Communication between distinct subunit interfaces of the cohesin complex promotes its topological entrapment of DNA
Figures
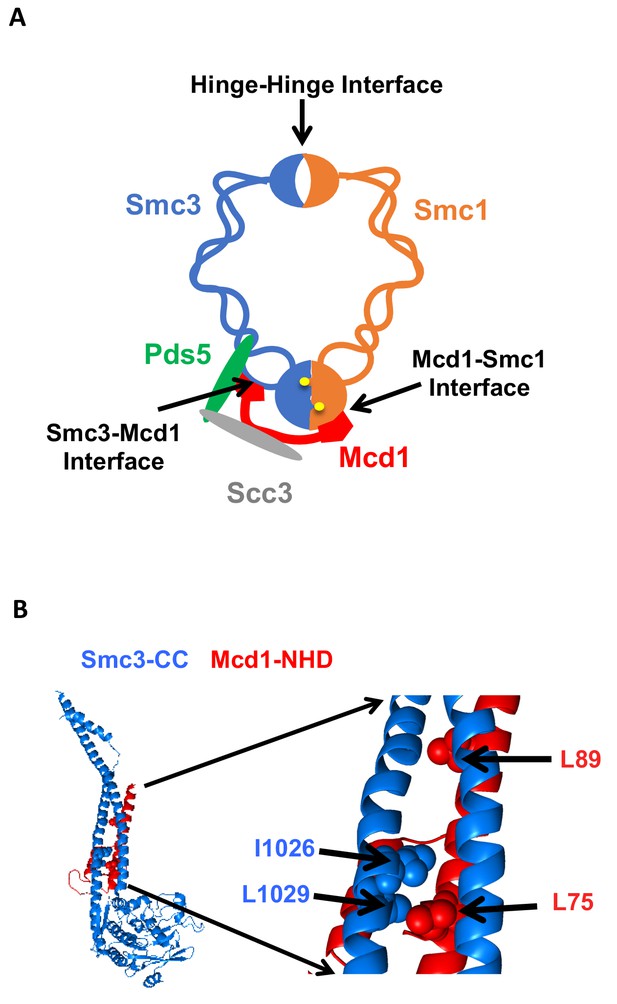
Cohesin structure.
(A) Cartoon showing the cohesin complex and Pds5p. The three interfaces that can be chemically crosslinked to trap DNA within the Smc1p-Smc3p-Mcd1p trimer are marked by arrows. These are 1) Smc3p Mcd1p interface, 2) Smc1p hinge Smc3p hinge dimer interface, and 3) Smc1p Mcd1p interface. (B) Crystal structure of the Smc3p coiled-coil and Mcd1p NHD domain interface. Crystal structure entry PDB 4U × 3 of the Smc3 head domain + short coiled-coil (Smc3 CC; blue) and Mcd1/Scc1 NHD (Mcd1 N; red) is shown. Right Side is enlargement showing essential residues in the interface as spheres.
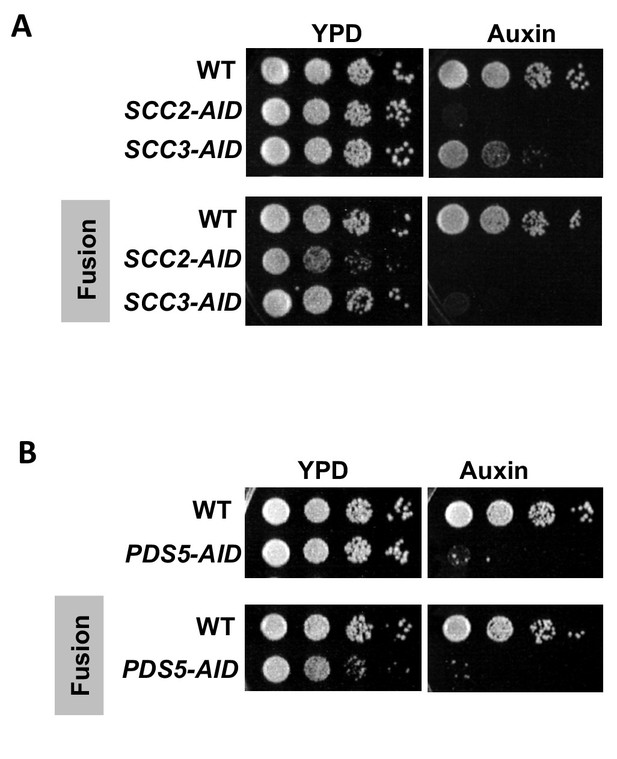
Cohesin regulators are required fusion cohesin function.
(A-B) Haploid strains with normal cohesin or fusion were grown to saturation, then plated at 10-fold serial dilution onto YPD alone or containing auxin (750 μM) and incubated at 23°C for 3 days. (A) Scc2p-AID and Scc3p-AID depletion causes in inviability in normal and fusion cohesion strains. Top panel shows strains with normal cohesin, WT (VG3620-4C), containing SCC2-3V5-AID (VG3630-7A) or SCC3-3V5-AID (VG3808-1A). Bottom panel shows strains with fusion cohesin, WT (VG3940-2D) or with SCC2-3V5-AID (VG3945-1A) or SCC3-3V5-AID (VG3946-7B). (B) Pds5p-AID depletion causes in inviability in normal and fusion cohesion strains. Top panel shows strains with normal cohesin, WT (VG3620-4C) or containing PDS5-3V5-AID2 (VG3954-10C). Bottom panel shows strains with fusion cohesin, WT (VG3940-2D) or containing PDS5-3V5-AID2 (VG3955-4D).
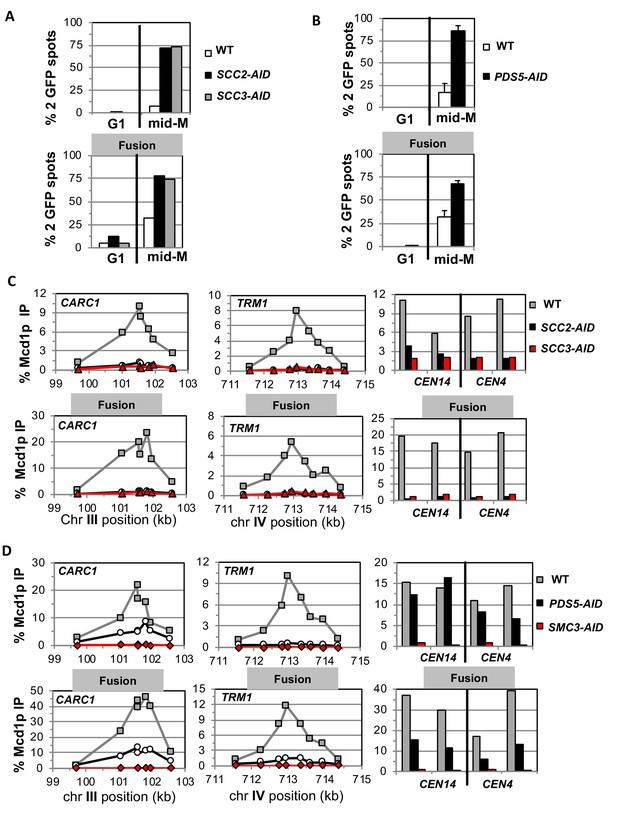
Fusion cohesin requires cohesin regulators for sister chromatid cohesion and cohesin binding to DNA.
Haploids strains with normal cohesin or fusion cohesin from Figure 2 were grown to mid-log phase then arrested in G1 using α factor, auxin added to induce loss of AID-tagged proteins then released into media containing nocodazole and auxin to arrest in mid-M phase under AID depletion conditions as described for synchronous mid-M phase arrest in Materials and methods. Cells were fixed and processed to monitor loss of sister chromatid cohesion and for ChIP to monitor cohesin DNA binding. (A-B) Cohesion loss monitored in mid-M phase cells. The number of GFP spots was scored in G1 arrested cells and mid-M phase cells. The percentage of cells with 2 GFP spots was plotted. 100–200 cells were scored for each data point and data was generated from two independent experiments. (A) Scc2p-AID or Scc3p-AID depletion induces cohesion loss in strains with normal or fusion cohesin. Top panel is strains with normal cohesin and bottom panel is fusion cohesin strains. WT (White), SCC2-AID (black) and SCC3-AID (gray). (B) Pds5p-AID depletion induces cohesion loss in strains with normal or fusion cohesin. Top panel is strains with normal cohesin and bottom panel is fusion cohesin strains. WT (White) and PDS5-AID (black). (C-D) mid-M phase arrested cells fixed and processed for ChIP using anti-Mcd1p antibodies as described in Materials and methods. Mcd1p binding was assessed by qPCR and presented as percentage of total DNA using the same primer pairs at each site. Left Panel is chromosome III peri-centric region (CARC1), middle panel is chromosome IV arm CAR region (TRM1) and right panel is regions immediately adjacent to CEN4 and CEN14. (C) Scc2p-AID or Scc3p-AID depletion induces loss of fusion cohesin and normal cohesin binding to DNA. Top panel is strains with normal cohesin and bottom panel is fusion cohesin strains. WT (gray), SCC2-AID (black) and SCC3-AID (red). (D) Pds5p-AID depletion reduces the amount of fusion cohesin and normal cohesin binding to DNA. Top panel is strains with normal cohesin and bottom panel is fusion cohesin strains. WT (gray), PDS5-AID (black) and SMC3-AID (red).
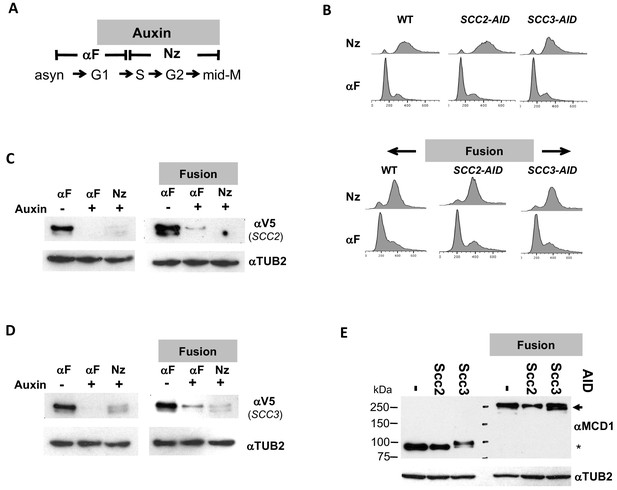
Characterizing the effect of Scc2p-AID or Scc3p-AID depletion in normal and fusion cohesin strains.
(A) Schematic of regimen used to synchronously arrest cells in mid-M phase. (B–D) Strains with normal cohesin or fusion cohesin alone or bearing Scc2p-AID or Scc3p-AID from Figure 2A were synchronously arrested in mid-M then processed to generate data presented in Figure 3A and C. (B) FACS to confirm arrest of cells. (C–D) Assessing depletion of Scc2p-AID and Scc3p-AID assessed. Protein extracts were made from G1 arrested cells before and after auxin treatment and from cells synchronous arrested in mid-M then subjected to SDS-PAGE and analyzed by western blot. 3V5-AID-tagged protein depletion was monitored using mouse antibodies against V5 (aV5) and rabbit anti-tubulin for a loading control (αTUB2). (C) Scc2p-3V5-AID depletion. (D) Scc3p-3V5-AID depletion (E) Mcd1p and Smc3pMcd1p fusion levels in mid-M arrested cells. Protein extracts from mid-M phase cells in C and D were analyzed by western blot. Mcd1p and fusion Smc3-Mcd1p fusion protein levels were monitored using rabbit antibodies against Mcd1p (aMCD1) and rabbit anti-tubulin for a loading control (αTUB2).
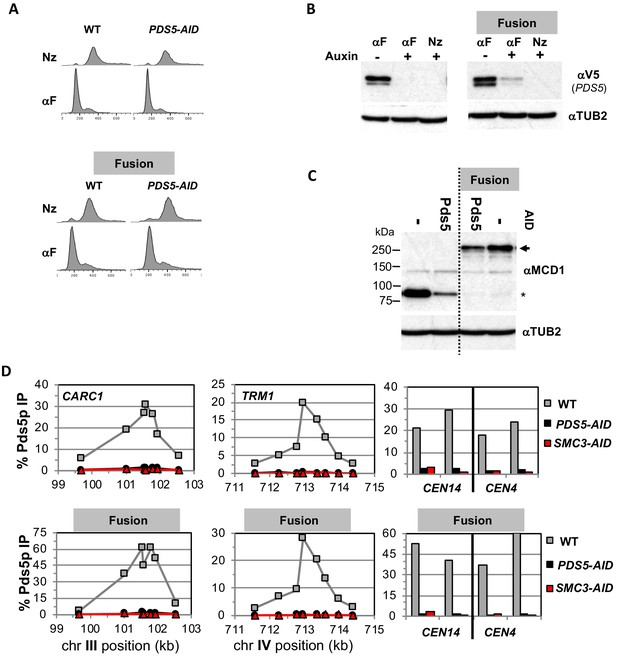
Characterizing the effects of Pds5p-AID depletion in normal and fusion cohesin strains.
Strains with normal cohesin or fusion cohesin alone or bearing Pds5p-AID from Figure 2B were synchronously arrested in mid-M then processed to generate data presented in described in Figure 3B and D. (A) FACS to confirm arrest of cells. (B) Assessing depletion of Pds5p-AID. Protein extracts were made from G1 arrested cells before and after auxin treatment and from synchronous arrest in mid-M were subjected to SDS-PAGE and analyzed by western blot. (B) Pds5p-3V5- AID depletion was monitored using mouse antibodies against V5 (αV5) and rabbit anti-tubulin for a loading control (aTUB2) (C) Mcd1p and Smc3p-Mcd1p fusion levels in mid-M arrested cells. Protein extracts from mid-M phase cells in B were analyzed by western blot. Mcd1p and Smc3-Mcd1p fusion protein levels were monitored using rabbit antiMcd1p antibodies (αMCD1) and rabbit anti-tubulin for a loading control (αTUB2). (D) Pds5p binding to chromosomes using ChIP. Mid-M phase cells from B-C were fixed and processed for ChIP to assess Pds5p binding after Pds5p depletion as described in Figure 3. Pds5p binding was assayed using rabbit anti-Pds5p antibodies (αPDS5). WT (gray), PDS5-AID (black) and SMC3-AID (red).
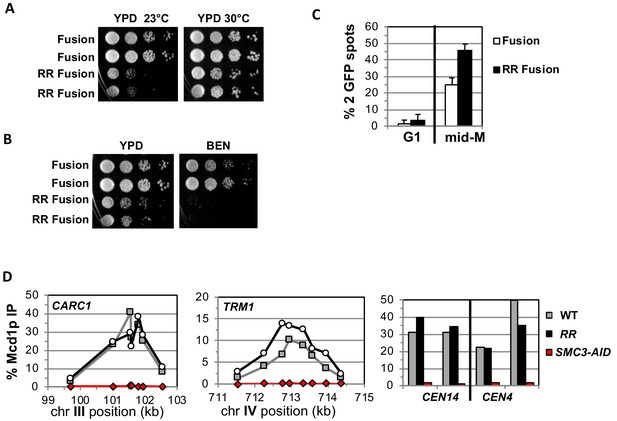
Fusion cohesin with the smc3 K112R K113R mutations are viable but have defects in growth defects in both growth and sister chromatid cohesion.
(A–B) Fusion cohesion bearing smc3-K112R, K113R (RR) mutations are viable but cold sensitive and benomyl sensitive. Haploid strains with wild-type fusion cohesin (VG3940-2D) or RR fusion (VG3930-5C) were grown to saturation at 30°C then plated at 10-fold serial dilution onto (A) YPD at either 23°C or 30°C and incubated for 3 days. (B) YPD alone or YPD containing benomyl at 12.5 μg/ml (BEN) then incubated for 23°C for 4 days. (C–D) Strains in A-B were synchronously arrested in mid-M as described in Figure 3 above except cells were grown at 30°C and auxin was omitted. Cells were fixed and processed to assess cohesion and for ChIP. (C) The RR mutation in fusion cohesin causes an increased defect in cohesion fusion. Cohesin was scored and plotted as described in Figure 3. Fusion cohesin (White) and RR fusion cohesin (black). (D) Fusion and RR fusion cohesin bind DNA at similar levels. mid-M phase arrested cells were fixed and processed for ChIP using anti-Mcd1p antibodies as described in Materials and methods and Figure 3. Left Panel is chromosome III peri-centric region (CARC1), middle panel is chromosome IV arm CAR region (TRM1) and right panel is regions immediately adjacent to CEN4 and CEN14. WT fusion (gray), RR fusion (black) and SMC3-AID (red).
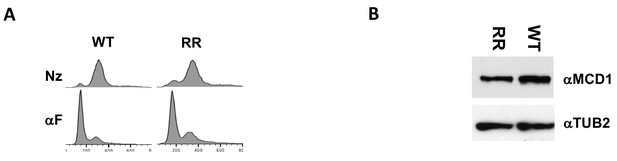
Characterization of fusion wild-type and smc3-K112R K113R (RR) mutations.
Strains with fusion cohesin or RR fusion cohesin in Figure 4 were grown treated as described. (A) FACS to confirm arrest of cells. (B) Smc3p-Mcd1p fusion levels in mid-M arrested cells. Protein extracts from mid-M phase cells were analyzed by western blot. Smc3- Mcd1p fusion protein levels were monitored using rabbit anti-Mcd1p antibodies (αMCD1) and rabbit anti-tubulin for a loading control (αTUB2).
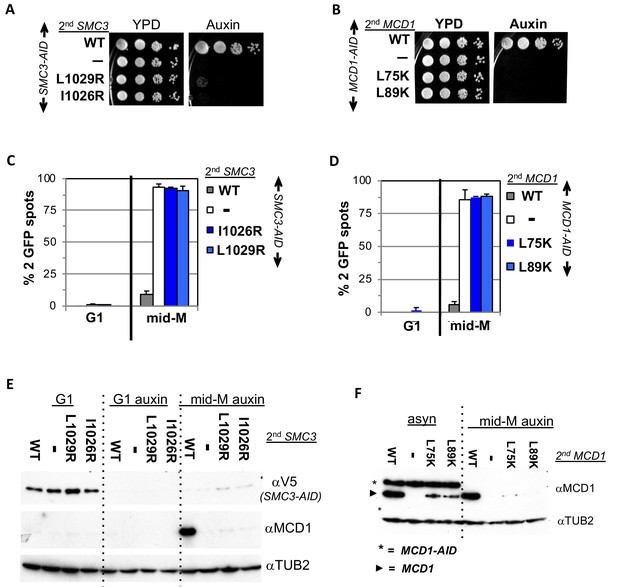
Smc3p coiled-coil and Mcd1p NHD interface residues are required for cohesin function and integrity.
(A) Viability loss of Smc3p interface mutant strains. Haploid SMC3-AID strain alone (VG3651-3D) or containing either wild-type (WT; BRY474), smc3-I1026R (VG3905-7A) or smc3-L1029R (BRY492) were grown and plated as described in Figure 2. (B) Viability loss of Mcd1p interface mutant strains. Haploid MCD1-AID strain alone (VG3902-3A) or containing, either wild-type (WT; VG3914-2C), mcd1-L75K (VG3916-5B) or mcd1-L89K (VG3918-9D) were grown and plated as described in Figure 2. (C–D) Haploids in A and B were synchronously arrested in mid-M phase as described in Figure 3. The number of GFP spots was scored in G1 arrested cells and mid-M phase cells. The percentage of cells with 2 GFP spots was plotted. 100–200 cells were scored for each data point and data was generated from two independent experiments. (C) Cohesion loss in smc3 interface mutant mid-M cells. SMC3-AID alone (white), or containing WT (gray), smc3-I1026R (dark blue) and smc3-L1029R (light blue). (D) Cohesion loss in mcd1 mutant mid-M cells. MCD1-AID strain alone (white) or containing, either WT (gray), mcd1-L75K (dark blue) or mcd1-L89K (light blue). (E–F) Protein extracts from the synchronous arrest regimen in A and B were made from G1 cells, auxin treated G1 cells and mid-M cells then subjected to western blot analysis. (E) Mcd1p is degraded in mid-M phase smc3 interface mutants. Top panel detects Smc3p-3V5-AID (αV5), middle panel detects Mcd1p (αMcd1). Tubulin (αTub2p; bottom panel) was used as a loading control. As a control we showed in (Figure 6—figure supplement 2) that the smc3-L1029R mutation did not affect its protein levels by comparing 6 HA-epitope-tagged Smc3p (VG3943-1C) and Smc3p-L1029R (VG3944-3D). (F) Mcd1p interface mutants are degraded in mid-M cells. Top panel detects Mcd1p (αMcd1) with star indicating Mcd1p-AID and arrow head Mcd1p WT and interface mutants, bottom panel detects Tubulin (αTub2p). .
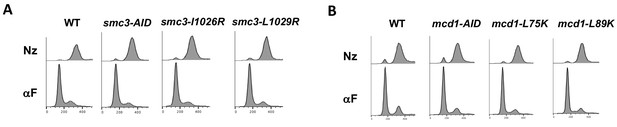
FACS analysis showing cell cycle arrest of normal cohesin strains bearing interface mutations.
Strains with normal cohesin alone or bearing interface mutants were synchronously arrested in mid-M as described in Figure 3. (A) smc3 interface mutants in normal cohesin strains from Figure 5. (B) mcd1 interface mutants in normal cohesin strains from Figure 5.
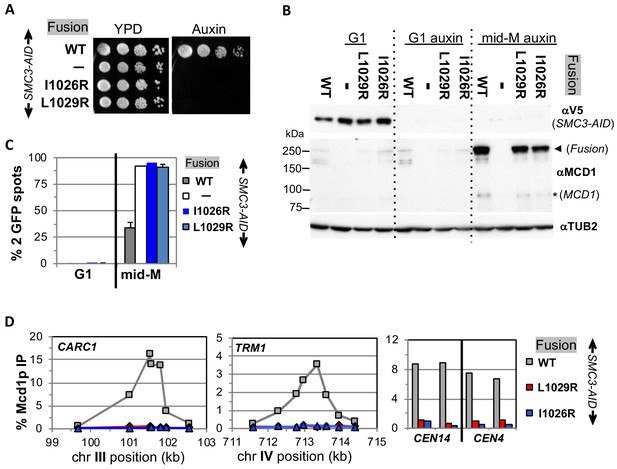
Fusion cohesin requires Smc3p coiled coil residues for function.
(A) Viability loss of Smc3p coiled coil mutant fusion strains. Haploid SMC3-AID strain alone (VG3651-3D) or containing fusion cohesin, either wild-type (WT; VG3694-7C), smc3-I1026R (VG3908-17B) or smc3-L1029R (VG3872-3B) were grown and plated as described in Figure 2. (B–D) Haploids in A were synchronously arrested in mid-M phase as described in Figure 3. (B) Fusion WT and smc3 interface mutant proteins are present in mid-M cells. Protein extracts from G1 and auxin treated G1 and mid-M phase arrested cells were subjected to western blot analysis using antibodies against V5 to detect Smc3p-3V5-AID (αV5; top panel) and against Mcd1p (αMcd1p; middle panel). Arrow head indicates fusion protein and star (*) indicates normal Mcd1p. Tubulin (αTub2p; bottom panel) was used as a loading control. (C) Cohesion is completely lost in smc3 mutant fusion mid-M phase arrested cells. The number of GFP spots was scored in G1 arrested cells and mid-M phase cells. The percentage of cells with 2 GFP spots was plotted. SMC3-AID alone (white), or containing fusion WT (gray), smc3-I1026R (dark blue) and smc3-L1029R (light blue). 100–200 cells were scored for each data point and data was generated from two independent experiments. (D) fusion smc3 interface mutant cohesin fails to bind DNA in mid-M phase. Fusion cohesin binding to chromosomes was assayed by ChIP using anti-Mcd1p antibodies as described in Figure 3. WT fusion (gray), smc3-I1026R fusion (blue) and smc3-L1029R fusion (red).
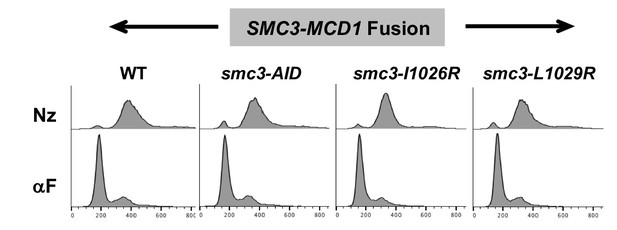
Confirming cell cycle arrest of fusion cohesin bearing smc3 interface mutants.
Strains with fusion cohesin alone or bearing smc3 interface mutants from Figure 6 were synchronously arrested in mid-M as described. FACS analysis was used to confirm arrest.
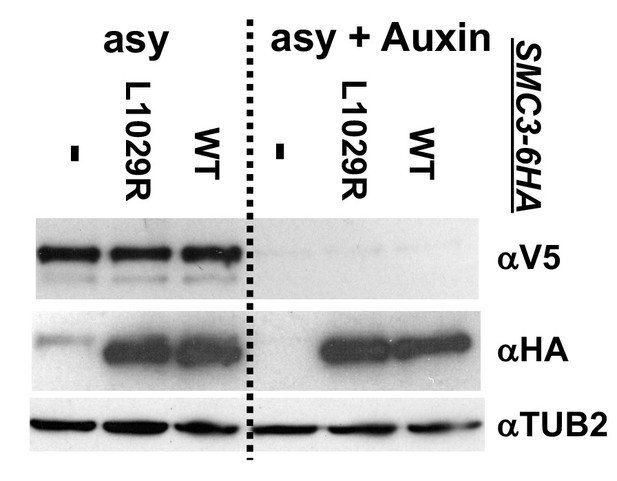
Western blot to assess the effect of smc3-L1029R interface mutant on cohesin levels.
Haploid normal cohesin strains bearing Smc3p-3V5-AID alone (VG3651-3D) or containing either Smc3p-6HA (VG3943-1C) or Smc3p-6HAL1029R (VG3944-3D) were grown asynchronously, auxin added and cells incubated 1 hr. Protein extracts were made before after auxin addition and analyzed by western blot. Smc3p-6HA tagged proteins were monitored using mouse anti-HA antibodies (αHA), Smc3p-3V5-AID depletion was using mouse anti-V5 antibodies (αV5) and rabbit antitubulin for a loading control (αTUB2).
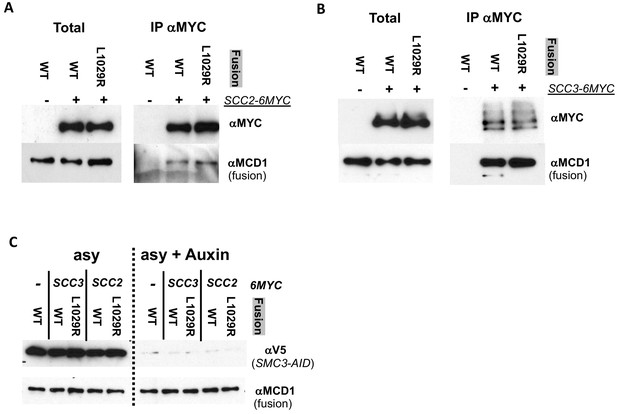
Fusion wild-type and fusion L1029 mutant coimmunoprecipitate with Scc2p and Scc3p equally well.
(A) Fusion wild-type and L1029R mutant cohesin both co-IP with Scc2p. Haploid strain Scc2p-6MYC, Smc3p-3V5-AID bearing wild-type fusion cohesin (VG3952-14C) or fusion L1029R mutant (VG3953-17D) were grown asynchronously then auxin added and cells incubated 1 hr. An co-IP control without any MYC-tagged proteins (VG3694-7C; wild-type fusion and Smc3p-3V5-AID) was also used. Protein extracts were made before after auxin addition. Mouse anti-MYC antibodies were used to immunoprecipitate MYC tagged proteins and analyzed by western blot. Mouse anti-MYC antibodies detected MYC tagged Scc2p (αMYC), rabbit anti-Mcd1p antibodies detected fusion cohesin co-IP detected (αMCD1). (B) Fusion wild-type and L1029R mutant cohesin both co-IP with Scc3p. Haploid strain Scc3p-6MYC, Smc3p-3V5-AID bearing wild-type fusion cohesin (VG3949-5C) or fusion L1029R mutant (VG3950-8D) as well as control without any MYC-tagged proteins (VG3694-7C; wild-type fusion and Smc3p-3V5-AID) were grown and treated as described in A. (C) Smc3p-3V5 depletion to confirm that fusion cohesin is sole cohesion in cells. Extracts from cells in A and B before and after auxin addition were analyzed by Western Blot. Smc3p-3V5-AID depletion was monitored using mouse anti-V5 antibodies (αV5) and fusion cohesin levels monitored using rabbit anti-Mcd1p antibodies (αMCD1).
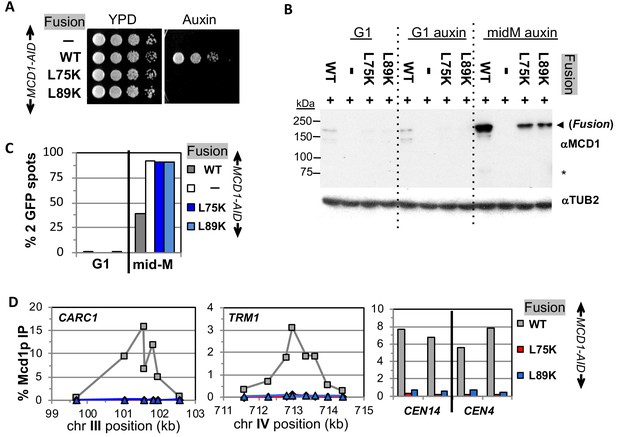
Fusion cohesin requires Mcd1p NHD residues for function.
(A) Viability loss of Mcd1p NHD mutant fusion strains. Haploid MCD1-AID strain alone (VG3902-3A) or containing fusion cohesin, either wild-type (WT; VG3937-2C), mcd1-L75K (VG3938-3A) or mcd1-L89K (VG3939-7B) were grown and plated as described in Figure 2. (B–D) Haploids in (A) were synchronously arrested in mid-M phase as described in Figure 3. (B) Fusion WT and mcd1 NHD mutant proteins are present in mid-M cells. Protein extracts from G1, auxin-treated G1 and mid-M phase cells were subjected to western blot analysis using antibodies against Mcd1p (αMcd1p; top panel). Arrow head indicates fusion protein and star (*) indicates normal Mcd1p. Tubulin (αTub2p; bottom panel) was used as a loading control. (C) Cohesion is completely lost in mcd1 mutant fusion mid-M cells. The number of GFP spots was scored in G1 arrested cells and mid-M phase cells. The percentage of cells with 2 GFP spots was plotted. Haploid MCD1-AID strain alone (white) or containing fusion cohesin, either WT (gray), mcd1-L75K (dark blue) or mcd1-L89K (light blue). 100–200 cells were scored for each data point. (D) fusion mcd1 NHD mutant cohesin fails to bind DNA in mid-M phase. Fusion cohesin binding to chromosomes was assayed by ChIP using anti-Mcd1p antibodies as described in Figure 3. WT (gray), mcd1-L75K (red) and mcd1-L89K (blue).
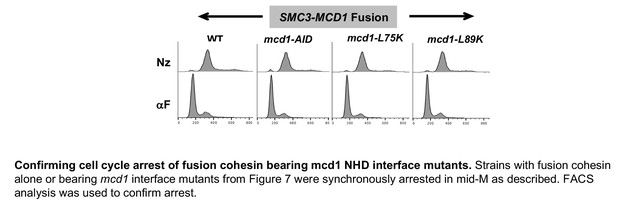
Confirming cell cycle arrest of fusion cohesin bearing mcd1 NHD interface mutants.
Strains with fusion cohesin alone or bearing mcd1 interface mutants from Figure 7 were synchronously arrested in mid-M as described. FACS analysis was used to confirm arrest.
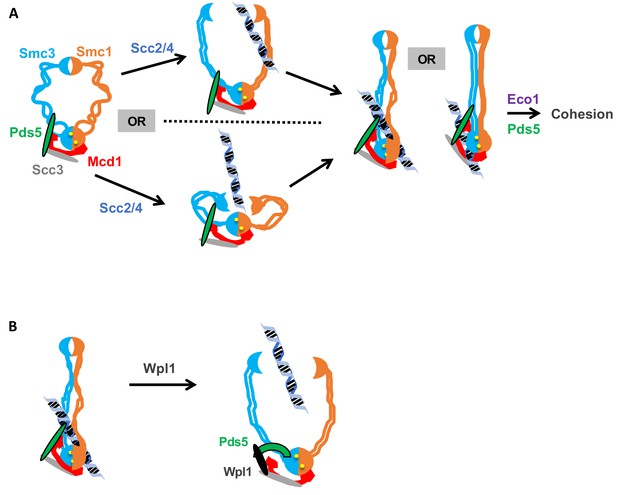
Model for how the Smc3p Mcd1p interface regulates the hinge dimer interface to control cohesin DNA binding.
(A) Left side: cohesin complex in an open ring conformation. Middle top: cohesin loader Scc2p/Scc4p binding at the Smc3p Mcd1p interface (not shown) triggers a conformation change that opens the hinge dimer (half-moons) interface, which allows DNA (blue-black helix) to enters cohesin. Alternatively, this conformational change could generate a folded ring to bring the hinges into close proximity to Scc2/Scc4 bound near the head, thereby enabling it to directly act on the hinges. Right side: DNA interaction with cohesin and Pds5p (green oval) binding at Smc3p Mcd1p interface triggers another conformational change that traps DNA either near the head (left) or in the small lumen formed by Mcd1p binding both the Smc1p head and the Smc3p coiled-coil (right). (B) Left side: Cohesin stably bound to DNA. Right side: Wpl1p (black oval) binds Pds5p (green arc) and cohesin at its Smc3p Mcd1p interface and triggers a conformation change that opens hinge dimer (half-moons) interface and allows DNA to escape. Pds5p has both positive and negative functions so it is depicted differently with the green oval showing its positive role in promoting DNA binding and the green arc showing its negative role as it binds Wpl1p. We note that recent in vitro evidence suggests DNA may first interact with cohesin at the Scc3p and Mcd1p region entry into the trimer. This earlier step is not depicted and has no bearing on our model.
Tables
| Reagent type or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (S. cerevisiae) | NCBITaxon:4932 | this paper | Yeast strains | Supplementary file 1 |
| qPCR primers | DNA primer | IDT | Supplementary file 2 | |
| Antibody | Rabbit polyclonal Anti-Mcd1p | V. Guacci (via Covance) | RbαMcd1p (555) | WB (1:10,000) ChIP (1:1,000) |
| Antibody | Rabbit polyclonal Anti-Pds5p | V. Guacci (via Covance) | RbαPds5p (556) | WB (1:20,000) ChIP (1:1,000) |
| Antibody | Mouse monoclonal Anti-MYC | Roche | MαMYC (9E10) Cat#116666006001 | WB (1:10,000) IP (1:667) |
| Antibody | Mouse monoclonal Anti-HA | Roche | MαHA (12CA5) Cat#11667203001 | WB (1:10,000) |
| Antibody | Mouse monoclonal Anti-V5 | Invitrogen | MαV5 Cat# 46–0705 | WB (1:10,000) |
| Antibody | Goat polyclonal HRP Anti-rabbit | Biorad | Cat# 170–6515 | WB (1:10,000) |
| Antibody | Goat polyclonal HRP Anti-mouse | Biorad | Cat# 170–6516 | WB (1:10,000) |
| Antibody | Rabbit polyclonal Anti-Tub2p | P. Meluh (via Covance) | RbαTUB2 | WB (1:40,000) |
| Dynabeads | Protein A | Invitrogen | Ref# 10002D | IP: Use 50 ul/IP |
| Dynabeads | Protein G | Invitrogen | Ref# 10004D | IP: Use 50 ul/IP |
| Chemical compound | Auxin (3-indole acetic acid) | Sigma | Cat# C9911 | 750 μM for plates 500 μM for liquid |
| Chemical compound | Alpha factor | Sigma | αF (αFactor) Cat# T6901 | |
| Chemical compound | Nocodazole | Sigma | Nz Cat# M1404 |
Additional files
-
Supplementary file 1
Yeast strains.
- https://doi.org/10.7554/eLife.46347.018
-
Supplementary file 2
Primers used for chromatin immunoprecipitation (ChIP).
- https://doi.org/10.7554/eLife.46347.019
-
Transparent reporting form
- https://doi.org/10.7554/eLife.46347.020





