to the Δloop ribosomes, consistent with the higher force and lower fraction folded protein seen for the WT ribosome data at the same L values (panel a).
Videos
The ribosome exit tunnel (mesh), as calculated for PDB 3JBU, uL23 Δloop and uL24 Δloop ribosomes by POVME.
See Figure 1A for coloring scheme. The b-hairpin loops deleted in uL23 Δloop and uL24 Δloop ribosomes are shown in yellow and light blue, respectively. To facilitate the visualization of the exit tunnel, spheres left outside the exit tunnel after POVME processing were manually removed.
Tables
Lonset, Lmax, and Lend values calculated from the fFL profiles in Figure 3.
https://doi.org/10.7554/eLife.36326.020| ADR1a | R16 | I27 | |||||||
|---|---|---|---|---|---|---|---|---|---|
| WT | uL23 Δloop | uL24 Δloop | WT | uL23 Δloop | uL24 Δloop | WT | uL23 Δloop | uL24 Δloop | |
| Lonset | 21 | 17 | 20 | 31 | 31 | 29 | 32 | 33 | 28 |
| Lmax | 25 | 22 | 25 | 35 | 35 | 33 | 35 | 35 | 35 |
| Lend | 27 | 26 | 27 | 42 | 39 | 40 | 38 | 38 | 41 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Escherichia coli) | HDB140, HDB143, HDB144, Strain background N281 | 10.1111/j.1365–2958 .2010.07325.x | NA | Strains used to isolate high-salt washed ribosomes in this study. |
| Antibody | uL23, uL24 | 10.1111/j.1365–2958 .2010.07325.x | NA | 1:8000 dilution used (incubated for one hour). Secondary antibody: Mouse (1:20,000 dilution incubated for one hour). Nitrocellulose membrane blocked with 5% Milk in TBS-T for an hour. |
| Peptide, recombinant protein | uL23 Δ loop, uL24 Δ loop | 10.1111/j.1365–2958 .2010.07325.x | UniProtKB- P0ADZ0 (rplW) UniProtKB-P60624 (rplX) | Referred to as HDB 143 and HDB 144 in original paper. Refers to genes rplWΔ65–74 and rplX Δ43–57 respectively. |
| Commercial assay or kit | GeneJET Plasmid miniprep kit | Thermo Fisher Scientific RRID:SCR_008452 | Cat no. K0502 | Used to purify plasmids |
| Commercial assay or kit | PURExpress Δ Ribosome kit | New England Biolabs | Cat no. E3313S | Kit was supplemented with ribosomes purified in the lab. Translation carried out at 37°C for 20 min. |
| Software, algorithm | EasyQuant | doi: 10.1038/ nsmb.2376 | NA | Used to quantify relative fraction full-length of translated protein from SDS-PAGE |
| Software, algorithm | cryoSPARC version v2 | Structura Biotechnology Inc | NA | Used for ab-initio reconstruction of uL23Δloop ribosomes. The following operations were carried out as part of the cryoSPARC workflow: 2D classification, Ab initio reconstruction, Homogeneous refinement, Sharpening and map flipping to correct for handedness, local resolution. |
| Software, algorithm | UCSF ChimeraX | SCR_015872 | Used to make Figure 2A | |
| Software, algorithm | UCSF Chimera v. 1.12 | J Comput Chem. 2004 Oct;25 (Nilsson et al., 2015): 1605–12. | SCR_004097 | Used to visualise the cryoEM map, fit PDB models 3JBU, 4YBB to check for differences in maps. Used to make figures. |
| Software, algorithm | Jalview v 2.10.4 | doi: 10.1093/ bioinformatics/btp033 | SCR_006459 | Use for generating multiple sequence alignments of uL23 and uL24 in the supplementary figures. |
| Chemical | Potassium acetate | Sigma-Aldrich (SCR_008988) | Cat no. P1190 | Source of potassium ions to stabilise ribosomes |
| Chemical | Magnesium acetate | Sigma-Aldrich (SCR_008988) | Cat no. M5661 | Source of Magnesium ions to stabilise ribosomes |
| Chemical | cOmplete protease inhibitor cocktail | Sigma-Aldrich (SCR_008988) | Cat no. 04693116001 | Used as a protease inhibitor during cell lysis to obtain ribosomes |
| Chemical | N,N,N′,N′-Tetrakis (2-pyridylmethyl) ethylenediamine | Sigma Aldrich (SCR_008988) | Cat no. P4413 | Used to chelate Zn for the -Zn reactions in the ADR1 plot. |
| Chemical | threo-1,4-Dimercapto-2, 3-butanediol DL-Dithiothreitol | Sigma Aldrich (SCR_008988) | Cat no. DTT-RO | Reducing agent added to buffers during ribosome purification and as a reductant for SDS-PAGE |
| Chemical | Tris Base | Sigma-Aldrich (SCR_008988) | Cat no. T1503 | Buffering agent during ribosome preparation |
| Chemical | 35S Methionine | Perkin-Elmer | Cat no. NEG009T001MC | 35S Methionine is incorporated into the protein during in vitro translation and aids detection by phosphorimaging. |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.36326.021





