HRI coordinates translation necessary for protein homeostasis and mitochondrial function in erythropoiesis
Figures
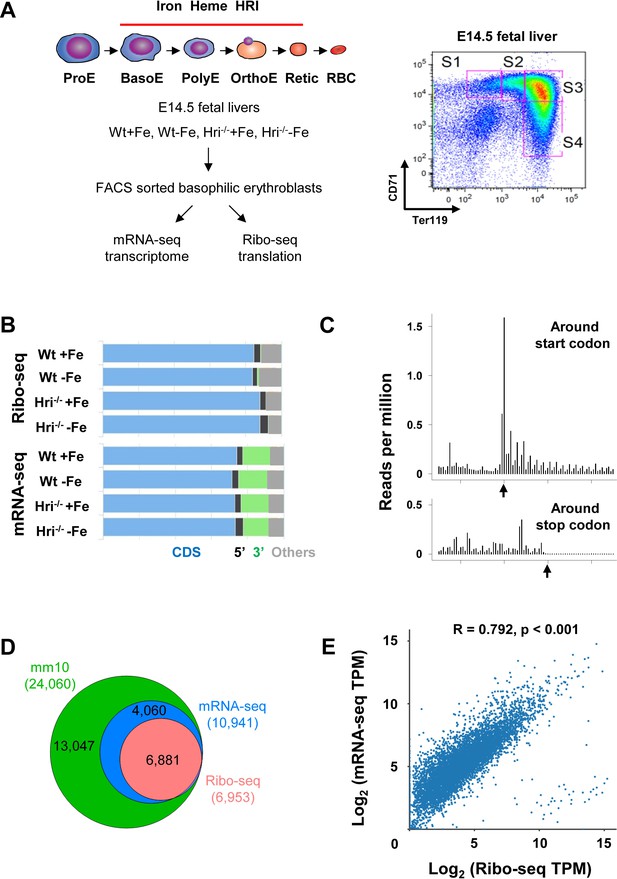
Overview of Ribo-seq and mRNA-seq data.
(A) Illustration of experimental designs. Basophilic erythroblasts (EBs) (S3) from E14.5 FLs of Wt +Fe, Wt –Fe, Hri–/– +Fe and Hri–/– –Fe embryos were sorted and subjected to Ribo-seq and mRNA-seq library preparations. (B) Distribution of the mapped reads from Ribo-seq and mRNA-seq from one replica. (C) A representative plot of the triplet periodicity of Ribo-seq from Wt –Fe EBs. Arrows indicate the start and stop codons. (D) Gene coverages of Ribo-seq and mRNA-seq data in the mouse genome (UCSC, mm10). (E) Scatter plot and correlation analysis of log2-transformed TPM (transcript per million) of Ribo-seq and mRNA-seq data from Wt +Fe EBs. E, Erythroblast; RBC, red blood cell; Retic, Reticulocyte.
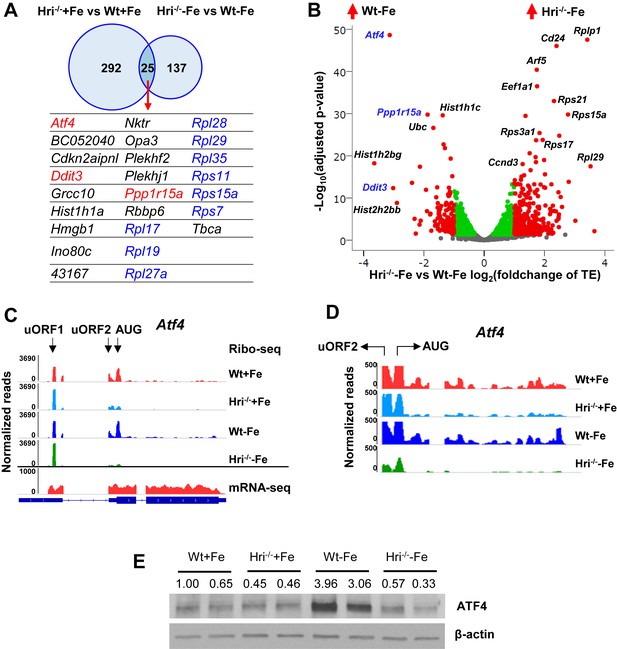
Differentially translated mRNAs in Hri–/– EBs and iron deficiency.
(A) The mRNAs that are significantly differentially translated between Hri–/– and Wt EBs in +Fe or –Fe conditions. (B) Volcano plot of mRNAs that are differentially translated between Hri–/– –Fe and Wt –Fe EBs. Red dots on the positive end of the X-axis indicate significantly differentially translated mRNAs that were upregulated in Hri–/– –Fe EBs, whereas red dots on the negative end of the X-axis indicate significantly differentially translated mRNAs that were upregulated in Wt –Fe EBs. Green and gray dots indicate mRNAs that have no significant difference in translation. TE, translational efficiency. (C) Ribosome occupancies, as visualized using Integrative Genomics Viewer (IGV), of Atf4 mRNA, with an enlarged view shown in (D). (E) ATF4 protein expression in E14.5 FL cells. Ratios of ATF4/β-actin expression are indicated.
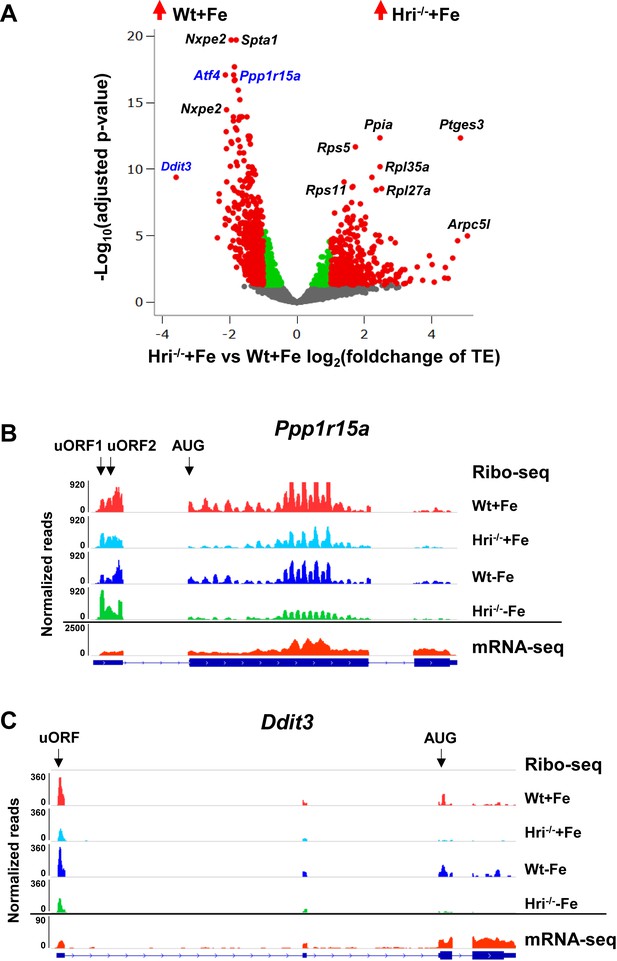
Translational regulation of ISR mRNAs by HRI.
(A) Increased translation of ISR mRNAs in Wt +Fe EBs compared to Hri–/– +Fe EBs. Red dots on the positive end of the X-axis indicate significantly differentially translated mRNAs that are upregulated in Hri–/– +Fe EBs compared to Wt +Fe EBs, whereas red dots on the negative end of the X-axis indicate mRNAs that are upregulated in Wt +Fe EBs compared to Hri–/– +Fe EBs. Green and gray dots indicate mRNAs that have no significant differences in translation. Genes labeled in blue indicate ISR mRNAs. TE, translational efficiency. (B–C) IGV-illustration of ribosome occupancies of the Ppp1r15a and Ddit3 mRNAs. Mapped reads of mRNA-seq data from Wt +Fe EBs only are shown because no significant change in mRNA levels was observed among four samples.
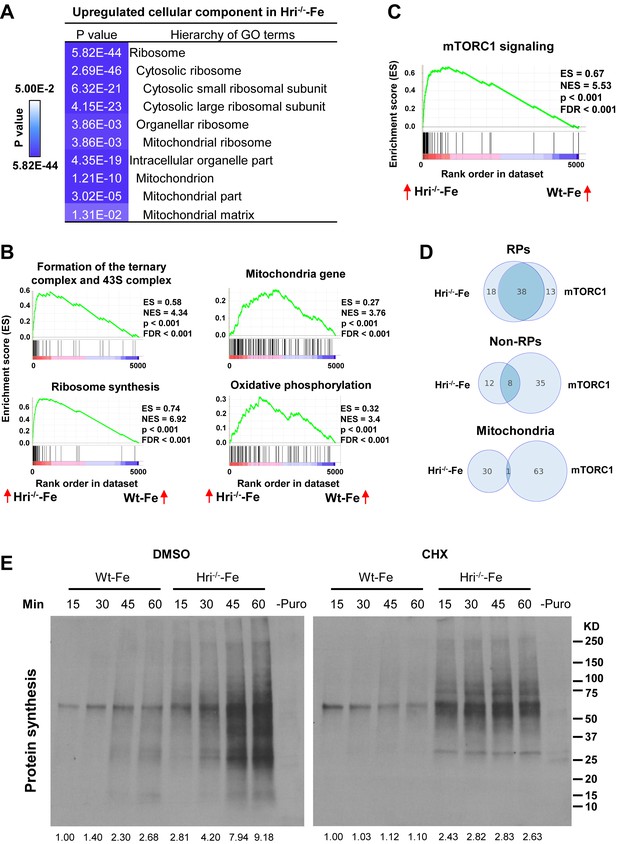
Analyses of the mRNAs that are differentially translated between Wt and Hri–/– EBs.
(A) GO analysis of the most highly translated mRNAs in Hri-/– –Fe EBs compared to Wt –Fe EBs. (B) Increased 43S initiation complex, ribosomal protein (RP) synthesis and mitochondrial pathways in Hri–/– –Fe EBs compared to Wt –Fe EBs by Gene Set Enrichment Analysis. (C) Increased mTORC1 signaling pathway in Hri–/– –Fe EBs compared to Wt –Fe EBs. (D) Venn diagrams comparing mRNAs that are differentially translated in Hri–/– –Fe EBs and Wt –Fe EBs, and that are known mTORC1 targets in the categories of RPs, non-RPs and mitochondrial proteins. (E) Protein synthesis (total (left) and in mitochondria (right)) of erythroid cells from the bone marrow (BM) of Wt –Fe and Hri–/– –Fe mice. Cells were treated with dimethyl sulfoxide (DMSO) or cycloheximide (CHX) for 15 min before the addition of puromycin for 15–60 min as indicated. Protein synthesis was determined by the nascent polypeptides covalently linked by puromycin using anti-puromycin antibody. Equal numbers of nucleated cells were loaded to each lane. The numbers of cells loaded for cytoplasmic protein synthesis were 30% of those loaded for mitochondrial protein synthesis. The results shown are from the same exposure time for developing the Western blot. Puromycin signals from polypeptides of the entire lane were quantified for protein synthesis activity and the protein synthesis in the first lane was define as 1. –Puro indicates cells without puromycin treatment, which were used as a negative control for Western signals from anti-puromycin antibody. FDR, false discovery rate; NES, normalized enrichment score.
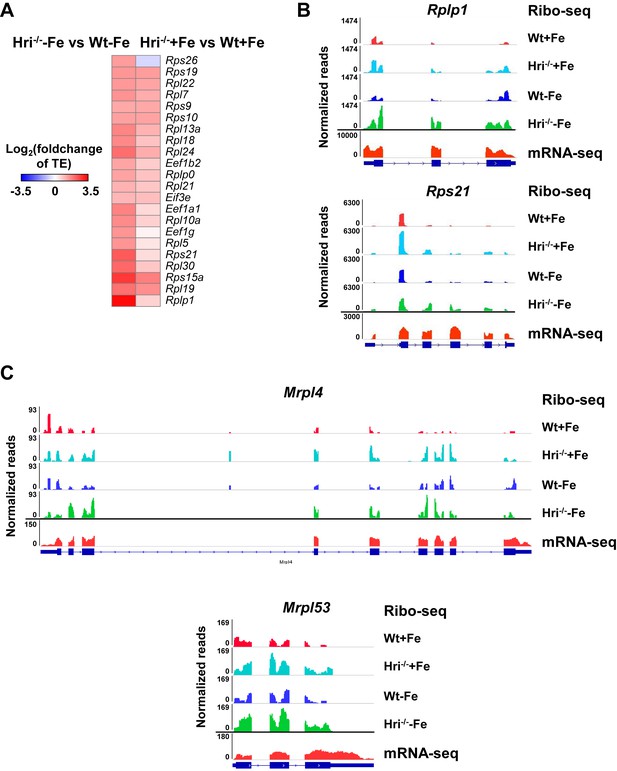
Differential translation of cytoplasmic and mitochondrial RP mRNAs between Wt and Hri–/– EBs.
(A) The heatmaps of significantly differentially translated 5’ TOP/TOP-like mRNAs, the mTORC1 translational targets. Positive values of Log2(foldchange of TE) indicate upregulated translation in Hri–/– –Fe EBs as compared to Wt –Fe EBs or in Hri–/– +Fe EBs as compared to Wt +Fe EBs. (B) IGV-illustration of ribosome occupancies of two representative cytoplasmic RP mRNAs, Rplp1 and Rps21, and (C) of the mitochondrial RP mRNAs, Mrpl4 and Mrpl53.
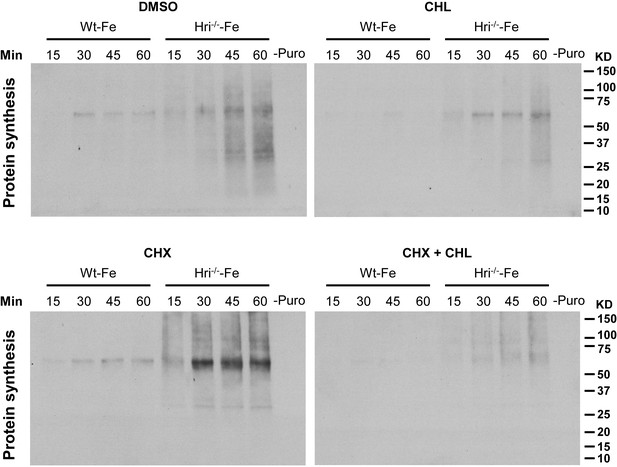
Inhibition of mitochondrial protein synthesis by chloramphenicol.
Protein Synthesis assays were performed in DMSO-treated, CHL-treated, CHX-treated and CHX + CHL-treated erythroid cells from the BM of Wt –Fe and Hri–/– –Fe mice. The DMSO-treated control is used to measure total protein synthesis in the cells, cytoplasm and mitochondria. In the presence of CHX, a cytoplasmic protein synthesis inhibitor, only mitochondrial protein synthesis was measured. In the presence of chloramphenicol (CHL), a mitochondrial protein synthesis inhibitor, only cytoplasmic protein synthesis was measured. In the presence of both CHX and CHL, protein synthesis in both the cytoplasm and mitochondria was inhibited. Cells were treated with DMSO and the inhibitors for 15 min before addition of puromycin for 15–60 min as indicated. Protein synthesis was determined by the nascent polypeptides covalently linked by puromycin using anti-puromycin antibody. Equal numbers of nucleated cells were loaded to each lane. The numbers of cells loaded for cytoplasmic protein synthesis were 30% of those loaded for mitochondrial protein synthesis. The results shown are from the same exposure time for developing the Western blots. –Puro indicates cells without puromycin treatment, which were used as a negative control for Western signals from the anti-puromycin antibody.
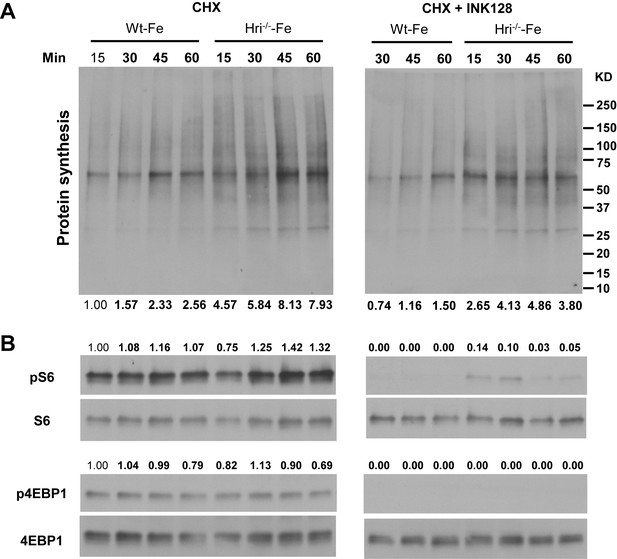
Inhibition of mitochondrial protein synthesis by INK128.
(A) Protein synthesis in the CHX-treated and CHX + INK128-treated erythroid cells from the BM of Wt –Fe and Hri–/– –Fe mice. In the presence of CHX, only mitochondrial protein synthesis was measured. In the presence of both CHX and INK128, a mTORC1 inhibitor, mTORC1-regulated mitochondrial protein synthesis was inhibited. Cells were treated with inhibitors for 15 min before the addition of puromycin for 15–60 min as indicated. Measurement of protein synthesis was as described in Figure 3—figure supplement 2. (B) Inhibition of mTORC1 activities as measured by pS6 and p4EBP1 levels. Signal from Wt –Fe with CHX treatment and 15 min treatment of puromycin was defined as 1. Equal numbers of nucleated cells were loaded into each lane. The results shown are from the same exposure time for developing the Western blot.
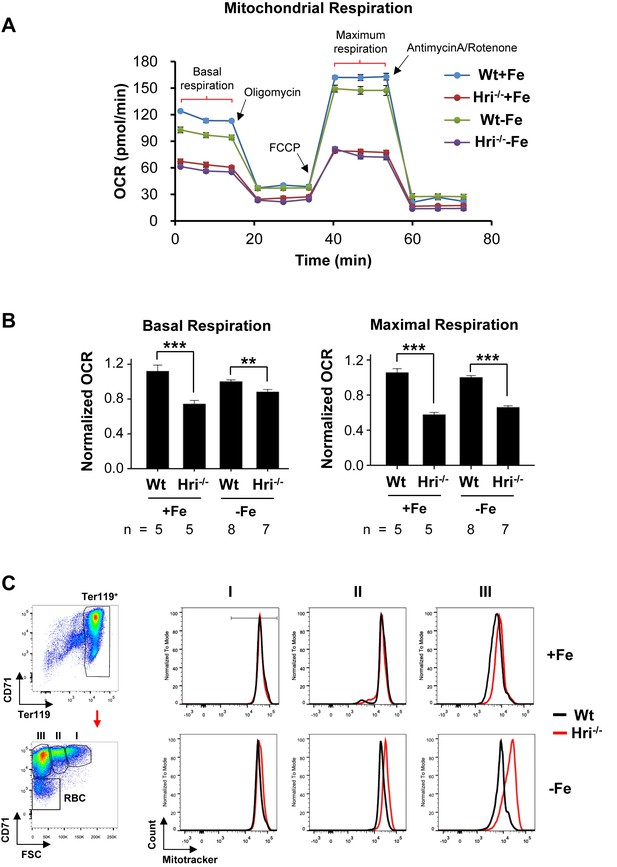
Decreased mitochondrial respiration in HRI deficiency.
(A) A representative result for the oxygen consumption rate (OCR) of Wt and Hri–/– erythroid cells. Erythroid cells were isolated from the BM of Wt and Hri-–/– mice in +Fe and –Fe conditions. Five technical replicas were performed for each condition. (B) Quantitative analysis of basal and maximal OCR from four separate experiments. The numbers of mice used in each condition are indicated. OCR of Wt –Fe erythroid cells was defined as 1. Data were presented as mean ± SE. **p<0.01, ***p<0.001. (C) Representative plots of mitochondrial mass of erythroid cells at different differentiation stages obtained from mitotracker flow cytometry analysis. A diagram of gating for the differentiation stages is shown on the left.
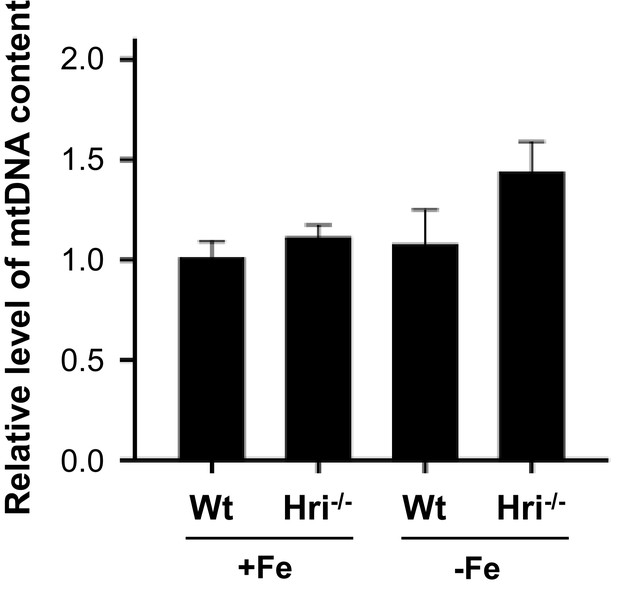
The relative mitochondrial DNA contents of erythroid cells.
The ratios of mitochondrial DNA (mtDNA) versus genomic DNA (gDNA) were determined by qPCR using the ΔΔCt method for erythroid cells isolated from bone marrow samples. The level in Wt +Fe was defined as 1 (n = 3). Data are presented as mean ± SE.
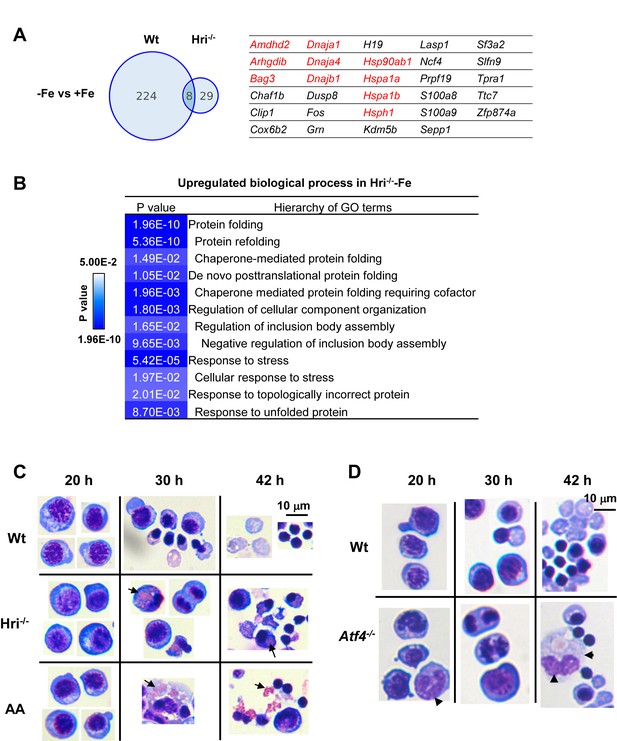
Increased expression of cytoplasmic chaperones, accumulation of unfolded protein inclusions and impaired FL differentiation in Hri–/– erythroid cells.
(A) Venn diagrams depicting the numbers of mRNAs that are significantly differentially expressed between –Fe and +Fe conditions in Wt or Hri–/– EBs. The table on the right lists mRNAs that were expressed at higher level only in Hri–/––Fe EBs. Results were obtained from three biological replicas. (B) GO terms related to protein folding which were most highly upregulated in Hri–/– –Fe EBs compared to Wt –Fe EBs from the Gene ontology analysis of significantly differentially expressed mRNAs. (C–D) Ex vivo erythroid differentiation from HRI-, eIF2αP-, and ATF4-deficient FL erythroid progenitors. The representative images are of cytospin slides stained with May-Grunwald/Giemsa staining. Cells at 20, 30 and 42 hr of erythroid differentiation are shown. AA, universal eIF2α Ala51/Ala51 knockin resulting in complete ablation of eIF2αP. Arrow in (C) indicates globin inclusions and arrowhead in (D) indicates myeloid cells. Numbers of FL differentiation performed were: n = 6 for Wt and Hri–/–; n = 4 for AA and n = 3 for Atf4–/–. Some of the cell images in (C) were cropped from the same slides and compiled together for illustrative purposes.
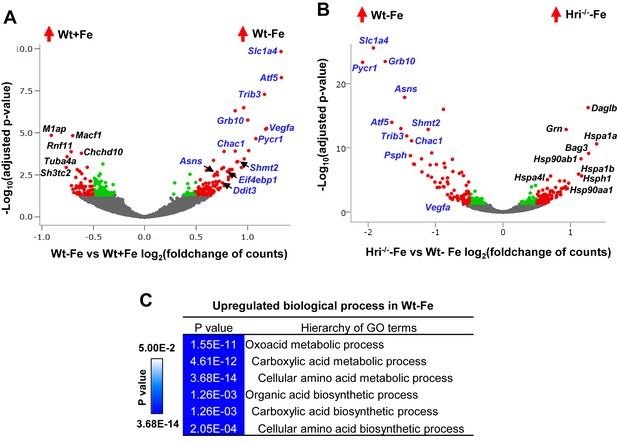
Analyses of differentially expressed mRNAs in iron and HRI deficiencies.
(A) Volcano plots of mRNAs that are differentially expressed between Wt –Fe and Wt +Fe or (B) between Hri–/– –Fe and Wt –Fe EBs. Red dots represent the significantly differentially expressed mRNAs. Green and gray dots indicate mRNAs whose expression does not differ significantly. ATF4 target genes are labeled in blue. (C) GO terms for amino acid metabolism that are most highly upregulated in Wt –Fe EBs compared to Hri–/– –Fe EBs.
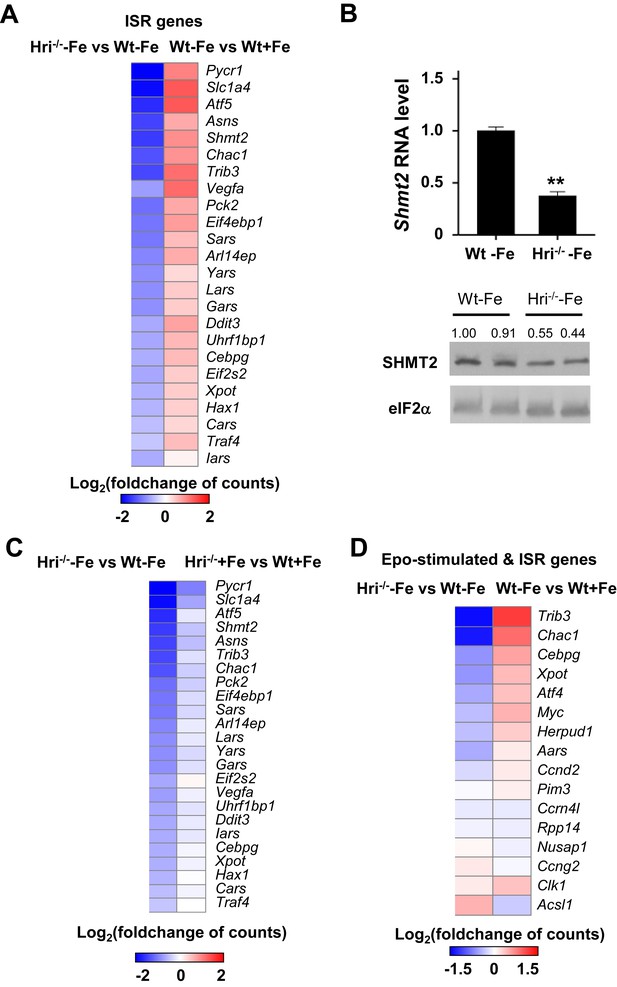
Analyses of differentially expressed ISR-target mRNAs.
(A) Heatmaps of ISR-target genes that have a significant difference in expression levels between Hri–/– –Fe and Wt –Fe EBs or between Wt –Fe and Wt +Fe EBs. (B) Expression of Shmt2 in E14.5 FL cells. The expression level in Wt –Fe was defined as 1 (n = 3). Data were presented as mean ± SE. **p<0.01. (C) The heatmaps of ISR-target genes that are differentially expressed between Hri–/– –Fe and Wt –Fe EBs or between Hri–/– +Fe and Wt +Fe EBs. (D) Epo-stimulated ISR-target genes that are differentially expressed between Hri–/– -Fe and Wt –Fe EBs or between Wt –Fe and Wt +Fe EBs.
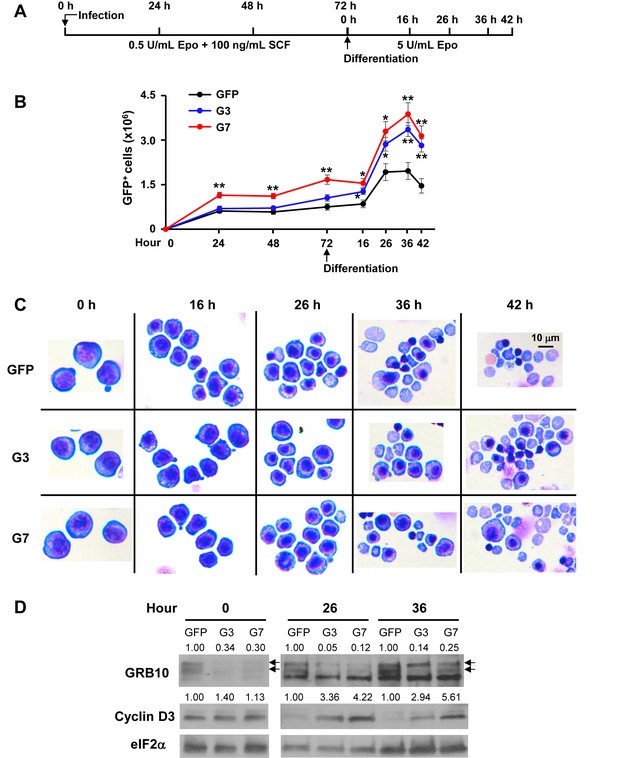
Knockdown of Grb10 increases proliferation and inhibits differentiation of FL erythroblasts.
(A) Designs of shRNA knockdown experiments using Lin–Ter119–CD71– FL erythroid progenitors. (B) Proliferation of control and Grb10 knockdown GFP+ cells. Three biological replicas were performed. Data are presented as means ± SEs. *p<0.05, **p<0.01. (C) Representative cell morphology of GFP control and Grb10 knockdown cells. (D) GRB10 and Cyclin D3 protein levels of GFP control and Grb10 knockdown cells during differentiation. Arrows indicate the doublets of GRB10. The green fluorescent protein (GFP) control was infected with retrovirus expressing GFP only; G3 and G7 were infected with retroviruses expressing shRNA_G3 and shRNA_G7, respectively.
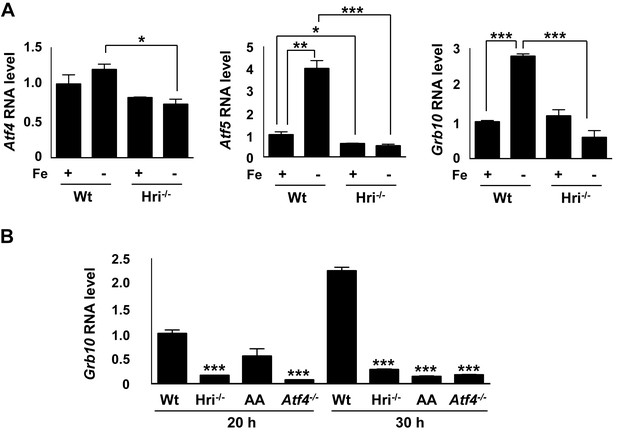
Expression of Atf4, Atf5 and Grb10 mRNAs in HRI-ISR defective erythroid cells.
(A) Atf4, Atf5 and Grb10 expression in sorted basophilic EBs as illustrated in Figure 1A. The expression level in Wt +Fe EBs was defined as 1 (n = 3). (B) Grb10 expression at 20 hr and 30 hr of ex vivo differentiation of Wt, Hri–/–, AA and Atf4 FL erythroid progenitors. The expression level in Wt EBs at 20 hr was defined as 1. For each time point, expression levels in Hri–/–, AA and Atf4 EBs were compared to those in Wt EBs (n = 3). Data are presented as means ± SEs. *p<0.05, **p<0.01, ***p<0.001.
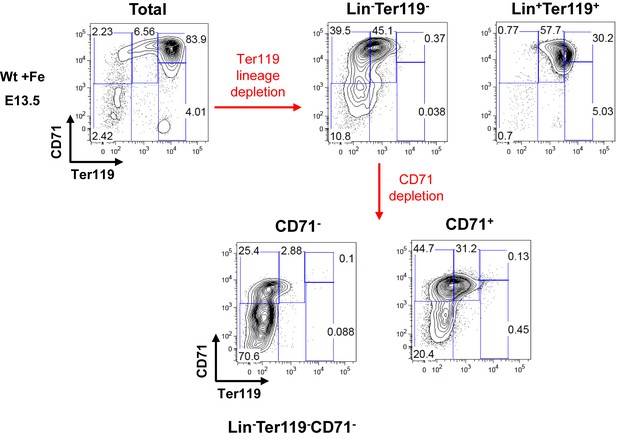
Enrichment of Lin–Ter119–CD71– erythroid progenitors from Wt +Fe E13.5 FLs for ex vivo experiments.
https://doi.org/10.7554/eLife.46976.017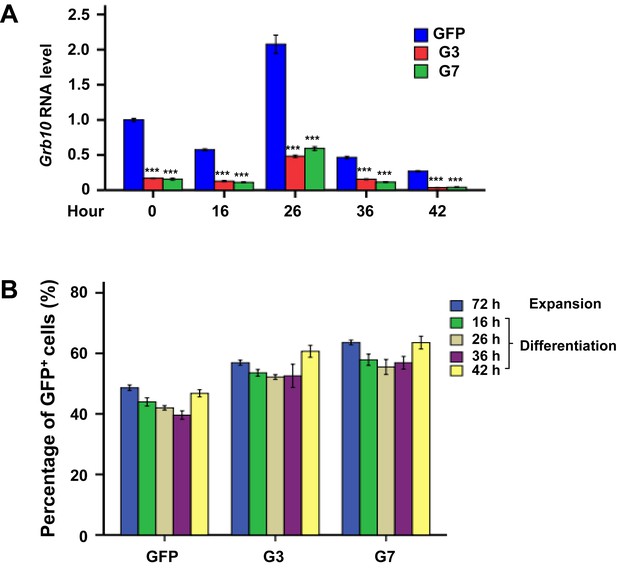
Knockdown efficiency of Grb10 in FL cells during ex vivo differentiation.
(A) Knockdown efficiency of Grb10 RNA by shRNA_G3 and shRNA_G7. Grb10 expression in the GFP control at 0 hr of differentiation is defined as 1. (B) Percentages of GFP+ cells during the expansion and differentiation phases. The GFP control was infected with retrovirus expressing GFP only. G3 and G7 were infected with retroviruses expressing shRNA_G3 and shRNA_G7, respectively. Data are presented as means ± SEs. ***p<0.001.
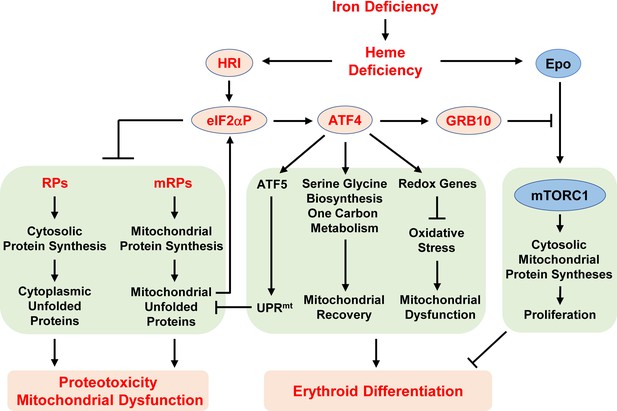
Proposed model of the regulation of erythropoiesis by heme and the HRI-ISR pathway during ID.
Diet-induced systemic ID results in heme deficiency, which activates HRI to phosphorylate its substrate eIF2α. The primary function of eIF2αP is to inhibit general protein synthesis in both the cytoplasm and mitochondria via decreased translation of ribosomal protein mRNAs from both cellular compartments. In the absence of HRI, continued protein synthesis results in the accumulation of unfolded proteins in both the cytoplasm and mitochondria, leading to proteotoxicity and mitochonfrial dysfunction. Second, eIF2αP also enhances the translation of Atf4 mRNA via regulation by uORFs. Atf4 mRNA is the mRNA that shows the greatest differences in translation levels between Wt and Hri–/– EBs. The ATF4 protein induces the expression of three genetic pathways to activate mitochondrial UPR (UPRmt), reprogram mitochondrial metabolism and reduce oxidative stress, all of which enables adaptation to ID and erythroid differentiation. Strikingly, the expression of ATF4 target genes is most highly activated in ID and requires HRI. Thus, assessment of the global genome-wide gene expression of primary EBs in vivo reveals that HRI-ISR contributes most significantly to adaptation to ID. We have shown previously that HRI-ISR suppressed mTORC1 signaling, which is activated by elevated Epo levels in ID (Zhang et al., 2018). Here, we provide evidence that GRB10 that is induced by ATF4 may be one of the molecules involved in the repression of mTORC1 signaling.
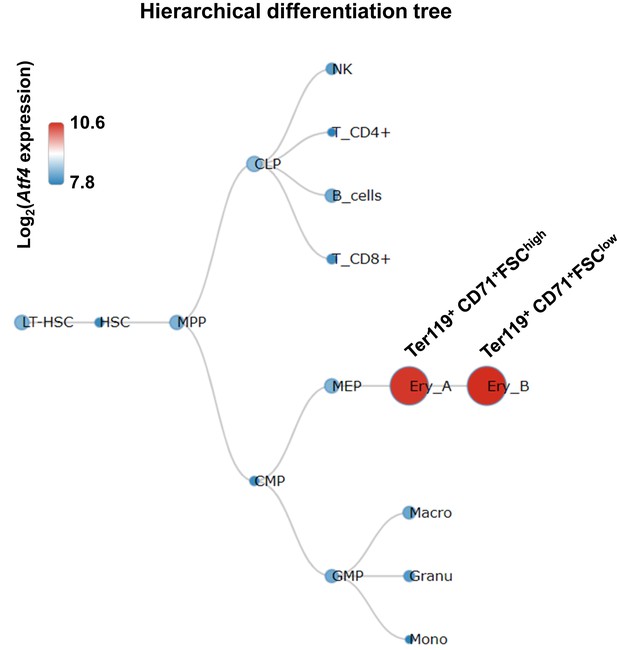
Expression of Atf4 mRNA in hematopoietic cells.
The plot of Atf4 expression levels in hematopoietic cells of murine bone marrow was obtained from the BloodSpot database (Bagger et al., 2016). The dataset used for ploting the tree was Mouse Normal RNA-seq, which was derived from the work of Lara-Astiaso et al. (2014). Expression level is log2-transformed and visualized by the size and color of the nodes.
Additional files
-
Supplementary file 1
List of diifferentially translated mRNAs by HRI and mTORC1.
Supplementary File 1a. Quality control and mapping of Ribo-seq and mRNA-seq. Supplementary File 1b. Complete gene list of differentially translated mRNAs. Supplementary File 1c. Eif2ak1, Ppp1r15a and Atf4 mRNAs are highly expressed in basophilic erythroblasts. Supplementary File 1d. Higher ribosome occupancy at the AUG of uORF1 in Atf4 mRNA.Supplementary File 1e. Complete gene list of differentially expressed mRNAs.
- https://doi.org/10.7554/eLife.46976.021
-
Supplementary file 2
List of mRNAs that are differentially translated between Hri–/– –Fe EBs and Wt –Fe EBs and known mTORC1 targets in the categories of (a) RPs and non-RPs and (b) mitochondrial proteins.
- https://doi.org/10.7554/eLife.46976.022
-
Supplementary file 3
List of RNA primers, antibodies and ShRNA oligonucleotides.
Supplementary File 3a. DNA sequence of qPCR primers. Supplementary File 3b. List of antibodies used for western blot analyses. Supplementary File 3c. DNA sequences of shRNA oligonucleotides.
- https://doi.org/10.7554/eLife.46976.023
-
Transparent reporting form
- https://doi.org/10.7554/eLife.46976.024





