Spliceosome factors target timeless (tim) mRNA to control clock protein accumulation and circadian behavior in Drosophila
Abstract
Transcription-translation feedback loops that comprise eukaryotic circadian clocks rely upon temporal delays that separate the phase of active transcription of clock genes, such as Drosophila period (per) and timeless (tim), from negative feedback by the two proteins. However, our understanding of the mechanisms involved is incomplete. Through an RNA interference screen, we found that pre-mRNA processing 4 (PRP4) kinase, a component of the U4/U5.U6 triple small nuclear ribonucleoprotein (tri-snRNP) spliceosome, and other tri-snRNP components regulate cycling of the molecular clock as well as rest:activity rhythms. Unbiased RNA-Sequencing uncovered an alternatively spliced intron in tim whose increased retention upon prp4 downregulation leads to decreased TIM levels. We demonstrate that the splicing of tim is rhythmic with a phase that parallels delayed accumulation of the protein in a 24 hr cycle. We propose that alternative splicing constitutes an important clock mechanism for delaying the daily accumulation of clock proteins, and thereby negative feedback by them.
Editorial note: This article has been through an editorial process in which the authors decide how to respond to the issues raised during peer review. The Reviewing Editor's assessment is that all the issues have been addressed (see decision letter).
https://doi.org/10.7554/eLife.39821.001Introduction
Circadian rhythms allow organisms to orchestrate behavioral and physiological outputs in anticipation of predictable diurnal changes in the environment. These rhythms are generated by endogenous molecular clocks that entrain to environmental cycles, predominantly light and temperature, and can maintain rhythms when released into constant conditions (free-run). A conserved mechanistic feature of circadian clocks is an auto-regulatory transcriptional feedback loop in which circadian proteins rhythmically regulate their own expression to generate a clock, and also drive a global program of cycling gene expression. Discovery of this clock mechanism was revolutionary and has received well-deserved recognition, but critical aspects of how the clock is sustained remain unclear.
PERIOD (PER) and TIMELESS (TIM) are the auto-regulating elements of the circadian clock in Drosophila (Allada and Chung, 2010; Hardin, 2011; Zheng and Sehgal, 2012). The expression of per and tim is driven by circadian transcription factors CLOCK (CLK) and CYCLE (CYC), and peaks around the early night. Relative to their mRNA peak, accumulation of PER and TIM proteins is delayed by ~6 hr. In the mid-to-late night, PER and TIM are predominantly nuclear, and once in the nucleus, they repress CLK-CYC activity to decrease per and tim expression. Degradation of TIM and then of PER in the morning resets the transcription cycle and restarts the loop. In order to maintain rhythmicity and set the proper pace of the circadian clock, both PER and TIM need to be dynamically regulated on multiple levels. For instance, a stable circadian molecular oscillator requires temporal delays to separate the phases of gene transcription and repression and thereby prevent these from reaching equilibrium (Zheng and Sehgal, 2012).
The overall levels and stability of TIM constitute a critical circadian modality. Although PER is the more important factor for transcriptional regulation, levels and activity of PER depend upon TIM (Price et al., 1995; Dubruille and Emery, 2008). TIM levels are acutely modulated by light, which promotes the degradation of TIM, and thereby PER, during the day and allows the rise in circadian transcription (Suri et al., 1999; Yang and Sehgal, 2001). Subsequently, TIM accumulation is necessary to stabilize PER and promote its nuclear accumulation (Jang et al., 2015). Thus, in the presence of light:dark cycles, light delays the accumulation of TIM-PER and so contributes to the lag in repression. These temporal relationships are largely preserved in constant darkness, and are also entrained by temperature cycles regardless of light cues, although the mechanisms under these conditions are not known.
While regulated protein stability and translation have been directly explored as mechanisms that could contribute to maintenance of the feedback loop (Dembinska et al., 1997; Chen et al., 1998 ;Lim and Allada, 2013; Zhang et al., 2013), and regulation of protein stability is indeed critical (Zheng and Sehgal, 2012), little investigation has focused on a potential role of alternative splicing. To date, the best-studied role for alternative splicing in Drosophila rhythms is in the temperature-dependence of the behavioral siesta (Majercak et al., 1999; Majercak et al., 2004; Collins et al., 2004). Splicing is driven by spliceosomes, dynamic RNA-protein complexes composed of five core small nuclear ribonucleoprotein particles (U1, U2, U4, U5, U6 snRNP) and >150 additional proteins specific for each snRNP complex (Wahl et al., 2009). In this study, we report a circadian role for Pre-mRNA Processing factor 4 (PRP4), a conserved component of the spliceosomal U4/U6.U5 triple small nuclear ribonucleoprotein (tri-snRNP) complex. We identified PRP4 in a screen for novel regulators of the free-running circadian period, and established that PRP4 is necessary in tim+ clock cells to maintain 24 hr period and robust rhythms of the circadian clock. In addition to prp4, downregulation of multiple tri-snRNP components affected circadian period length and rhythmicity, which led us to implicate this entire spliceosomal complex in circadian regulation. Using unbiased RNA-Sequencing, we characterized the splicing events regulated by PRP4 and identified a novel intron retention event in tim. We show that alternative splicing of this intron in tim represents an important mechanism to time the daily accumulation of TIM, in constant darkness following entrainment to light:dark cycles and also in temperature cycles. Together, these findings identify a mechanism contributing to the maintenance of clock function.
Results
Pre-mRNA splicing factor four is a new regulator of the circadian clock
In a screen for kinases that affect circadian period length of Drosophila rest:activity rhythms in constant dark:dark (DD) conditions, we identified Pre-mRNA Processing factor 4 (PRP4), a splicing factor that also has kinase activity (Kojima et al., 2001; Schneider et al., 2010). RNA interference (RNAi)-mediated knockdown of prp4 in tim+ clock neurons with the Tim-UAS-Gal4 (TUG) Gal4 driver resulted in consistently long free-running circadian periods (Figure 1A–B). Two independent RNAi lines (GD and KK) were used to confirm these findings.
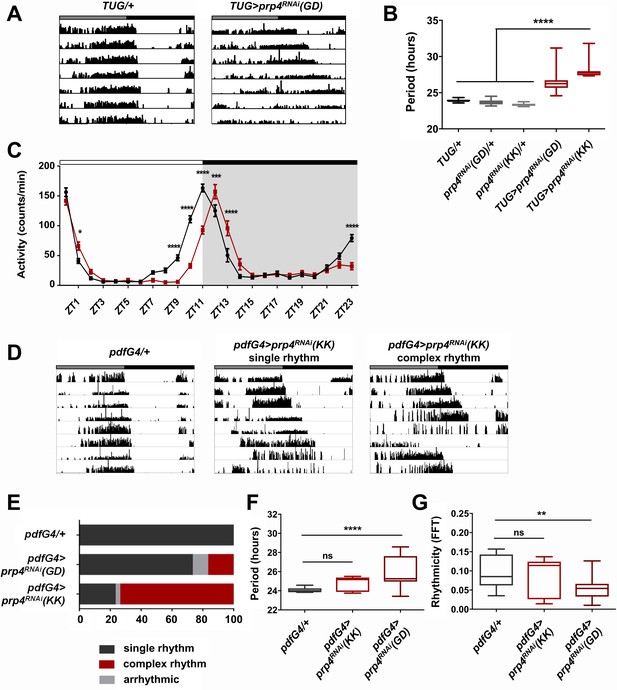
Pre-mRNA splicing factor four is a new regulator of the circadian clock.
(A) Representative activity records of free-running fly behavior upon downregulation of prp4 in tim+ neurons. Dicer2 (Dcr2) was co-expressed with the RNAi transgenes to increase the knockdown efficiency. Genotypes are indicated on top of each panel. The gray and black bars indicate the subjective day and night, respectively. (B) The lengthening of free-running circadian period is significant for two independent prp4 RNAi lines (GD and KK). ****p ≤ 0.0001 relative to heterozygous controls by one-way ANOVA and Tukey’s post hoc test, n = 8–24. (C) Downregulation of prp4 in tim+ cells affects morning and evening anticipation in 12 hr:12 hr light:dark (LD) conditions. The activity profile of control (TUG; Dcr2/+) flies is in black, while the activity profile of experimental (TUG; Dcr2 >prp4RNAi(GD)) flies is in red. The white and black bars indicate light and dark conditions, respectively. *p ≤ 0.05, ***p ≤ 0.001,****p ≤ 0.0001 to control (TUG; Dcr2/+) for each ZT range by two-way ANOVA and Sidak’s post hoc test. Data represent mean ±SEM (n = 31–32). (D) Activity records demonstrate 7 days of representative free-running rhythms of prp4 knockdown flies. Dicer2 (Dcr2) was co-expressed with the RNAi transgenes to increase the knockdown efficiency. Genotypes are indicated on top of each panel. The gray and black bars indicate the subjective day and night, respectively. (E) Knockdown of prp4 in LNv pacemaker cells causes complex behavioral periods (p < 0.0001 by χ2 analysis, n = 19–38). (F, G) Knockdown of prp4 in LNvs lengthens circadian period and decreases rhythm strength. n.s., not significant at the 0.05 level, **p ≤ 0.01, ****p ≤ 0.0001 to control (pdfG4; Dcr2/+) by one-way ANOVA and Holm-Sidak’s post hoc test. Only rhythmic flies (FFT > 0.01) were analyzed (n = 9–22). In panels (B), (F) and (G), the boxes extend from the 25th to 75th percentiles, the line within the box is plotted at the median and whiskers extend from the lowest to highest value.
The length of the free-running circadian period is an important parameter that also affects the daily distribution of fly activity in light:dark (LD) cycles. Thus, we determined the effect of prp4 knockdown on fly diurnal activity in LD conditions (Figure 1C). Control (TUG; Dcr2/+) flies displayed characteristic bimodal activity in LD, with activity peaks that precede the onset of light (ZT0) and the onset of dark (ZT12). Compared to controls, the flies with circadian-cell-specific prp4 knockdown (TUG; Dcr2 >prp4RNAi(GD)) exhibited a delayed evening activity peak as well as a slightly delayed morning peak. These findings are consistent with the longer endogenous period that we report for prp4 knockdown flies (Figure 1A–B).
As the small lateral ventral neurons (s-LNvs) are the most relevant clock cells for driving rest:activity rhythms under constant dark conditions, we asked if effects of PRP4 were mediated in these cells (Helfrich-Förster, 1998). To test this hypothesis, we downregulated prp4 in both s-LNvs and large lateral ventral neurons (l-LNvs) with a Gal4 transgene driven by a peptide expressed specifically in these cells, Pigment Dispersing Factor (PDF). Knockdown of prp4 with pdf-Gal4 driving the weaker RNAi line (GD) led to a modest yet significant lengthening of free-running circadian period (Figure 1F). Interestingly, the knockdown of prp4 with a stronger RNAi (KK) led to complex rhythms (Figure 1E). These complex rhythms were generally characterized by changes in the behavioral pattern, often manifest as phase shifts during day 4 or 5 of constant darkness (see ~6 hr shift in the record shown), in the midst of an otherwise rhythmic record (Figure 1D). Such a complex rhythm phenotype could reflect uncoupling of period into two components of pdf+ and pdf- cell oscillators (Yao et al., 2016). Overall, these findings suggest that PRP4 action in LNvs contributes to maintaining clock function.
PRP4 is required for robust TIM and PER cycling
Since our data indicated that PRP4 is necessary in LNvs for proper circadian rhythmicity (Figure 1), we next examined the circadian cycling of PER and TIM in s-LNvs at regular intervals around the clock (Figure 2A–C). Cycling of both PER and TIM was affected in s-LNvs upon prp4 knockdown with the clock-specific TUG driver (Figure 2A–C). Total TIM levels, as quantified by corrected total cell fluorescence (CTCF) image analysis, were decreased during the night (Figure 2B–C). PER levels were also lower in the late night (ZT20), and especially so in the early morning (ZT2), in prp4 knockdown flies (Figure 2A,C). Additionally, nuclear accumulation of PER was delayed beyond ~ZT20, the time point at which PER was already partially localized to the nucleus in control flies (Figure 2A). To further characterize the effect on nuclear expression, we profiled relative nuclear PER expression upon prp4 knockdown with each of the two RNAi lines (GD and KK) in s-LNvs around the time of nuclear accumulation of PER (ZT18-ZT22). We found that the nuclear accumulation of PER was slower in flies with decreased prp4 (both GD and KK RNAi lines) than in controls (TUG; Dcr2/+) at these time points (Figure 2—figure supplement 1). Overall, our immunohistochemistry data point to a distinct molecular clock phenotype upon prp4 downregulation. Because TIM is necessary for nuclear accumulation of PER (Vosshall et al., 1994; Saez and Young, 1996; Jang et al., 2015), reduced TIM accumulation could account for the delay and overall reduced nuclear expression of PER in s-LNvs (Figure 2, Figure 2—figure supplement 1), which together could account for the longer free-running period.
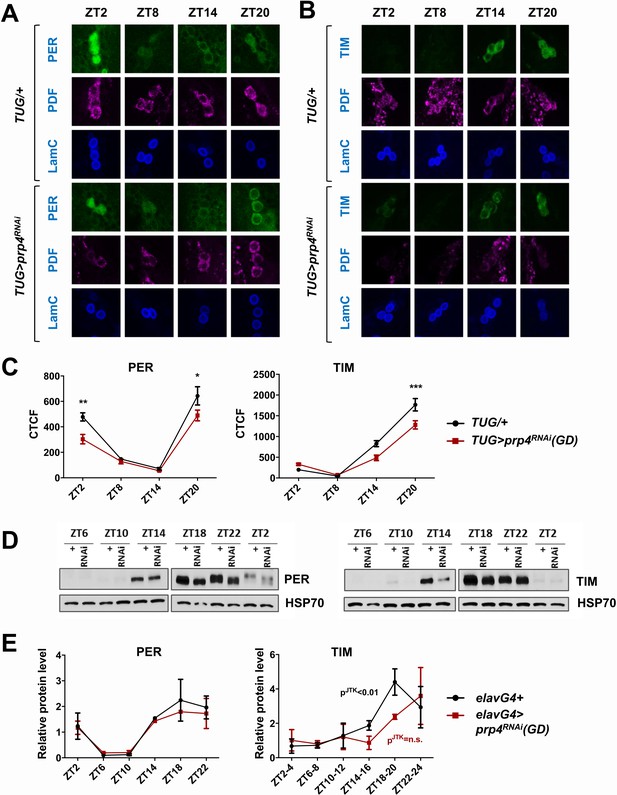
PRP4 is required for robust TIM and PER cycling.
(A, B) Cycling of PER and TIM is disrupted in s-LNvs of prp4 knockdown flies. Adult brains were dissected at time points indicated and immunostained with PER or TIM (green), PDF (magenta) and LaminC (blue) antibodies. Dicer2 (Dcr2) was co-expressed with the prp4 RNAi transgene to increase its knockdown efficiency. Genotypes are indicated on the sides of each panel. The displayed images are representative of 2–3 independent experiments. (C) Corrected Total Cell Fluorescence (CTCF) was used to quantitatively assess the change in levels of PER and TIM in s-LNvs. The signal from both the nucleus and the cytoplasm was used to calculate CTCF. 10–21 cells from 5 to 8 brains were analyzed for ZT2, ZT14 and ZT20 and 8–10 cells from 3 to 5 brains for ZT8. Images were taken with identical confocal settings. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001 to control (TUG;Dcr2/+) for each ZT time point as determined by two-way ANOVA and Sidak’s post hoc test. Error bars = ± SEM. (D) PER phosphorylation is decreased and cycling of TIM is blunted in fly heads with pan-neuronal knockdown of prp4. Dicer2 (Dcr2) was co-expressed with the prp4 RNAi transgene to increase its knockdown efficiency. Adult fly heads were collected at indicated zeitgeber (ZT) time points in a 12 hr:12 hr light:dark cycle. Representative western blots probed for PER (right) or TIM (left) are shown. HSP70 was used as a loading control. (E) Total PER and TIM levels were quantified from western blots in (D). JTK cycle analysis identified significant cycling (pJTK ≤ 0.01) for control (elavGal4; Dcr2/+) and not significant (n.s at the 0.05 level) cycling for samples with pan-neuronal prp4 knockdown. Data represent mean ±SEM (n = 2–3 independent experiments).
As biochemical analysis of PER and TIM oscillations requires large amounts of tissue, we conducted pan-neuronal knockdown of prp4 with the elavGal4 driver. The pan-neuronal manipulation targeted prp4 more broadly, allowing us to verify the efficiency of the knockdown through RNA analysis of whole heads. We consistently observed ~50% reduction in prp4 RNA in head lysates of flies where it was knocked down pan-neuronally (Figure 2—figure supplement 2). This broad manipulation also allowed us to test the effects of prp4 reduction on PER and TIM levels using western blotting of whole head lysates. The results obtained with this relatively crude method agreed well with our immunohistochemistry profiling of s-LNvs. First, we observed a strong effect of prp4 downregulation on total TIM levels, particularly during the initial TIM accumulation phase (ZT10-18) (Figure 2D–E). Quantification of our western blot data across multiple experiments suggested that prp4 knockdown also blunted the circadian cycling of TIM protein (Figure 2E). While the total levels of PER were not changed upon prp4 downregulation, its phosphorylation profile was dampened. In control (elavGal4; Dcr2/+) flies, as previously reported (Edery et al., 1994), PER was increasingly phosphorylated in the late night and early morning up until its degradation. In flies with reduced prp4 levels, PER migrated at a lower molecular weight on western blots (ZT18-ZT2). We speculate that the dampened phosphorylation profile of PER reflects, at least in part, the defect in nuclear accumulation of PER because a number of sites on PER get phosphorylated only when PER starts to accumulate in the nucleus (after ZT20) (Chiu et al., 2011). Alternatively, prp4 depletion could have a TIM-independent effect on PER to regulate its phosphorylation profile.
Circadian rhythmicity is modulated by tri-snRNP levels
As noted above, PRP4 is associated with kinase activity, but it is also a component of the spliceosome (Kojima et al., 2001; Schneider et al., 2010). To address if the circadian role of PRP4 was spliceosome-dependent, we assayed circadian behavior following knockdown of additional tri-snRNP components. Strikingly, clock-specific downregulation of pre-mRNA processing factor 3 (prp3) and pre-mRNA processing factor 31 (prp31), which are associated primarily with U4/U6 snRNP, as well as pre-mRNA processing factor 8 (prp8) and bad response to refrigeration 2 (brr2), which are associated with U5 snRNP, caused pronounced defects in free-running circadian behavior (Table 1). The phenotypes ranged from period lengthening to complete arrhythmicity, suggesting that the overall levels and/or stability of tri-snRNP regulate circadian rhythms.
Free-running locomotor behavior of flies expressing RNAi against tri-snRNP components
https://doi.org/10.7554/eLife.39821.006| Genotype | N | rhythmicity* (%) | Period (hours ± SEM) | Power (FFT ± SEM) |
|---|---|---|---|---|
| TUG; Dcr2/+ | 35 | 100% | 23.80 (0.06) | 0.11 (0.01) |
| pdfGal4, Dcr2/+ | 19 | 100% | 24.06 (0.06) | 0.10 (0.01) |
| elavGal4; Dcr2/+ | 30 | 97% | 23.59 (0.33) | 0.06 (0.03) |
| +/prp4RNAi(GD) | 16 | 100% | 23.62 (0.06) | 0.09 (0.01) |
| +/prp4RNAi(KK) | 16 | 100% | 23.32 (0.05) | 0.13 (0.01) |
| +/prp3RNAi(GD) | 10 | 100% | 23.73 (0.06) | 0.10 (0.01) |
| +/prp3RNAi(KK) | 15 | 100% | 23.45 (0.06) | 0.12 (0.01) |
| +/prp8RNAi(GD) | 15 | 100% | 23.64 (0.07) | 0.11 (0.01) |
| +/prp31RNAi(KK) | 13 | 100% | 23.35 (0.06) | 0.12 (0.04) |
| +/brr2RNAi(KK) | 16 | 100% | 23.47 (0.06) | 0.11 (0.01) |
| TUG; Dcr2 > prp4RNAi(GD) | 34 | 71%† | 26.55 (0.29)‡ | 0.09 (0.01) |
| TUG; Dcr2 > prp4RNAi(KK) | 40 | 45%† | 29.47 (0.48)‡ | 0.09 (0.01) |
| TUG; Dcr2 > prp3RNAi(GD) | 11 | 100% | 27.45 (0.49)‡ | 0.08 (0.02) |
| TUG; Dcr2 > prp3RNAi(KK) | 33 | 0%† | - | - |
| TUG; Dcr2 > prp8RNAi(GD) | 30 | 23%† | 28.00 (1.02)‡ | 0.02 (0.01)‡ |
| TUG;Dcr2 > prp31RNAi(KK) | 17 | 88% | 25.33 (0.19)‡ | 0.06 (0.01)¶ |
| TUG;Dcr2 > brr2RNAi(KK) | 24 | 0%† | - | - |
| pdfGal4, Dcr2 > prp4RNAi(GD) | 30 | 90% | 25.85 (0.32)‡ | 0.06 (0.01)|| |
| pdfGal4, Dcr2 > prp4RNAi(KK) | 38 | 97% | 24.81 (0.32)‡ | 0.09 (0.02) |
| elavGal4; Dcr2 > prp4RNAi(GD) | 27 | 52%† | 24.54 (0.14)§ | 0.03 (0.02)# |
-
* Flies with FFT value >0.01 are considered to be rhythmic.
†p < 0.001 compared to both of the heterozygous controls, by χ2 analysis.
-
‡p < 0.001 compared to both of the heterozygous controls, by Student’s t test.
§p < 0.001 compared to RNAi control but not significant (p > 0.05) compared to elavGal4; Dcr2/+ control, by
-
Student’s t test.
¶p < 0.01 compared to TUG; Dcr2/+ control but not significant (p > 0.05) compared to RNAi control, by Student’s t test.
-
||p < 0.01 compared to pdfGal4; Dcr2/+ control and p < 0.05 compared to RNAi control, by Student’s t test.
#p < 0.05 compared to RNAi control but not significant (p > 0.05) compared to elavGal4; Dcr2/+ control, by Student’s t test.
To determine if the molecular signatures of prp4 knockdown and other tri-snRNP downregulation phenotypes were similar, we performed s-LNv-specific analysis of PER and TIM levels with a few randomly selected RNAi lines. We found that the s-LNvs of flies with downregulated prp8 had lower TIM levels at night, and also showed decreased nuclear accumulation at ZT20 relative to control flies (Figure 3A). These data allow us to conclude that circadian oscillations of PER (data not shown) and TIM are sensitive to changes in the levels of multiple tri-snRNP components.
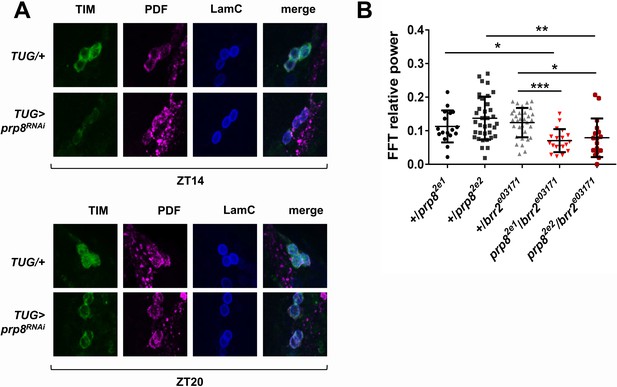
PRP8 regulates TIM levels and the strength of rest:activity rhythms.
(A) TIM levels are decreased in s-LNvs of prp8 knockdown flies. Adult brains were dissected at ZT14 and ZT20 on the 4th day in LD cycle and immunostained with TIM (green), PDF (magenta) and LaminC (blue) antibodies. Genotypes are indicated on the sides of each panel. Displayed images are representative of two independent experiments. (C) Trans-heterozygous prp8/brr2 mutants have weaker circadian rhythms compared to their heterozygous controls, p* ≤ 0.05, p** ≤ 0.01, p*** ≤ 0.001 by one-way ANOVA, Tukey post hoc test. All FFT value were used in the analysis, including the arrhythmic ones (FFT < 0.01). Error bars represent mean ±SEM (n = 17–36). Figure 3 is related to Table 1.
We further examined tri-snRNP function in circadian regulation by utilizing some of the previously characterized mutants of prp8 and brr2 (Coelho et al., 2005; Bivik et al., 2015). Because tri-snRNP components are essential genes with no homozygous viable knockout mutants described to date, we tested circadian behavior in the viable transheterozygous prp8/brr2 mutants. The heterozygous prp8/+ and brr2/+ mutants displayed normal circadian rhythms, readily explained by the haplosufficiency of these two genes (Figure 3B). Transheterozygous prp8/brr2 mutant flies had normal circadian period (data not shown), but displayed decreased rhythm strength compared to the prp8/+ and brr2/+ heterozygous flies, confirming a circadian function of the tri-snRNP (Figure 3B).
PRP4 regulates tim splicing
Next, we attempted to identify the mechanism by which PRP4 regulates circadian rhythms, starting with the broad hypothesis that downregulation of prp4 leads to the aberrant splicing of one or more core clock transcripts. Analysis of clock transcripts indicated an increase in tim levels in the late night in flies with reduced prp4 (Figure 4—figure supplement 1), but in order to apply an unbiased approach, we performed RNA-Sequencing (RNA-Seq) analysis of fly heads in which prp4 had been knocked down. As clock protein expression in the eye contributes a majority of the signal in head assays, we used an eye-specific Glass Multiple Promoter (GMR-Gal4) driver for prp4 knockdown. Overall gene expression was dramatically influenced by prp4 downregulation (433 down, 310 up at FDR < 0.05) (Supplementary file 2). Pathway enrichment analysis using DAVID identified changes in folate biosynthesis as well as in other broad pathway categories such as protein export, protein processing in endoplasmic reticulum and drug metabolism (Supplementary file 1). Interestingly, components of folate metabolism have been previously implicated in circadian clock regulation in human cells (Zhang et al., 2009). Despite the fact that PRP4 is a component of the core spliceosome required for constitutive splicing, we did not detect dramatic effects on global splicing. Using the Comprehensive AS Hunting (CASH) method, which assays for splicing events, our analysis identified 45 genes exhibiting differential splicing upon prp4 downregulation, with FDR ≤ 0.05 (Wu, W., et al., 2017) (Supplementary file 3).
An intron retention event in tim that was significantly upregulated upon prp4 knockdown was of particular interest due to its potential clock function (Figure 4A–B). Our initial splicing analysis was performed with CASH, but we additionally ran the Cufflinks-2.2 pipeline to obtain psi (percent spliced in) information for differentially spliced isoforms (Supplementary file 4). Importantly, the same retention event in tim, at position chr2L:3499764–3500000 (hereafter we refer to this intron as tim-tiny for simplicity), was consistently identified as significantly upregulated with both analyses (Figure 4A). For comparison, differential alternative splicing in only four other genes (trol, pex7, Npc1a, vsg) was consistently identified with both of the algorithms (Figure 4A). Moreover, the RNA-Seq reads across the tiny-tiny junction normalized to its neighboring junction (2L:3500362–3500422), which is spliced out in all of the tim isoforms, were significantly increased in the prp4 knockdown samples (GMR >prp4RNAi) compared to the controls (GMR/+), further pointing to the retention of tim-tiny. To estimate how common tim-tiny retention was in control flies (GMR/+), we quantified the ratio of isoforms that contain tim-tiny (tim-RM and tim-RS) to those that do not contain this intron using our Cufflinks-2.2 output (Supplementary file 4). This analysis indicated that the isoforms retaining tim-tiny were twice as abundant as other isoforms at ZT 8 (when all of the RNA-Seq samples were collected) (Supplementary file 5).
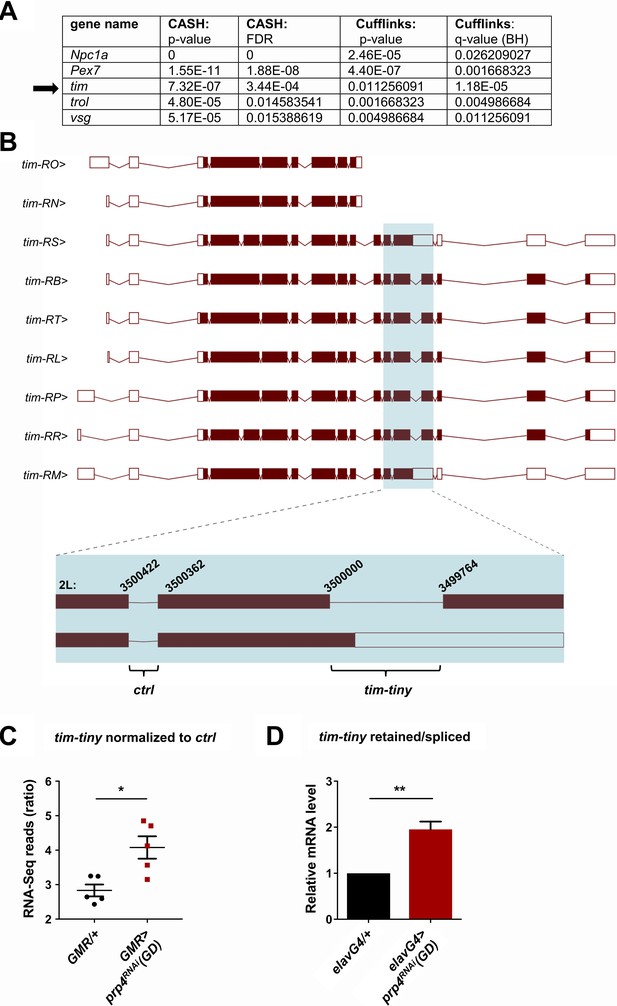
PRP4 regulates tim splicing.
(A) Only five genes were identified as differentially spliced upon prp4 downregulation with both CASH and Cufflinks/differential psi (percent spliced in) pipelines. For each gene, the corresponding p-value and False Discovery Rate (FDR) or q-value as determined by Benjamini-Hochberg (BH) procedure are reported. (B) tim isoforms (image adapted from Ensembl Fruitfly release 92, genome assembly BDGP6) are displayed. The boxes indicate exons, with filled boxes (brown) representing protein-coding sequences. The region of interest is enlarged (blue box) and depicts a constitutively spliced intron (‘ctrl’) and the intron that gets retained upon prp4 knockdown (‘tim-tiny’). The chromosomal coordinates of these introns are indicated at their respective exon-intron junctions. (C) tim-tiny retention was revealed by RNA-Seq analysis in samples with prp4 downregulated (GMR > prp4 RNAi). The number of RNA-Seq reads across the tim-tiny intron normalized to the number of reads across the ctrl intron is higher in prp4 knockdown flies (GMR > prp4RNAi) compared to controls (GMR/+). Data represent five independent biological replicates. Error bars represent mean RNAiSEM. *p ≤ 0.0001 as determined by CASH (refer to panel A). (D) An increase in intron retention in flies with pan-neuronal prp4 knockdown was confirmed with qPCR analysis. Dicer2 (Dcr2) was co-expressed with the prp4 RNAi transgene to increase its knockdown efficiency. Data represent four independent biological replicates, with technical triplicates performed during the qPCR step for each replicate. **p ≤ 0.01 to control (elavGal4; Dcr2/+) as determined by Student’s t test. Data represent mean ±SEM.
Motivated by the RNA-Seq data, we sought to verify the retention of the tim-tiny intron upon pan-neuronal prp4 knockdown. For this purpose, we designed primers that would amplify only the transcript containing the retained intron or only the spliced transcript respectively (Figure 4—figure supplement 2). The ratio of retained to spliced signal would then indicate the relative intron retention. Using this approach, increased tim-tiny retention was consistently detected upon pan-neuronal prp4 knockdown (Figure 4D). Additionally, by performing a control set of experiments, the signal from the ‘retained’ primer set targeting tim-tiny was verified not to reflect genomic DNA contamination (Figure 4—figure supplement 2A–B). First, retained intron levels were normalized to total tim mRNA. As total tim mRNA was amplified with primers that do not span junctions, but rather bind within sequences of a different exon (‘exon’), they are also expected to detect any contaminating DNA in addition to mRNA. As with the tim-tiny retained/spliced ratio, the retained/exon ratio indicated increased tim-tiny retention upon prp4 knockdown, in particular at ZT12, a time point used for our initial RNA-Seq analysis (Figure 4—figure supplement 2B). Additionally, we amplified RNA using primers that span an exon-exon junction at a different part of tim (‘mRNA’), which hence should only detect mRNA, and found that the ratio of tim ‘exon’ to ‘mRNA’ was not different between the control (elavGal4; Dcr2/+) and the prp4 knockdown flies (elavGal4; Dcr2 > prp4RNAi(GD)), indicating that residual DNA contamination does not contribute to the increased detection of tim-tiny with prp4 knockdown (Figure 4—figure supplement 2B).
Intron retention in tim decreases TIM levels and affects circadian behavior
To assay the influence of prp4-dependent tim-tiny intron retention on TIM levels, we generated tim cDNA constructs that lacked the intron (‘tim-spliced‘), included the intron (‘tim-retained’) or included the intron along with a silent T > A mutation in the 5’ donor splice site at 2L:3499999 (‘tim-retained-ssM’) (Figure 5A). All constructs were transfected into S2 cells and assayed for their ability to drive expression of TIM protein. Intron inclusion had a drastic effect on the total levels of TIM, ranging from highest in the condition when no intron was present to no detectable full size TIM in the condition when splicing was blocked with the mutation (Figure 5B). Expression of the construct with the 5’ splice site mutation led to low level production of a shorter TIM isoform that we called TIMtiny. This isoform was entirely absent when tim cDNA lacking the tim-tiny intron was expressed but was produced upon expression of the cDNA construct that included tim-tiny. Therefore, TIMtiny is a truncated TIM isoform generated from the tim mRNA carrying an unspliced tim-tiny intron. Total mRNA levels were not different between these three experimental conditions, pointing to the post-transcriptional regulation of TIM protein abundance (data not shown).
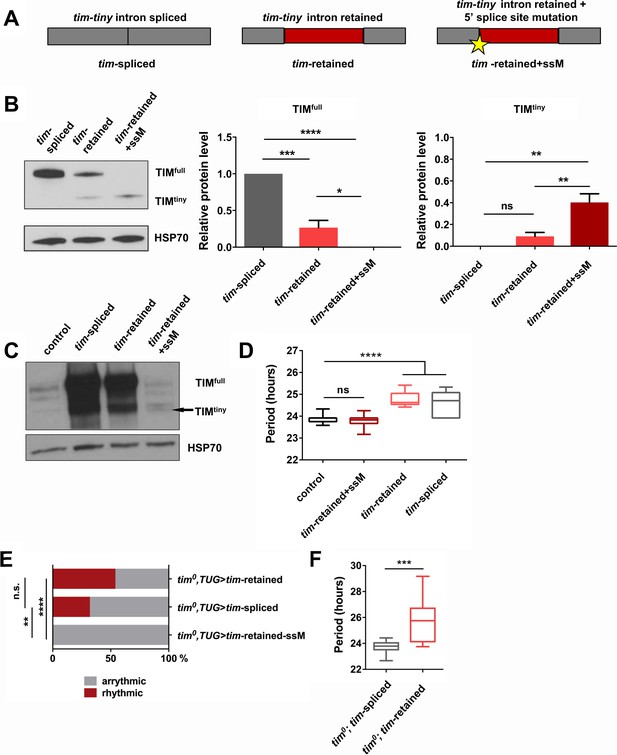
Intron retention in tim decreases TIM levels and affects circadian behavior.
(A) Schematic depiction of three tim cDNA constructs used to assess the effect of tim-tiny retention (red block) on TIM levels. (B) Retention of tim-tiny intron decreases full-length TIM and leads to production of a minor TIMtiny isoform. S2 cells were transfected with constructs described in (A) and western blots of cell lysates were probed with TIM antibody. Western blots are representative of 3 independent experiments. In the panels on the left, total levels of TIM isoforms upon expression of splice-specific cDNA constructs were quantified. TIM levels were normalized to HSP70 and expressed relative to the TIMfull levels in cells overexpressing a fully spliced (tim-spliced) construct. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001 by one-way ANOVA and Holm-Sidak’s post hoc test. Data represent mean ±SEM (n = 3). (C) Western blots of head lysates of flies overexpressing tim cDNA with a 5’ splice site mutation in tim-tiny (TUG; Dcr2 > tim-retained+ssM) reveal production of TIMtiny (arrow) and decrease in TIMfull compared to flies overexpressing intronless tim cDNA (TUG; Dcr2 > timspliced) or tim cDNA that includes tim-tiny ((TUG; Dcr2 > tim retained). All flies were collected at ZT10, when endogenous TIM levels are low in control flies (TUG; Dcr2/+). Western blots are representative of 4 independent experiments. (D) Flies overexpressing tim cDNA constructs with 5’ splice site mutation (TUG; Dcr2 >tim-retained+ssM) do not lengthen circadian period. n = 6–26; n.s., not significant at the 0.05 level; ****p ≤ 0.0001 to control (TUG; Dcr2/+) by one-way ANOVA and Tukey’s post hoc test. (E) TUG-driven expression of tim cDNA, with both ‘tim-spliced’ and ‘tim-retained’ constructs, rescues circadian rhythms in tim0 flies. n.s., not significant at the 0.05 level, **p ≤ 0.01, ****p ≤ 0.0001 by pairwise Fischer’s exact test (n = 28–41). (F) TUG-driven rescue of tim0 circadian rhythms with tim cDNA lacking tim-tiny (tim0,TUG > tim spliced) results in shorter periods than with tim cDNA that includes tim-tiny (tim0,TUG > tim retained). ***p ≤ 0.001 by Student’s t test, n = 10–22.
Next, we overexpressed tim cDNA transgenes (Figure 5A) in flies, using a circadian cell driver (TUG). Normally, overexpression of tim using the Gal4-UAS system reduces rhythmicity and increases free-running periods, likely due to prolonged expression of the excess protein coupled with little negative feedback to downregulate production (Yang and Sehgal, 2001). Overexpression of ‘tim-spliced’ and ‘tim-retained’ cDNAs caused lengthening of free-running periods (Figure 5D). However, the circadian behavior of flies overexpressing the cDNA construct with the 5’ splice site mutation in tim-tiny (TUG; Dcr2 > tim-retained + ssM) was not different from that of the controls (TUG; Dcr2/+). This would fit with our observations from S2 cells, which suggested a loss of full-sized TIM (TIMfull) expression when tim-tiny was primed for selective retention. Thus, we hypothesized that overexpression of the construct with the 5’ splice site mutation did not produce TIMfull in flies. We assayed TIM levels in flies overexpressing different tim cDNA constructs through western blots of fly head lysates collected at ZT10 when the endogenous TIM levels are low in controls (TUG; Dcr2/+) (Figure 5C). As predicted, flies that overexpressed ‘tim-retained-ssM’ cDNA did not have increased TIMfull relative to controls, although both ‘tim-spliced’ and ‘tim-retained’ transgenes expressed abundant amounts of protein. Interestingly, we detected a band, likely corresponding to TIMtiny, in TUG; Dcr2 > tim-retained + ssM flies, which was absent in the TUG; Dcr2/+ controls. These data further corroborate our S2 cell findings and suggest that selective tim-tiny retention acts to reduce overall TIM levels.
To further understand the effect of tim-tiny splicing on circadian behavior, we overexpressed ‘tim-spliced’, ‘tim-retained’ and ‘tim-retained-ssM’ using the TUG driver in the tim0 homozygous background. We hypothesized that differential splicing of tim-tiny is necessary for the maintenance of circadian rhythmicity and so the ‘tim-retained’ construct would rescue the behavioral rhythm most efficiently because it allows for both the splicing and retention of the intron. Additionally, because prp4 knockdown increases the retention of tim-tiny (Figure 4) and prolongs the rhythm (Figure 1), we speculated that the ‘tim-spliced’ construct would rescue with a shorter period than the ‘tim-retained’ construct. As expected, the expression of tim cDNA with a 5’ splice site mutation in tim-tiny (‘tim-retained + ssM’) did not restore rhythms in tim0 flies (Figure 5E). The other two cDNA constructs, ‘tim-retained’ and ‘tim-spliced’ rescued rhythms in 54% and 32% of tim0 flies, respectively. Importantly, there was a significant difference in period length between the flies that were rhythmic (Figure 5F). As discussed above, the UAS-GAL4 system typically over-expresses proteins and so rescues per/tim mutants with longer periods (Yang and Sehgal, 2001), which we observed for the ‘tim-retained’ isoform (~26 hr). Shorter periods were seen with the ‘tim-spliced’ version (Figure 5F), supporting the idea that tim-tiny retention promotes clock delays.
tim splicing is regulated by the clock and by temperature
We next asked if the splicing of tim is under circadian regulation. For this purpose, we generated relative intron retention profiles across regular intervals under both LD and DD conditions (Figure 6A–B). In LD conditions, intron retention of tim-tiny did not display a significant cycle (by JTK analysis) but nevertheless showed a peak at ZT8. In DD conditions, interestingly, we observed robust cycling of tim-tiny with a crest at CT8. It was previously reported that the last (~850 bp) intron in tim (also known as tim-cold) is also sometimes retained (Boothroyd et al., 2007). To determine if the retention of tim-cold is rhythmic, as is the retention of tim-tiny, we profiled tim-cold intron retention across a 24 hr cycle under different conditions. We found that tim-cold retention cycled in both LD and DD with similar phases, such that peak intron retention was during the day in LD and the subjective day in DD. To further verify that the cycling we detected for tim splicing was under clock control, we assayed tim-tiny and tim-cold intron retention profiles in per01 mutant flies. In per01 flies under both LD and DD conditions, the cycling of both tim-tiny and tim-cold intron retention was abolished, indicating that the circadian clock drives circadian oscillations in tim splicing. We suggest that retention of the tim-tiny intron during the day serves to delay the accumulation of TIM protein, in particular when light is not available to degrade TIM.
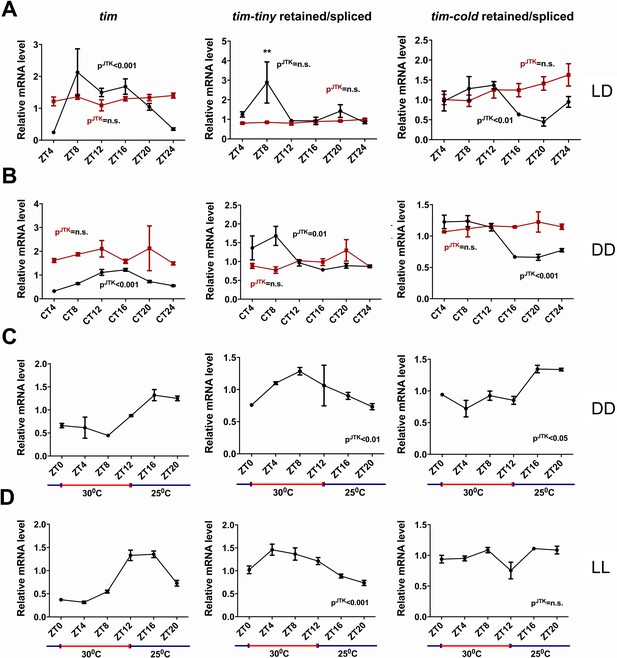
tim splicing is regulated by the clock and by temperature.
Flies were entrained for at least 3 days in 12 hr:12 hr light:dark (LD) conditions and collected in LD (A) or on the first day of transfer to constant darkness (B) at indicated ZT or CT time points, respectively. Splicing of tim-tiny and tim-cold introns was quantified as a ratio of retained to spliced levels using qPCR analysis. Three independent qPCR experiments were performed in triplicate, normalized to rp49 and analyzed using the ∆∆Ct method. pJTK indicates cycling as assessed by JTK cycle analysis for wild-type iso31 flies (black) and per01 circadian mutants (red). tim-tiny intron retention is increased in LD at ZT8 (A) as calculated by two-way ANOVA and Sidak’s post hoc test, p** ≤ 0.01. Wild-type iso31 flies were collected at indicated ZT time points after at least four full days of entrainment in 12 hr:12 hr 30C:25C temperature cycles in constant dark (C) or constant light (D) conditions. Splicing of tim-tiny and tim-cold introns was quantified as a ratio of retained to spliced levels using qPCR analysis. Three independent qPCR experiments were performed in triplicate, normalized to rp49 and analyzed using the ∆∆Ct method. pJTK indicates cycling as assayed by JTK cycle analysis. Error bars = SEM.
It was previously reported that tim-cold retention is regulated by temperature and peaks at colder temperatures (Boothroyd et al., 2007). To determine if the splicing of tim-tiny was similarly sensitive to temperature, we used a temperature entrainment paradigm (12 hr:12 hr 30°C:25°C) under constant photic conditions (Figure 6C–D). In constant dark, temperature cycles were able to drive tim expression as previously reported (Glaser, F.T., and Stanewsky, R., 2005). Additionally, the profile of tim-cold retention was cyclic with the highest retention levels during the colder (25°C) temperatures. Interestingly, tim-tiny retention was robustly rhythmic under these temperature conditions, yet, unlike tim-cold, intron retention was increased by higher temperatures and decreased by lower ones. Thus, higher temperatures, which are typically associated with daytime hours, may reduce TIM levels by regulating the splicing of tim-tiny, while light does so through TIM degradation (Hunter-Ensor et al., 1996; Myers et al., 1996; Zeng et al., 1996; Naidoo et al., 1999). We observed the same relationship and even more robust cycling of tim-tiny under temperature cycles (12 hr:12 hr 30°C:25°C) in constant light conditions (Figure 6D). In these conditions, tim-cold did not cycle, further indicating different regulation of these two splicing events in tim.
Discussion
In this study, we identify a novel alternative splicing (AS) mechanism that affects the pace of endogenous circadian oscillations and implicates PRP4 and other tri-snRNP components in circadian clock regulation. Importantly, our findings contribute to the understanding of a longstanding question in the circadian field, specifically how negative feedback by clock proteins is delayed in order to permit distinct phases, and therefore oscillations, of transcriptional activation and repression. In Drosophila, PER translation does not appear to be delayed relative to its mRNA production in a daily cycle (Chen et al., 1998), but the protein is initially destabilized through its phosphorylation by the DOUBLETIME (DBT) kinase (Price et al., 1998). TIM stabilizes PER by alleviating this effect of DBT, and so accumulation of TIM, which only occurs after dark as TIM is degraded by light, determines the rise of PER. How this mechanism persists when light is not a cycling cue, for instance in constant darkness or in temperature cycles in constant light, was not known. Alternative splicing of the tim-tiny intron may be critical under these conditions.
The circadian profile of tim-tiny intron retention (i.e. high during the daytime) is consistent with it delaying the accumulation of TIM protein. Indeed, this mechanism is particularly robust in DD (Figure 6) and also in temperature cycles in constant light conditions (Glaser and Stanewsky, 2005; Yoshii et al., 2005). The retention of tim-tiny peaks at high temperatures, which correlates with low TIM levels (Glaser, F.T., and Stanewsky, R., 2005; Yoshii et al., 2005). We propose a model (Figure 7) whereby the splice choice at the tim-tiny locus modulates the rate of TIM accumulation, which in turn affects the total levels of nuclear PER and TIM. The intron retention of tim-tiny, therefore, likely contributes to a delay between initial tim expression and accumulation of TIM, ensuring stability and robustness of the circadian oscillator. At the same time, the response of this splicing event to temperature promotes flexibility of the clock.
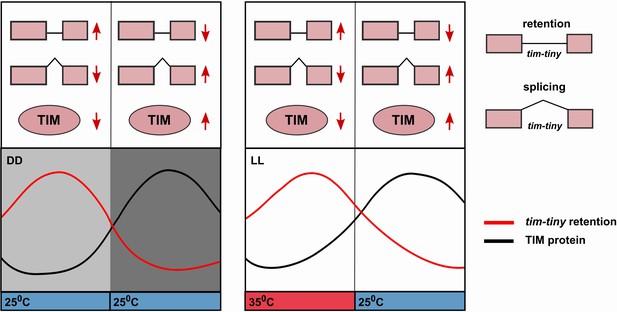
Alternative splicing of the tim-tiny intron promotes oscillations of TIM levels.
The model depicts how retention of the tim-tiny intron, which is increased upon downregulation of prp4, regulates TIM cycling. Both the circadian clock (A) and temperature cycles (B) regulate retention of the tim-tiny intron. (A) Increased retention of tim-tiny during the subjective day (light gray) in dark:dark (DD) conditions serves to decrease TIM levels and delay the accumulation of TIM in the absence of light. (B) Higher temperatures, typically associated with daytime hours, increase tim-tiny retention. Temperature cycles can maintain clock function under constant light conditions (LL), which would otherwise disrupt the clock. Entrainment by temperature appears to be driven by a reduction of TIM protein at the higher temperature (Yoshii, T., et al., 2005). We propose that under temperature cycles, retention of tim-tiny sets the levels of TIM and contributes to maintenance of the molecular clock.
Traditionally, it has been accepted that AS is driven by a set of auxiliary splicing factors, such as serine/arginine-rich (SR) proteins and heterogeneous nuclear ribonucleoproteins (hnRNPs), that act on cis-regulatory splicing modules and recruit U1 and U2 snRNP machinery (Matera and Wang, 2014; Bradley et al., 2015; Han et al., 2011). However, studies from a range of organisms increasingly point to the involvement of the core spliceosomal machinery not only in AS execution but also in splice junction selection (Brooks et al., 2015; Burckin et al., 2005; Clark et al., 2002; Park et al., 2004; Pleiss et al., 2007). Our RNA-seq findings further support the idea that the abundance/stability of at least some components of the core spliceosome has an effect on a select subset of AS events, while constitutive splicing seems to be unperturbed (Supplementary file 3 and 4). A functional network of spliceosomal proteins has been proposed on the basis of their knockdown phenotypes (Papasaikas et al., 2015). According to this model, PRP4 resides within the tri-snRNP regulatory module, which largely coincides with the previously reported physical interactions between the components of the tri-snRNP (Dellaire et al., 2002; Bottner et al., 2005; Schneider et al., 2010). Our findings of similar circadian behavioral effects caused by loss of any of several tri-snRNP components strongly implicate the tri-snRNP complex in circadian regulation (Table 1). It remains to be established how the decreased abundance of tri-snRNP components triggers changes in alternative splicing. We hypothesize that tri-snRNP level/activity is limiting for a subset of AS reactions, likely the ones with weaker splice sites. This hypothesis is supported by two recent studies reporting that (1) Prp4 in fission yeast is necessary to recognize and splice the introns with weak splice sites (Eckert et al., 2016) and (2) decreased availability of mammalian Prpf8 leads to the selective retention of introns that harbor weak 5’ splice sites (Wickramasinghe, V.O., et al., 2015).
We speculate that tri-snRNP components constitute a well-conserved regulatory module for circadian clocks. Conservation of the circadian role of tri-snRNP is suggested by several findings. First, the human homolog of PRP4 was identified as a hit in a genome-wide RNAi screen for regulators of the circadian clock (Zhang et al., 2009). Secondly, SM-like (LSM) proteins that are associated with U6 snRNP were recently shown to regulate circadian rhythmicity in both Arabidopsis and in mammalian cell culture (Perez-Santángelo et al., 2014). Finally, some tri-snRNP components (Prpf8, Prpf31 and SART1) physically associate with mammalian PER2 complexes, further highlighting potential cross-talk between central clock components and the tri-snRNP (Kim et al., 2014).
Although AS has been implicated in the regulation of circadian clocks, it has not been linked to clock function in the manner we report here. In Neurospora crassa, the ratio of alternatively spliced frequency (frq) isoforms determines the robustness of circadian rhythmicity and fine-tunes the period length (Garceau et al., 1997; Liu et al., 1997). In Arabidopsis thaliana, AS regulates the circadian clock by multiple mechanisms, such as the production of new isoforms that competitively inhibit functional clock proteins (Seo et al., 2012) and the modulation of clock RNA levels via the nonsense-mediated decay (NMD) pathway (James et al., 2012; Kwon et al., 2014). In Drosophila, the alternative splicing regulator SR-related matrix protein of 160 kDa (SRm160) modulates PER levels locally in the pacemaker neurons to regulate circadian rhythms (Beckwith et al., 2017). The key theme that emerges from these studies is that AS acts directly on core clock components to set their levels. Alternatively, splicing mechanisms could regulate diurnal rhythmicity of neuronal excitability, as proposed for splicing of BK channels in the suprachiasmatic nucleus (Shelley et al., 2013).
Our study identifies a novel splicing event in tim that can regulate TIM levels in both cell culture and in flies (Figure 5). How does the splicing affect TIM levels? It is unlikely that the retention of tim-tiny leads to NMD-mediated RNA decrease because the flies with prp4 downregulated do not have reduced tim levels (Figure 4—figure supplement 1). On the contrary, overall tim RNA levels tend to be increased in those flies. Additionally, expression of tim cDNA constructs with constitutively retained tim-tiny (Figure 5) decreases TIM levels without altering tim mRNA levels (data not shown). Therefore, we suggest that it is either the tim mRNA translation step or the stability of the truncated TIM isoform, TIMtiny, produced by tim-tiny that is sub-optimal. The 267 amino acids at the C-terminus of TIM that are predicted to be lost in TIMtiny include a putative cytoplasmic localization signal (Saez et al., 2011) as well as a predicted threonine phosphorylation site (Bodenmiller et al., 2007), both of which could significantly change stability and function of TIM. Notably, TIMtiny is typically not detected in western blots of TIM expression in flies, so it appears that tim-tiny retention serves only to reduce the amount of tim RNA that can effectively produce protein.
While studying the effect of tim-tiny retention in isolation can give us a quick snapshot of its importance, overall splicing of tim is considerably more complex. In parallel to tim-tiny, in this study we examined the splicing profile of a previously reported tim-cold intron (Boothroyd et al., 2007). Cycles of tim-tiny and tim-cold intron retention are similar under light:dark and constant dark conditions (Figure 6), but temperature cycles have different effects on the splicing of these two introns. tim-tiny intron retention increases with hot temperature, whereas tim-cold intron retention increases at the onset of the cold cycle (Figure 6). Also, tim-cold retention does not cycle in temperature cycles in LL, suggesting that it does not contribute to rhythmicity under these conditions. tim-tiny intron is upstream of tim-cold, which means that tim-tiny retention should lead to TIM downregulation regardless of the splicing decision at the tim-cold locus. Following differential splicing at tim-tiny locus, tim-cold could get either retained or spliced, introducing an additional regulatory layer. The interplay of AS at the level of these two introns can produce a range of TIM isoforms, the roles of which remain to be elucidated, particularly with respect to temperature entrainment.
In Drosophila, splicing of D. melanogaster per intron 8 (dmip8), the intron located in the 3’ untranslated region (UTR) of per, is regulated by both light and temperature (Collins et al., 2004; Majercak et al., 2004; Majercak et al., 1999). This splicing mechanism allows flies to delay their evening behavior during long photoperiods and/or high temperatures, and might play a role in seasonal adaptation. On a molecular level, dmip8 retention delays per mRNA and PER protein accumulation. While dmip8 retention was not identified in our initial RNA-Seq data (Supplementary file 3 and 4), our follow up qPCR splicing analysis suggested that PRP4 modestly regulates per splicing (Figure 4—figure supplement 2C). dmip8 retention has a small effect on the free-running period length (~25 hr period) (Cheng et al., 1998; Majercak et al., 1999), but it cannot fully account for the period lengthening phenotype we report for flies with downregulated prp4 (Figure 1A–B; Table 1).
Studies in Arabidopsis suggest that the expression of a number of tri-snRNP components is regulated by the circadian clock (Perez-Santángelo et al., 2013). In Drosophila, recent profiling of mRNA cycling in different neuronal clusters detected prp4 mRNA oscillations in DN1 clock neurons and brr2 cycling in LNvs (Abruzzi et al., 2017). While these RNA-Seq findings have not been verified through other approaches, they lead us to hypothesize that diurnal oscillations in tri-snRNP components drive circadian splicing of tim (Figure 6) and potentially other circadian output genes locally in specific clock neurons. This hypothesis is further strengthened by another recent report (Wang et al., 2018) that suggests heterogeneous alternative splicing profiles for different circadian neuronal groups. In addition to changes in total levels of PRP4, circadian regulation of its kinase activity could contribute to differential splicing of tim over the course of the day. We establish a role for PRP4 in LNvs, the key pacemaker cluster necessary for the maintenance of circadian cycles under constant dark conditions (Figure 1). However, based on our findings that splicing of tim is regulated by temperature and persists in constant light (Figure 6), we speculate that PRP4 also functions in DN1s, clock cells implicated in temperature sensing and entrainment (Yadlapalli et al., 2018; Zhang et al., 2010). In summary, while much of the focus in the circadian field has been on transcriptional or post-translational control, our findings indicate a critical role for alternative splicing, perhaps in a cell-type-specific manner.
Materials and methods
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Drosophila melanogaster) | prp4 | NA | FLYB:FBgn0027587 | |
| Gene (Drosophila melanogaster) | timeless (tim) | NA | FLYB:FBgn0014396 | |
| Gene (Drosophila melanogaster) | period (per) | NA | FLYB:FBgn0003068 | |
| Gene (Drosophila melanogaster) | prp8 | NA | FLYB:FBgn0033688 | |
| Gene (Drosophila melanogaster) | brr2 | NA | FLYB:FBgn0263599 | also known as l(3)72Ab |
| Gene (Drosophila melanogaster) | prp3 | NA | FLYB:FBgn0036915 | |
| Gene (Drosophila melanogaster) | prp31 | NA | FLYB:FBgn0036487 | |
| Strain, strain background (Drosophila melanogaster) | iso31 | from laboratory stocks | NA | |
| Genetic reagent (Drosophila melanogaster) | per01 | Bloomington Drosophila Stock Center (BDSC) | FLYB:FBal0013649 | |
| Genetic reagent (Drosophila melanogaster) | tim0 | BDSC | FLYB:FBal0035778 | |
| Genetic reagent (Drosophila melanogaster) | TUG (Tim-UAS-Gal4) | BDSC | FLYB:FBtp0011839 | |
| Genetic reagent (Drosophila melanogaster) | pdfGal4; pdfG4 | BDSC | FLYB:FBtp0011844 | |
| Genetic reagent (Drosophila melanogaster) | elavGal4; elavG4 | BDSC | BDSC:25750 | |
| Genetic reagent (Drosophila melanogaster) | GMRGal4; GMR | BDSC | FLYB:FBti0002994 | |
| Genetic reagent (Drosophila melanogaster) | prp4RNAi(GD) | Vienna Drosophila Resource Center (VDRC) | VDRC:27808 | |
| Genetic reagent (Drosophila melanogaster) | prp4RNAi(KK) | VDRC | VDRC:107042 | |
| Genetic reagent (Drosophila melanogaster) | prp8RNAi(GD) | VDRC | VDRC:18565 | |
| Genetic reagent (Drosophila melanogaster) | prp3RNAi(GD) | VDRC | VDRC:25547 | |
| Genetic reagent (Drosophila melanogaster) | prp3RNAi(KK) | VDRC | VDRC:103628 | |
| Genetic reagent (Drosophila melanogaster) | prp31RNAi(KK) | VDRC | VDRC:103721 | |
| Genetic reagent (Drosophila melanogaster) | brr2RNAi(KK) | VDRC | VDRC:110666 | |
| Genetic reagent (Drosophila melanogaster) | prp82e1 | BDSC | FLYB: FBal0190235; BDSC:25905 | |
| Genetic reagent (Drosophila melanogaster) | prp82e2 | BDSC | FLYB:FBal0190015; BDSC:25912 | |
| Genetic reagent (Drosophila melanogaster) | brr2e03171 | BDSC | FLYB:FBti0041681; BDSC:18127 | |
| Genetic reagent (Drosophila melanogaster) | UAS-Dicer2; Dcr2 | BDSC | FLYB:FBtp0036672 | |
| Genetic reagent (Drosophila melanogaster) | UAS-tim-spliced; tim-spliced | this paper | NA | generated by the site-specific PhiC31 Integration System (Rainbow Transgenics) using the attP on the 3rd chromosome; pUAST-tim-spliced plasmid was used for injection |
| Genetic reagent (Drosophila melanogaster) | UAS-tim-retained; tim-retained | this paper | NA | generated by the site-specific PhiC31 Integration System (Rainbow Transgenics) using the attP on the 3rd chromosome; pUAST-tim- retained plasmid was used for injection |
| Genetic reagent (Drosophila melanogaster) | UAS-tim-retained+ssM; tim-retained+ ssM | this paper | NA | generated by the site-specific PhiC31 Integration System (Rainbow Transgenics) using the attP on the 3rd chromosome; pUAST-tim- retained+ssM plasmid was used for injection |
| Cell line (Drosophila melanogaster) | S2 | ATCC (Manassas, VA) | FLYB:FBtc0000181; RRID:CVCL:Z992 | |
| Antibody | guinea pig anti- PER (UP1140) | Garbe et al., 2013 | NA | 1:1000 |
| Antibody | rat anti- TIM (UPR42) | Jang et al., 2015 | NA | 1:1000 |
| Antibody | rabbit anti-PDF (HH74) | Garbe et al., 2013 | NA | 1:500 |
| Antibody | mouse anti-LaminC | Developmental Studies Hybridoma Bank (DSHB) | LC28.26 | 1:500 |
| Antibody | mouse anti-HSP70 | Sigma | Cat# H5147 | 1:5000 |
| Recombinant DNA reagent | pIZ/V5-His plasmid | ThermoFischer | Cat# V800001 | backbone |
| Recombinant DNA reagent | pBluescript-tim | lab collection | NA | tim sequence contained tim-tiny; used for subcloning |
| Recombinant DNA reagent | tim-spliced; pIZ-tim-spliced | this paper | NA | tim cDNA was subcloned into pIZ-V5 plasmids |
| Recombinant DNA reagent | tim-retained; pIZ-tim-retained | this paper | NA | tim-tiny intron was subcloned into pIZ-tim- spliced vector from pBluescript-tim plasmid |
| Recombinant DNA reagent | tim-retained+ ssM; pIZ-tim-retained+ssM | this paper | NA | generated by mutagenesis of the 5’ splice donor site of tim-tiny intron from pIZ-tim-retained plasmid |
| Recombinant DNA reagent | tim-spliced; pUAST-tim-spliced | this paper | NA | tim cDNA was subcloned from pIZ-tim- spliced into pUAST -attB vector |
| Recombinant DNA reagent | tim-retained; pUAST-tim-retained | this paper | NA | tim cDNA was subcloned from pIZ-tim-retained into pUAST-attB vector |
| Recombinant DNA reagent | tim-retained + ssM; pUAST-tim-retained+ssM | this paper | NA | tim cDNA was subcloned from pIZ-tim- retained+ssM into pUAST-attB vector |
| Sequence- based reagent | tim PP11542 (‘mRNA’) _F | ATGGACTGGTTACTAGCAACTCC | ||
| Sequence- based reagent | tim PP11542 (‘mRNA’) _R | GGTCCTCATAGGTGAGCTTGT | ||
| Sequence- based reagent | per_F | CGTCAATCC ATGGTCCCG | ||
| Sequence- based reagent | per_R | CCTGAAAGACGCGATGGTG | ||
| Sequence- based reagent | clk_F | GGATGCCAATGCCTACGAGT | ||
| Sequence- based reagent | clk_R | ACCTACGAAAGTAGCCCACG | ||
| Sequence- based reagent | prp4_F | CACAAGCAGCATCTTTGTATGG | ||
| Sequence- based reagent | prp4_R | TGTGGAGTCCCACATTCTTG | ||
| Sequence- based reagent | tim-tiny_retained_F | AAACGTGAGTTAAAGTCAACC | ||
| Sequence- based reagent | tim-tiny_retained_R | GAGAGGCACACAGCATATC | ||
| Sequence- based reagent | tim-tiny_spliced_F | CCGCTGGACAAACTCAACCTC | ||
| Sequence- based reagent | tim-tiny_spliced_R | TCGGTATCGCCGAGATCCACG | ||
| Sequence- based reagent | tim-cold_retained_F | GGCTCATGATCATTGCAGCAGC | ||
| Sequence- based reagent | tim-cold_retained_R | ATAGTGGGGCACCCGGATCTC | ||
| Sequence- based reagent | tim-cold_spliced_F | TTAAACAGCGACAATGTCTCTTTGG | ||
| Sequence- based reagent | tim-cold_spliced_R | GAATTGGATCCTCAGTGATAGTGGG | ||
| Sequence- based reagent | tim_non_spanning ('exon')_F | GAAGAACAACGATATTGTGGGAAAG | ||
| Sequence- based reagent | tim_non_spanning ('exon')_R | AGTGGGAGTTGTCAGCAAAG | ||
| Sequence- based reagent | per_retained_F | GAGGACCAGACACAGCACGG | ||
| Sequence- based reagent | per_retained_R | CGGAGGCAATTGCTCACTCGT | ||
| Sequence- based reagent | per_spliced_F | GAGGACCAGACACAGCACGG | ||
| Sequence- based reagent | per_spliced_R | TCGCGTTGATTCGAAGAATCGTT | ||
| Sequence- based reagent | rp49_F | GACGCTTCAAGGGACAGTATCTG | ||
| Sequence- based reagent | rp49_R | AAACGCGGTTCTGCATGAG | ||
| Sequence- based reagent | tim_tinySSdonorT > A_F | CTGGACAAACGAGAGTTAAAGTCAACC | ||
| Sequence- based reagent | tim_tinySSdonorT > A_R | CGGTCCCAGCTTTTTGGC | ||
| Commercial assay or kit | RNeasy Plus Mini Kit | Qiagen | Cat# 74134 | |
| Commercial assay or kit | Superscript II Reverse Transcriptase | ThermoFischer | Cat# 18064014 | |
| Commercial assay or kit | TRIzol Reagent | ThermoFischer | Cat# 15596026 | |
| Commercial assay or kit | Q5 Site-Directed Mutagenesis Kit | NEB | Cat# E0554S | |
| Commercial assay or kit | Effectene Transfection Reagent | Qiagen | Cat# 301425 | |
| Software, algorithm | Graphpad Prism v7 | Graphpad Software | https://www.graphpad.com/ | |
| Software, algorithm | JTK_CYCLE v3 | Hughes et al., 2010 | NA | |
| Software, algorithm | ImageJ | NIH | https://imagej.nih.gov/ij/ | |
| Software, algorithm | ClockLab Software | Actimetrics (Wilmette, IL) | https://actimetrics.com/products/clocklab |
Fly husbandry and stocks
Request a detailed protocolFly stocks and crosses were maintained at room temperature or at 18°C on standard cornmeal molasses medium. Stocks for RNAi overexpression were obtained from VRDC. brr2e03171, prp82e1, prp82e2 mutants were obtained from the Bloomington stock center. iso31, per01 and Gal4 stocks were from the Sehgal lab stock collection. Transgenic lines for tim cDNA overexpression were generated by the site-specific PhiC31 Integration System (Rainbow Transgenics) using the attP on the 3rd chromosome. The DNA for fly embryo injections contained tim cDNA constructs (Figure 5A) subcloned into pUASTattB vectors.
Circadian behavior analysis
Request a detailed protocolFor free-running circadian analysis, male flies were entrained to 12 hr:12 hr light:dark (LD) cycles at 25°C for at least three complete cycles. Flies were loaded into TriKinetics Drosophila Activity Monitor (DAM) system (Trikinetics, Waltham, MA), released into constant darkness and recorded for at least 7 days. Circadian parameters (period and rhythm strength) were determined using Clocklab Software (Actimetrics, Wilmette, IL). Period length was determined with χ2 periodogram analysis. Rhythm strength was determined using Fast Fourier Transform (FFT) values. A fly was considered rhythmic if the FFT value was greater than 0.01.
For analysis of circadian behavior in LD, flies were stably entrained in LD at 25°C for 3 days and their behavior was recorded as described above for the next three subsequent days. The analysis of activity counts was performed using Insomniac3 Software (RP Metrix).
Immunohistochemistry and confocal microscopy
Request a detailed protocolFly brains were dissected in 4% paraformaldehyde (PFA) in 1x phosphate-buffered saline (PBS), fixed in PFA for 20 min at room temperature (RT) and then additionally trimmed of the air sacs and other contaminating tissues in 1xPBST. Dissected brains from each time point were stored in 1xPBST at 4°C until all of the time points were collected (never for more than 8 hr). Once all dissections and fixations were completed, brains were washed in 1xPBST for 20 min at RT, blocked with 5% Donkey Serum (DS) in 1xPBST for 20 min at RT, and incubated with the primary antibodies diluted in 5% DS overnight with gentle shaking at 4°C. The primary antibodies included rat anti-TIM (UPR42, 1:1000), guinea pig anti-PER (UP1140, 1:1000), rabbit anti-PDF (HH74, 1:500) and mouse anti-LaminC (LC28.26 from Developmental Studies Hybridoma Bank, 1:500). After a 30 min wash in 1xPBST at RT, brains were incubated with secondary antibodies diluted in 5% DS at RT. Secondary antibodies were used at 1:500 dilution and included FITC donkey-anti-guinea pig (Rockland), Alexa555 donkey-anti-rabbit (Jackson Immuno), Alexa647 donkey-anti-mouse (Jackson Immuno). Samples were washed for 30 min in 1xPBST at RT, and mounted in VectaShield. Slides were imaged with a Leica SP5 confocal microscope using a 40x oil-immersion objective and a 0.5 μM step size. The signal was adjusted to be not saturated as determined by QLUT parameters. After this original adjustment, all the settings were kept constant for a given experimental set. ImageJ software was used for analysis.
Western blot analysis
Request a detailed protocolWestern blot assays with fly heads were performed as previously described (Garbe et al., 2013). Briefly, 7–10 fly heads per genotype/condition were lysed in 1x Passive Lysis Buffer (Promega), supplemented with protease and phosphatase inhibitors. For S2 cell extracts, 48 hr after transfection, cells were collected and lysed in the same buffer as described for fly heads. The following primary antibodies were used: anti-PER (UP1140, 1:1000), anti-TIM (UPR42, 1:1000) and anti-HSP70 (Sigma, 1:5000).
Plasmids and S2 cell culture
Request a detailed protocolFor cell culture expression, tim cDNA was subcloned into pIZ-V5 plasmids. Using standard restriction enzyme cloning technique, the tim-tiny intron was subcloned into pIZ-V5 vectors carrying intronless tim cDNA from the pBluescript-tim plasmid. Mutagenesis of the 5’ splice donor of tim-tiny intron site was performed with primers catalogued in the Key resources table using Q5 Site-Directed Mutagenesis Kit (NEB).
S2 cells were cultured in a standard Schneider medium (Invitrogen) supplemented with 10% FBS (Sigma). Transfection was done using an Effectene kit (Qiagen) according to the manufacturer’s protocol.
Quantitative RT-PCR
Request a detailed protocolFlies were collected on dry ice at indicated time points and stored at −80°C until all of the time points were collected (within 24 hr). Fly heads were then collected on dry ice and homogenized with TRIzol (ThermoFischer) on ice using standard protocols. Following the phase separation, the aqueous phase was transferred into a new tube, mixed with an equal volume of 70% ethanol and loaded directly onto the RNeasy mini kit columns (Qiagen). The rest of the RNA isolation was done according to the manufacturer’s protocol. On-column DNase digestion (Qiagen) for 15 min at RT was always included. cDNA was generated with Superscript II (Invitrogen), according to the manufacturer’s protocol. Quantitative RT-PCR reaction was performed in a ViiA7 Real-Time PCR system (Applied Biosystems) using SYBR Green Master Mix (Applied Biosystems) with gene specific primers. Relative gene expression was calculated using the ΔΔCt method with rp49 as normalization control.
RNA-Sequencing and data analysis
Request a detailed protocolRNA extraction was performed as described in the ‘Quantitative RT-PCR’ section above. Tapestation (Agilent) was used to ensure that all of RNA samples were of high-quality (RIN >8). five samples per each genotype were selected and prepared with Lexogen’s SENSE mRNA-Seq library Prep Kit. Illumina Next-Generation Sequencing (NextSeq 500) with 300pb paired-end high output run (at ~50M reads per sample) was performed by the Genomics Facility at the Wistar Institute, Philadelphia, PA.
The RNA-seq reads were aligned to the Drosophila genome (dm6.BDGP6.v88) using STAR version 2.5.3a (Dobin et al., 2013). Normalization and quantification were performed with the PORT version 0.8.2a-beta pipeline (Grant, 2018) which first removes reads that map to ribosomal RNA sequences or mitochondrial DNA and then uses a read re-sampling strategy for normalization to minimize unwanted variance such as differences in sequencing depth among the samples. PORT normalization was performed before quantification, at the aligned read level. After normalization, the quantification of features (genes, exons, introns, junctions) was done with respect to the Ensemblv88 annotation. The differential expression analysis was performed using the R Bioconductor package limma-voom (Ritchie et al., 2015). General pathway enrichment analyses were performed using DAVID (https://david.ncifcrf.gov/). The top 743 differentially expressed genes upon prp4 downregulation, corresponding to FDR ≤ 0.05, were used for pathway enrichment analysis. Differential splicing analysis for prp4 knockdown samples (with respect to the control) was performed using CASH (Wu et al., 2018). Full transcript quantification was done using Cufflinks-2.2, and differential psi (percent spliced in) analysis was performed for various gene isoforms to identify alternate splicing events (Trapnell et al., 2013; Trapnell et al., 2010).
Quantification and statistical analysis
Request a detailed protocolThe statistical parameters are included in the legends of each figure. JTK_CYCLEv3.1 was run in R for circadian statistical analyses. GraphPad Prism was used for all other statistical tests.
Data deposition
Request a detailed protocolThe RNA-Seq data generated in this work are freely available at the Gene Expression Omnibus (GEO) standard repository (accession # GSE115163).
Data availability
Sequencing data have been deposited in GEO under accession code GSE115163.
-
NCBI Gene Expression OmnibusID GSE115163. RNAseq of prp4 knockdown in Drosophila.
References
-
Circadian organization of behavior and physiology in DrosophilaAnnual Review of Physiology 72:605–624.https://doi.org/10.1146/annurev-physiol-021909-135815
-
Regulation of alternative splicing in Drosophila by 56 RNA binding proteinsGenome Research 25:1771–1780.https://doi.org/10.1101/gr.192518.115
-
Exploring functional relationships between components of the gene expression machineryNature Structural & Molecular Biology 12:175–182.https://doi.org/10.1038/nsmb891
-
Alterations of per RNA in noncoding regions affect periodicity of circadian behavioral rhythmsJournal of Biological Rhythms 13:364–379.https://doi.org/10.1177/074873098129000192
-
Mammalian PRP4 kinase copurifies and interacts with components of both the U5 snRNP and the N-CoR deacetylase complexesMolecular and Cellular Biology 22:5141–5156.https://doi.org/10.1128/MCB.22.14.5141-5156.2002
-
Circadian cycling of a PERIOD-beta-galactosidase fusion protein in Drosophila: evidence for cyclical degradationJournal of Biological Rhythms 12:157–172.https://doi.org/10.1177/074873049701200207
-
STAR: ultrafast universal RNA-seq alignerBioinformatics 29:15–21.https://doi.org/10.1093/bioinformatics/bts635
-
A plastic clock: how circadian rhythms respond to environmental cues in DrosophilaMolecular Neurobiology 38:129–145.https://doi.org/10.1007/s12035-008-8035-y
-
Temperature synchronization of the Drosophila circadian clockCurrent Biology 15:1352–1363.https://doi.org/10.1016/j.cub.2005.06.056
-
Pre-mRNA splicing: where and when in the nucleusTrends in Cell Biology 21:336–343.https://doi.org/10.1016/j.tcb.2011.03.003
-
Molecular genetic analysis of circadian timekeeping in DrosophilaAdvances in genetics 74:141–173.https://doi.org/10.1016/B978-0-12-387690-4.00005-2
-
Robust circadian rhythmicity of drosophila melanogaster requires the presence of lateral neurons: a brain-behavioral study of disconnected mutantsJournal of Comparative Physiology A: Sensory, Neural, and Behavioral Physiology 182:435–453.https://doi.org/10.1007/s003590050192
-
JTK_CYCLE: an efficient nonparametric algorithm for detecting rhythmic components in genome-scale data setsJournal of Biological Rhythms 25:372–380.https://doi.org/10.1177/0748730410379711
-
Cloning of human PRP4 reveals interaction with Clk1Journal of Biological Chemistry 276:32247–32256.https://doi.org/10.1074/jbc.M103790200
-
Splicing of the period gene 3'-terminal intron is regulated by light, circadian clock factors, and phospholipase CMolecular and Cellular Biology 24:3359–3372.https://doi.org/10.1128/MCB.24.8.3359-3372.2004
-
A day in the life of the spliceosomeNature Reviews Molecular Cell Biology 15:108–121.https://doi.org/10.1038/nrm3742
-
Conserved regions of the timeless (tim) clock gene in Drosophila analyzed through phylogenetic and functional studiesGenetics 148:815–825.
-
Genomic analysis reveals novel connections between alternative splicing and circadian regulatory networksBriefings in Functional Genomics 12:13–24.https://doi.org/10.1093/bfgp/els052
-
Human PRP4 kinase is required for stable tri-snRNP association during spliceosomal B complex formationNature Structural & Molecular Biology 17:216–221.https://doi.org/10.1038/nsmb.1718
-
Phosphorylation of a constitutive serine inhibits BK channel variants containing the alternate exon "SRKR"The Journal of General Physiology 142:585–598.https://doi.org/10.1085/jgp.201311072
-
Differential analysis of gene regulation at transcript resolution with RNA-seqNature Biotechnology 31:46–53.https://doi.org/10.1038/nbt.2450
-
CASH: a constructing comprehensive splice site method for detecting alternative splicing eventsBriefings in Bioinformatics 19:905–917.https://doi.org/10.1093/bib/bbx034
-
Temperature cycles drive Drosophila circadian oscillation in constant light that otherwise induces behavioural arrhythmicityEuropean Journal of Neuroscience 22:1176–1184.https://doi.org/10.1111/j.1460-9568.2005.04295.x
-
Speed control: cogs and gears that drive the circadian clockTrends in Neurosciences 35:574–585.https://doi.org/10.1016/j.tins.2012.05.007
Article and author information
Author details
Funding
NIH Clinical Center (UL1TR001817)
- Gregory R Grant
National Institute of Neurological Disorders and Stroke (R37NS048471)
- Amita Sehgal
The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Acknowledgements
We thank David Garbe for his assistance with the original kinase screen. The work was supported by an NIH grant, R37NS048471, to AS and UL1TR001817 (Institutional Clinical and Translational Science Award), to Garret Fitzgerald.
Copyright
© 2018, Shakhmantsir et al.
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 2,069
- views
-
- 313
- downloads
-
- 41
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Citations by DOI
-
- 41
- citations for umbrella DOI https://doi.org/10.7554/eLife.39821






